Abstract
Clubroot, caused by Plasmodiophora brassicae, is an economically important soil-borne disease that threatens Brassicaceae crops worldwide. In recent years, the incidence area of Chinese cabbage (Brassica rapa ssp. pekinensis) clubroot disease has increased, which severely affects the yield and quality of Chinese cabbage. The resistance of varieties harboring the single clubroot-resistance (CR) gene is easily broken through by P. brassicae pathotypes. CRa and CRd, genetically identified in B. rapa, are CR genes known to be highly resistant to different P. brassicaea pathotypes. In our study, we perform the gene pyramiding of CRa and CRd in Chinese cabbages through marker-assisted selection (MAS), and develop homozygous pyramided lines. The newly generated pyramided lines exhibit greater resistance to six different pathotypes than that of two parental lines carrying a single CR gene. This study provides new CR-gene-pyramided lines for the development of clubroot-resistant Brassica varieties for future breeding programs.
1. Introduction
Chinese cabbage (B. rapa L. ssp. pekinensis), belonging to the Brassica subspecies of the Brassicaceae family, is a type of leafy vegetable with a long history of cultivation. Clubroot is one of the most destructive diseases of cruciferous crops worldwide. In recent years, the incidence of clubroot disease has become increasingly severe, and the quality and yield of Chinese cabbage have been affected by it. Breeding clubroot-resistant (CR) varieties is the most effective and environmentally friendly control method. At present, several CR varieties with high-resistance properties have been developed, but their control effect of clubroot remains inadequate. An increasing number of resistance genes are demanded to aggregate in a single CR variety to meet the diversity of P. brassicae physiological races. Therefore, the pyramiding of multiple CR genes is of great significance to improve the resistance of these products to clubroot.
Clubroot is an obligate, parasitic, soil-borne disease caused by P. brassicae, which is commonly pathogenic to cruciferous crops [1,2]. Due to the long survival time of P. brassicae in soil, soil carrying dormant spores can easily lead to the persistence of the disease, causing severe damage that is difficult to control by physical, chemical, or biological practices [3,4,5]. If the plant itself is resistant to P. brassicae, the reliance on these controls would be reduced and more cost-effective. Consequently, disease resistance is an important trait in plant breeding, and cultivating resistant varieties using CR genes is the best way to control clubroot disease in Chinese cabbage [6]. At present, several CR-gene loci have been reported in B. rapa, namely, Crr1, Crr2, Crr3, Crr4, CRa, CRb, CRc, CRd, CRk, PbBa3.1, PbBa3.2, PbBa8.1, Rcr1, Rcr4, Rcr8, and Rcr9 [4,7,8,9,10,11,12,13,14,15,16]. Among them, Crr1, PbBa8.1, and Rcr9 were located in the A8 linkage group; Crr2 in the A1 linkage group; Crr4 in the A6 linkage group; CRc and Rcr8 in the A2 linkage group; and Crr3, CRa, CRb, CRd, CRk, Rcr2, Rcr4, PbBa3.1, and PbBa3.2 in the A3 linkage group. The identification of these CR loci and their corresponding linkage markers allowed us to pyramid multiple CR genes into a variety by marker-assisted selection (MAS), which could accelerate the screening process of resistance sources and the identification of resistance genes, additionally improving the efficiency of breeding selection.
Gene pyramiding is the process of aggregating two or more favorable genes through the marker-assisted-selection breeding pathway to develop excellent varieties [17]. The pyramiding of different target genes in the same crop can not only improve crop yield and quality, but also improve the agronomic traits and enhance resistance durability [18,19,20,21,22]. In terms of crop disease resistance, because of the differentiation of the physiological race of pathogens, varieties containing a single disease-resistance gene are no longer adequate enough to resist diseases and also lose their resistance in a short period of time, whereas pyramiding multiple disease-resistance genes can improve the durability of crop disease resistance and provide a greater economic value [23,24,25]. Watson and Singh first developed the process of gene pyramiding in wheat crops for rust resistance [26]. Hittalmani et al. developed rice lines carrying two or three blight-resistance genes: Pi1, Piz5 and Pita. The disease-resistance test proved that the combination of resistance genes results in a greater resistance to leaf blight [27]. Pradhan et al. [28] conducted breeding work for rice bacterial blight resistance and eventually produced new varieties with pyramiding Xa21, xa13, and xa5 genes, which are highly resistant to bacterial blight disease. In Brassica crops, Matsumoto et al. pyramided three CR genes of CRa, CRc, and CRk by combining MAS with conventional breeding in Chinese cabbages, and the results showed that the disease resistance of pyramided lines were well-improved [24]. Similarly, the pyramiding of a major and minor gene has been reported to increase the resistance of Brassica napus and Brassica oleracea to a variety of P. brassicae pathogenic types [29,30]. In our study, the CRd and CRa genes are pyramided in Chinese cabbage varieties by MAS combined with disease-resistance identification. Following multiple generations of backcrossing and self-crossing practices, novel varieties resistant to multiple P. brassicae pathotypes are generated in this study, which is of great significance for the development of CR-breeding practices for Chinese cabbages.
