The Role of the N-Terminal Domains of Bacterial Initiator DnaA in the Assembly and Regulation of the Bacterial Replication Initiation Complex
Abstract
:1. Introduction
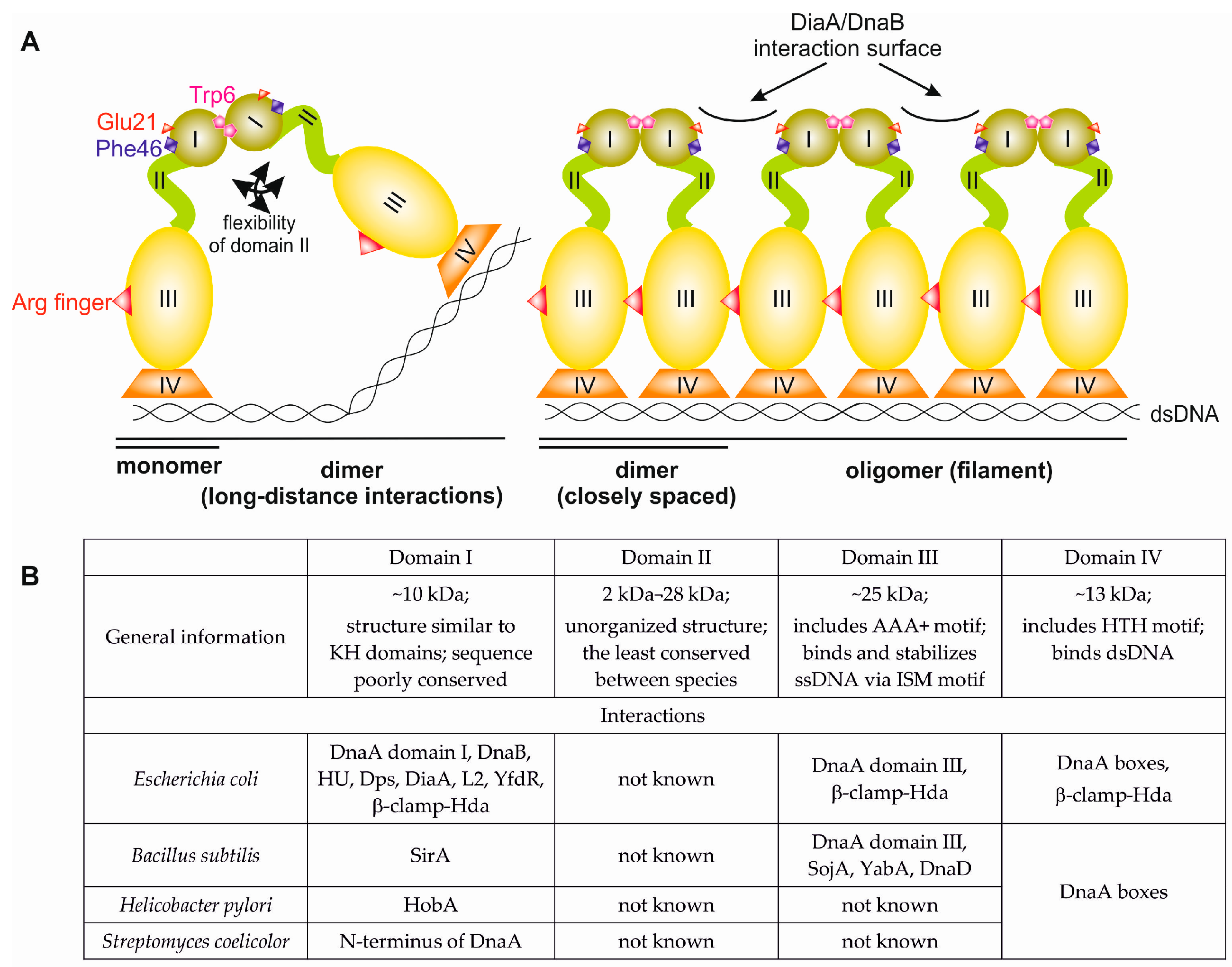
2. Bacterial DnaA—General Overview of the Structure and Function
3. N-Terminus of Bacterial DnaA
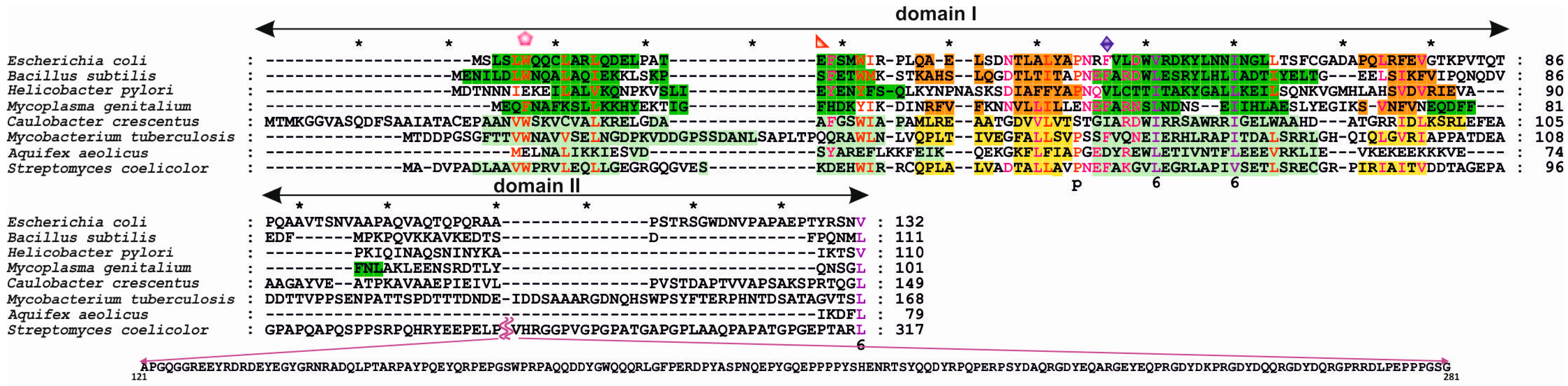
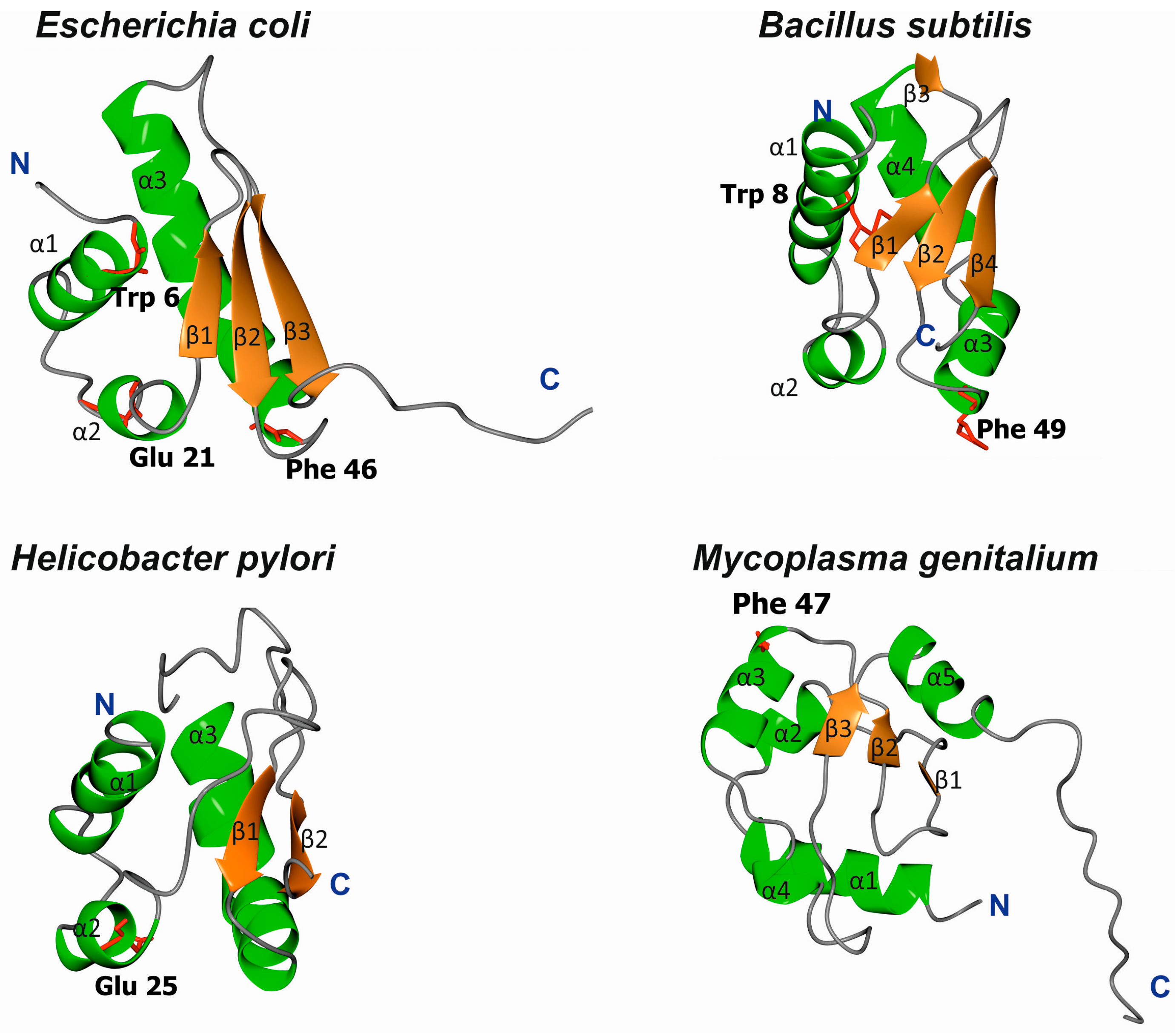
3.1. Structures of Bacterial DnaA Domains I and II
3.2. Escherichia Coli DnaA Domain I
3.3. Bacillus Subtilis DnaA Domain I
3.4. Helicobacter Pylori DnaA Domain I
3.5. Streptomyces Coelicolor DnaA Domain I
3.6. DnaA Domain II
4. Conclusions and Perspectives
Acknowledgments
Conflicts of Interest
References
- Leonard, A.C.; Méchali, M. DNA replication origins. Cold Spring Harb. Perspect. Biol. 2013, 5, a010116. [Google Scholar] [CrossRef] [PubMed]
- Kawakami, H.; Katayama, T. DnaA, ORC, and Cdc6: Similarity beyond the domains of life and diversity. Biochem. Cell Biol. Biochim. Biol. Cell. 2010, 88, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Costa, A.; Hood, I.V.; Berger, J.M. Mechanisms for initiating cellular DNA replication. Annu. Rev. Biochem. 2013, 82, 25–54. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, M.; Langston, L.; Stillman, B. Principles and concepts of DNA replication in bacteria, archaea, and eukarya. Cold Spring Harb. Perspect. Biol. 2013, 5. [Google Scholar] [CrossRef] [PubMed]
- Deegan, T.D.; Diffley, J.F. MCM: One ring to rule them all. Curr. Opin. Struct. Biol. 2016, 37, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Katayama, T.; Ozaki, S.; Keyamura, K.; Fujimitsu, K. Regulation of the replication cycle: Conserved and diverse regulatory systems for DnaA and oriC. Nat. Rev. Microbiol. 2010, 8, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Hwang, D.S.; Kornberg, A. Opening of the replication origin of Escherichia coli by DnaA protein with protein HU or IHF. J. Biol. Chem. 1992, 267, 23083–23086. [Google Scholar] [PubMed]
- Mott, M.L.; Berger, J.M. DNA replication initiation: Mechanisms and regulation in bacteria. Nat. Rev. Microbiol. 2007, 5, 343–354. [Google Scholar] [CrossRef] [PubMed]
- Wolański, M.; Donczew, R.; Zawilak-Pawlik, A.; Zakrzewska-Czerwińska, J. oriC-encoded instructions for the initiation of bacterial chromosome replication. Front. Microbiol. 2014, 5, 735. [Google Scholar] [PubMed]
- Chodavarapu, S.; Kaguni, J.M. Replication Initiation in Bacteria. The Enzymes 2016, 39, 1–30. [Google Scholar] [PubMed]
- Smith, J.L.; Grossman, A.D. In Vitro Whole Genome DNA Binding Analysis of the Bacterial Replication Initiator and Transcription Factor DnaA. PLoS Genet. 2015, 11, e1005258. [Google Scholar] [CrossRef] [PubMed]
- Messer, W.; Weigel, C. DnaA as a transcription regulator. Methods Enzymol. 2003, 370, 338–349. [Google Scholar] [PubMed]
- Erzberger, J.P.; Mott, M.L.; Berger, J.M. Structural basis for ATP-dependent DnaA assembly and replication-origin remodeling. Nat. Struct. Mol. Biol. 2006, 13, 676–683. [Google Scholar] [CrossRef] [PubMed]
- Ran, L.; Larsson, J.; Vigil-Stenman, T.; Nylander, J.A.A.; Ininbergs, K.; Zheng, W.-W.; Lapidus, A.; Lowry, S.; Haselkorn, R.; Bergman, B. Genome Erosion in a Nitrogen-Fixing Vertically Transmitted Endosymbiotic Multicellular Cyanobacterium. PLoS ONE 2010, 5, e11486. [Google Scholar] [CrossRef]
- Akman, L.; Yamashita, A.; Watanabe, H.; Oshima, K.; Shiba, T.; Hattori, M.; Aksoy, S. Genome sequence of the endocellular obligate symbiont of tsetse flies, Wigglesworthia glossinidia. Nat. Genet. 2002, 32, 402–407. [Google Scholar] [CrossRef] [PubMed]
