Abstract
To generate gametes, sexually reproducing organisms need to achieve a reduction in ploidy, via meiosis. Several mechanisms are set in place to ensure proper reductional chromosome segregation at the first meiotic division (MI), including chromosome remodeling during late prophase I. Chromosome remodeling after crossover formation involves changes in chromosome condensation and restructuring, resulting in a compact bivalent, with sister kinetochores oriented to opposite poles, whose structure is crucial for localized loss of cohesion and accurate chromosome segregation. Here, we review the general processes involved in late prophase I chromosome remodeling, their regulation, and the strategies devised by different organisms to produce bivalents with configurations that promote accurate segregation.
1. Chromosome Remodeling
Sexual reproduction relies on the production of haploid gametes (i.e., eggs and sperm) from diploid germ cells during meiosis. The meiotic cell division program reduces the diploid chromosome number by half, by following a single round of DNA replication, with two rounds of cell division. In the first meiotic division (MI), homologous chromosomes segregate away from each other; whereas, in the second meiotic division (MII), sister chromatids separate []. During meiotic prophase I, homologous chromosomes organize into pairs, connected by crossovers and sister chromatid cohesion (bivalents) []. Bivalents are restructured during meiotic prophase I, to achieve a configuration that promotes their proper attachment to opposite poles of the MI spindle, and subsequent accurate meiotic chromosome segregation; therefore, errors in bivalent structural remodeling result in improper chromosome segregation, embryonic lethality, and birth defects [].
Chromosome remodeling is a key feature of meiosis. During early prophase, events such as the formation of programmed DNA double-strand breaks (DSBs), repair of a subset of these DSBs as interhomolog crossover recombination events, and assembly of the zipper-like scaffold, known as the synaptonemal complex (SC), between homologs require both localized as well as global changes in the chromosome structure. During late prophase, the structural remodeling of chromosomes involves changes in chromosome condensation, disassembly of the SC, and restructuring around the crossover site. Here, we will focus on the mechanisms underlying proper chromosome remodeling during late prophase I (pachytene to diakinesis transition), because of their importance in achieving a configuration, observed by late diakinesis/pro-metaphase I for bivalents, that promotes bipolar spindle attachment and localized loss of cohesion, summarizing the current knowledge for different organisms.
2. Crossover Formation Triggers Chromosome Remodeling in Late Prophase I
Meiotic crossovers (COs) are formed via the homology-directed repair of programmed DSBs, and both their frequency and distribution are tightly regulated. For example, every pair of homologous chromosomes must undergo at least one CO (obligate crossover), and when two or more COs are formed along a chromosome, they do not tend to occur near one another (crossover interference) [,]. Although there are exceptions (reviewed in []), COs are essential for accurate chromosome segregation in most organisms. Specifically, COs in conjunction with sister chromatid cohesion (SCC) result in structures referred to as “chiasmata”, which tether homolog pairs, thereby facilitating their correct orientation on the MI spindle [,,].
In most organisms, for CO formation, chromosomes are required to locate and pair with their homologs, and stabilize this association via assembly of the synaptonemal complex (SC) []. The SC is a proteinaceous structure whose organization is widely conserved. As revealed by electron microscopy, it consists of two lateral elements, formed by axis-associated proteins along each homolog, that flank a central region with proteins bridging both axes, resulting in a zipper-like structure [,,]. The SC is required to stabilize the interactions between pairs of homologous chromosomes until COs are formed, as shown in budding yeast, worms, flies, and mice [,,,,,,,,], and to prevent the formation of closely spaced COs in Arabidopsis [,].
CO formation, in turn, triggers chromosome remodeling in late prophase I, which involves the coordinated loss, gain, retention and/or relocalization of several important proteins along different chromosome subdomains. The end result of this chromosome remodeling is a configuration that will ensure the regulated removal of SCC, bipolar attachment at the spindle, and subsequent accurate chromosome segregation at MI (Figure 1). After CO formation, the SC begins to disassemble in an asymmetric manner, and SC proteins are either retained at specific chromosome subdomains until late prophase I (worms, budding yeast, flies, and mice) or completely removed from the chromosomes (plants).
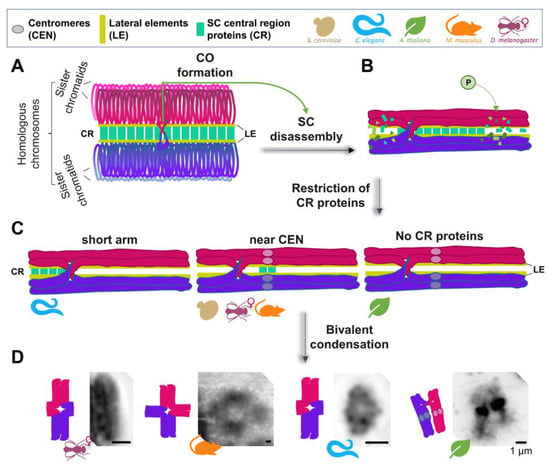
Figure 1.
