Regulation of Pseudomonas aeruginosa Virulence by Distinct Iron Sources
Abstract
:1. Introduction
2. Iron and Host–Pathogen Interactions
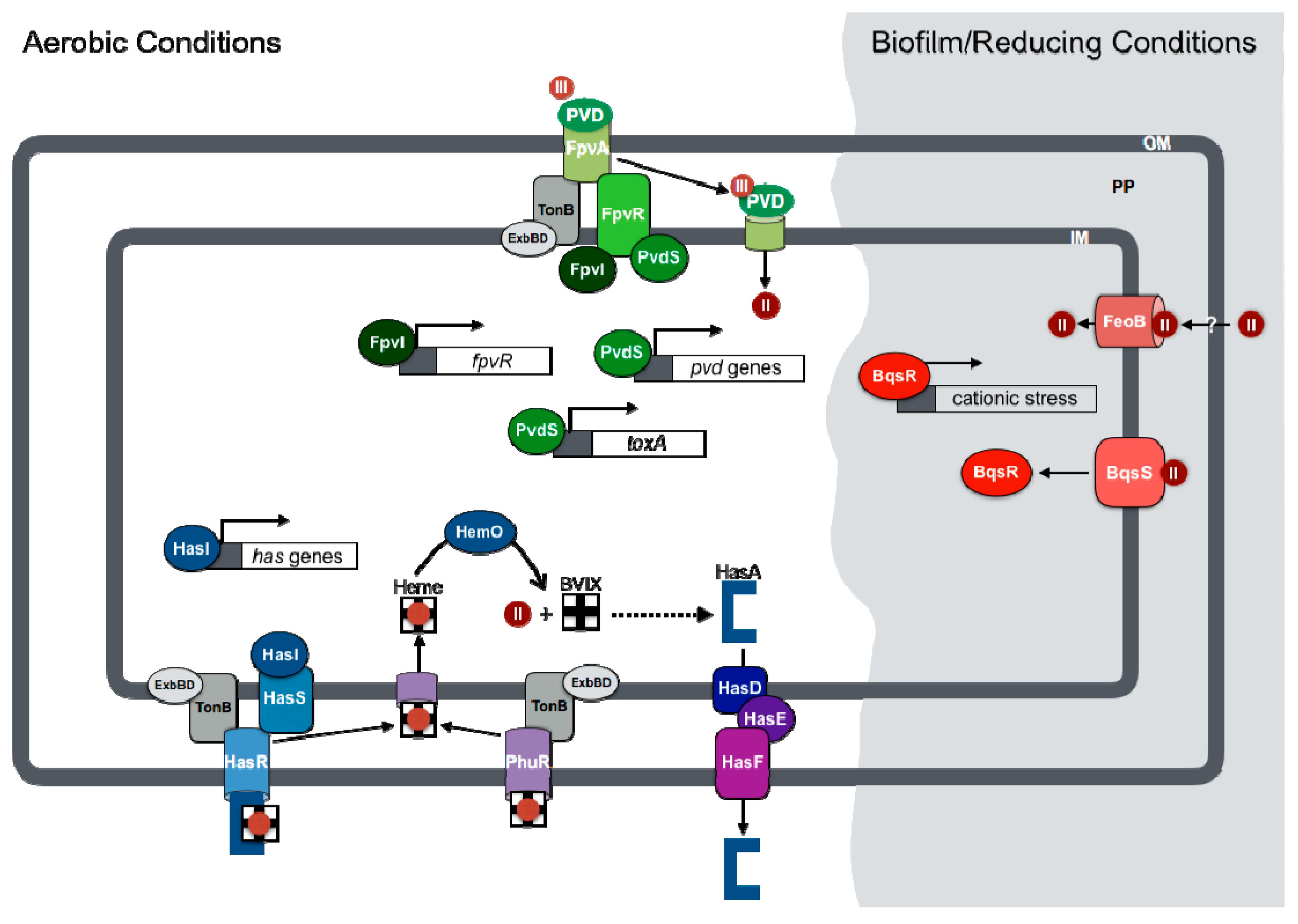
3. Coordinated Regulation of Virulence and Iron Uptake
3.1. Pyoverdine-Mediated Regulation of Virulence
3.2. Ferrous Iron Regulation of Biofilm Formation and Antibiotic Resistance
3.3. Heme-Dependent Regulation of Gene Expression
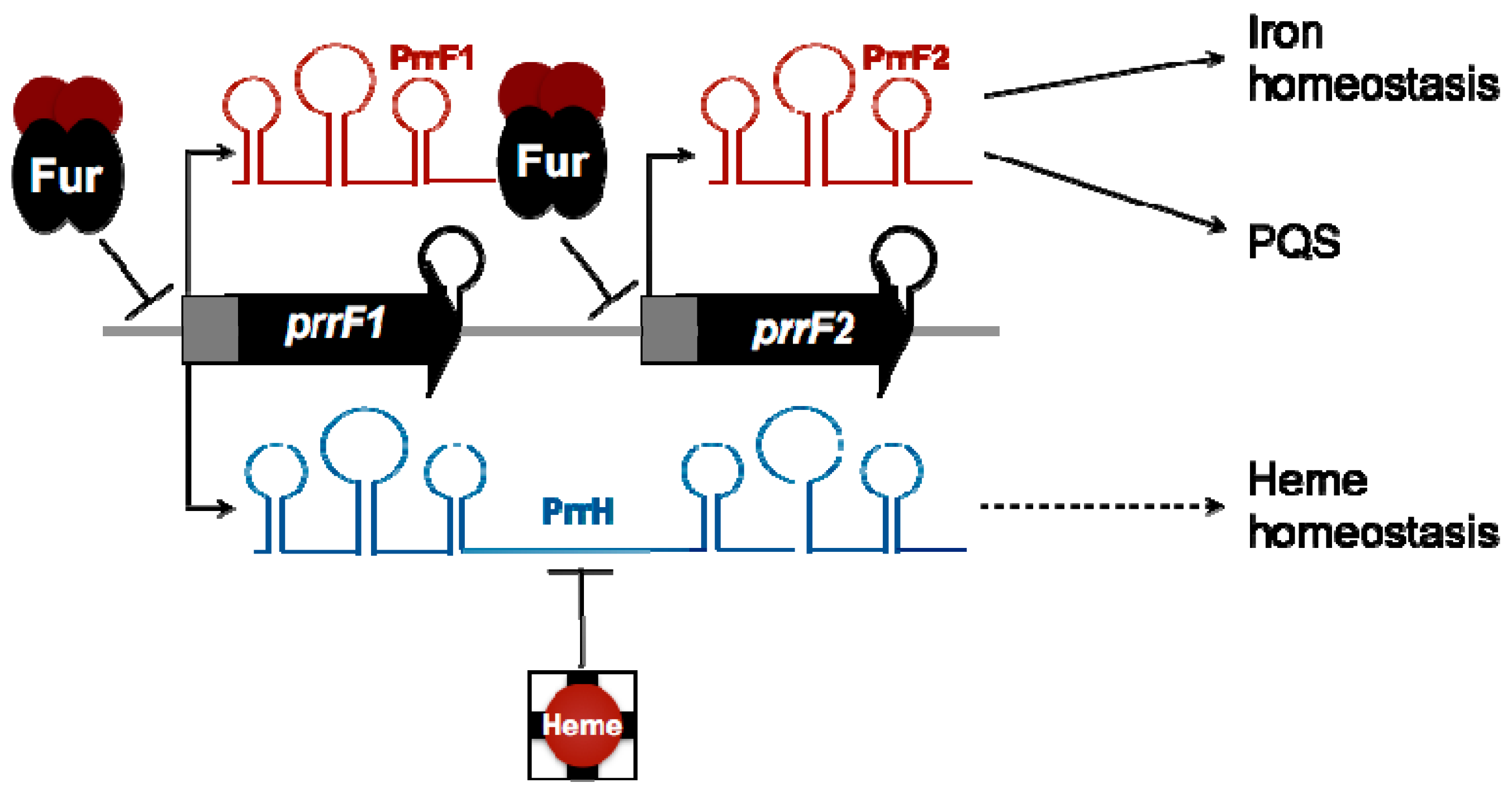
4. Post-Transcriptional Regulation by Iron-Responsive Small RNAs
5. Iron Regulation of Biofilm Formation
6. Future Outlooks
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Vento, S.; Cainelli, F.; Temesgen, Z. Lung infections after cancer chemotherapy. Lancet Oncol. 2008, 9, 982–992. [Google Scholar] [CrossRef]
- Klastersky, J.; Ameye, L.; Maertens, J.; Georgala, A.; Muanza, F.; Aoun, M.; Ferrant, A.; Rapoport, B.; Rolston, K.; Paesmans, M. Bacteraemia in febrile neutropenic cancer patients. Int. J. Antimicrob. Agents 2007, 30, S51–S59. [Google Scholar] [CrossRef] [PubMed]
- Chatzinikolaou, I.; Abi-Said, D.; Bodey, G.P.; Rolston, K.V.; Tarrand, J.J.; Samonis, G. Recent experience with Pseudomonas aeruginosa bacteremia in patients with cancer: Retrospective analysis of 245 episodes. Arch. Intern. Med. 2000, 160, 501–509. [Google Scholar] [CrossRef] [PubMed]
- NNIS. National Nosocomial Infections Surveillance (NNIS) System Report, data summary from January 1992 through June 2004, issued October 2004. Am. J. Infect. Control 2004, 32, 470–485. [Google Scholar]
- FitzSimmons, S.C. The changing epidemiology of cystic fibrosis. J. Pediatr. 1993, 122, 1–9. [Google Scholar] [CrossRef]
- Gjodsbol, K.; Christensen, J.J.; Karlsmark, T.; Jorgensen, B.; Klein, B.M.; Krogfelt, K.A. Multiple bacterial species reside in chronic wounds: A longitudinal study. Int. Wound J. 2006, 3, 225–231. [Google Scholar] [CrossRef] [PubMed]
- Jeffcoate, W.J.; Harding, K.G. Diabetic foot ulcers. Lancet 2003, 361, 1545–1551. [Google Scholar] [CrossRef]
- Vasil, M.L. Pseudomonas aeruginosa: Biology, mechanisms of virulence, epidemiology. J. Pediatr. 1986, 108, 800–805. [Google Scholar] [CrossRef]
- Cornelis, P.; Dingemans, J. Pseudomonas aeruginosa adapts its iron uptake strategies in function of the type of infections. Front. Cell. Infect. Microbiol. 2013, 3, 75. [Google Scholar] [CrossRef] [PubMed]
- Gellatly, S.L.; Hancock, R.E. Pseudomonas aeruginosa: New insights into pathogenesis and host defenses. Pathog. Dis. 2013, 67, 159–173. [Google Scholar] [CrossRef] [PubMed]
- Shaver, C.M.; Hauser, A.R. Relative contributions of Pseudomonas aeruginosa ExoU, ExoS, and ExoT to virulence in the lung. Infect. Immun. 2004, 72, 6969–6977. [Google Scholar] [CrossRef] [PubMed]
- Schulert, G.S.; Feltman, H.; Rabin, S.D.; Martin, C.G.; Battle, S.E.; Rello, J.; Hauser, A.R. Secretion of the toxin ExoU is a marker for highly virulent Pseudomonas aeruginosa isolates obtained from patients with hospital-acquired pneumonia. J. Infect. Dis. 2003, 188, 1695–1706. [Google Scholar] [CrossRef] [PubMed]
- Jain, M.; Ramirez, D.; Seshadri, R.; Cullina, J.F.; Powers, C.A.; Schulert, G.S.; Bar-Meir, M.; Sullivan, C.L.; McColley, S.A.; Hauser, A.R. Type III secretion phenotypes of Pseudomonas aeruginosa strains change during infection of individuals with cystic fibrosis. J. Clin. Microbiol. 2004, 42, 5229–5237. [Google Scholar] [CrossRef] [PubMed]
