Transcriptome Analysis of Paeonia ostii ‘Fengdan’ Seeds Uncovers Starch and Sucrose Metabolism Conferring High Yield Under Brassinosteroid Treatment
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Trait Measurements
2.3. RNA Extraction, cDNA Library Construction, High-Throughput Sequencing, Raw Data Processing, mappingRNA Extraction, Library Preparation, Sequencing, and Bioinformatic Analysis
2.4. Quantification of Gene Expression, Gene Ontology, and KEGG Pathway Enrichment Analysis
2.5. Weighted Gene Co-Expression Network Analysis
2.6. RT-qPCR Analysis of Genes Involved in Starch and Sucrose Metabolic Pathways
3. Results
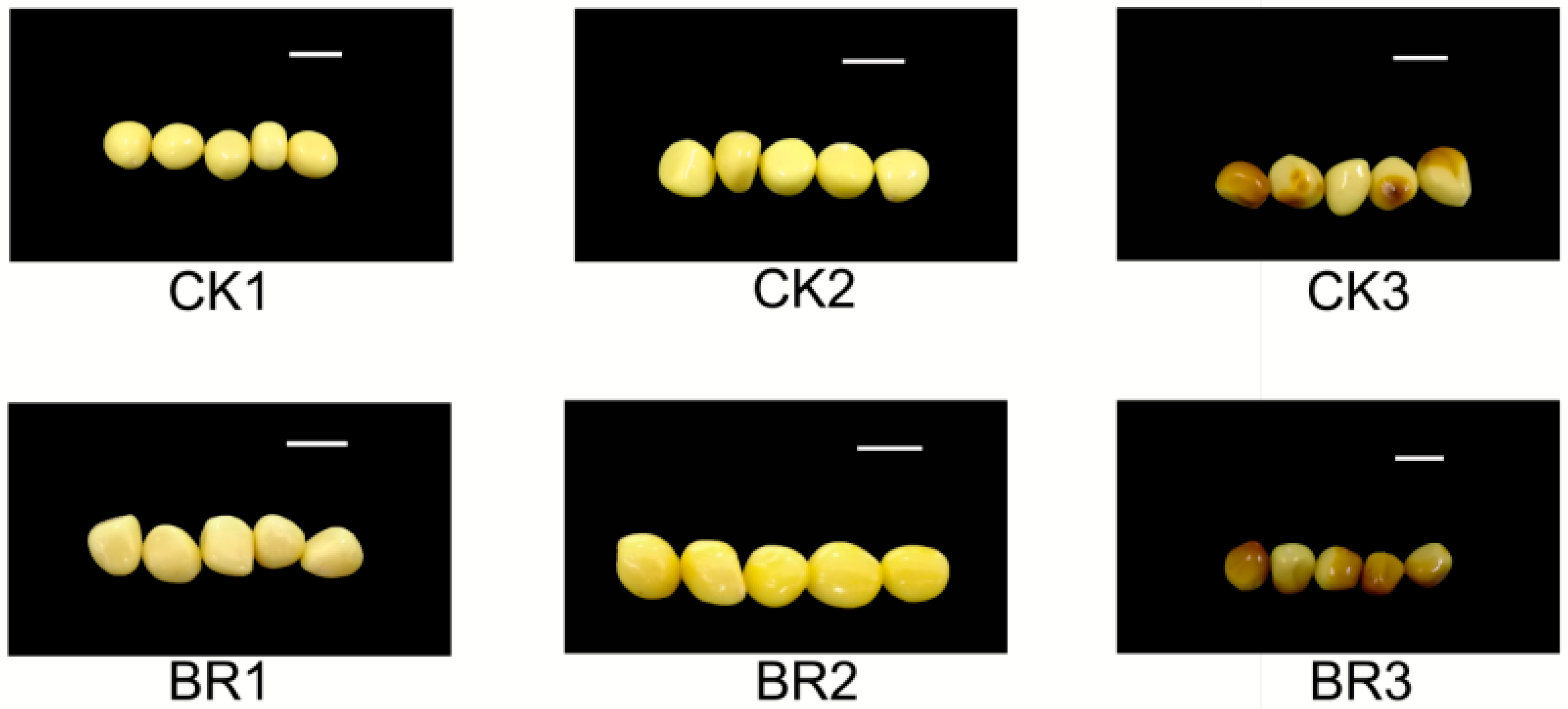
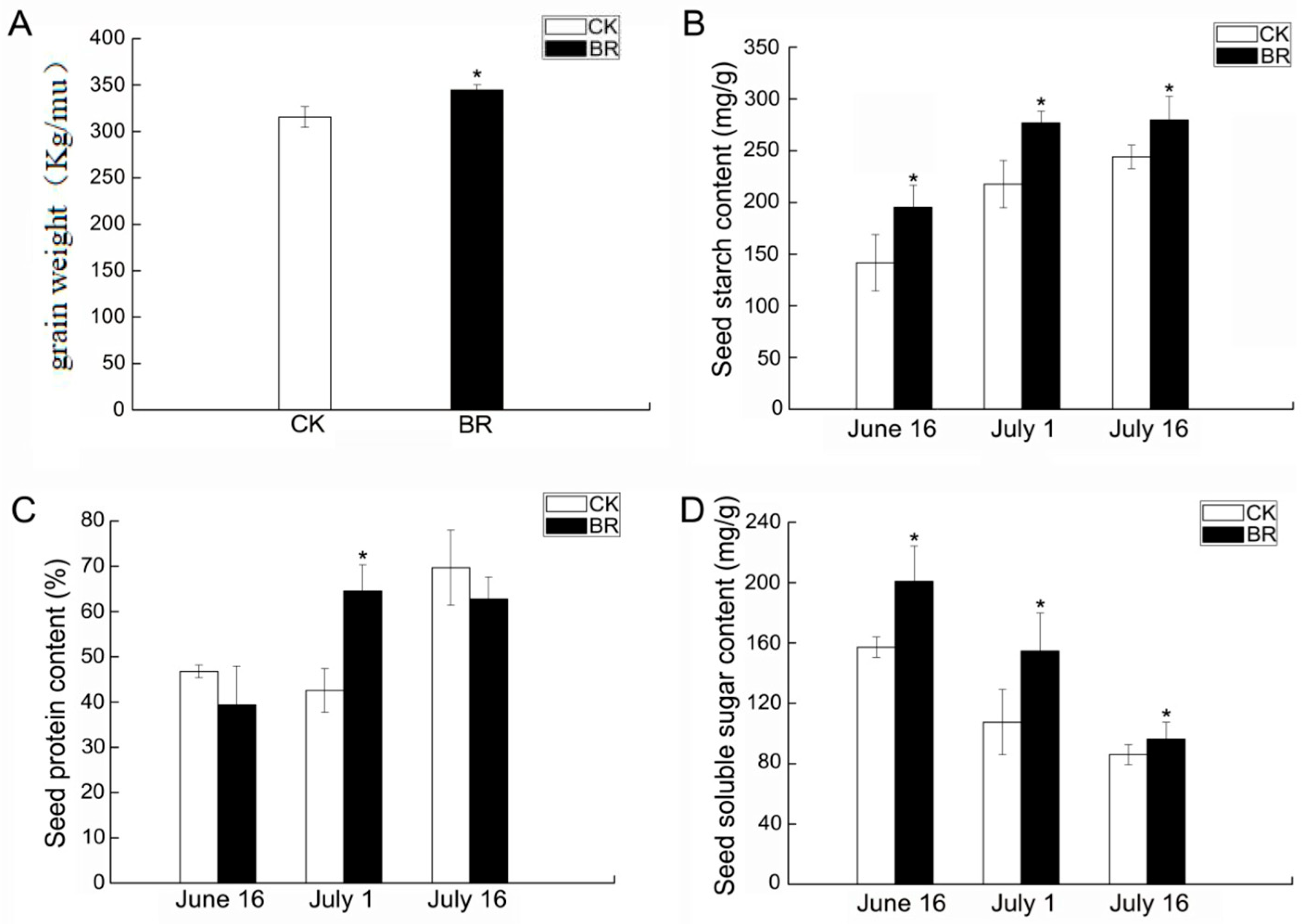
3.1. Impact of Exogenous Brassinosteroids on Seed Yield and Content of P. ostii ‘Fengdan’
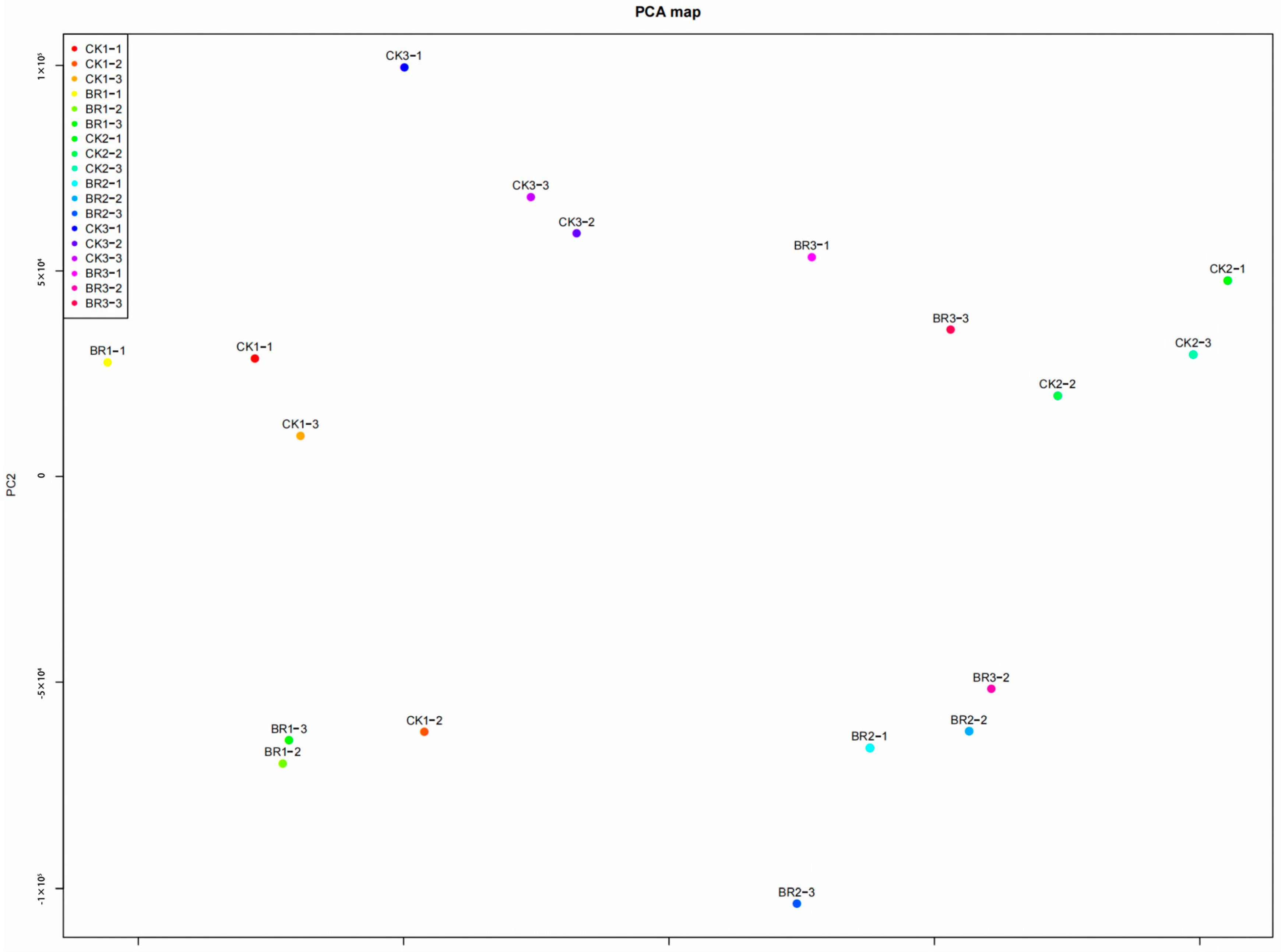
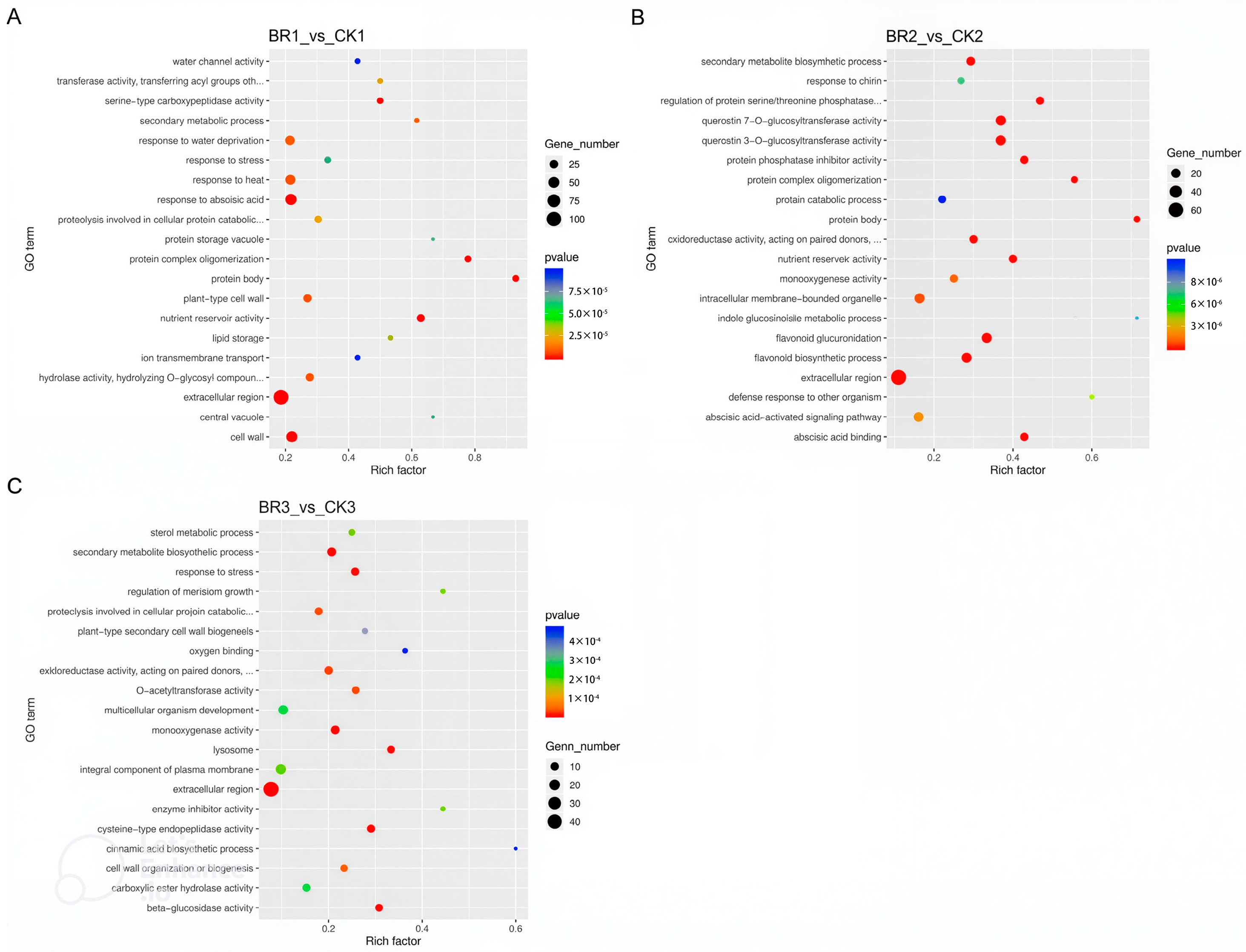
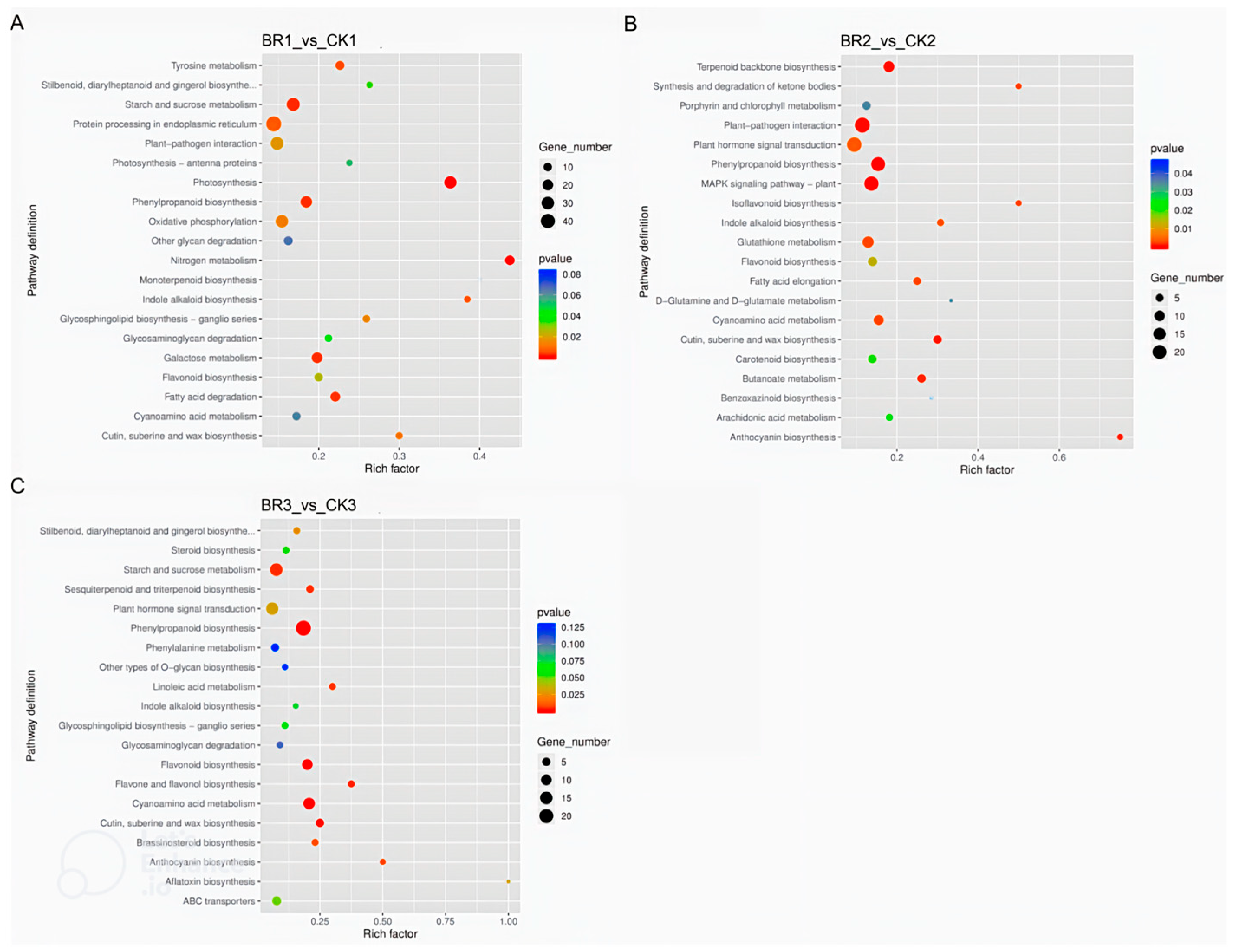
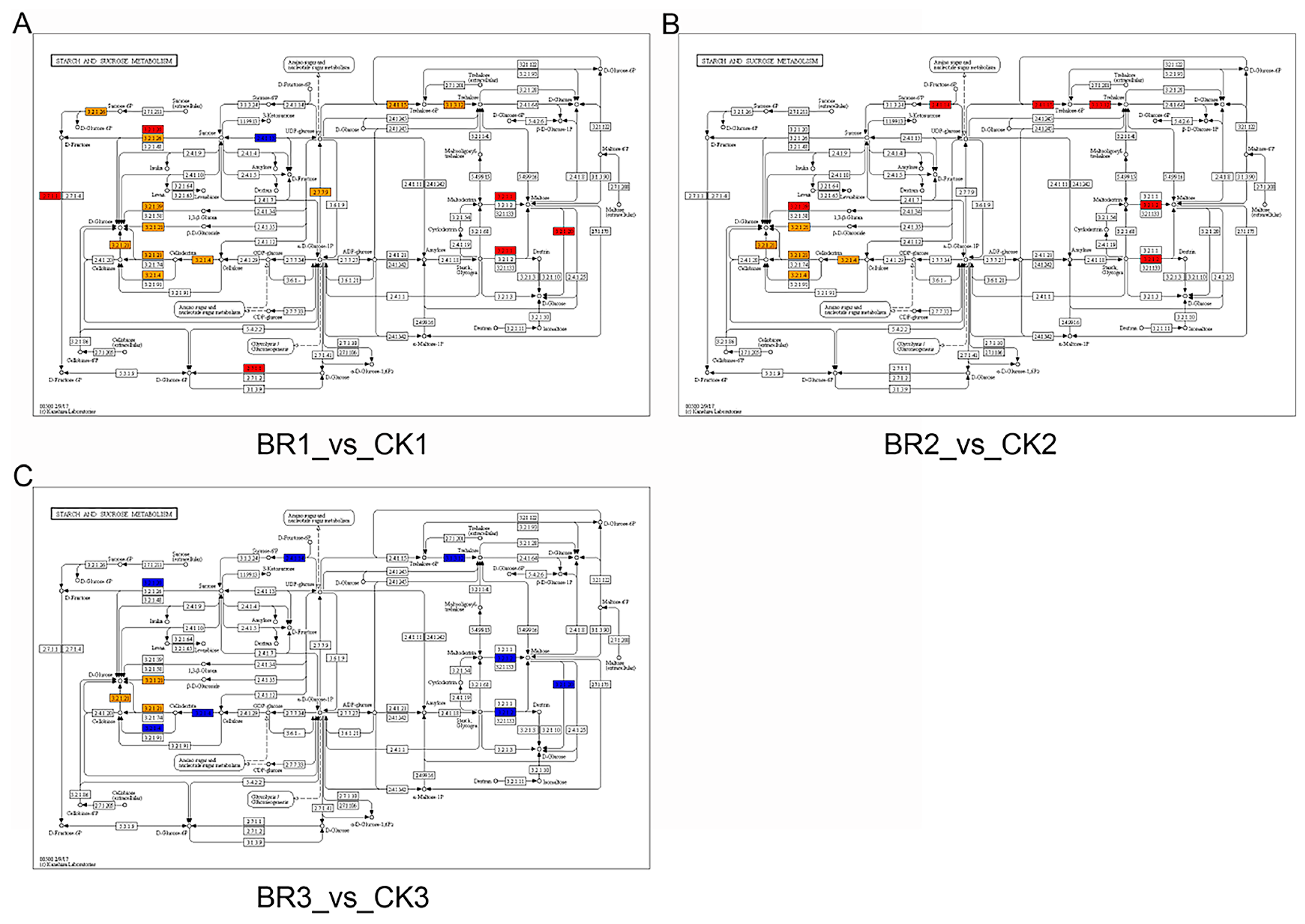
3.2. RNA-Seq Analysis of BR-Treated P. ostii ‘Fengdan’
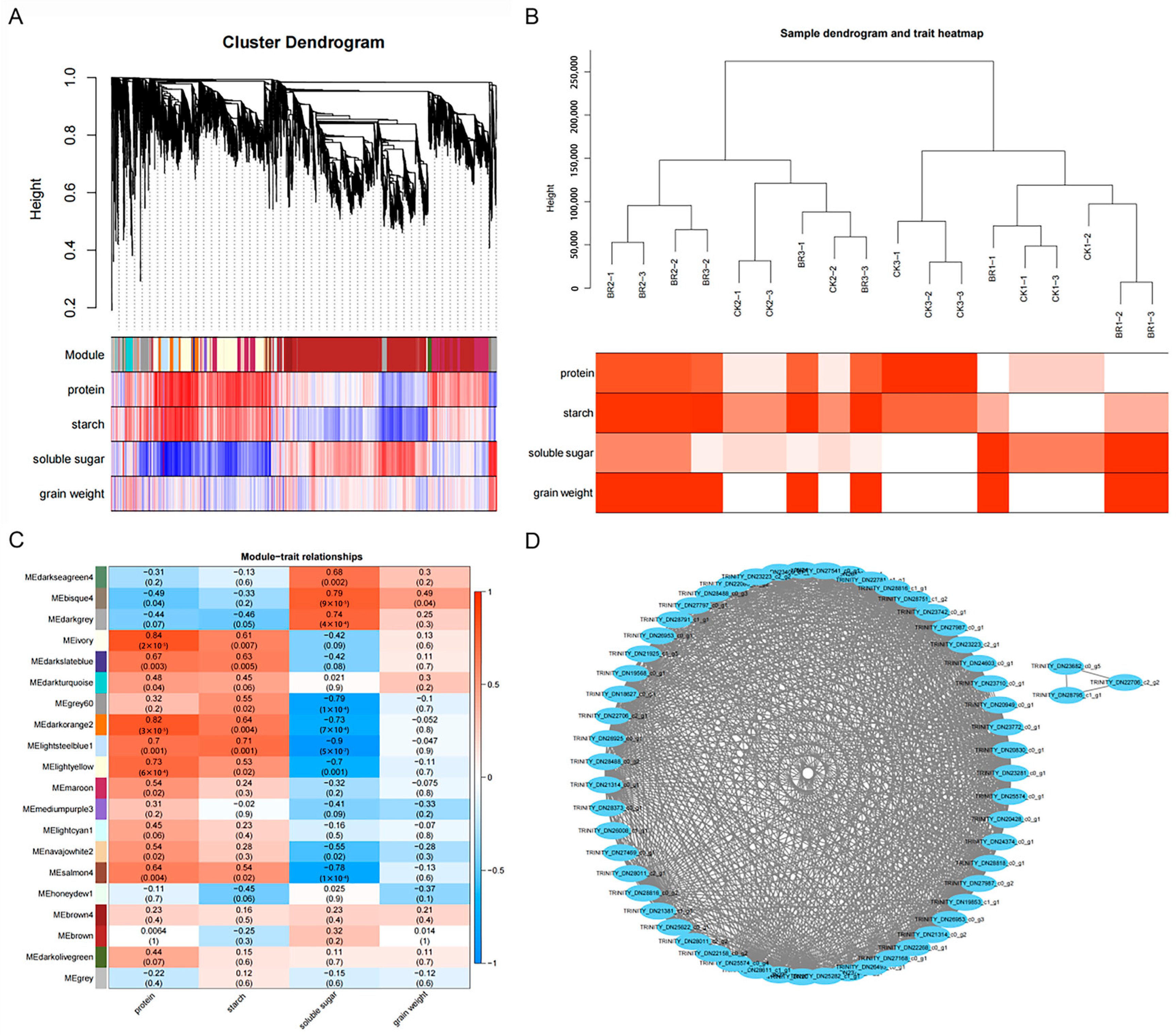
3.3. Gene Expression Network Related to BR Response Identified by WGCNA
3.4. Expression Levels of Key Pathway Genes in P. ostii ‘Fengdan’ Under BR Treatment
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
| Transcript_Id | Forward Primer (5′ → 3′) | Reverse Primer (5′ → 3′) |
|---|---|---|