Meiotic bivalent remodeling in different model organisms. (A) A pair of fully synapsed homologous chromosomes (in magenta and purple) is shown, with only the lateral element (LE) and central region (CR) proteins of the synaptonemal complex (SC) being highlighted for simplicity. (B) Upon interhomolog crossover (CO) formation, different types of regulation may trigger SC disassembly, starting in late pachytene (shown is the phosphorylation of SC proteins). (C) SC disassembly can result in either retention/restriction of the CR proteins of the SC to a particular subregion of the bivalents (such as the “short arm” in Caenorhabditis elegans, or near the centromeres in several other organisms), or the complete removal of CR proteins, such as in Arabidopsis thaliana. The process of late prophase I bivalent remodeling includes changes in chromatin condensation and compaction, to produce the final bivalent structure, characteristic of diakinesis/prometaphase I. (D) Representative images of DAPI-stained bivalents at this stage are shown for Drosophila melanogaster (courtesy of Stacie Hughes and Scott Hawley, Stowers Institute for Medical Research), Mus musculus (courtesy of Tegan S. Horan and Paula Cohen, Cornell University), C. elegans, and A. thaliana (courtesy of Monica Pradillo, Universidad Complutense de Madrid), to provide examples of the conservation and diversity of bivalent morphology.
In C. elegans, which carry holocentric chromosomes (centromere activity is distributed throughout the chromosome length) and exhibit absolute CO interference, a single CO is formed per homologous chromosome pair []. Furthermore, since this CO usually occurs in an off-center position along the length of the chromosome, late prophase remodeling results in six cruciform-shaped asymmetric bivalents, with distinct long and short arms [,,]. Following CO formation, chromosome remodeling is initiated with the asymmetric disassembly of the SC. Central region proteins (SYP-1/2/3/4/5/6) are lost along the longest distance from the site of the CO to a chromosome end (the long arm of the bivalent), and persist on the short arms of the bivalent [,,]. In contrast, the HORMA domain, containing the axis-associated proteins HTP-1 and HTP-2, is lost from the short arms of the bivalent and retained on the long arms []. The axis-associated components HTP-3 and HIM-3 retain a relatively uniform distribution along the length of the chromosome axes, and remain associated with both the long and short arms of the bivalent [,]. In late diakinesis, the disassembly of the SC proteins on the short arm requires the conserved nuclear protein Akirin (AKIR-1 in worms), which acts through a mechanism that is still unknown []. The asymmetric disassembly of the SC central region components, dictated by the position of the CO, was proposed to play an important role in regulating the loss of SCC at MI in C. elegans []. This was directly tested through DSB induction at specific genomic positions, by Mos1 transposon excision in spo-11 mutants lacking endogenous meiotic DSB formation, which revealed that the CO position in early meiotic prophase must be tightly regulated for proper designation of chromosome subdomains and chromosome remodeling, resulting in the regulated loss of sister chromatid cohesion and accurate chromosome segregation in C. elegans [].
In organisms with monocentric chromosomes (with a discrete or confined centromere), such as yeast and flies, the SC also disassembles asymmetrically, and persists at centromeres until late prophase [,,]. Similarly, in mouse spermatocytes, the SC is retained at paired centromeres, after it is lost along the length of the chromosomes [,]. In budding yeast, as the SC disassembles, the central region component Zip1 is preferentially retained at the centromeres, where it remains associated until the centromeres are attached to microtubules. Furthermore, evidence suggests that this localized retention promotes centromere biorientation, to ensure homologous chromosome segregation to opposite poles at meiosis I []. In flies, the SC starts to progressively disassemble along the chromosome arms during mid prophase I, and is completely disassembled from the arms by late prophase I. This loss of the SC coincides temporally with the loading of condensin complex components []. Interestingly, SC components, such as the central region proteins C(3)G and Cona, as well as the cohesin subunit SMC1, remain associated with the centromeres beyond euchromatic SC disassembly []. In mouse spermatocytes, the SC proteins that persist at the chiasmata, after SC disassembly, are proposed to regulate local remodeling of homologous chromosome axes, thereby promoting centromere and chiasma functions. In mice, instead of triggering global chromosome remodeling, COs are proposed to generate local changes in SCC and chromatin condensation []. CO-triggered local changes/destabilization of chromosome axes have been described in other organisms. In rat spermatocytes at diplotene, the bridges between homologs, observed at chiasma sites, are not stained by antibodies that recognize the meiosis-specific cohesin REC8; however, REC8 is present on the axes flanking the chiasma []. Additionally, in Sordaria, it has been demonstrated, by EM, that sister chromatid axes are found to be locally separated at CO sites []. Plants, such as A. thaliana, undergo a complete loss of the SC central region proteins from the bivalents during late prophase I remodeling [], while the ASY1 lateral element protein can persist [], occasionally in strange “tinsel-like” diplotene structures, such as in wheat and barley [].
3. Restructuring of the Bivalent to Form a Compact Structure
3.1. Regulation of SC Disassembly
Cell cycle kinases and post-translational regulation of SC components play an important role in regulating SC disassembly (reviewed in [,]). Yeast requires Cdc5 (Polo-like kinase), Ipl1 (Aurora B), DDK (Dbf4-dependent Cdc7 kinase), and CDK1 for SC disassembly [,,]. However, their targets that lead to SC disassembly remain unclear. PLK1 and Aurora B are also required for SC disassembly in mice. In this case, PLK1 directly phosphorylates the SC central element proteins SYCP1, TEX12, and SYCE1 during meiotic prophase, helping with the disassembly of the central region and lateral element proteins of the SC []. Aurora B and the inner centromere protein (INCEP) also localize to the meiotic chromosomes during prophase []. However, unlike PLK1, Aurora kinase activity is required exclusively for disassembly of the lateral elements []. CDK1–cyclin B is also involved in this process [,,]. Currently, there is no evidence suggesting that cell cycle kinases are involved in SC disassembly in flies. However, many of these kinases are present during late prophase I female meiosis in flies, so it is possible that their role in disassembly is yet to be uncovered. The involvement of mitogen-activated protein (MAP) kinase has been revealed recently in C. elegans. Specifically, ECT-2, the mammalian Rho guanine nucleotide exchange factor homolog, working through the conserved RAS/ERK MAP kinase signaling pathway, regulates the disassembly of the SC, via direct phosphorylation of the central region component SYP-2 []. Additionally, regulation of the HIM-3 phosphorylation status is required for timely disassembly of SC central region proteins from the long arm, and for proper timing of HTP-1 and HTP-2 dissociation from the short arm [].