- Boucher, J.C.; Yu, H.; Mudd, M.H.; Deretic, V. Mucoid Pseudomonas aeruginosa in cystic fibrosis: Characterization of muc mutations in clinical isolates and analysis of clearance in a mouse model of respiratory infection. Infect. Immun. 1997, 65, 3838–3846. [Google Scholar] [PubMed]
- Otto, B.R.; Verweij-van Vught, A.M.; MacLaren, D.M. Transferrins and heme-compounds as iron sources for pathogenic bacteria. Crit. Rev. Microbiol. 1992, 18, 217–233. [Google Scholar] [CrossRef] [PubMed]
- Nairz, M.; Schroll, A.; Sonnweber, T.; Weiss, G. The struggle for iron—A metal at the host-pathogen interface. Cell. Microbiol. 2010, 12, 1691–1702. [Google Scholar] [CrossRef] [PubMed]
- Vasil, M. How we learnt about iron acquisition in Pseudomonas aeruginosa: A series of very fortunate events. BioMetals 2007, 20, 587–601. [Google Scholar] [CrossRef] [PubMed]
- Meyer, J.M.; Neely, A.; Stintzi, A.; Georges, C.; Holder, I.A. Pyoverdine is essential for virulence of Pseudomonas aeruginosa. Infect. Immun. 1996, 64, 518–523. [Google Scholar] [PubMed]
- Takase, H.; Nitanai, H.; Hoshino, K.; Otani, T. Impact of siderophore production on Pseudomonas aeruginosa infections in immunosuppressed mice. Infect. Immun. 2000, 68, 1834–1839. [Google Scholar] [CrossRef] [PubMed]
- Nadal Jimenez, P.; Koch, G.; Papaioannou, E.; Wahjudi, M.; Krzeslak, J.; Coenye, T.; Cool, R.H.; Quax, W.J. Role of PvdQ in Pseudomonas aeruginosa virulence under iron-limiting conditions. Microbiology 2010, 156, 49–59. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.Q.; Vasil, M.L.; Johnson, Z.; Ochsner, U.A.; Bayer, A.S. The oxygen- and iron-dependent sigma factor pvdS of Pseudomonas aeruginosa is an important virulence factor in experimental infective endocarditis. J. Infect. Dis. 2000, 181, 1020–1026. [Google Scholar] [CrossRef] [PubMed]
- Takase, H.; Nitanai, H.; Hoshino, K.; Otani, T. Requirement of the Pseudomonas aeruginosa tonB gene for high-affinity iron acquisition and infection. Infect. Immun. 2000, 68, 4498–4504. [Google Scholar] [CrossRef] [PubMed]
- Cox, C.D. Effect of pyochelin on the virulence of Pseudomonas aeruginosa. Infect. Immun. 1982, 36, 17–23. [Google Scholar] [PubMed]
- Vasil, M.L.; Ochsner, U.A. The response of Pseudomonas aeruginosa to iron: Genetics, biochemistry and virulence. Mol. Microbiol. 1999, 34, 399–413. [Google Scholar] [CrossRef] [PubMed]
- Porcheron, G.; Dozois, C.M. Interplay between iron homeostasis and virulence: Fur and RyhB as major regulators of bacterial pathogenicity. Vet. Microbiol. 2015, 179, 2–14. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.W.; Helmann, J.D. Functional specialization within the Fur family of metalloregulators. Biometals 2007, 20, 485–499. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, B.M.; Whitmire, J.M.; Merrell, D.S. This is not your mother’s repressor: The complex role of Fur in pathogenesis. Infect. Immun. 2009, 77, 2590–2601. [Google Scholar] [CrossRef] [PubMed]
- Poole, K.; McKay, G.A. Iron acquisition and its control in Pseudomonas aeruginosa: Many roads lead to Rome. Front. Biosci. 2003, 8, d661–d686. [Google Scholar] [CrossRef] [PubMed]
- Cornelis, P.; Matthijs, S.; Van Oeffelen, L. Iron uptake regulation in Pseudomonas aeruginosa. BioMetals 2009, 22, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Worwood, M. Estimation of Body Iron Stores. In Iron Physiology and Pathophysiology in Humans; Anderson, G.J., McLaren, G.D., Eds.; Nutrition and Health Series; Springer: New York, NY, USA, 2012; pp. 499–528. [Google Scholar]