2. Materials and Methods
2.1. Plant and P. brassicae Materials
Two Chinese cabbage, clubroot-resistant (CR), inbred lines, CR252 and 85-74, were used for pyramiding resistance genes. CR252 is an inbred line with good commercial traits and possesses a resistance locus, CRa, which is used as a recurrent parent. The donor parent, 85-74, contains the CRd resistance locus [13].
Six field isolates of P. brassicae, namely, LNND-2, LNXM-1, SCXC-60, YNTC-75, LAB-5, and CQPL-99, were used for the resistance evaluation of the pyramiding lines, which have been classified as pathogens Pb3, Pb4, Pb5, Pb8, Pb9, and Pb12, according to the Sinitic clubroot differential set [31], respectively. The P. brassicae isolates were maintained on the roots of a susceptible Chinese cabbage variety, 91-12, and stored in a −40℃ refrigerator belonging to the Vegetable Molecular Biology Group, Shenyang Agricultural University.
2.2. Breeding Program for Gene Pyramiding
To pyramid CRd and CRa genes, crossing and selection for resistance characteristics were conducted from 2017 to 2020, as presented in Figure 1. The two paternal lines were crossed to produce the F1 hybrid, and then continuously backcrossed with CR252 as the recurrent parent to produce BC4F1 accompanied with marker-assisted selection (MAS) in each generation. Then, the selected BC4F1 lines were self-crossed to generate BC4F2. BC4F2 generation was screened for genotype identification, and homozygous lines with resistance-gene loci were then selected for the self-crossing process. Finally, homozygous, stable, BC4F3 lines with a high recovery rate of genetic background and high resistance to clubroot were obtained.
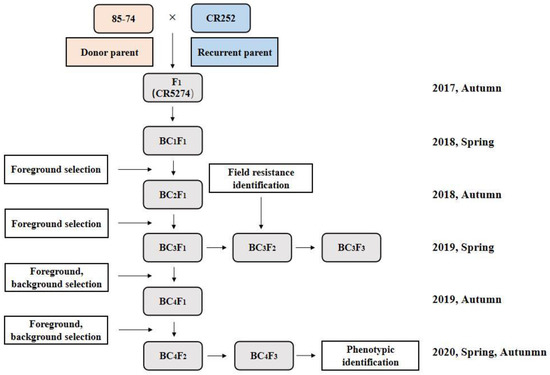
Figure 1.
Schematic diagram of marker-assisted gene pyramiding. CR252: recurrent parent; 85-74: donor parent.
2.3. Development and Validation of Polymorphic Markers between Parents
Seven pairs of flanking SSR markers closely linked to the CRd gene and three markers based on the CRa gene sequence were selected for polymorphism screening as foreground markers between the parents (Table S1). The DNA of the parents and lines of each generation was extracted by the modified CTAB (cetyltrimethylammonium bromide) method. The genomic DNA was amplified by PCR using a 10 μL reaction system containing 2 μL genomic DNA, 1 μL forward and reverse primers, 5 μL of 10× PCR buffer, and 2 μL of ddH2O. The PCR products of CRa gene-related markers were observed by using 1% agarose gel electrophoresis. A total of 6% polyacrylamide gel electrophoresis was used to detect the SSR markers designed for the CRd gene [13].