| TRINITY_DN23567_c1_g4 | GGCTTGCAAGAATACACCGCA | AAGGGGCTCCACGTTCACAA |
| TRINITY_DN19592_c1_g1 | CGAAGTTCGCGGGTCAATGG | ACACCTTGACCTTCCCACCC |
| TRINITY_DN23401_c0_g1 | GGTGGCACCCGGTGGTTATA | TCAGGGCTGATGGGTGACAA |
| TRINITY_DN23738_c0_g5 | AGATTCTGCCCGTGGGTGTT | GCTCCTGCCGTTGCTGTAAG |
References
- Li, S.S.; Yuan, R.Y.; Chen, L.G.; Wang, L.S.; Hao, X.H.; Wang, L.J.; Zheng, X.C.; Du, H. Systematic qualitative and quantitative assessment of fatty acids in the seeds of 60 tree peony (Paeonia section Moutan DC.) cultivars by GC-MS. Food Chem. 2015, 173, 133–140. [Google Scholar] [CrossRef]
- Zhang, X.X.; Zhang, Y.L.; Niu, L.X.; Sun, J.Y.; Li, L.H.; Zhang, J.; Li, J. Chemometric classification of different tree peony species native to China based on the assessment of major fatty acids of seed oil and phenotypic characteristics of the seeds. Chem. Biodivers. 2017, 14, e1600111. [Google Scholar] [CrossRef]
- Murumalla, R.K.; Gunasekaran, M.K.; Padhan, J.K.; Bencharif, K.; Gence, L.; Festy, F.; Césari, M.; Roche, R.; Hoareau, L. Fatty acids do not pay the toll: Effect of SFA and PUFA on human adipose tissue and mature adipocytes inflammation. Lipids Health Dis. 2012, 11, 175. [Google Scholar] [CrossRef]
- Perng, W.; Villamor, E.; Mora-Plazas, M.; Marin, C.; Baylin, A. Alpha-linolenic acid (ALA) is inversely related to development of adiposity in school-age children. Eur. J. Clin. Nutr. 2015, 69, 167–172. [Google Scholar] [CrossRef]