- Gil, R.; Silva, F.J.; Zientz, E.; Delmotte, F.; González-Candelas, F.; Latorre, A.; Rausell, C.; Kamerbeek, J.; Gadau, J.; Hölldobler, B.; et al. The genome sequence of Blochmannia floridanus: comparative analysis of reduced genomes. Proc. Natl. Acad. Sci. USA 2003, 100, 9388–9393. [Google Scholar] [CrossRef] [PubMed]
- Erzberger, J.P.; Pirruccello, M.M.; Berger, J.M. The structure of bacterial DnaA: implications for general mechanisms underlying DNA replication initiation. EMBO J. 2002, 21, 4763–4773. [Google Scholar] [CrossRef] [PubMed]
- Duderstadt, K.E.; Berger, J.M. A structural framework for replication origin opening by AAA+ initiation factors. Curr. Opin. Struct. Biol. 2013, 23, 144–153. [Google Scholar] [CrossRef] [PubMed]
- Martinez, M.P.; Jones, J.M.; Bruck, I.; Kaplan, D.L. Origin DNA Melting-An Essential Process with Divergent Mechanisms. Genes 2017, 8, 26. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, M.; Noguchi, Y.; Sakiyama, Y.; Kawakami, H.; Katayama, T.; Takada, S. Near-atomic structural model for bacterial DNA replication initiation complex and its functional insights. Proc. Natl. Acad. Sci. USA 2016, 113, E8021–E8030. [Google Scholar] [CrossRef] [PubMed]
- Duderstadt, K.E.; Chuang, K.; Berger, J.M. DNA stretching by bacterial initiators promotes replication origin opening. Nature 2011, 478, 209–213. [Google Scholar] [CrossRef] [PubMed]
- Ozaki, S.; Kawakami, H.; Nakamura, K.; Fujikawa, N.; Kagawa, W.; Park, S.-Y.; Yokoyama, S.; Kurumizaka, H.; Katayama, T. A common mechanism for the ATP-DnaA-dependent formation of open complexes at the replication origin. J. Biol. Chem. 2008, 283, 8351–8362. [Google Scholar] [CrossRef] [PubMed]
- Ozaki, S.; Katayama, T. Highly organized DnaA-oriC complexes recruit the single-stranded DNA for replication initiation. Nucleic Acids Res. 2012, 40, 1648–1665. [Google Scholar] [CrossRef] [PubMed]
- Ozaki, S.; Noguchi, Y.; Hayashi, Y.; Miyazaki, E.; Katayama, T. Differentiation of the DnaA-oriC Subcomplex for DNA Unwinding in a Replication Initiation Complex. J. Biol. Chem. 2012, 287, 37458–37471. [Google Scholar] [CrossRef] [PubMed]
- Carr, K.M.; Kaguni, J.M. Stoichiometry of DnaA and DnaB protein in initiation at the Escherichia coli chromosomal origin. J. Biol. Chem. 2001, 276, 44919–44925. [Google Scholar] [CrossRef] [PubMed]
- Carr, K.M.; Kaguni, J.M. Escherichia coli DnaA protein loads a single DnaB helicase at a DnaA box hairpin. J. Biol. Chem. 2002, 277, 39815–39822. [Google Scholar] [CrossRef] [PubMed]
- Mott, M.L.; Erzberger, J.P.; Coons, M.M.; Berger, J.M. Structural synergy and molecular crosstalk between bacterial helicase loaders and replication initiators. Cell 2008, 135, 623–634. [Google Scholar] [CrossRef] [PubMed]
- Smits, W.K.; Goranov, A.I.; Grossman, A.D. Ordered association of helicase loader proteins with the Bacillus subtilis origin of replication in vivo. Mol. Microbiol. 2010, 75, 452–461. [Google Scholar] [CrossRef] [PubMed]
- Soultanas, P. Loading mechanisms of ring helicases at replication origins: Helicase loading. Mol. Microbiol. 2012, 84, 6–16. [Google Scholar] [CrossRef] [PubMed]
- Brézellec, P.; Vallet-Gely, I.; Possoz, C.; Quevillon-Cheruel, S.; Ferat, J.-L. DciA is an ancestral replicative helicase operator essential for bacterial replication initiation. Nat. Commun. 2016, 7, 13271. [Google Scholar] [CrossRef] [PubMed]
- Su’etsugu, M.; Harada, Y.; Keyamura, K.; Matsunaga, C.; Kasho, K.; Abe, Y.; Ueda, T.; Katayama, T. The DnaA N-terminal domain interacts with Hda to facilitate replicase clamp-mediated inactivation of DnaA. Environ. Microbiol. 2013, 15, 3183–3195. [Google Scholar] [CrossRef] [PubMed]