3.2. Changes in Chromosome Condensation and Compaction
The late prophase I chromosome remodeling process involves changes in chromosome condensation and compaction, by which meiotic chromosomes are transformed from the elongated linear structures visible at mid-pachytene into the condensed, highly asymmetrical cross-shaped bivalent structures present at diakinesis (worms and mice) and prometaphase I (flies), or into “rings” or “rods” (plants). The multi-subunit condensin complex is responsible for chromosome condensation in multiple organisms. At least two different types of condensin complexes, known as condensin I and II, exist among eukaryotes []. The two complexes share the same pair of SMC2 and SMC4 subunits, both belonging to the structural maintenance of chromosomes (SMC) family of chromosomal ATPases. Additionally, each complex has a unique set of three non-SMC subunits (i.e., CAP-D2, CAP-G, and CAP-H for condensin I, and CAPD3, CAP-G2, and CAP-H2 for condensin II) (reviewed in []).
In Saccharomyces cerevisiae, the single condensin complex localizes to the axial core of pachytene chromosomes and contributes to their axial compaction []. Chromosome bridges are often observed during anaphase I in condensin mutants, indicating that condensin function is required for proper chromosome segregation. Condensin has also been shown to help recruit Polo/Cdc5 to meiotic chromosomes, which, in turn, contributes to cohesin release, through its direct phosphorylation []. A four-subunit complex, known as monopolin, plays a central role in co-orienting sister kinetochores during meiosis I [,]. Condensin contributes to properly localizing a monopolin subunit (Mam1) at kinetochores, thereby ensuring co-orientation [].
Studies in A. thaliana suggest that condensin I might be sufficient to support meiotic chromosome condensation, and, therefore, proper chromosome segregation, in plants [,,]. Among the three condensin complexes present in C. elegans, only condensin II associates with sister chromatids during diplotene and diakinesis, and mediates their compaction and resolution, contributing to the formation of compact and well-resolved cruciform bivalents []. Additionally, AKIR-1 affects the condensation of the bivalents, independently of condensin []. In Drosophila, CAP-G, a condensin I subunit, is required for SC disassembly, and to keep chromosomes in their metaphase I configuration in female meiosis []. Based on the results from antibody injection experiments, it has been proposed that condensins I and II play distinct roles during the construction of bivalent chromosomes in mouse oocytes; condensin II is involved in resolving sister chromatid axes, whereas condensin I may assemble or reinforce a unique centromeric structure, contributing to the monopolar attachment of sister kinetochores []. In addition to condensin complexes, other axis-associated components, such as topoisomerase II, also seem to participate in the compaction of bivalents during late prophase I remodeling in several organisms [,,].
3.3. Regulation of Loss of Cohesion
The loss, gain, and retention of proteins in the newly established subdomains, generated during late prophase I chromosome remodeling, is critical for the regulated stepwise loss of SCC [,,,]. In monocentric organisms, such as yeast, flies, Arabidopsis, and mice, both the SC and cohesin are lost along the chromosome arms, but persist at the centromere to ensure that sister chromatids segregate together, probably by promoting proper centromere biorientation (reviewed in []). Once homolog biorientation has occurred, separase becomes active. Separase cleaves the meiosis-specific cohesin Rec8 at the chromosome arms only, while, in centromeric regions, Rec8 is protected from separase activity by the presence of shugoshin. Rec8 phosphorylation is essential for its cleavage by separase [,,,,]. At the pericentromere, members of the shugoshin/Mei-S332 protein family counteract this phosphorylation by recruiting a specific form of the protein phosphatase 2A [,,], retaining cohesion in this region until meiosis II. In C. elegans, which has holocentric chromosomes, upon loss of the residual SC central region proteins, localized at the short arms of the bivalent at late diakinesis, the Aurora B kinase homolog AIR-2 localizes to the short arm, where it phosphorylates REC-8, allowing its cleavage by separase at anaphase I, and subsequent segregation of homologous chromosomes to opposite spindle poles [,]. LAB-1, the functional analog of shugoshin, which, during earlier prophase, was localized throughout the length of the chromosomes, is restricted to the long arms by late prophase I, protecting SCC along this domain from AIR-2 phosphorylation and premature removal, by targeting GSP-1 and GSP-2 (PPI protein phosphatase homologs) to this chromosomal region [,,]. In addition, the chromatin-associated protein HIM-17 has been implicated in preventing the loading of AIR-2 on the long arm of the bivalent, by regulating the expression, localization, and possibly the phosphorylation of GSP-1 and GSP-2 [].