- Ochsner, U.A.; Johnson, Z.; Vasil, M.L. Genetics and regulation of two distinct haem-uptake systems, phu and has, in Pseudomonas aeruginosa. Microbiology 2000, 146, 85–198. [Google Scholar] [CrossRef] [PubMed]
- Hood, M.I.; Skaar, E.P. Nutritional immunity: Transition metals at the pathogen-host interface. Nat. Rev. Microbiol. 2012, 10, 525–537. [Google Scholar] [CrossRef] [PubMed]
- Youard, Z.A.; Wenner, N.; Reimmann, C. Iron acquisition with the natural siderophore enantiomers pyochelin and enantio-pyochelin in Pseudomonas species. BioMetals 2011, 24, 513–522. [Google Scholar] [CrossRef] [PubMed]
- Hunter, R.C.; Asfour, F.; Dingemans, J.; Osuna, B.L.; Samad, T.; Malfroot, A.; Cornelis, P.; Newman, D.K. Ferrous iron is a significant component of bioavailable iron in cystic fibrosis airways. mBio 2013, 4, e00557-13. [Google Scholar] [CrossRef] [PubMed]
- Marshall, B.; Stintzi, A.; Gilmour, C.; Meyer, J.M.; Poole, K. Citrate-mediated iron uptake in Pseudomonas aeruginosa: Involvement of the citrate-inducible FecA receptor and the FeoB ferrous iron transporter. Microbiology 2009, 155, 305–315. [Google Scholar] [CrossRef] [PubMed]
- Hassett, D.J.; Sokol, P.A.; Howell, M.L.; Ma, J.F.; Schweizer, H.T.; Ochsner, U.; Vasil, M.L. Ferric uptake regulator (Fur) mutants of Pseudomonas aeruginosa demonstrate defective siderophore-mediated iron uptake, altered aerobic growth, and decreased superoxide dismutase and catalase activities. J. Bacteriol. 1996, 178, 3996–4003. [Google Scholar] [CrossRef] [PubMed]
- Ochsner, U.A.; Vasil, A.I.; Johnson, Z.; Vasil, M.L. Pseudomonas aeruginosa fur overlaps with a gene encoding a novel outer membrane lipoprotein, OmlA. J. Bacteriol. 1999, 181, 1099–1109. [Google Scholar] [PubMed]
- Wong, S.M.; Mekalanos, J.J. Genetic footprinting with mariner-based transposition in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2000, 97, 10191–10196. [Google Scholar] [CrossRef] [PubMed]
- Wilderman, P.J.; Vasil, A.I.; Johnson, Z.; Wilson, M.J.; Cunliffe, H.E.; Lamont, I.L.; Vasil, M.L. Characterization of an endoprotease (PrpL) encoded by a PvdS-regulated gene in Pseudomonas aeruginosa. Infect. Immun. 2001, 69, 5385–5394. [Google Scholar] [CrossRef] [PubMed]
- Ochsner, U.A.; Vasil, A.I.; Vasil, M.L. Role of the ferric uptake regulator of Pseudomonas aeruginosa in the regulation of siderophores and exotoxin A expression: Purification and activity on iron-regulated promoters. J. Bacteriol. 1995, 177, 7194–7201. [Google Scholar] [CrossRef] [PubMed]
- Ochsner, U.A.; Vasil, M.L. Gene repression by the ferric uptake regulator in Pseudomonas aeruginosa: Cycle selection of iron-regulated genes. Proc. Natl. Acad. Sci. USA 1996, 93, 4409–4414. [Google Scholar] [CrossRef] [PubMed]
- Bjorn, M.J.; Iglewski, B.H.; Ives, S.K.; Sadoff, J.C.; Vasil, M.L. Effect of iron on yields of exotoxin A in cultures of Pseudomonas aeruginosa PA-103. Infect. Immun. 1978, 19, 785–791. [Google Scholar] [PubMed]
- Pollack, M. The role of exotoxin A in pseudomonas disease and immunity. Rev. Infect. Dis. 1983, 5 (Suppl. 5), S979–S984. [Google Scholar] [CrossRef] [PubMed]