- Wargachuk, R.; Marczynski, G.T. The Caulobacter crescentus Homolog of DnaA (HdaA) Also Regulates the Proteolysis of the Replication Initiator Protein DnaA. J. Bacteriol. 2015, 197, 3521–3532. [Google Scholar] [CrossRef] [PubMed]
- Scholefield, G.; Errington, J.; Murray, H. Soj/ParA stalls DNA replication by inhibiting helix formation of the initiator protein DnaA. EMBO J. 2012, 31, 1542–1555. [Google Scholar] [CrossRef] [PubMed]
- Scholefield, G.; Murray, H. YabA and DnaD inhibit helix assembly of the DNA replication initiation protein DnaA. Mol. Microbiol. 2013, 90, 147–159. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.; Ogasawara, N.; Ishikawa, S. The functional analysis of YabA, which interacts with DnaA and regulates initiation of chromosome replication in Bacillus subtils. Genes Genet. Syst. 2008, 83, 111–125. [Google Scholar] [CrossRef] [PubMed]
- Sutton, M.D.; Carr, K.M.; Vicente, M.; Kaguni, J.M. Escherichia coli DnaA protein. The N-terminal domain and loading of DnaB helicase at the E. coli chromosomal origin. J. Biol. Chem. 1998, 273, 34255–34262. [Google Scholar] [CrossRef] [PubMed]
- Abe, Y.; Jo, T.; Matsuda, Y.; Matsunaga, C.; Katayama, T.; Ueda, T. Structure and function of DnaA N-terminal domains: Specific sites and mechanisms in inter-DnaA interaction and in DnaB helicase loading on oriC. J. Biol. Chem. 2007, 282, 17816–17827. [Google Scholar] [CrossRef] [PubMed]
- Lowery, T.J.; Pelton, J.G.; Chandonia, J.-M.; Kim, R.; Yokota, H.; Wemmer, D.E. NMR structure of the N-terminal domain of the replication initiator protein DnaA. J. Struct. Funct. Genomics 2007, 8, 11–17. [Google Scholar] [CrossRef] [PubMed]
- Simossis, V.A.; Heringa, J. PRALINE: A multiple sequence alignment toolbox that integrates homology-extended and secondary structure information. Nucleic Acids Res. 2005, 33, W289–W294. [Google Scholar] [CrossRef] [PubMed]
- Natrajan, G.; Noirot-Gros, M.F.; Zawilak-Pawlik, A.; Kapp, U.; Terradot, L. The structure of a DnaA/HobA complex from Helicobacter pylori provides insight into regulation of DNA replication in bacteria. Proc. Natl. Acad. Sci. USA 2009, 106, 21115–21120. [Google Scholar] [CrossRef] [PubMed]
- Jameson, K.H.; Rostami, N.; Fogg, M.J.; Turkenburg, J.P.; Grahl, A.; Murray, H.; Wilkinson, A.J. Structure and interactions of the Bacillus subtilis sporulation inhibitor of DNA replication, SirA, with domain I of DnaA. Mol. Microbiol. 2014, 93, 975–991. [Google Scholar] [CrossRef] [PubMed]
- Simmons, L.A.; Felczak, M.; Kaguni, J.M. DnaA Protein of Escherichia coli: oligomerization at the E. coli chromosomal origin is required for initiation and involves specific N-terminal amino acids. Mol. Microbiol. 2003, 49, 849–858. [Google Scholar] [CrossRef] [PubMed]
- Felczak, M.M.; Simmons, L.A.; Kaguni, J.M. An essential tryptophan of Escherichia coli DnaA protein functions in oligomerization at the E. coli replication origin. J. Biol. Chem. 2005, 280, 24627–24633. [Google Scholar] [CrossRef] [PubMed]
- Keyamura, K.; Abe, Y.; Higashi, M.; Ueda, T.; Katayama, T. DiaA dynamics are coupled with changes in initial origin complexes leading to helicase loading. J. Biol. Chem. 2009, 284, 25038–25050. [Google Scholar] [CrossRef] [PubMed]
- Valverde, R.; Edwards, L.; Regan, L. Structure and function of KH domains. FEBS J. 2008, 275, 2712–2726. [Google Scholar] [CrossRef] [PubMed]
- Richardson, T.T.; Harran, O.; Murray, H. The bacterial DnaA-trio replication origin element specifies single-stranded DNA initiator binding. Nature 2016, 534, 412–416. [Google Scholar] [CrossRef] [PubMed]