For the genetic background selection, a total of 1200 SSR primers covering 10 chromosomes of B. rapa were used to screen the polymorphisms between parents. Subsequently, the selected markers with good polymorphism properties, clear bands, and an even distribution on 10 chromosomes were used to screen the genomic background from the BC3F1 generation. The degree of backcross offspring recovery to recurrent parent was reflected by the value of the background recovery rate. The SSR marker information, PCR reaction, and amplification conditions used for background selection were referred to by Li et al. [32] and Ge et al. [33].
2.4. Investigation of the Phenotypic Characteristics
P. brassicae fluid was prepared for the clubroot disease-resistance identification of the parental lines, backcross generation, and pyramided lines. The swollen, infected roots were ground with a water solution, and then filtered through a gauze. The filtrate was collected into a 50 mL centrifuge tube, and the supernatant was discarded following repeated centrifugation. The precipitation was retained and dissolved in sterile water, and finally stored in a 4 °C refrigerator. The resting spores’ concentration was diluted to 1 × 107 spores·mL−1. Inoculation was performed by injecting 2 mL of spore solution around the roots for each plant in a 72-well seeding tray at the stage when 3–4 true leaves had developed whilst in a greenhouse. A disease-resistance survey was conducted to record the incidence rate and grade at 45 days following P. brassicae inoculation. The disease symptoms were scored as follows: level 0: roots not visible, no clubs symbol, and plant growth normal; level 1; a few small clubs on lateral roots; level 2: larger clubs on lateral roots or small clubs on main roots; and level 3: main and lateral roots present significant swelling. The disease index (DI) value was calculated according to the formula DI = [(n1 + 2n2 + 3n3)/NT × 3] × 100, where ni represents the number of plants with the symptom of i and NT is the total number of plants tested.
In the autumn of 2020 and 2021, two selected pyramided lines (BC4F3) were planted in a field in the Xinmin district of Shenyang, a clubroot-disease epidemic area, for field-resistance testing. The pathogen LNXM-1 collected in this field broke through the resistance of the CRa gene, but not the CRd locus.
The main agronomic traits of the pyramided lines and recurrent parent were investigated in the harvest period, including the outer-leaf length, outer-leaf width, outer-leaf number, outer-petiole length, outer-petiole width, petiole color, head weight, head length, head width, head shape, head solidity, head color, plant height, plant width, and plant weight (Figure 2). Twelve plants for each line were used for leaf and head related traits investigation. The pod traits were also investigated, including the pod length, pod width, number of seeds per pod, and hundred-grain weight. Moreover, the flowering time of the BC4F2 generation was measured, that is, the days from the seedling stage to the first flower’s emergence. An independent-samples t-test analysis was performed between pyramided line and recurrent parental line at p ≤ 0.05 significant level for all agronomic traits. If p value ≤ 0.05, the agronomic traits of the pyramided line would be considered to be significantly different from the recurrent parental line.
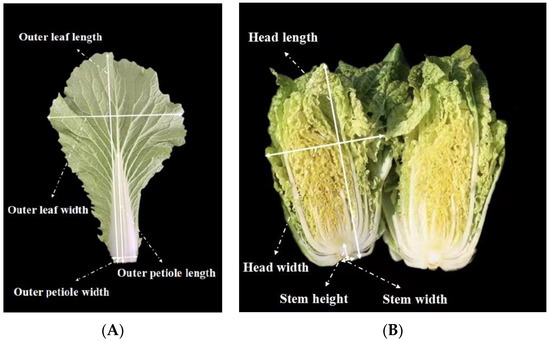
Figure 2.
(A) The measurement standard of outer-leaf length, outer-leaf width, outer-petiole length, and outer-petiole width. (B) The measurement standard of head length, head width, Stem height, and Stem width.
3. Results
3.1. Polymorphic-Marker Screening for Foreground Selection
Among the seven SSR markers designed for the CRd locus, yau106 and yau115 closely linked to CRd presented polymorphisms between CR252 and 85–74 (Figure 3). Marker CRa full-length, designed for cloning the full-length sequence of CRa, presented a polymorphism between two parental lines. CRa full-length gene sequence was only amplified in parental line CR252. These three markers were used for the foreground selection of each generation. The details of the markers are presented in Table 1.