- Prince, R.W.; Cox, C.D.; Vasil, M.L. Coordinate regulation of siderophore and exotoxin A production: Molecular cloning and sequencing of the Pseudomonas aeruginosa fur gene. J. Bacteriol. 1993, 175, 2589–2598. [Google Scholar] [CrossRef] [PubMed]
- Ochsner, U.A.; Johnson, Z.; Lamont, I.L.; Cunliffe, H.E.; Vasil, M.L. Exotoxin A production in Pseudomonas aeruginosa requires the iron-regulated pvdS gene encoding an alternative sigma factor. Mol. Microbiol. 1996, 21, 1019–1028. [Google Scholar] [CrossRef] [PubMed]
- Poole, K.; Zhao, Q.; Neshat, S.; Heinrichs, D.E.; Dean, C.R. The Pseudomonas aeruginosa tonB gene encodes a novel TonB protein. Microbiology 1996, 142, 1449–1458. [Google Scholar] [CrossRef] [PubMed]
- Cunliffe, H.E.; Merriman, T.R.; Lamont, I.L. Cloning and characterization of pvdS, a gene required for pyoverdine synthesis in Pseudomonas aeruginosa: PvdS is probably an alternative sigma factor. J. Bacteriol. 1995, 177, 2744–2750. [Google Scholar] [CrossRef] [PubMed]
- Vasil, M.L.; Ochsner, U.A.; Johnson, Z.; Colmer, J.A.; Hamood, A.N. The fur-regulated gene encoding the alternative sigma factor PvdS is required for iron-dependent expression of the LysR-type regulator ptxR in Pseudomonas aeruginosa. J. Bacteriol. 1998, 180, 6784–6788. [Google Scholar] [PubMed]
- Gaines, J.M.; Carty, N.L.; Tiburzi, F.; Davinic, M.; Visca, P.; Colmer-Hamood, J.A.; Hamood, A.N. Regulation of the Pseudomonas aeruginosa toxA, regA and ptxR genes by the iron-starvation sigma factor PvdS under reduced levels of oxygen. Microbiology 2007, 153, 4219–4233. [Google Scholar] [CrossRef] [PubMed]
- Shigematsu, T.; Fukushima, J.; Oyama, M.; Tsuda, M.; Kawamoto, S.; Okuda, K. Iron-mediated regulation of alkaline proteinase production in Pseudomonas aeruginosa. Microbiol. Immunol. 2001, 45, 579–590. [Google Scholar] [CrossRef] [PubMed]
- Minandri, F.; Imperi, F.; Frangipani, E.; Bonchi, C.; Visaggio, D.; Facchini, M.; Pasquali, P.; Bragonzi, A.; Visca, P. Role of iron uptake systems in Pseudomonas aeruginosa virulence and airway infection. Infect. Immun. 2016, 84, 2324–2335. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wilks, J.C.; Danhorn, T.; Ramos, I.; Croal, L.; Newman, D.K. Phenazine-1-carboxylic acid promotes bacterial biofilm development via ferrous iron acquisition. J. Bacteriol. 2011, 193, 3606–3617. [Google Scholar] [CrossRef] [PubMed]
- Ochsner, U.A.; Wilderman, P.J.; Vasil, A.I.; Vasil, M.L. GeneChip expression analysis of the iron starvation response in Pseudomonas aeruginosa: Identification of novel pyoverdine biosynthesis genes. Mol. Microbiol. 2002, 45, 1277–1287. [Google Scholar] [CrossRef] [PubMed]
- Kreamer, N.N.; Wilks, J.C.; Marlow, J.J.; Coleman, M.L.; Newman, D.K. BqsR/BqsS constitute a two-component system that senses extracellular Fe(II) in Pseudomonas aeruginosa. J. Bacteriol. 2012, 194, 1195–1204. [Google Scholar] [CrossRef] [PubMed]
- Dong, Y.H.; Zhang, X.F.; An, S.W.; Xu, J.L.; Zhang, L.H. A novel two-component system BqsS-BqsR modulates quorum sensing-dependent biofilm decay in Pseudomonas aeruginosa. Commun. Integr. Biol. 2008, 1, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Kreamer, N.N.; Costa, F.; Newman, D.K. The ferrous iron-responsive BqsRS two-component system activates genes that promote cationic stress tolerance. mBio 2015, 6, e02549. [Google Scholar] [CrossRef] [PubMed]