- Sutton, M.D.; Kaguni, J.M. Novel alleles of the Escherichia coli dnaA gene. J. Mol. Biol. 1997, 271, 693–703. [Google Scholar] [CrossRef] [PubMed]
- Weigel, C.; Schmidt, A.; Seitz, H.; Tüngler, D.; Welzeck, M.; Messer, W. The N-terminus promotes oligomerization of the Escherichia coli initiator protein DnaA. Mol. Microbiol. 1999, 34, 53–66. [Google Scholar] [CrossRef] [PubMed]
- Messer, W.; Blaesing, F.; Majka, J.; Nardmann, J.; Schaper, S.; Schmidt, A.; Seitz, H.; Speck, C.; Tüngler, D.; Wegrzyn, G.; Weigel, C.; Welzeck, M.; Zakrzewska-Czerwinska, J. Functional domains of DnaA proteins. Biochimie 1999, 81, 819–825. [Google Scholar] [CrossRef]
- Messer, W. The bacterial replication initiator DnaA. DnaA and oriC, the bacterial mode to initiate DNA replication. FEMS Microbiol. Rev. 2002, 26, 355–374. [Google Scholar] [PubMed]
- Messer, W.; Blaesing, F.; Jakimowicz, D.; Krause, M.; Majka, J.; Nardmann, J.; Schaper, S.; Seitz, H.; Speck, C.; Weigel, C.; et al. Bacterial replication initiator DnaA. Rules for DnaA binding and roles of DnaA in origin unwinding and helicase loading. Biochimie 2001, 83, 5–12. [Google Scholar] [CrossRef]
- Miller, D.T.; Grimwade, J.E.; Betteridge, T.; Rozgaja, T.; Torgue, J.J.-C.; Leonard, A.C. Bacterial origin recognition complexes direct assembly of higher-order DnaA oligomeric structures. Proc. Natl. Acad. Sci. USA 2009, 106, 18479–18484. [Google Scholar] [CrossRef] [PubMed]
- Seitz, H.; Weigel, C.; Messer, W. The interaction domains of the DnaA and DnaB replication proteins of Escherichia coli. Mol. Microbiol. 2000, 37, 1270–1279. [Google Scholar] [CrossRef] [PubMed]
- Zawilak-Pawlik, A.; Donczew, R.; Szafrański, S.; Mackiewicz, P.; Terradot, L.; Zakrzewska-Czerwińska, J. DiaA/HobA and DnaA: A pair of proteins co-evolved to cooperate during bacterial orisome assembly. J. Mol. Biol. 2011, 408, 238–251. [Google Scholar] [CrossRef] [PubMed]
- Keyamura, K.; Fujikawa, N.; Ishida, T.; Ozaki, S.; Su’etsugu, M.; Fujimitsu, K.; Kagawa, W.; Yokoyama, S.; Kurumizaka, H.; Katayama, T. The interaction of DiaA and DnaA regulates the replication cycle in E. coli by directly promoting ATP DnaA-specific initiation complexes. Genes Dev. 2007, 21, 2083–2099. [Google Scholar] [CrossRef] [PubMed]
- Ishida, T.; Akimitsu, N.; Kashioka, T.; Hatano, M.; Kubota, T.; Ogata, Y.; Sekimizu, K.; Katayama, T. DiaA, a novel DnaA-binding protein, ensures the timely initiation of Escherichia coli chromosome replication. J. Biol. Chem. 2004, 279, 45546–45555. [Google Scholar] [CrossRef] [PubMed]
- Kaguni, J.M. Replication initiation at the Escherichia coli chromosomal origin. Curr. Opin. Chem. Biol. 2011, 15, 606–613. [Google Scholar] [CrossRef] [PubMed]
- Su’etsugu, M.; Shimuta, T.-R.; Ishida, T.; Kawakami, H.; Katayama, T. Protein associations in DnaA-ATP hydrolysis mediated by the Hda-replicase clamp complex. J. Biol. Chem. 2005, 280, 6528–6536. [Google Scholar] [CrossRef] [PubMed]
- Keyamura, K.; Katayama, T. DnaA protein DNA-binding domain binds to Hda protein to promote inter-AAA+ domain interaction involved in regulatory inactivation of DnaA. J. Biol. Chem. 2011, 286, 29336–29346. [Google Scholar] [CrossRef] [PubMed]
- Fujimitsu, K.; Senriuchi, T.; Katayama, T. Specific genomic sequences of E. coli promote replicational initiation by directly reactivating ADP-DnaA. Genes Dev. 2009, 23, 1221–1233. [Google Scholar] [CrossRef] [PubMed]
- Saxena, R.; Fingland, N.; Patil, D.; Sharma, A.K.; Crooke, E. Crosstalk between DnaA Protein, the Initiator of Escherichia coli Chromosomal Replication, and Acidic Phospholipids Present in Bacterial Membranes. Int. J. Mol. Sci. 2013, 14, 8517–8537. [Google Scholar] [CrossRef] [PubMed]
