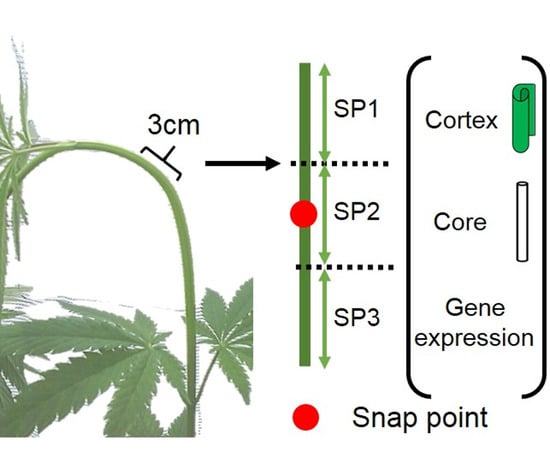
Molecular Investigation of the Stem Snap Point in Textile Hemp
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. RNA Extraction and RT-qPCR
3. Results and Discussion
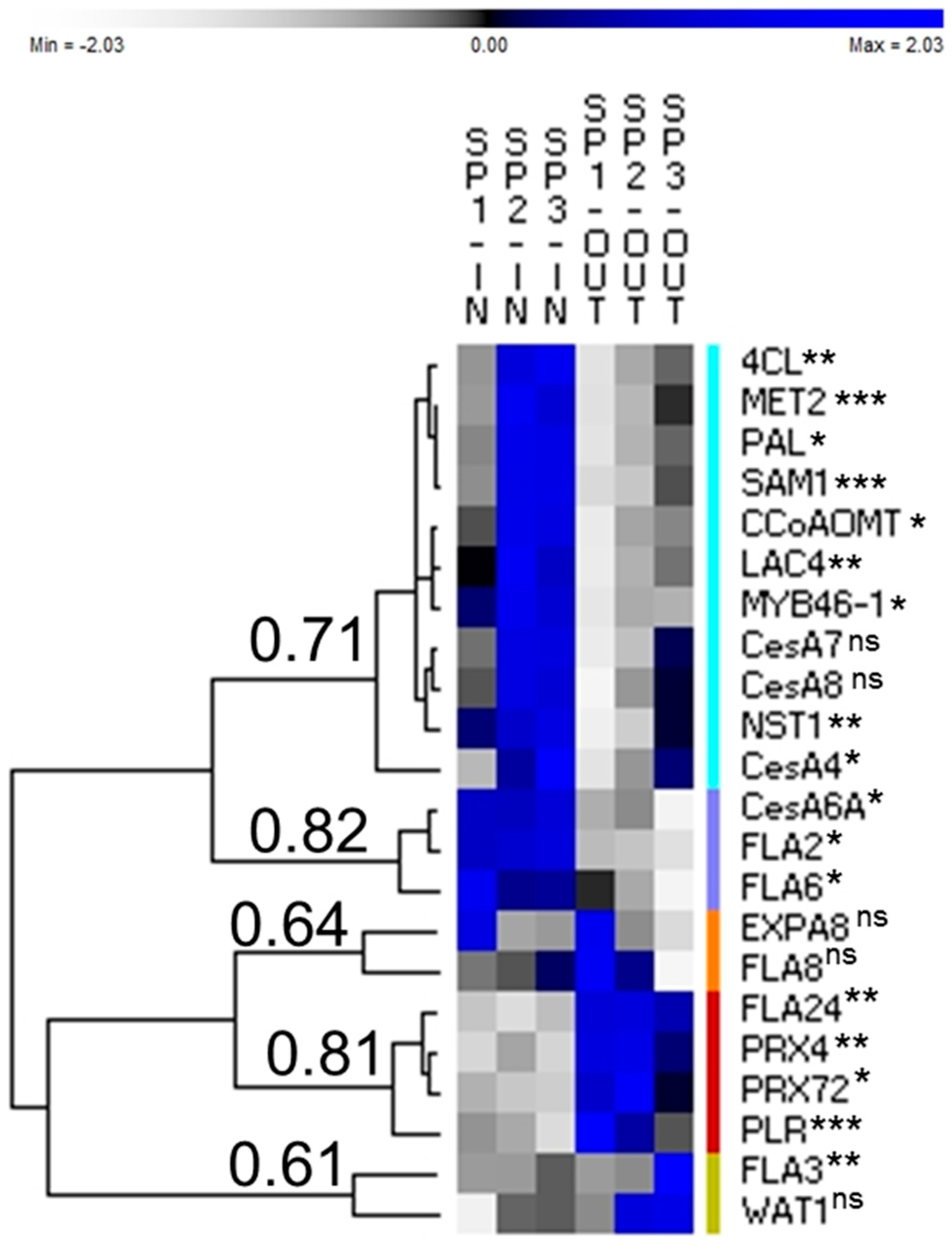
3.1. Genes Preferentially Expressed in the Inner Tissues
3.2. Genes Preferentially Expressed in the Outer Tissues
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Guerriero, G.; Sergeant, K.; Hausman, J.-F. Integrated-omics: A powerful approach to understanding the heterogeneous lignification of fibre crops. Int. J. Mol. Sci. 2013, 14, 10958–10978. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Sergeant, K.; Hausman, J.F. Wood biosynthesis and typologies: A molecular rhapsody. Tree Physiol. 2014, 34, 839–855. [Google Scholar] [CrossRef] [PubMed]
- Mellerowicz, E.J.; Gorshkova, T.A. Tensional stress generation in gelatinous fibres: A review and possible mechanism based on cell-wall structure and composition. J. Exp. Bot. 2012, 63, 551–565. [Google Scholar] [CrossRef] [PubMed]
- Snegireva, A.; Chernova, T.; Ageeva, M.; Lev-Yadun, S.; Gorshkova, T. Intrusive growth of primary and secondary phloem fibres in hemp stem determines fibre-bundle formation and structure. AoB Plants 2015, 7, plv061. [Google Scholar] [CrossRef] [PubMed]
- Gorshkova, T.A.; Sal’nikov, V.V.; Chemikosova, S.B.; Ageeva, M.V.; Pavlencheva, N.V.; van Dam, J.E.G. The snap point: A transition point in linum usitatissimum bast fiber development. Ind. Crops Prod. 2003, 18, 213–221. [Google Scholar] [CrossRef]