- Giske, C.G.; Monnet, D.L.; Cars, O.; Carmeli, Y. Clinical and economic impact of common multidrug-resistant gram-negative bacilli. Antimicrob. Agents Chemother. 2008, 52, 813–821. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, A.T.; O’Neill, M.J.; Watts, A.M.; Robson, C.L.; Lamont, I.L.; Wilks, A.; Oglesby-Sherrouse, A.G. Adaptation of iron homeostasis pathways by a Pseudomonas aeruginosa pyoverdine mutant in the cystic fibrosis lung. J. Bacteriol. 2014, 196, 2265–2276. [Google Scholar] [CrossRef] [PubMed]
- Marvig, R.L.; Damkiaer, S.; Khademi, S.M.; Markussen, T.M.; Molin, S.; Jelsbak, L. Within-host evolution of Pseudomonas aeruginosa reveals adaptation toward iron acquisition from hemoglobin. mBio 2014, 5, e00966-14. [Google Scholar] [CrossRef] [PubMed]
- Konings, A.F.; Martin, L.W.; Sharples, K.J.; Roddam, L.F.; Latham, R.; Reid, D.W.; Lamont, I.L. Pseudomonas aeruginosa uses multiple pathways to acquire iron during chronic infection in cystic fibrosis lungs. Infect. Immun. 2013, 81, 2697–2704. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.D.; Wilks, A. Differential contributions of the outer membrane receptors PhuR and HasR to heme acquisition in Pseudomonas aeruginosa. J. Biol. Chem. 2015, 290, 7756–7766. [Google Scholar] [CrossRef] [PubMed]
- Ghigo, J.M.; Letoffe, S.; Wandersman, C. A new type of hemophore-dependent heme acquisition system of Serratia marcescens reconstituted in Escherichia coli. J. Bacteriol. 1997, 179, 3572–3579. [Google Scholar] [CrossRef] [PubMed]
- Biville, F.; Cwerman, H.; Letoffe, S.; Rossi, M.S.; Drouet, V.; Ghigo, J.M.; Wandersman, C. Haemophore-mediated signalling in Serratia marcescens: A new mode of regulation for an extra cytoplasmic function (ECF) sigma factor involved in haem acquisition. Mol. Microbiol. 2004, 53, 1267–1277. [Google Scholar] [CrossRef] [PubMed]
- Lansky, I.B.; Lukat-Rodgers, G.S.; Block, D.; Rodgers, K.R.; Ratliff, M.; Wilks, A. The cytoplasmic heme-binding protein (PhuS) from the heme uptake system of Pseudomonas aeruginosa is an intracellular heme-trafficking protein to the δ-regioselective heme oxygenase. J. Biolog. Chem. 2006, 281, 13652–13662. [Google Scholar] [CrossRef] [PubMed]
- Ratliff, M.; Zhu, W.; Deshmukh, R.; Wilks, A.; Stojiljkovic, I. Homologues of neisserial heme oxygenase in gram-negative bacteria: Degradation of heme by the product of the pigA gene of Pseudomonas aeruginosa. J. Bacteriol. 2001, 183, 6394–6403. [Google Scholar] [CrossRef] [PubMed]
- Friedman, J.; Lad, L.; Li, H.; Wilks, A.; Poulos, T.L. Structural basis for novel δ-regioselective heme oxygenation in the opportunistic pathogen Pseudomonas aeruginosa. Biochemistry 2004, 43, 5239–5245. [Google Scholar] [CrossRef] [PubMed]
- Mourino, S.; Giardina, B.J.; Reyes-Caballero, H.; Wilks, A. Metabolite Driven Regulation of Heme Uptake by the Biliverdin IXβ/δ Selective HemO of Pseudomonas aeruginosa. J. Biol. Chem. 2016, 291, 20503–20515. [Google Scholar] [CrossRef] [PubMed]
- Hom, K.; Heinzl, G.A.; Eakanunkul, S.; Lopes, P.E.; Xue, F.; Mackerell, A.D., Jr.; Wilks, A. Small Molecule Antivirulents Targeting the Iron-Regulated Heme Oxygenase (HemO) of P. aeruginosa. J. Med. Chem. 2013, 56, 2097–2109. [Google Scholar] [CrossRef] [PubMed]
- Wilderman, P.J.; Sowa, N.A.; FitzGerald, D.J.; FitzGerald, P.C.; Gottesman, S.; Ochsner, U.A.; Vasil, M.L. Identification of tandem duplicate regulatory small RNAs in Pseudomonas aeruginosa involved in iron homeostasis. Proc. Natl. Acad. Sci. USA 2004, 101, 9792–9797. [Google Scholar] [CrossRef] [PubMed]