- Aranovich, A.; Braier-Marcovitz, S.; Ansbacher, E.; Granek, R.; Parola, A.H.; Fishov, I. N-terminal-mediated oligomerization of DnaA drives the occupancy-dependent rejuvenation of the protein on the membrane. Biosci. Rep. 2015, 35. [Google Scholar] [CrossRef] [PubMed]
- Aranovich, A.; Gdalevsky, G.Y.; Cohen-Luria, R.; Fishov, I.; Parola, A.H. Membrane-catalyzed nucleotide exchange on DnaA. Effect of surface molecular crowding. J. Biol. Chem. 2006, 281, 12526–12534. [Google Scholar] [CrossRef] [PubMed]
- Macvanin, M.; Adhya, S. Architectural organization in E. coli nucleoid. Biochim. Biophys. Acta 2012, 1819, 830–835. [Google Scholar] [CrossRef] [PubMed]
- Ryan, V.T.; Grimwade, J.E.; Nievera, C.J.; Leonard, A.C. IHF and HU stimulate assembly of pre-replication complexes at Escherichia coli oriC by two different mechanisms. Mol. Microbiol. 2002, 46, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Chodavarapu, S.; Felczak, M.M.; Yaniv, J.R.; Kaguni, J.M. Escherichia coli DnaA interacts with HU in initiation at the E. coli replication origin. Mol. Microbiol. 2008, 67, 781–792. [Google Scholar] [CrossRef] [PubMed]
- Claret, L.; Rouviere-Yaniv, J. Variation in HU composition during growth of Escherichia coli: The heterodimer is required for long term survival. J. Mol. Biol. 1997, 273, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Chodavarapu, S.; Gomez, R.; Vicente, M.; Kaguni, J.M. Escherichia coli Dps interacts with DnaA protein to impede initiation: A model of adaptive mutation. Mol. Microbiol. 2008, 67, 1331–1346. [Google Scholar] [CrossRef] [PubMed]
- Noguchi, Y.; Katayama, T. The Escherichia coli Cryptic Prophage Protein YfdR Binds to DnaA and Initiation of Chromosomal Replication Is Inhibited by Overexpression of the Gene Cluster yfdQ-yfdR-yfdS-yfdT. Front. Microbiol. 2016, 7, 239. [Google Scholar] [CrossRef] [PubMed]
- Chodavarapu, S.; Felczak, M.M.; Kaguni, J.M. Two forms of ribosomal protein L2 of Escherichia coli that inhibit DnaA in DNA replication. Nucleic Acids Res. 2011, 39, 4180–4191. [Google Scholar] [CrossRef] [PubMed]
- Calhoun, L.N.; Kwon, Y.M. Structure, function and regulation of the DNA-binding protein Dps and its role in acid and oxidative stress resistance in Escherichia coli: A review. J. Appl. Microbiol. 2011, 110, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Higgins, D.; Dworkin, J. Recent progress in Bacillus subtilis sporulation. FEMS Microbiol. Rev. 2012, 36, 131–148. [Google Scholar] [CrossRef] [PubMed]
- Tan, I.S.; Ramamurthi, K.S. Spore formation in Bacillus subtilis. Environ. Microbiol. Rep. 2014, 6, 212–225. [Google Scholar] [CrossRef] [PubMed]
- Boonstra, M.; de Jong, I.G.; Scholefield, G.; Murray, H.; Kuipers, O.P.; Veening, J.-W. Spo0A regulates chromosome copy number during sporulation by directly binding to the origin of replication in Bacillus subtilis. Mol. Microbiol. 2013, 87, 925–938. [Google Scholar] [CrossRef] [PubMed]
- Castilla-Llorente, V.; Muñoz-Espín, D.; Villar, L.; Salas, M.; Meijer, W.J.J. Spo0A, the key transcriptional regulator for entrance into sporulation, is an inhibitor of DNA replication. EMBO J. 2006, 25, 3890–3899. [Google Scholar] [CrossRef] [PubMed]
- Moriya, S.; Atlung, T.; Hansen, F.G.; Yoshikawa, H.; Ogasawara, N. Cloning of an autonomously replicating sequence (ars) from the Bacillus subtilis chromosome. Mol. Microbiol. 1992, 6, 309–315. [Google Scholar] [CrossRef] [PubMed]
- Moriya, S.; Imai, Y.; Hassan, A.K.; Ogasawara, N. Regulation of initiation of Bacillus subtilis chromosome replication. Plasmid 1999, 41, 17–29. [Google Scholar] [CrossRef] [PubMed]