4. Concluding Remarks/Open Questions
Here, we summarized the progress made in elucidating the mechanisms regulating chromosome remodeling during late prophase I. Although different organisms use distinct strategies, late prophase I chromosome remodeling is a conserved and crucial process that ensures the regulated loss of sister chromatid cohesion, bipolar attachment of homologs at the metaphase I spindle, and accurate chromosome segregation, avoiding aneuploidy. One of the main differences among species derives from whether their chromosomes are monocentric or holocentric. In monocentric organisms, the centromere serves as the focal point for mechanisms that serve to shape the remodeling process, such as co-orienting sister chromatids and promoting the local protection of cohesion to allow for the two-step release of SCC. Holocentric organisms apply strategies, such as using the CO position as the symmetry breaking point that subdivides the bivalent into distinct subdomains with different fates, that can promote the localized removal of SCC.
Although some of the basic mechanisms underlying late prophase I chromosome remodeling are understood, there are still many unanswered important questions, including the following: What kind of signals are used to indicate CO formation and trigger post-translational modifications of SC proteins and asymmetric SC disassembly? What mechanisms assess the distance between a CO and the ends of the chromosomes, to determine subdomains in holocentric chromosomes? What other proteins/mechanisms of regulation are implicated in asymmetric SC disassembly? How is SC disassembly connected with changes in chromosome condensation? Further studies in different organisms will reveal additional levels of regulation and identify new proteins, contributing to a better understanding of chromosome remodeling in late prophase I.
Author Contributions
Writing—original draft preparation, L.I.L.-L. and M.P.C.; writing—review and editing, L.I.L.-L., M.M.-G. and M.P.C.; visualization, M.M.-G.; supervision, M.P.C.; funding acquisition, M.P.C. All authors have read and agreed to the published version of the manuscript.
Funding
This work was funded by a National Institutes of Health grant R01GM072551 to M.P.C.
Acknowledgments
We are grateful to be part of a very collaborative environment in the meiosis field. We specially thank the following researchers, who provided us with images: Stacie Hughes and Scott Hawley (Drosophila), Tegan S. Horan and Paula Cohen (mouse), and Monica Pradillo (Arabidopsis). We also thank Brianna Alico for critically reading this manuscript and for providing helpful suggestions.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Láscarez-Lagunas, L.; Martinez-Garcia, M.; Colaiácovo, M. SnapShot: Meiosis—Prophase I. Cell 2020, 181, 1442. [Google Scholar] [CrossRef]
- Zickler, D.; Kleckner, N. Meiotic chromosomes: Integrating structure and function. Annu. Rev. Genet. 1999, 33, 603–754. [Google Scholar] [CrossRef]
- Hassold, T.; Hunt, P. To err (meiotically) is human: The genesis of human aneuploidy. Nat. Rev. Genet. 2001, 2, 280–291. [Google Scholar] [CrossRef] [PubMed]
- Jones, G.H. The control of chiasma distribution. Symp. Soc. Exp. Biol. 1984, 38, 293–320. [Google Scholar]
- Jones, G.H.; Franklin, F.C.H. Meiotic crossing-over: Obligation and interference. Cell 2006, 126, 246–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wolf, K.W. How meiotic cells deal with non-exchange chromosomes. Bioessays 1994, 16, 107–114. [Google Scholar] [CrossRef]
- Hunter, N. Meiotic recombination: The essence of heredity. Cold Spring Harb. Perspect. Biol. 2015, 7, a016618. [Google Scholar] [CrossRef] [Green Version]
- Gray, S.; Cohen, P.E. Control of meiotic ccrossovers: From double-strand break formation to designation. Annu. Rev. Genet. 2016, 50, 175–210. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pazhayam, N.M.; Turcotte, C.A.; Sekelsky, J. Meiotic crossover patterning. Front. Cell Dev. Biol. 2021, 9, 681123. [Google Scholar] [CrossRef] [PubMed]
- Colaiácovo, M.P. The many facets of SC function during C. elegans meiosis. Chromosoma 2006, 115, 195–211. [Google Scholar] [CrossRef]
- Page, S.L.; Hawley, R.S. The genetics and molecular biology of the synaptonemal complex. Annu. Rev. Cell Dev. Biol. 2004, 20, 525–558. [Google Scholar] [CrossRef] [PubMed]