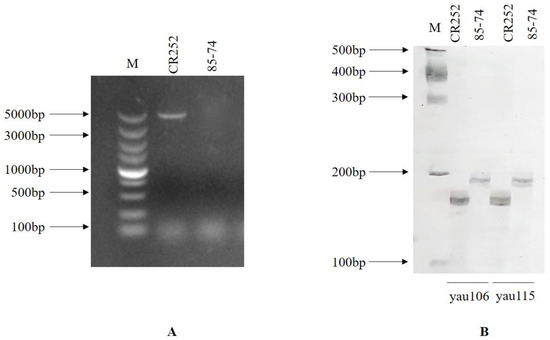
Figure 3.
Screening results for polymorphic markers for foreground selection. (A) polymorphism of CRa full-length marker between CR252 and 85–74. (B) polymorphism of CRd linked markers yau106 and yau115 between CR252 and 85–74, respectively.

Table 1.
Molecular marker details for foreground selection of two CR genes.
3.2. Polymorphic Markers’ Screening for Background Selection
A total of 1200 genomic SSR markers designed based on the B. rapa genome sequence [32,33] were used for polymorphism screening between two parental lines. Finally, 95 polymorphic markers evenly distributed on 10 chromosomes were selected for background screening from the BC3F1 generation, of which the numbers of markers on chromosomes A1 to A10 were 13, 9, 11, 7, 9, 10, 10, 9, 9, and 8. The total physical length covered was 205.7 Mb. The average interval of markers was 2.6 Mb (Table S2), and the distribution of these markers in the linkage groups is presented in Figure 4.

Figure 4.
Distribution of background-selection markers in B. rapa chromosomes.
3.3. Foreground and Background Selection for Each Generation
Foreground selection was performed for each generation of backcross hybrids from BC1F1 to BC4F1 using the CRa and CRd linked markers previously screened. From the BC3F1 generation, genomic background selection was performed on the plants that met the selection results of the foreground marker.
For the foreground selection, twenty heterozygous lines were preliminarily screened with two CRd linkage markers, yau106 and yau115, and screened further with the CRa full-length marker out of 60 BC1F1 plants. Finally, two BC1F1 robust lines with two resistance loci were selected for further backcross. The 152 BC2F1 plants were screened by the same method, and 6 lines were used for further backcross.
In the BC3F1 generation, 76 plants that carried both CRd and CRa loci were screened out of 197 lines, with background recovery rates ranging from 69.47% to 85.26% (Figure 5). Among them, 10 lines with the highest recovery rate and good growth conditions were selected for self-crossing and backcrossing to obtain BC3F2and BC4F1. According to the results of the foreground marker screening, 26 lines with both CRd and CRa loci were obtained from 116 BC4F1 lines. Twenty-nine markers that had not recovered in recurrent parental genotypes in the BC3F1 generation were selected for the background selection of these 26 lines. The background recovery rates of 26 BC4F1 lines ranged from 78.95% to 97.89%, with one exceeding the expected value by 96.88% (Figure 5). Eight BC4F1 plants were selected for self-crossing to generate a BC4F2 population. Finally, six lines of BC4F3 were obtained from 11 BC4F2 lines with homozygous CRa and CRd loci and a 100% background recovery rate.
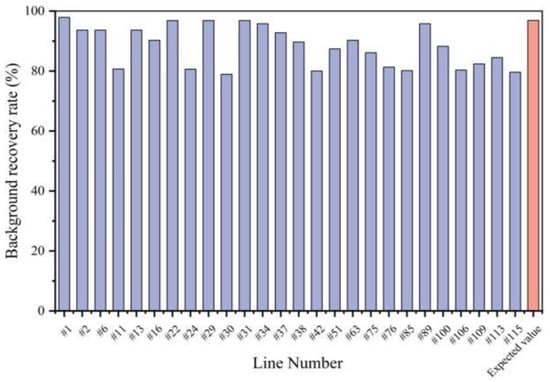
Figure 5.
Background recovery rate of 26 BC4F1 individual plants (#1~#115).
3.4. Clubroot-Resistance Identification and Morphological-Trait Evaluation of Pyramided Line
The clubroot-resistance identification of the parents and F1 generations was conducted by the inoculation of the P. brassica field isolate LNXM-1. The donor parent, ‘85-74′, was all-resistant to clubroot; the recipient parent, ‘CR252′, was all-susceptible; and 60 F1 (CR5274) plants showed all-resistance traits (Table 2).