- Sun, X.; Zhang, Q.; Zhang, H.; Niu, L.; Zhang, M.; Zhang, Y. A set of artificial pollination technical measures: Improved seed yields and active ingredients of seeds in oil tree peonies. Plants 2024, 13, 1194. [Google Scholar] [CrossRef]
- Li, W.; Yang, B. Translational genomics of grain size regulation in wheat. Theor. Appl. Genet. 2017, 130, 1765–1771. [Google Scholar] [CrossRef]
- Hu, Y.; Liu, J.; Lin, Y.; Xu, X.; Xia, Y.; Bai, J.; Yu, Y.; Xiao, F.; Ding, Y.; Ding, C.; et al. Sucrose nonfermenting-1-related protein kinase 1 regulates sheath-to-panicle transport of nonstructural carbohydrates during rice grain filling. Plant Physiol. 2022, 189, 1694–1714. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Liu, J.; Cao, B.; Liu, B.; Zhang, X.; Chen, Z.; Dong, C.; Liu, X.; Zhang, Z.; Wang, W.; et al. Reducing brassinosteroid signalling enhances grain yield in semi-dwarf wheat. Nature 2023, 617, 118–124. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Bai, M.Y.; Chong, K. Brassinosteroid-mediated regulation of agronomic traits in rice. Plant Cell Rep. 2014, 33, 683–696. [Google Scholar] [CrossRef]