- Masse, E.; Gottesman, S. A small RNA regulates the expression of genes involved in iron metabolism in Escherichia coli. Proc. Natl. Acad. Sci. USA 2002, 99, 4620–4625. [Google Scholar] [CrossRef] [PubMed]
- Afonyushkin, T.; Vecerek, B.; Moll, I.; Blasi, U.; Kaberdin, V.R. Both RNase E and RNase III control the stability of sodB mRNA upon translational inhibition by the small regulatory RNA RyhB. Nucl. Acids Res. 2005, 33, 1678–1689. [Google Scholar] [CrossRef] [PubMed]
- Masse, E.; Escorcia, F.E.; Gottesman, S. Coupled degradation of a small regulatory RNA and its mRNA targets in Escherichia coli. Genes Dev. 2003, 17, 2374–2383. [Google Scholar] [CrossRef] [PubMed]
- Jacques, J.F.; Jang, S.; Prevost, K.; Desnoyers, G.; Desmarais, M.; Imlay, J.; Masse, E. RyhB small RNA modulates the free intracellular iron pool and is essential for normal growth during iron limitation in Escherichia coli. Mol. Microbiol. 2006, 62, 1181–1190. [Google Scholar] [CrossRef] [PubMed]
- Oglesby-Sherrouse, A.G.; Murphy, E.R. Iron-responsive bacterial small RNAs: Variations on a theme. Metallomics 2013, 5, 276–286. [Google Scholar] [CrossRef] [PubMed]
- Oglesby, A.G.; Farrow, J.M., 3rd; Lee, J.H.; Tomaras, A.P.; Greenberg, E.P.; Pesci, E.C.; Vasil, M.L. The influence of iron on Pseudomonas aeruginosa physiology: A regulatory link between iron and quorum sensing. J. Biol. Chem. 2008, 283, 15558–15567. [Google Scholar] [CrossRef] [PubMed]
- Deziel, E.; Gopalan, S.; Tampakaki, A.P.; Lepine, F.; Padfield, K.E.; Saucier, M.; Xiao, G.; Rahme, L.G. The contribution of MvfR to Pseudomonas aeruginosa pathogenesis and quorum sensing circuitry regulation: Multiple quorum sensing-regulated genes are modulated without affecting lasRI, rhlRI or the production of N-acyl-l-homoserine lactones. Mol. Microbiol. 2005, 55, 998–1014. [Google Scholar] [CrossRef] [PubMed]
- Calfee, M.W.; Coleman, J.P.; Pesci, E.C. Interference with Pseudomonas quinolone signal synthesis inhibits virulence factor expression by Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 2001, 98, 11633–11637. [Google Scholar] [CrossRef] [PubMed]
- Farrow, J.M., 3rd; Pesci, E.C. Two distinct pathways supply anthranilate as a precursor of the Pseudomonas quinolone signal. J. Bacteriol. 2007, 189, 3425–3433. [Google Scholar] [CrossRef] [PubMed]
- Oglesby-Sherrouse, A.G.; Vasil, M.L. Characterization of a heme-regulated non-coding RNA encoded by the prrF locus of Pseudomonas aeruginosa. PLoS ONE 2010, 5, e9930. [Google Scholar] [CrossRef] [PubMed]
- Kawasaki, S.; Arai, H.; Kodama, T.; Igarashi, Y. Gene cluster for dissimilatory nitrite reductase (nir) from Pseudomonas aeruginosa: Sequencing and identification of a locus for heme d1 biosynthesis. J. Bacteriol. 1997, 179, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Alexandria, A.A.; Powell, D.A.; Nguyen, A.T.; O’Neill, M.; Djapgne, L.; Wilks, A.; Ernst, R.K.; Oglesby-Sherrouse, A.G. The prrF-encoded Small Regulatory RNAs Are Required for Iron Homeostasis and Virulence of Pseudomonas aeruginosa. Infect. Immun. 2015, 83, 863–875. [Google Scholar]
- Llamas, M.A.; van der Sar, A.; Chu, B.C.; Sparrius, M.; Vogel, H.J.; Bitter, W. A novel extracytoplasmic function (ECF) sigma factor regulates virulence in Pseudomonas aeruginosa. PLoS Pathog. 2009, 5, e1000572. [Google Scholar]