- Krause, M.; Rückert, B.; Lurz, R.; Messer, W. Complexes at the replication origin of Bacillus subtilis with homologous and heterologous DnaA protein. J. Mol. Biol. 1997, 274, 365–380. [Google Scholar] [CrossRef] [PubMed]
- shigo-Oka, D.; Ogasawara, N.; Moriya, S. DnaD protein of Bacillus subtilis interacts with DnaA, the initiator protein of replication. J. Bacteriol. 2001, 183, 2148–2150. [Google Scholar] [CrossRef] [PubMed]
- Wagner, J.K.; Marquis, K.A.; Rudner, D.Z. SirA enforces diploidy by inhibiting the replication initiator DnaA during spore formation in Bacillus subtilis. Mol. Microbiol. 2009, 73, 963–974. [Google Scholar] [CrossRef] [PubMed]
- Rahn-Lee, L.; Gorbatyuk, B.; Skovgaard, O.; Losick, R. The conserved sporulation protein YneE inhibits DNA replication in Bacillus subtilis. J. Bacteriol. 2009, 191, 3736–3739. [Google Scholar] [CrossRef] [PubMed]
- Rahn-Lee, L.; Merrikh, H.; Grossman, A.D.; Losick, R. The sporulation protein SirA inhibits the binding of DnaA to the origin of replication by contacting a patch of clustered amino acids. J. Bacteriol. 2011, 193, 1302–1307. [Google Scholar] [CrossRef] [PubMed]
- Duan, Y.; Huey, J.D.; Herman, J.K. The DnaA inhibitor SirA acts in the same pathway as Soj (ParA) to facilitate oriC segregation during Bacillus subtilis sporulation. Mol. Microbiol. 2016, 102, 530–544. [Google Scholar] [CrossRef] [PubMed]
- Xenopoulos, P.; Piggot, P.J. Regulation of growth of the mother cell and chromosome replication during sporulation of Bacillus subtilis. J. Bacteriol. 2011, 193, 3117–3126. [Google Scholar] [CrossRef] [PubMed]
- Briggs, G.S.; Smits, W.K.; Soultanas, P. Chromosomal replication initiation machinery of low-g+c-content firmicutes. J. Bacteriol. 2012, 194, 5162–5170. [Google Scholar] [CrossRef] [PubMed]
- Velten, M.; McGovern, S.; Marsin, S.; Ehrlich, S.D.; Noirot, P.; Polard, P. A two-protein strategy for the functional loading of a cellular replicative DNA helicase. Mol. Cell 2003, 11, 1009–1020. [Google Scholar] [CrossRef]
- Davey, M.J.; O’Donnell, M. Replicative helicase loaders: Ring breakers and ring makers. Curr. Biol. 2003, 13, R594–R596. [Google Scholar] [CrossRef]
- Atherton, J.C.; Blaser, M.J. Coadaptation of Helicobacter pylori and humans: Ancient history, modern implications. J. Clin. Investig. 2009, 119, 2475–2487. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.; Tompkins, L.S.; Amieva, M.R. Helicobacter pylori usurps cell polarity to turn the cell surface into a replicative niche. PLoS Pathog. 2009, 5, e1000407. [Google Scholar] [CrossRef] [PubMed]
- Tomb, J.F.; White, O.; Kerlavage, A.R.; Clayton, R.A.; Sutton, G.G.; Fleischmann, R.D.; Ketchum, K.A.; Klenk, H.P.; Gill, S.; Dougherty, B.A.; et al. The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature 1997, 388, 539–547. [Google Scholar] [CrossRef] [PubMed]
- Danielli, A.; Scarlato, V. Regulatory circuits in Helicobacter pylori: Network motifs and regulators involved in metal-dependent responses. FEMS Microbiol. Rev. 2010, 34, 738–752. [Google Scholar] [CrossRef] [PubMed]
- Donczew, R.; Weigel, C.; Lurz, R.; Zakrzewska-Czerwinska, J.; Zawilak-Pawlik, A. Helicobacter pylori oriC—The first bipartite origin of chromosome replication in Gram-negative bacteria. Nucleic Acids Res. 2012, 40, 9647–9660. [Google Scholar] [CrossRef] [PubMed]
- Donczew, R.; Mielke, T.; Jaworski, P.; Zakrzewska-Czerwińska, J.; Zawilak-Pawlik, A. Assembly of Helicobacter pylori initiation complex is determined by sequence-specific and topology-sensitive DnaA-oriC interactions. J. Mol. Biol. 2014, 426, 2769–2782. [Google Scholar] [CrossRef] [PubMed]