- Sakamoto, T.; Morinaka, Y.; Ohnishi, T.; Sunohara, H.; Fujioka, S.; Ueguchi-Tanaka, M.; Mizutani, M.; Sakata, K.; Takatsuto, S.; Yoshida, S.; et al. Erect leaves caused by brassinosteroid deficiency increase biomass production and grain yield in rice. Nat. Biotechnol. 2006, 24, 105–109. [Google Scholar] [CrossRef] [PubMed]
- Morinaka, Y.; Sakamoto, T.; Inukai, Y.; Agetsuma, M.; Kitano, H.; Ashikari, M.; Matsuoka, M. Morphological alteration caused by brassinosteroid insensitivity increases the biomass and grain production of rice. Plant Physiol. 2006, 141, 924–931. [Google Scholar] [CrossRef] [PubMed]
- Divi, U.K.; Krishna, P. Brassinosteroid: A biotechnological target for enhancing crop yield and stress tolerance. New Biotechnol. 2009, 26, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Schröder, F.; Lisso, J.; Müssig, C. EXORDIUM-LIKE1 promotes growth during low carbon availability in Arabidopsis. Plant Physiol. 2011, 156, 1620–1630. [Google Scholar] [CrossRef]
- Oh, M.H.; Sun, J.; Oh, D.H.; Zielinski, R.E.; Clouse, S.D.; Huber, S.C. Enhancing Arabidopsis leaf growth by engineering the brassinosteroid insensitive1 receptor kinase. Plant Physiol. 2011, 157, 120–131. [Google Scholar] [CrossRef]
- Laxmi, A.; Paul, L.K.; Peters, J.L.; Khurana, J.P. Arabidopsis constitutive photomorphogenic mutant, bls1, displays altered brassinosteroid response and sugar sensitivity. Plant Mol. Biol. 2004, 56, 185–201. [Google Scholar] [CrossRef]