- O’Neill, M.J.; Wilks, A. The P. aeruginosa Heme Binding Protein PhuS Is a Heme Oxygenase Titratable Regulator of Heme Uptake. ACS Chem. Biol. 2013, 8, 1794–1802. [Google Scholar] [CrossRef] [PubMed]
- Patriquin, G.M.; Banin, E.; Gilmour, C.; Tuchman, R.; Greenberg, E.P.; Poole, K. Influence of quorum sensing and iron on twitching motility and biofilm formation in Pseudomonas aeruginosa. J. Bacteriol. 2008, 190, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Banin, E.; Vasil, M.L.; Greenberg, E.P. Iron and Pseudomonas aeruginosa biofilm formation. Proc. Natl. Acad. Sci. USA 2005, 102, 11076–11081. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, Y.; Thoendel, M.; Olakanmi, O.; Britigan, B.E.; Singh, P.K. The transition metal gallium disrupts Pseudomonas aeruginosa iron metabolism and has antimicrobial and antibiofilm activity. J. Clin. Investig. 2007, 117, 877–888. [Google Scholar] [CrossRef] [PubMed]
- Glick, R.; Gilmour, C.; Tremblay, J.; Satanower, S.; Avidan, O.; Deziel, E.; Greenberg, E.P.; Poole, K.; Banin, E. Increase in rhamnolipid synthesis under iron-limiting conditions influences surface motility and biofilm formation in Pseudomonas aeruginosa. J. Bacteriol. 2010, 192, 2973–2980. [Google Scholar] [CrossRef] [PubMed]
- Yu, S.; Wei, Q.; Zhao, T.; Guo, Y.; Ma, L.Z. A novel survival strategy of Pseudomonas aeruginosa: Using exopolysaccharides to sequester and store iron to stimulate Psl-dependent biofilm formation. Appl. Environ. Microbiol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Wiens, J.R.; Vasil, A.I.; Schurr, M.J.; Vasil, M.L. Iron-regulated expression of alginate production, mucoid phenotype, and biofilm formation by Pseudomonas aeruginosa. mBio 2014, 5, e01010-13. [Google Scholar] [CrossRef] [PubMed]
- De Vos, D.; De Chial, M.; Cochez, C.; Jansen, S.; Tummler, B.; Meyer, J.M.; Cornelis, P. Study of pyoverdine type and production by Pseudomonas aeruginosa isolated from cystic fibrosis patients: Prevalence of type II pyoverdine isolates and accumulation of pyoverdine-negative mutations. Arch. Microbiol. 2001, 175, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Moreau-Marquis, S.; Bomberger, J.M.; Anderson, G.G.; Swiatecka-Urban, A.; Ye, S.; O’Toole, G.A.; Stanton, B.A. The DeltaF508-CFTR mutation results in increased biofilm formation by Pseudomonas aeruginosa by increasing iron availability. Am. J. Physiol. Lung Cell Mol. Physiol. 2008, 295, L25–L37. [Google Scholar] [CrossRef] [PubMed]
- Moreau-Marquis, S.; O’Toole, G.A.; Stanton, B.A. Tobramycin and FDA-approved iron chelators eliminate Pseudomonas aeruginosa biofilms on cystic fibrosis cells. Am. J. Respir. Cell Mol. Biol. 2009, 41, 305–313. [Google Scholar] [CrossRef] [PubMed]
- Oglesby-Sherrouse, A.G.; Djapgne, L.; Nguyen, A.T.; Vasil, A.I.; Vasil, M.L. The complex interplay of iron, biofilm formation, and mucoidy affecting antimicrobial resistance of Pseudomonas aeruginosa. Pathog. Dis. 2014, 70, 307–320. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Yang, L.; Molin, S. Synergistic activities of an efflux pump inhibitor and iron chelators against Pseudomonas aeruginosa growth and biofilm formation. Antimicrob. Agents Chemother. 2010, 54, 3960–3963. [Google Scholar] [CrossRef] [PubMed]
- Banin, E.; Lozinski, A.; Brady, K.M.; Berenshtein, E.; Butterfield, P.W.; Moshe, M.; Chevion, M.; Greenberg, E.P.; Banin, E. The potential of desferrioxamine-gallium as an anti-Pseudomonas therapeutic agent. Proc. Natl. Acad. Sci. USA 2008, 105, 16761–16766. [Google Scholar] [CrossRef] [PubMed]