- Koziel, S.P. Genetic Analysis of Lignification and Secondary Wall Development in Bast Fibers of Industrial Hemp (Cannabis sativa). Master’s Thesis, University of Alberta, Edmonton, AB, Canada, 2010. [Google Scholar]
- Guerriero, G.; Behr, M.; Legay, S.; Mangeot-Peter, L.; Zorzan, S.; Ghoniem, M.; Hausman, J.-F. Transcriptomic profiling of hemp bast fibres at different developmental stages. Sci. Rep. 2017, 7, 4961. [Google Scholar] [CrossRef] [PubMed]
- Behr, M.; Legay, S.; Zizková, E.; Motyka, V.; Dobrev, P.I.; Hausman, J.F.; Lutts, S.; Guerriero, G. Studying secondary growth and bast fiber development: The hemp hypocotyl peeks behind the wall. Front. Plant Sci. 2016, 7, 1733. [Google Scholar] [CrossRef] [PubMed]
- Guerriero, G.; Mangeot-Peter, L.; Legay, S.; Behr, M.; Lutts, S.; Siddiqui, K.S.; Hausman, J.-F. Identification of fasciclin-like arabinogalactan proteins in textile hemp (Cannabis sativa L.): In silico analyses and gene expression patterns in different tissues. BMC Genom. 2017, 18, 741. [Google Scholar] [CrossRef] [PubMed]
- Mangeot-Peter, L.; Legay, S.; Hausman, J.F.; Esposito, S.; Guerriero, G. Identification of reference genes for RT-qPCR data normalization in Cannabis sativa stem tissues. Int. J. Mol. Sci. 2016, 17, 1556. [Google Scholar] [CrossRef] [PubMed]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. Qbase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef] [PubMed]
- Medicinal plant genomics resource. Available online: http://medicinalplantgenomics.msu.edu/index.shtml (accessed on 17 October 2017).