- Zawilak, A.; Durrant, M.C.; Jakimowicz, P.; Backert, S.; Zakrzewska-Czerwińska, J. DNA binding specificity of the replication initiator protein, DnaA from Helicobacter pylori. J. Mol. Biol. 2003, 334, 933–947. [Google Scholar] [CrossRef] [PubMed]
- Zawilak-Pawlik, A.; Kois, A.; Majka, J.; Jakimowicz, D.; Smulczyk-Krawczyszyn, A.; Messer, W.; Zakrzewska-Czerwińska, J. Architecture of bacterial replication initiation complexes: orisomes from four unrelated bacteria. Biochem. J. 2005, 389, 471–481. [Google Scholar] [CrossRef] [PubMed]
- Rain, J.C.; Selig, L.; De Reuse, H.; Battaglia, V.; Reverdy, C.; Simon, S.; Lenzen, G.; Petel, F.; Wojcik, J.; Schächter, V.; Chemama, Y.; Labigne, A.; Legrain, P. The protein-protein interaction map of Helicobacter pylori. Nature 2001, 409, 211–215. [Google Scholar] [CrossRef] [PubMed]
- Zawilak-Pawlik, A.; Kois, A.; Stingl, K.; Boneca, I.G.; Skrobuk, P.; Piotr, J.; Lurz, R.; Zakrzewska-Czerwińska, J.; Labigne, A. HobA—A novel protein involved in initiation of chromosomal replication in Helicobacter pylori. Mol. Microbiol. 2007, 65, 979–994. [Google Scholar] [CrossRef] [PubMed]
- Natrajan, G.; Hall, D.R.; Thompson, A.C.; Gutsche, I.; Terradot, L. Structural similarity between the DnaA-binding proteins HobA (HP1230) from Helicobacter pylori and DiaA from Escherichia coli. Mol. Microbiol. 2007, 65, 995–1005. [Google Scholar] [CrossRef] [PubMed]
- Bazin, A.; Cherrier, M.V.; Gutsche, I.; Timmins, J.; Terradot, L. Structure and primase-mediated activation of a bacterial dodecameric replicative helicase. Nucleic Acids Res. 2015, 43, 8564–8576. [Google Scholar] [CrossRef] [PubMed]
- Stelter, M.; Gutsche, I.; Kapp, U.; Bazin, A.; Bajic, G.; Goret, G.; Jamin, M.; Timmins, J.; Terradot, L. Architecture of a dodecameric bacterial replicative helicase. Struct. Lond. Engl. 1993 2012, 20, 554–564. [Google Scholar] [CrossRef] [PubMed]
- Zakrzewska-Czerwińska, J.; Schrempf, H. Characterization of an autonomously replicating region from the Streptomyces lividans chromosome. J. Bacteriol. 1992, 174, 2688–2693. [Google Scholar] [CrossRef] [PubMed]
- Majka, J.; Messer, W.; Schrempf, H.; Zakrzewska-Czerwińska, J. Purification and characterization of the Streptomyces lividans initiator protein DnaA. J. Bacteriol. 1997, 179, 2426–2432. [Google Scholar] [CrossRef] [PubMed]
- Jakimowicz, D.; Majka, J.; Messer, W.; Speck, C.; Fernandez, M.; Martin, M.C.; Sanchez, J.; Schauwecker, F.; Keller, U.; Schrempf, H.; Zakrzewska-Czerwińska, J. Structural elements of the Streptomyces oriC region and their interactions with the DnaA protein. Microbiol. Read. Engl. 1998, 144 (Pt 5), 1281–1290. [Google Scholar] [CrossRef] [PubMed]
- Majka, J.; Jakimowicz, D.; Messer, W.; Schrempf, H.; Lisowski, M.; Zakrzewska-Czerwińska, J. Interactions of the Streptomyces lividans initiator protein DnaA with its target. Eur. J. Biochem. FEBS 1999, 260, 325–335. [Google Scholar] [CrossRef]
- Majka, J.; Zakrzewska-Czerwiñska, J.; Messer, W. Sequence recognition, cooperative interaction, and dimerization of the initiator protein DnaA of Streptomyces. J. Biol. Chem. 2001, 276, 6243–6252. [Google Scholar] [CrossRef] [PubMed]
- Jakimowicz, D.; Majkadagger, J.; Konopa, G.; Wegrzyn, G.; Messer, W.; Schrempf, H.; Zakrzewska-Czerwińska, J. Architecture of the Streptomyces lividans DnaA protein-replication origin complexes. J. Mol. Biol. 2000, 298, 351–364. [Google Scholar] [CrossRef] [PubMed]