- Nag, D.K.; Scherthan, H.; Rockmill, B.; Bhargava, J.; Roeder, G.S. Heteroduplex DNA formation and homolog pairing in yeast meiotic mutants. Genetics 1995, 141, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Storlazzi, A.; Xu, L.; Schwacha, A.; Kleckner, N. Synaptonemal Complex (SC) component Zip1 plays a role in meiotic recombination independent of SC polymerization along the chromosomes. Proc. Natl. Acad. Sci. USA 1996, 93, 9043–9048. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Page, S.L.; Hawley, R.S. C(3)G encodes a Drosophila synaptonemal complex protein. Genes Dev. 2001, 15, 3130–3143. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- MacQueen, A.J.; Colaiácovo, M.P.; McDonald, K.; Villeneuve, A.M. Synapsis-dependent and -independent mechanisms stabilize homolog pairing during meiotic prophase in C. elegans. Genes Dev. 2002, 16, 2428–2442. [Google Scholar] [CrossRef] [Green Version]
- Colaiácovo, M.P.; MacQueen, A.J.; Martinez-Perez, E.; McDonald, K.; Adamo, A.; La Volpe, A.; Villeneuve, A.M. Synaptonemal complex assembly in C. elegans is dispensable for loading strand-exchange proteins but critical for proper completion of recombination. Dev. Cell 2003, 5, 463–474. [Google Scholar] [CrossRef] [Green Version]
- De Vries, F.A.T.; de Boer, E.; van den Bosch, M.; Baarends, W.M.; Ooms, M.; Yuan, L.; Liu, J.-G.; van Zeeland, A.A.; Heyting, C.; Pastink, A. Mouse Sycp1 functions in synaptonemal complex assembly, meiotic recombination, and XY body formation. Genes Dev. 2005, 19, 1376–1389. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Smolikov, S.; Eizinger, A.; Schild-Prufert, K.; Hurlburt, A.; McDonald, K.; Engebrecht, J.; Villeneuve, A.M.; Colaiácovo, M.P. SYP-3 restricts synaptonemal complex assembly to bridge paired chromosome axes during meiosis in Caenorhabditis elegans. Genetics 2007, 176, 2015–2025. [Google Scholar] [CrossRef] [Green Version]
- Smolikov, S.; Eizinger, A.; Hurlburt, A.; Rogers, E.; Villeneuve, A.M.; Colaiácovo, M.P. Synapsis-defective mutants reveal a correlation between chromosome conformation and the mode of double-strand break repair during Caenorhabditis elegans meiosis. Genetics 2007, 176, 2027–2033. [Google Scholar] [CrossRef] [Green Version]
- Smolikov, S.; Schild-Prüfert, K.; Colaiácovo, M.P. A yeast two-hybrid screen for SYP-3 interactors identifies SYP-4, a component required for synaptonemal complex assembly and chiasma formation in Caenorhabditis elegans Meiosis. PLoS Genet. 2009, 5, e1000669. [Google Scholar] [CrossRef] [Green Version]
- France, M.G.; Enderle, J.; Röhrig, S.; Puchta, H.; Franklin, F.C.H.; Higgins, J.D. ZYP1 is required for obligate cross-over formation and cross-over interference in Arabidopsis. Proc. Natl. Acad. Sci. USA 2021, 118, e2021671118. [Google Scholar] [CrossRef] [PubMed]
- Capilla-Pérez, L.; Durand, S.; Hurel, A.; Lian, Q.; Chambon, A.; Taochy, C.; Solier, V.; Grelon, M.; Mercier, R. The synaptonemal complex imposes crossover interference and heterochiasmy in Arabidopsis. Proc. Natl. Acad. Sci. USA 2021, 118, e2023613118. [Google Scholar] [CrossRef] [PubMed]
- Meneely, P.M.; Farago, A.F.; Kauffman, T.M. Crossover distribution and high interference for both the X Chromosome and an autosome during oogenesis and spermatogenesis in Caenorhabditis elegans. Genetics 2002, 162, 1169–1177. [Google Scholar] [CrossRef]
- Barnes, T.M.; Kohara, Y.; Coulson, A.; Hekimi, S. Meiotic recombination, noncoding DNA and genomic organization in Caenorhabditis elegans. Genetics 1995, 141, 159–179. [Google Scholar] [CrossRef] [PubMed]
- Schvarzstein, M.; Wignall, S.M.; Villeneuve, A.M. Coordinating cohesion, co-orientation, and congression during meiosis: Lessons from holocentric chromosomes. Genes Dev. 2010, 24, 219–228. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lui, D.Y.; Colaiácovo, M.P. Meiotic development in Caenorhabditis elegans. In Germ Cell Development in C. elegans; Schedl, T., Ed.; Springer: New York, NY, USA, 2013; Volume 757, pp. 133–170. ISBN 978-1-4614-4014-7. [Google Scholar]
- Nabeshima, K.; Villeneuve, A.M.; Colaiácovo, M.P. Crossing over is coupled to late meiotic prophase bivalent differentiation through asymmetric disassembly of the SC. J. Cell Biol. 2005, 168, 683–689. [Google Scholar] [CrossRef] [Green Version]
- Hurlock, M.E.; Čavka, I.; Kursel, L.E.; Haversat, J.; Wooten, M.; Nizami, Z.; Turniansky, R.; Hoess, P.; Ries, J.; Gall, J.G.; et al. Identification of novel synaptonemal complex components in C. elegans. J. Cell Biol. 2020, 219, e201910043. [Google Scholar] [CrossRef] [Green Version]
- Zhang, Z.; Xie, S.; Wang, R.; Guo, S.; Zhao, Q.; Nie, H.; Liu, Y.; Zhang, F.; Chen, M.; Liu, L.; et al. Multivalent weak interactions between assembly units drive synaptonemal complex formation. J. Cell Biol. 2020, 219, e201910086. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinez-Perez, E.; Schvarzstein, M.; Barroso, C.; Lightfoot, J.; Dernburg, A.F.; Villeneuve, A.M. Crossovers trigger a remodeling of meiotic chromosome axis composition that is linked to two-step loss of sister chromatid cohesion. Genes Dev. 2008, 22, 2886–2901. [Google Scholar] [CrossRef] [Green Version]