- Caraux, G.; Pinloche, S. Permutmatrix: A graphical environment to arrange gene expression profiles in optimal linear order. Bioinformatics 2005, 21, 1280–1281. [Google Scholar] [CrossRef] [PubMed]
- Persson, S.; Wei, H.; Milne, J.; Page, G.P.; Somerville, C.R. Identification of genes required for cellulose synthesis by regression analysis of public microarray data sets. Proc. Natl. Acad. Sci. USA 2005, 102, 8633–8638. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Ye, Z.H. Secondary cell walls: Biosynthesis, patterned deposition and transcriptional regulation. Plant Cell Physiol. 2015, 56, 195–214. [Google Scholar] [CrossRef] [PubMed]
- Zhong, R.; Lee, C.; Zhou, J.; McCarthy, R.L.; Ye, Z.H. A battery of transcription factors involved in the regulation of secondary cell wall biosynthesis in Arabidopsis. Plant Cell 2008, 20, 2763–2782. [Google Scholar] [CrossRef] [PubMed]
- Van den Broeck, H.C.; Maliepaard, C.; Ebskamp, M.J.M.; Toonen, M.A.J.; Koops, A.J. Differential expression of genes involved in C1 metabolism and lignin biosynthesis in wooden core and bast tissues of fibre hemp (Cannabis sativa L.). Plant Sci. 2008, 174, 205–220. [Google Scholar] [CrossRef]
- Gonneau, M.; Desprez, T.; Guillot, A.; Vernhettes, S.; Höfte, H. Catalytic subunit stoichiometry within the cellulose synthase complex. Plant Physiol. 2014, 166, 1709–1712. [Google Scholar] [CrossRef] [PubMed]
- Mokshina, N.; Gorshkov, O.; Ibragimova, N.; Chernova, T.; Gorshkova, T. Cellulosic fibres of flax recruit both primary and secondary cell wall cellulose synthases during deposition of thick tertiary cell walls and in the course of graviresponse. Funct. Plant Biol. 2017, 44, 820–831. [Google Scholar] [CrossRef]
- Mokshina, N.; Gorshkova, T.; Deyholos, M.K. Chitinase-like and cellulose synthase gene expression in gelatinous-type cellulosic walls of flax (Linum usitatissimum L.) bast fibers. PLoS ONE 2014, 9, e97949. [Google Scholar] [CrossRef] [PubMed]
- Schwager, K.M.; Calderon-Villalobos, L.I.A.; Dohmann, E.M.N.; Willige, B.C.; Knierer, S.; Nill, C.; Schwechheimer, C. Characterization of the VIER F-BOX PROTEINE genes from Arabidopsis reveals their importance for plant growth and development. Plant Cell 2007, 19, 1163–1178. [Google Scholar] [CrossRef] [PubMed]
- Oh, E.; Zhu, J.Y.; Bai, M.Y.; Arenhart, R.A.; Sun, Y.; Wang, Z.Y. Cell elongation is regulated through a central circuit of interacting transcription factors in the Arabidopsis hypocotyl. eLife 2014, 3, e03031. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Rodríguez, C.; Rubio-Somoza, I.; Sibout, R.; Persson, S. Phytohormones and the cell wall in Arabidopsis during seedling growth. Trends Plant Sci. 2010, 15, 291–301. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.B.; Cosgrove, D.J. Xyloglucan and its interactions with other components of the growing cell wall. Plant Cell Physiol. 2015, 56, 180–194. [Google Scholar] [CrossRef] [PubMed]
- De Pauw, M.A.; Vidmar, J.J.; Collins, J.; Bennett, R.A.; Deyholos, M.K. Microarray analysis of bast fibre producing tissues of Cannabis sativa identifies transcripts associated with conserved and specialised processes of secondary wall development. Funct. Plant Biol. 2007, 34, 737–749. [Google Scholar] [CrossRef]
- Huis, R.; Morreel, K.; Fliniaux, O.; Lucau-Danila, A.; Fénart, S.; Grec, S.; Neutelings, G.; Chabbert, B.; Mesnard, F.; Boerjan, W.; et al. Natural hypolignification is associated with extensive oligolignol accumulation in flax stems. Plant Physiol. 2012, 158, 1893–1915. [Google Scholar] [CrossRef] [PubMed]
- Yamauchi, S.; Ichikawa, H.; Nishiwaki, H.; Shuto, Y. Evaluation of plant growth regulatory activity of furofuran lignan bearing a 7,9′:7′,9-diepoxy structure using optically pure (+)- and (−)-enantiomers. J. Agric. Food Chem. 2015, 63, 5224–5228. [Google Scholar] [CrossRef] [PubMed]
- EMBOSS Needle suite. Available online: https://www.ebi.ac.uk/Tools/psa/emboss_needle/ (accessed on 17 October 2017).