- Molt, K.L.; Sutera, V.A.; Moore, K.K.; Lovett, S.T. A role for nonessential domain II of initiator protein, DnaA, in replication control. Genetics 2009, 183, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Boeneman, K.; Fossum, S.; Yang, Y.; Fingland, N.; Skarstad, K.; Crooke, E. Escherichia coli DnaA forms helical structures along the longitudinal cell axis distinct from MreB filaments. Mol. Microbiol. 2009, 72, 645–657. [Google Scholar] [CrossRef] [PubMed]
- Nozaki, S.; Niki, H.; Ogawa, T. Replication initiator DnaA of Escherichia coli changes its assembly form on the replication origin during the cell cycle. J. Bacteriol. 2009, 191, 4807–4814. [Google Scholar] [CrossRef] [PubMed]
- Nozaki, S.; Ogawa, T. Determination of the minimum domain II size of Escherichia coli DnaA protein essential for cell viability. Microbiol. Read. Engl. 2008, 154, 3379–3384. [Google Scholar] [CrossRef] [PubMed]
- Leonard, A.C.; Grimwade, J.E. The orisome: Structure and function. Front. Microbiol. 2015, 6, 545. [Google Scholar] [CrossRef] [PubMed]
- Terradot, L.; Zawilak-Pawlik, A. Structural insight into Helicobacter pylori DNA replication initiation. Gut Microbes 2010, 1, 330–334. [Google Scholar] [CrossRef] [PubMed]
- Zawilak, A.; Cebrat, S.; Mackiewicz, P.; Król-Hulewicz, A.; Jakimowicz, D.; Messer, W.; Gosciniak, G.; Zakrzewska-Czerwinska, J. Identification of a putative chromosomal replication origin from Helicobacter pylori and its interaction with the initiator protein DnaA. Nucleic Acids Res. 2001, 29, 2251–2259. [Google Scholar] [CrossRef] [PubMed]
- Jaworski, P.; Donczew, R.; Mielke, T.; Thiel, M.; Oldziej, S.; Weigel, C.; Pawlik, A.M. Unique and universal features of Epsilonproteobacterial origins of chromosome replication and DnaA-DnaA box interactions. Evol. Genomic Microbiol. 2016, 7, 1555. [Google Scholar] [CrossRef] [PubMed]
- Ulrich, L.E.; Zhulin, I.B. The MiST2 database: A comprehensive genomics resource on microbial signal transduction. Nucleic Acids Res. 2010, 38, D401–D407. [Google Scholar] [CrossRef] [PubMed]
- Collier, J. Regulation of chromosomal replication in Caulobacter crescentus. Plasmid 2012, 67, 76–87. [Google Scholar] [CrossRef] [PubMed]
- Robinson, A.; Causer, R.J.; Dixon, N.E. Architecture and conservation of the bacterial DNA replication machinery, an underexploited drug target. Curr. Drug Targets 2012, 13, 352–372. [Google Scholar] [CrossRef] [PubMed]
- van Eijk, E.; Wittekoek, B.; Kuijper, E.J.; Smits, W.K. DNA replication proteins as potential targets for antimicrobials in drug-resistant bacterial pathogens. J. Antimicrob. Chemother. 2017, 72, 1275–1284. [Google Scholar] [CrossRef] [PubMed]

© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zawilak-Pawlik, A.; Nowaczyk, M.; Zakrzewska-Czerwińska, J. The Role of the N-Terminal Domains of Bacterial Initiator DnaA in the Assembly and Regulation of the Bacterial Replication Initiation Complex. Genes 2017, 8, 136. https://doi.org/10.3390/genes8050136
Zawilak-Pawlik A, Nowaczyk M, Zakrzewska-Czerwińska J. The Role of the N-Terminal Domains of Bacterial Initiator DnaA in the Assembly and Regulation of the Bacterial Replication Initiation Complex. Genes. 2017; 8(5):136. https://doi.org/10.3390/genes8050136
Chicago/Turabian StyleZawilak-Pawlik, Anna, Małgorzata Nowaczyk, and Jolanta Zakrzewska-Czerwińska. 2017. "The Role of the N-Terminal Domains of Bacterial Initiator DnaA in the Assembly and Regulation of the Bacterial Replication Initiation Complex" Genes 8, no. 5: 136. https://doi.org/10.3390/genes8050136
APA StyleZawilak-Pawlik, A., Nowaczyk, M., & Zakrzewska-Czerwińska, J. (2017). The Role of the N-Terminal Domains of Bacterial Initiator DnaA in the Assembly and Regulation of the Bacterial Replication Initiation Complex. Genes, 8(5), 136. https://doi.org/10.3390/genes8050136