- Fillion, P.; Desjardins, A.; Sayasith, K.; Lagace, J. Encapsulation of DNA in negatively charged liposomes and inhibition of bacterial gene expression with fluid liposome-encapsulated antisense oligonucleotides. Biochim. Biophys. Acta 2001, 1515, 44–54. [Google Scholar] [CrossRef]
- Harth, G.; Zamecnik, P.C.; Tang, J.Y.; Tabatadze, D.; Horwitz, M.A. Treatment of Mycobacterium tuberculosis with antisense oligonucleotides to glutamine synthetase mRNA inhibits glutamine synthetase activity, formation of the poly-l-glutamate/glutamine cell wall structure, and bacterial replication. Proc. Natl. Acad. Sci. USA 2000, 97, 418–423. [Google Scholar] [CrossRef] [PubMed]
- Meng, J.; Wang, H.; Hou, Z.; Chen, T.; Fu, J.; Ma, X.; He, G.; Xue, X.; Jia, M.; Luo, X. Novel anion liposome-encapsulated antisense oligonucleotide restores susceptibility of methicillin-resistant Staphylococcus aureus and rescues mice from lethal sepsis by targeting mecA. Antimicrob. Agents Chemother. 2009, 53, 2871–2878. [Google Scholar] [CrossRef] [PubMed]
- Sterzenbach, T.; Nguyen, K.T.; Nuccio, S.P.; Winter, M.G.; Vakulskas, C.A.; Clegg, S.; Romeo, T.; Baumler, A.J. A novel CsrA titration mechanism regulates fimbrial gene expression in Salmonella typhimurium. EMBO J. 2013, 32, 2872–2883. [Google Scholar] [CrossRef] [PubMed]
- Gruber, C.C.; Sperandio, V. Posttranscriptional control of microbe-induced rearrangement of host cell actin. mBio 2014, 5, e01025-13. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.H.; Wang, C.K.; Peng, H.L.; Wu, C.C.; Chen, Y.T.; Hong, Y.M.; Lin, C.T. Role of the small RNA RyhB in the Fur regulon in mediating the capsular polysaccharide biosynthesis and iron acquisition systems in Klebsiella pneumoniae. BMC Microbiol. 2012, 12, 148. [Google Scholar] [CrossRef] [PubMed]
- Porcheron, G.; Habib, R.; Houle, S.; Caza, M.; Lepine, F.; Daigle, F.; Masse, E.; Dozois, C.M. The small RNA RyhB contributes to siderophore production and virulence of uropathogenic Escherichia coli. Infect. Immun. 2014, 82, 5056–5068. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.R.; Payne, S.M. RyhB, an iron-responsive small RNA molecule, regulates Shigella dysenteriae virulence. Infect. Immun. 2007, 75, 3470–3477. [Google Scholar] [CrossRef] [PubMed]
- Oglesby, A.G.; Murphy, E.R.; Iyer, V.R.; Payne, S.M. Fur regulates acid resistance in Shigella flexneri via RyhB and ydeP. Mol. Microbiol. 2005, 58, 1354–1367. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Reinhart, A.A.; Oglesby-Sherrouse, A.G. Regulation of Pseudomonas aeruginosa Virulence by Distinct Iron Sources. Genes 2016, 7, 126. https://doi.org/10.3390/genes7120126
Reinhart AA, Oglesby-Sherrouse AG. Regulation of Pseudomonas aeruginosa Virulence by Distinct Iron Sources. Genes. 2016; 7(12):126. https://doi.org/10.3390/genes7120126
Chicago/Turabian StyleReinhart, Alexandria A., and Amanda G. Oglesby-Sherrouse. 2016. "Regulation of Pseudomonas aeruginosa Virulence by Distinct Iron Sources" Genes 7, no. 12: 126. https://doi.org/10.3390/genes7120126
APA StyleReinhart, A. A., & Oglesby-Sherrouse, A. G. (2016). Regulation of Pseudomonas aeruginosa Virulence by Distinct Iron Sources. Genes, 7(12), 126. https://doi.org/10.3390/genes7120126