- BLASTP. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 17 October 2017).
- Fernández-Pérez, F.; Vivar, T.; Pomar, F.; Pedreno, M.Á.; Novo-Uzal, E. Peroxidase 4 is involved in syringyl lignin formation in Arabidopsis thaliana. J. Plant Physiol. 2015, 175, 86–94. [Google Scholar] [CrossRef] [PubMed]
- Herrero, J.; Fernández-Pérez, F.; Yebra, T.; Novo-Uzal, E.; Pomar, F.; Pedreno, M.-Á.; Cuello, J.; Guéra, A.; Esteban-Carrasco, A.; Zapata, J.M. Bioinformatic and functional characterization of the basic peroxidase 72 from Arabidopsis thaliana involved in lignin biosynthesis. Planta 2013, 237, 1599–1612. [Google Scholar] [CrossRef] [PubMed]
- Andersson-Gunneras, S.; Mellerowicz, E.J.; Love, J.; Segerman, B.; Ohmiya, Y.; Coutinho, P.M.; Nilsson, P.; Henrissat, B.; Moritz, T.; Sundberg, B. Biosynthesis of cellulose-enriched tension wood in Populus: Global analysis of transcripts and metabolites identifies biochemical and developmental regulators in secondary wall biosynthesis. Plant J. 2006, 45, 144–165. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Jin, Y.; Wang, C.; Li, B.; Jiang, C.; Sun, Z.; Zhang, Z.; Kong, F.; Zhang, H. Fasciclin-like arabinogalactan proteins, PtFLAs, play important roles in GA-mediated tension wood formation in Populus. Sci. Rep. 2017, 7, 6182. [Google Scholar] [CrossRef] [PubMed]
- MacMillan, C.P.; Taylor, L.; Bi, Y.; Southerton, S.G.; Evans, R.; Spokevicius, A. The fasciclin-like arabinogalactan protein family of Eucalyptus grandis contains members that impact wood biology and biomechanics. New Phytol. 2015, 206, 1314–1327. [Google Scholar] [CrossRef] [PubMed]
- Ranocha, P.; Dima, O.; Nagy, R.; Felten, J.; Corratgé-Faillie, C.; Novák, O.; Morreel, K.; Lacombe, B.; Martinez, Y.; Pfrunder, S.; et al. Arabidopsis WAT1 is a vacuolar auxin transport facilitator required for auxin homoeostasis. Nat. Commun. 2013, 4, 2625. [Google Scholar] [CrossRef] [PubMed]
- Cassan-Wang, H.; Goué, N.; Saidi, M.N.; Legay, S.; Sivadon, P.; Goffner, D.; Grima-Pettenati, J. Identification of novel transcription factors regulating secondary cell wall formation in Arabidopsis. Front. Plant Sci. 2013, 4, 189. [Google Scholar] [CrossRef] [PubMed]
- Ranocha, P.; Denancé, N.; Vanholme, R.; Freydier, A.; Martinez, Y.; Hoffmann, L.; Köhler, L.; Pouzet, C.; Renou, J.-P.; Sundberg, B.; et al. Walls are thin 1 (WAT1), an Arabidopsis homolog of Medicago truncatula NODULIN21, is a tonoplast-localized protein required for secondary wall formation in fibers. Plant J. 2010, 63, 469–483. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Behr, M.; Legay, S.; Hausman, J.-F.; Lutts, S.; Guerriero, G. Molecular Investigation of the Stem Snap Point in Textile Hemp. Genes 2017, 8, 363. https://doi.org/10.3390/genes8120363
Behr M, Legay S, Hausman J-F, Lutts S, Guerriero G. Molecular Investigation of the Stem Snap Point in Textile Hemp. Genes. 2017; 8(12):363. https://doi.org/10.3390/genes8120363
Chicago/Turabian StyleBehr, Marc, Sylvain Legay, Jean-Francois Hausman, Stanley Lutts, and Gea Guerriero. 2017. "Molecular Investigation of the Stem Snap Point in Textile Hemp" Genes 8, no. 12: 363. https://doi.org/10.3390/genes8120363
APA StyleBehr, M., Legay, S., Hausman, J.-F., Lutts, S., & Guerriero, G. (2017). Molecular Investigation of the Stem Snap Point in Textile Hemp. Genes, 8(12), 363. https://doi.org/10.3390/genes8120363