- Chan, R.C.; Severson, A.F.; Meyer, B.J. Condensin restructures chromosomes in preparation for meiotic divisions. J. Cell Biol. 2004, 167, 613–625. [Google Scholar] [CrossRef]
- Clemons, A.M.; Brockway, H.M.; Yin, Y.; Kasinathan, B.; Butterfield, Y.S.; Jones, S.J.M.; Colaiácovo, M.P.; Smolikove, S. Akirin is required for diakinesis bivalent structure and synaptonemal complex disassembly at meiotic Prophase I. Mol. Biol. Cell 2013, 24, 1053–1067. [Google Scholar] [CrossRef] [PubMed]
- Altendorfer, E.; Láscarez-Lagunas, L.I.; Nadarajan, S.; Mathieson, I.; Colaiácovo, M.P. Crossover position drives chromosome remodeling for accurate meiotic chromosome segregation. Curr. Biol. 2020, 30, 1329–1338.e7. [Google Scholar] [CrossRef] [PubMed]
- Gladstone, M.N.; Obeso, D.; Chuong, H.; Dawson, D.S. The synaptonemal complex protein Zip1 promotes bi-orientation of centromeres at Meiosis I. PLoS Genet. 2009, 5, e1000771. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Newnham, L.; Jordan, P.; Rockmill, B.; Roeder, G.S.; Hoffmann, E. The synaptonemal complex protein, Zip1, promotes the segregation of nonexchange chromosomes at Meiosis I. Proc. Natl. Acad. Sci. USA 2010, 107, 781–785. [Google Scholar] [CrossRef] [Green Version]
- Takeo, S.; Lake, C.M.; Morais-de-Sá, E.; Sunkel, C.E.; Hawley, R.S. Synaptonemal complex-dependent centromeric clustering and the initiation of synapsis in drosophila oocytes. Curr. Biol. 2011, 21, 1845–1851. [Google Scholar] [CrossRef] [Green Version]
- Qiao, H.; Chen, J.K.; Reynolds, A.; Höög, C.; Paddy, M.; Hunter, N. Interplay between synaptonemal complex, homologous recombination, and centromeres during mammalian meiosis. PLoS Genet. 2012, 8, e1002790. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bisig, C.G.; Guiraldelli, M.F.; Kouznetsova, A.; Scherthan, H.; Höög, C.; Dawson, D.S.; Pezza, R.J. Synaptonemal complex components persist at centromeres and are required for homologous centromere pairing in mouse spermatocytes. PLoS Genet. 2012, 8, e1002701. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Resnick, T.D.; Dej, K.J.; Xiang, Y.; Hawley, R.S.; Ahn, C.; Orr-Weaver, T.L. Mutations in the chromosomal passenger complex and the condensin complex differentially affect synaptonemal complex disassembly and metaphase I configuration in drosophila female meiosis. Genetics 2009, 181, 875–887. [Google Scholar] [CrossRef] [Green Version]
- Eijpe, M.; Offenberg, H.; Jessberger, R.; Revenkova, E.; Heyting, C. Meiotic cohesin REC8 marks the axial elements of rat synaptonemal complexes before cohesins SMC1beta and SMC3. J. Cell Biol. 2003, 160, 657–670. [Google Scholar] [CrossRef] [Green Version]
- Storlazzi, A.; Tesse, S.; Ruprich-Robert, G.; Gargano, S.; Pöggeler, S.; Kleckner, N.; Zickler, D. Coupling meiotic chromosome axis integrity to recombination. Genes Dev. 2008, 22, 796–809. [Google Scholar] [CrossRef] [Green Version]
- Osman, K.; Higgins, J.D.; Sanchez-Moran, E.; Armstrong, S.J.; Franklin, F.C.H. Pathways to meiotic recombination in Arabidopsis Thaliana: Tansley review. New Phytol. 2011, 190, 523–544. [Google Scholar] [CrossRef] [PubMed]
- Chelysheva, L.; Vezon, D.; Chambon, A.; Gendrot, G.; Pereira, L.; Lemhemdi, A.; Vrielynck, N.; Le Guin, S.; Novatchkova, M.; Grelon, M. The Arabidopsis HEI10 is a new ZMM protein related to Zip3. PLoS Genet. 2012, 8, e1002799. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Colas, I.; Darrier, B.; Arrieta, M.; Mittmann, S.U.; Ramsay, L.; Sourdille, P.; Waugh, R. Observation of extensive chromosome axis remodeling during the “diffuse-phase” of meiosis in large genome cereals. Front. Plant Sci. 2017, 8, 1235. [Google Scholar] [CrossRef] [Green Version]
- Cahoon, C.K.; Hawley, R.S. Regulating the construction and demolition of the synaptonemal complex. Nat. Struct. Mol. Biol. 2016, 23, 369–377. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Colaiácovo, M.P. Zipping and unzipping: Protein modifications regulating synaptonemal complex dynamics. Trends Genet. 2018, 34, 232–245. [Google Scholar] [CrossRef] [PubMed]
- Sourirajan, A.; Lichten, M. Polo-like kinase Cdc5 drives exit from pachytene during budding yeast meiosis. Genes Dev. 2008, 22, 2627–2632. [Google Scholar] [CrossRef] [Green Version]
- Jordan, P.; Copsey, A.; Newnham, L.; Kolar, E.; Lichten, M.; Hoffmann, E. Ipl1/Aurora B kinase coordinates synaptonemal complex disassembly with cell cycle progression and crossover formation in budding yeast meiosis. Genes Dev. 2009, 23, 2237–2251. [Google Scholar] [CrossRef] [Green Version]