- Lisso, J.; Altmann, T.; Müssig, C. Metabolic changes in fruits of the tomato dx mutant. Phytochemistry 2006, 67, 2232–2238. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Xu, H.; Li, B.; Shang, Y.; Wei, M.; Zhang, S.; Zhao, C.; Qin, R.; Cui, F.; Wu, Y. The brassinosteroid biosynthesis gene, ZmD11, increases seed size and quality in rice and maize. Plant Physiol. Biochem. 2021, 160, 281–293. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef]
- Yu, X.; Lin, J.; Zack, D.J.; Qian, J. Computational analysis of tissue-specific combinatorial gene regulation: Predicting interaction between transcription factors in human tissues. Nucleic Acids Res. 2006, 34, 4925–4936. [Google Scholar] [CrossRef]
- Ruan, Y.L. Sucrose metabolism: Gateway to diverse carbon use and sugar signaling. Annu. Rev. Plant Biol. 2014, 65, 33–67. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Mukherjee, S.; Ayele, B.T. Molecular aspects of sucrose transport and its metabolism to starch during seed development in wheat: A comprehensive review. Biotechnol. Adv. 2018, 36, 954–967. [Google Scholar] [CrossRef] [PubMed]
- Nadolska-Orczyk, A.; Rajchel, I.K.; Orczyk, W.; Gasparis, S. Major genes determining yield-related traits in wheat and barley. Theor. Appl. Genet. 2017, 130, 1081–1098. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.Y.; Trieu, A.; Radhakrishnan, P.; Kwok, S.F.; Harris, S.; Zhang, K.; Wang, J.; Wan, J.; Zhai, H.; Takatsuto, S.; et al. Brassinosteroids regulate grain filling in rice. Plant Cell 2008, 20, 2130–2145. [Google Scholar] [CrossRef] [PubMed]