Table 2.
Disease-resistance verification results of parental lines and F1 generation by inoculation of LNXM-1 P. brassicae isolate.
Indoor disease-resistance identification was also performed for BC1F1 and BC4F2 generations. The chi-squared tests conducted for BC1F1 and BC4F2 generations were consistent with the Mendelian genetic segregation ratios of 1:1 and 3:1, respectively (Table 3). To better verify the resistance of the backcross progeny to the LNXM-1 pathotype, the BC4F2 generation and five pyramided homologous BC4F3 lines (CR5274①-4, CR5274⑥-2, CR5274⑥-4, CR5274⑥-8 and CR5274⑩-4) were planted in the infected field in the Xinmin area of Shenyang. The separation ratio of resistant to susceptible plants of 67 BC4F2 was 3:1, according to the chi-squared test (χ2 = 0.12) (Table 4). All five gene-pyramided BC4F3 lines presented strong resistance in the infected field (Table 4).

Table 3.
Disease resistance of various generations of CR5274.

Table 4.
Disease statistics of different populations in the field.
To further validate the disease resistance of gene-pyramiding lines to different physiological races, the lines of 85-74, CR252, CR5274⑥-4, and CR5274⑥-8 were selected for inoculation. The parent CR252 carrying the CRa gene was resistant to Pb3, but susceptible to Pb4, Pb5, Pb8, and Pb11. Parental line 85–74 harboring the CRd gene was resistant to Pb4, Pb5, Pb8, and Pb11, but susceptible to Pb3. Compared to the two parental lines carrying a single disease-resistance gene, the gene-pyramided lines were resistant to multiple physiological races (Table 5).

Table 5.
Resistance identification of gene-pyramided lines inoculated with different P. brassicae races.
The agronomic traits of the parents and two pyramided lines, CR5274⑥-4 and CR5274⑥-8, were also investigated, including the growth period and 16 heading-related agronomical characters. In addition, eight lines of the BC4F2 generation and the recurrent parent CR252 were used for the evaluation of the pod characters. The results show that there are no differences in any of the heading-related agronomic traits and pod-related traits between the progenies and recurrent parent, which confirms that the backcross progenies restore the genome of the recurrent parent. The agronomical and pod characters of the progeny and recurrent parents are presented in Table 6 and Table 7.

Table 6.
Agronomic-trait investigation of recurrent parent and pyramided lines.

Table 7.
Differences in main, botanical characters of offspring and recurrent parent.
4. Discussion
The marker-assisted backcross breeding that introgresses two or more genes for targeted traits into the elite cultivars has been widely used in several crops. The study showed that plants pyramided with multiple resistance genes improved their resistance properties compared to the parents, and the agronomic traits and qualities were equivalent or better than those of the parents, such as rice [34,35,36,37], soybean [38,39], and tomato [40]. A representative example for rice is the breeding of rice varieties resistant to bacterial blight (BB) and sheath blight disease. At present, lines introgressed with several BB-resistance genes (xa4, xa5, xa7, xa13, xa21 have been successfully cultivated using MAS and exhibit greater resistance than the lines with only one or two genes [36,37,41,42]. Ramalingam et al. (2020) improved two rice cultivars through introgression of three BB-resistance, one blast resistance, and three sheath blight resistance genes/QTLs. The pyramided lines of rice exhibited a high degree of resistance to these three diseases [36]. Compared to the field crop such as rice, wheat and maize, there are fewer reports on gene pyramiding in Chinese cabbage. To date, several CR genes/QTL have been identified from diverse Brassica resistant sources, of which CRa, CRb, and PbBa8.1 are well utilized in clubroot resistance breeding programs. In Brassica crops, most of the CR cultivars of B. rapa and B. napus harboring a single CR gene can easily be broken down by the new P. brassicae pathotypes. CRb and PbBa8.1 are two resistance genes genetically identified from B. rapa, and pyramided in B. napus through MAS [29,43]. The CRb and PbBa8.1 pyramided B. napus line exhibited high levels of resistance to most P. brassicae isolates than that of a single CR gene [29]; however, the pyramided line was susceptible to an isolate collected from the Xinmin area in Liaoning province, which was identified as Pb4 by the SCD system [31]. Tomita et al. (2013) accumulated multiple CR-QTLs in B. oleracea conferred broad-spectrum clubroot resistance against six P. brassicae isolates [30]. In Chinese cabbage, Matsumoto et al. (2012) pyramided three clubroot resistant genes (CRa, CRk, and CRc) through MAS to improve the resistance. The CRd was a new locus identified from a Chinese cabbage line, 85-74, and presented great resistance to Pb4, but was susceptible to Pb3 [31], whereas the CR252 parental line was resistant to Pb3. Following the pyramiding process, the pyramided lines exhibited great resistance to multiple P. brassicae isolates. In the future, the improved pyramided lines could be tested in multiple clubroot infected areas and used as a potential donor in hybridization breeding to develop clubroot resistant cultivars in Chinese cabbage. In addition, we previously developed a pair of CRb near isogenic lines and used for transcriptome analysi to reveal the molecular mechanism of CRb gene of B. rapa against P. brassicae infection [44]. Thus, the new generated CRa and CRd gene pyramided homozygouse lines could be also utilized for multi-omics analysis and gene interaction studies.