- Argunhan, B.; Leung, W.-K.; Afshar, N.; Terentyev, Y.; Subramanian, V.V.; Murayama, Y.; Hochwagen, A.; Iwasaki, H.; Tsubouchi, T.; Tsubouchi, H. Fundamental cell cycle kinases collaborate to ensure timely destruction of the synaptonemal complex during meiosis. EMBO J. 2017, 36, 2488–2509. [Google Scholar] [CrossRef]
- Jordan, P.W.; Karppinen, J.; Handel, M.A. Polo-like Kinase is required for synaptonemal complex disassembly and phosphorylation in mouse spermatocytes. J. Cell Sci. 2012, 125, 5061–5072. [Google Scholar] [CrossRef] [Green Version]
- Parra, M.T.; Viera, A.; Gómez, R.; Page, J.; Carmena, M.; Earnshaw, W.C.; Rufas, J.S.; Suja, J.A. Dynamic relocalization of the chromosomal passenger complex proteins inner centromere protein (INCENP) and aurora-B kinase during male mouse meiosis. J. Cell Sci. 2003, 116, 961–974. [Google Scholar] [CrossRef] [Green Version]
- Sun, F.; Handel, M.A. Regulation of the meiotic Prophase I to Metaphase I transition in mouse spermatocytes. Chromosoma 2008, 117, 471–485. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Allen, J.W.; Dix, D.J.; Collins, B.W.; Merrick, B.A.; He, C.; Selkirk, J.K.; Poorman-Allen, P.; Dresser, M.E.; Eddy, E.M. HSP70-2 is part of the synaptonemal complex in mouse and hamster spermatocytes. Chromosoma 1996, 104, 414–421. [Google Scholar] [CrossRef] [PubMed]
- Dix, D.J.; Allen, J.W.; Collins, B.W.; Poorman-Allen, P.; Mori, C.; Blizard, D.R.; Brown, P.R.; Goulding, E.H.; Strong, B.D.; Eddy, E.M. HSP70-2 is required for desynapsis of synaptonemal complexes during meiotic Prophase in juvenile and adult mouse spermatocytes. Development 1997, 124, 4595–4603. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.; Dix, D.J.; Eddy, E.M. HSP70-2 is required for CDC2 kinase activity in meiosis I of mouse spermatocytes. Development 1997, 124, 3007–3014. [Google Scholar] [CrossRef] [PubMed]
- Nadarajan, S.; Mohideen, F.; Tzur, Y.B.; Ferrandiz, N.; Crawley, O.; Montoya, A.; Faull, P.; Snijders, A.P.; Cutillas, P.R.; Jambhekar, A.; et al. The MAP kinase pathway coordinates crossover designation with disassembly of synaptonemal complex proteins during meiosis. eLife 2016, 5, e12039. [Google Scholar] [CrossRef] [PubMed]
- Sato-Carlton, A.; Nakamura-Tabuchi, C.; Li, X.; Boog, H.; Lehmer, M.K.; Rosenberg, S.C.; Barroso, C.; Martinez-Perez, E.; Corbett, K.D.; Carlton, P.M. Phosphoregulation of HORMA domain protein HIM-3 promotes asymmetric synaptonemal complex disassembly in meiotic Prophase in Caenorhabditis elegans. PLoS Genet. 2020, 16, e1008968. [Google Scholar] [CrossRef] [PubMed]
- Hirano, T. At the heart of the chromosome: SMC proteins in action. Nat. Rev. Mol. Cell Biol. 2006, 7, 311–322. [Google Scholar] [CrossRef]
- Hirano, T. Condensins: Universal organizers of chromosomes with diverse functions. Genes Dev. 2012, 26, 1659–1678. [Google Scholar] [CrossRef] [Green Version]
- Yu, H.-G.; Koshland, D.E. Meiotic condensin is required for proper chromosome compaction, SC assembly, and resolution of recombination-dependent chromosome linkages. J. Cell Biol. 2003, 163, 937–947. [Google Scholar] [CrossRef] [Green Version]
- Yu, H.-G.; Koshland, D. Chromosome morphogenesis: Condensin-dependent cohesin removal during meiosis. Cell 2005, 123, 397–407. [Google Scholar] [CrossRef]
- Rabitsch, K.P.; Petronczki, M.; Javerzat, J.P.; Genier, S.; Chwalla, B.; Schleiffer, A.; Tanaka, T.U.; Nasmyth, K. Kinetochore recruitment of two nucleolar proteins is required for homolog segregation in Meiosis I. Dev. Cell 2003, 4, 535–548. [Google Scholar] [CrossRef] [Green Version]
- Corbett, K.D.; Yip, C.K.; Ee, L.-S.; Walz, T.; Amon, A.; Harrison, S.C. The monopolin complex crosslinks kinetochore components to regulate chromosome-microtubule attachments. Cell 2010, 142, 556–567. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brito, I.L.; Yu, H.-G.; Amon, A. Condensins promote coorientation of sister chromatids during Meiosis I in budding yeast. Genetics 2010, 185, 55–64. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siddiqui, N.U.; Stronghill, P.E.; Dengler, R.E.; Hasenkampf, C.A.; Riggs, C.D. Mutations in Arabidopsis condensin genes disrupt embryogenesis, meristem organization and segregation of homologous chromosomes during meiosis. Development 2003, 130, 3283–3295. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Siddiqui, N.U.; Rusyniak, S.; Hasenkampf, C.A.; Riggs, C.D. Disruption of the Arabidopsis SMC4 gene, AtCAP-C, compromises gametogenesis and embryogenesis. Planta 2006, 223, 990–997. [Google Scholar] [CrossRef]