- Neuhaus, H.E.; Emes, M.J. Nonphotosynthetic metabolism in plastids. Annu Rev. Plant Biol. 2000, 51, 111–140. [Google Scholar] [CrossRef]
- Hou, J.; Huang, X.; Sun, W.; Du, C.; Wang, C.; Xie, Y.; Ma, D. Accumulation of water-soluble carbohydrates and gene expression in wheat stems correlates with drought resistance. J. Plant Physiol. 2018, 231, 182–191. [Google Scholar] [CrossRef]



| Sample | Raw_Reads | Raw_Bases | Valid_Reads | Valid_Bases | Valid% | Q20% | Q30% |
|---|---|---|---|---|---|---|---|
| CK1_1 | 69,940,236 | 9.86 G | 67,510,596 | 9.27 G | 96.53 | 96.82 | 91.48 |
| CK1_2 | 78,634,194 | 11.09 G | 74,233,766 | 10.28 G | 94.40 | 97.20 | 92.02 |
| CK1_3 | 59,538,838 | 8.39 G | 57,568,478 | 7.93 G | 96.69 | 96.80 | 91.25 |
| BR1_1 | 62,783,122 | 8.85 G | 60,346,868 | 8.26 G | 96.12 | 96.39 | 90.67 |
| BR1_2 | 80,168,182 | 11.30 G | 77,484,644 | 10.73 G | 96.65 | 96.81 | 91.11 |
| BR1_3 | 74,773,970 | 10.54 G | 72,013,958 | 9.96 G | 96.31 | 96.69 | 90.86 |
| CK2_1 | 73,030,808 | 10.30 G | 70,241,888 | 9.71 G | 96.18 | 96.83 | 91.11 |
| CK2_2 | 62,725,162 | 8.84 G | 60,350,984 | 8.35 G | 96.21 | 96.81 | 91.06 |
| CK2_3 | 71,055,804 | 10.02 G | 68,583,564 | 9.49 G | 96.52 | 96.88 | 91.30 |
| BR2_1 | 80,086,264 | 11.29 G | 76,328,078 | 10.61 G | 95.31 | 97.05 | 91.53 |
| BR2_2 | 75,388,476 | 10.63 G | 72,120,646 | 10.01 G | 95.67 | 96.88 | 91.18 |
| BR2_3 | 76,141,626 | 10.74 G | 73,096,770 | 10.14 G | 96.00 | 96.83 | 91.13 |
| CK3_1 | 85,975,682 | 12.12 G | 78,414,912 | 10.74 G | 91.21 | 96.64 | 90.93 |
| CK3_2 | 80,145,924 | 11.30 G | 74,536,402 | 10.27 G | 93.00 | 96.70 | 90.89 |
| CK3_3 | 71,487,958 | 10.08 G | 65,966,458 | 9.06 G | 92.28 | 96.66 | 90.88 |
| BR3_1 | 62,049,466 | 8.75 G | 58,120,858 | 8.01 G | 93.67 | 96.79 | 91.18 |
| BR3_2 | 76,144,758 | 10.74 G | 72,021,480 | 10.00 G | 94.58 | 97.07 | 91.59 |
| BR3_3 | 57,815,302 | 8.15 G | 54,521,084 | 7.53 G | 94.30 | 96.77 | 91.10 |
| Index | All | GC% | Min Length | Median Length | Max Length | Total Assembled Bases | N50 |
|---|---|---|---|---|---|---|---|
| Transcript | 85,214 | 41.01 | 201 | 691.00 | 16,336 | 83,855,058 | 1472 |
| Gene | 29,262 | 41.66 | 201 | 837.00 | 16,336 | 32,760,180 | 1661 |
| Module | Gene Number |
|---|---|
| bisque4 | 74 |
| brown | 4220 |
| brown4 | 183 |
| darkgrey | 298 |
| darkolivegreen | 107 |
| darkorange2 | 371 |
| darkseagreen4 | 37 |
| darkslateblue | 69 |
| darkturquoise | 175 |
| grey | 365 |
| grey60 | 218 |
| honeydew1 | 130 |
| ivory | 80 |
| lightcyan1 | 81 |
| lightsteelblue1 | 438 |