- Sakamoto, T.; Inui, Y.T.; Uraguchi, S.; Yoshizumi, T.; Matsunaga, S.; Mastui, M.; Umeda, M.; Fukui, K.; Fujiwara, T. Condensin II alleviates DNA damage and is essential for tolerance of boron overload stress in Arabidopsis. Plant Cell 2011, 23, 3533–3546. [Google Scholar] [CrossRef] [Green Version]
- Lee, J.; Ogushi, S.; Saitou, M.; Hirano, T. Condensins I and II are essential for construction of bivalent chromosomes in mouse oocytes. Mol. Biol. Cell 2011, 22, 3465–3477. [Google Scholar] [CrossRef]
- Cobb, J.; Miyaike, M.; Kikuchi, A.; Handel, M.A. Meiotic events at the centromeric heterochromatin: Histone H3 phosphorylation, Topoisomerase II alpha localization and chromosome condensation. Chromosoma 1999, 108, 412–425. [Google Scholar] [CrossRef]
- Agnieszka, W. Etoposide interferes with the process of chromatin condensation during alga Chara Vulgaris spermiogenesis. Micron 2014, 65, 45–50. [Google Scholar] [CrossRef]
- Martinez-Garcia, M.; Schubert, V.; Osman, K.; Darbyshire, A.; Sanchez-Moran, E.; Franklin, F.C.H. TOPII and chromosome movement help remove interlocks between entangled chromosomes during Meiosis. J. Cell Biol. 2018, 217, 4070–4079. [Google Scholar] [CrossRef]
- De Carvalho, C.E.; Zaaijer, S.; Smolikov, S.; Gu, Y.; Schumacher, J.M.; Colaiacovo, M.P. LAB-1 antagonizes the Aurora B kinase in C. elegans. Genes Dev. 2008, 22, 2869–2885. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marston, A.L. Shugoshins: Tension-sensitive pericentromeric adaptors safeguarding chromosome segregation. Mol. Cell. Biol. 2015, 35, 634–648. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Clyne, R.K.; Katis, V.L.; Jessop, L.; Benjamin, K.R.; Herskowitz, I.; Lichten, M.; Nasmyth, K. Polo-like kinase Cdc5 promotes chiasmata formation and cosegregation of sister centromeres at Meiosis I. Nat. Cell Biol. 2003, 5, 480–485. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.H.; Amon, A. Role of polo-like kinase CDC5 in programming Meiosis I chromosome segregation. Science 2003, 300, 482–486. [Google Scholar] [CrossRef] [PubMed]
- Brar, G.A.; Kiburz, B.M.; Zhang, Y.; Kim, J.-E.; White, F.; Amon, A. Rec8 phosphorylation and recombination promote the step-wise loss of cohesins in meiosis. Nature 2006, 441, 532–536. [Google Scholar] [CrossRef]
- Katis, V.L.; Lipp, J.J.; Imre, R.; Bogdanova, A.; Okaz, E.; Habermann, B.; Mechtler, K.; Nasmyth, K.; Zachariae, W. Rec8 phosphorylation by casein kinase 1 and Cdc7-Dbf4 kinase regulates cohesin cleavage by separase during meiosis. Dev. Cell 2010, 18, 397–409. [Google Scholar] [CrossRef] [Green Version]
- Ishiguro, T.; Tanaka, K.; Sakuno, T.; Watanabe, Y. Shugoshin–PP2A counteracts casein-kinase-1-dependent cleavage of Rec8 by separase. Nat. Cell Biol. 2010, 12, 500–506. [Google Scholar] [CrossRef] [PubMed]
- Riedel, C.G.; Katis, V.L.; Katou, Y.; Mori, S.; Itoh, T.; Helmhart, W.; Gálová, M.; Petronczki, M.; Gregan, J.; Cetin, B.; et al. Protein phosphatase 2A protects centromeric sister chromatid cohesion during Meiosis I. Nature 2006, 441, 53–61. [Google Scholar] [CrossRef]
- Kitajima, T.S.; Sakuno, T.; Ishiguro, K.; Iemura, S.; Natsume, T.; Kawashima, S.A.; Watanabe, Y. Shugoshin collaborates with protein phosphatase 2A to protect cohesin. Nature 2006, 441, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Cetin, B.; Anger, M.; Cho, U.S.; Helmhart, W.; Nasmyth, K.; Xu, W. Structure and function of the PP2A-shugoshin interaction. Mol. Cell 2009, 35, 426–441. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rogers, E.; Bishop, J.D.; Waddle, J.A.; Schumacher, J.M.; Lin, R. The Aurora kinase AIR-2 functions in the release of chromosome cohesion in Caenorhabditis elegans meiosis. J. Cell Biol. 2002, 157, 219–229. [Google Scholar] [CrossRef] [PubMed]
- Tzur, Y.B.; Egydio de Carvalho, C.; Nadarajan, S.; Van Bostelen, I.; Gu, Y.; Chu, D.S.; Cheeseman, I.M.; Colaiácovo, M.P. LAB-1 targets PP1 and restricts Aurora B kinase upon entrance into meiosis to promote sister chromatid cohesion. PLoS Biol. 2012, 10, e1001378. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ferrandiz, N.; Barroso, C.; Telecan, O.; Shao, N.; Kim, H.-M.; Testori, S.; Faull, P.; Cutillas, P.; Snijders, A.P.; Colaiácovo, M.P.; et al. Spatiotemporal regulation of Aurora B recruitment ensures release of cohesion during C. elegans oocyte meiosis. Nat. Commun. 2018, 9, 834. [Google Scholar] [CrossRef] [PubMed]
- Nadarajan, S.; Altendorfer, E.; Saito, T.T.; Martinez-Garcia, M.; Colaiácovo, M.P. HIM-17 regulates the position of recombination events and GSP-1/2 localization to establish short arm identity on bivalents in meiosis. Proc. Natl. Acad. Sci. USA 2021, 118, e2016363118. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).