| lightyellow | 1381 |
| maroon | 1457 |
| mediumpurple3 | 90 |
| navajowhite2 | 63 |
| salmon4 | 163 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, S.; Xiao, R.; Li, X.; Li, R.; Song, C.; Li, Y.; Zhao, J.; Hou, X. Transcriptome Analysis of Paeonia ostii ‘Fengdan’ Seeds Uncovers Starch and Sucrose Metabolism Conferring High Yield Under Brassinosteroid Treatment. Genes 2025, 16, 1424. https://doi.org/10.3390/genes16121424
Yu S, Xiao R, Li X, Li R, Song C, Li Y, Zhao J, Hou X. Transcriptome Analysis of Paeonia ostii ‘Fengdan’ Seeds Uncovers Starch and Sucrose Metabolism Conferring High Yield Under Brassinosteroid Treatment. Genes. 2025; 16(12):1424. https://doi.org/10.3390/genes16121424
Chicago/Turabian StyleYu, Shixi, Ruixue Xiao, Xiaopeng Li, Renjie Li, Chengwei Song, Yuying Li, Jingyi Zhao, and Xiaogai Hou. 2025. "Transcriptome Analysis of Paeonia ostii ‘Fengdan’ Seeds Uncovers Starch and Sucrose Metabolism Conferring High Yield Under Brassinosteroid Treatment" Genes 16, no. 12: 1424. https://doi.org/10.3390/genes16121424
APA StyleYu, S., Xiao, R., Li, X., Li, R., Song, C., Li, Y., Zhao, J., & Hou, X. (2025). Transcriptome Analysis of Paeonia ostii ‘Fengdan’ Seeds Uncovers Starch and Sucrose Metabolism Conferring High Yield Under Brassinosteroid Treatment. Genes, 16(12), 1424. https://doi.org/10.3390/genes16121424