Background selection is performed to feed back the extent to which the backcrossing progeny is recovered to the recurrent genome. Individuals with the highest ratio of recurrent parent genomes in each backcross generation were selected by marker-assisted selection as the parent of the subsequent backcross, and the recurrent parent genomes could be completely recovered within three backcross cycles, while conventional breeding required six generations [45]. In our study, a total of 95 SSR markers generated polymorphisms between the parental genotypes, ‘CR252′ and ‘85-74′, which were used to estimate the recovery rates of the recurrent parental genome in the pyramided BC3F1 genotypes. Chen et al. [46] observed that when selecting the genomes of recurrent parents, following three backcross cycles, approximately 98.8% of the genetic background information was obtained from recurrent parents. However, in this experiment, the background-marker selection in the BC3F1 generation of ‘CR5274′ did not achieve the desired recovery rate, and only plants that exceeded the expected recovery rate appeared in the BC4 generation. In general, our expectations were not met. This may have been caused by the fact that we used a small population of plants, performed late background-screening generations, or the presence of linkage drag. If the screening of background markers was performed at a later stage of breeding, the population size was necessary, and the number of populations needed to be expanded to meet the requirements for breeding. In addition, the linkage drag occurring during backcross-breeding process can lead to the introduction of other linkage genes in addition to the target gene, thus affecting the final effect [47].
In conclusion, two clubroot-resistant genes, CRd and CRa, were pyramided into Chinese cabbage samples by MAS, and the new Chinese cabbage pyramided material presenting excellent agronomic characters was obtained by the backcross-breeding method. The cultivated pyramided line recovered the phenotypic traits of the recurrent parents and strengthened clubroot resistance in B. rapa. The present study is highly significant for the development of breeding practices to develop a greater resistance to clubroot for Chinese cabbages.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/genes13122414/s1, Table S1: Molecular markers linked with CRd and CRa loci used for polymorphism screening; Table S2: The distribution of background markers on 10 linkage groups to CR5274.
Author Contributions
Conceptualization, X.L. and Y.W.; writing—original draft preparation, X.L., Y.W. and Y.M.; methodology, G.C., S.M. and T.Z.; data curation, Z.Z.; project administration, Z.P. Z.P. conceived the project, designed the research and manuscript drafting. Y.M., G.C., S.M. and T.Z. performed the marker development, backcross strategies, and field trial investigation. Z.Z. conceived the project and designed the research. All authors have read and agreed to the published version of the manuscript.
Funding
This study was supported by the LiaoNing Revitalization Talents Program (XLYC2002034) and the China Agriculture Research System of MOF and MARA (CARS-12). This study was also supported by the National Natural Science Foundation of China (No. 31772326).
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data presented in this study are available upon requestfrom the corresponding author.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Crute, I.R.; Gray, A.R.; Crisp, P.; Buczacki, S.T. Variation in Plasmodiophora brassicae and resistance to clubroot disease in brassicas and allied crops—A critical review. Plant Breed Abstr. 1980, 50, 91–104. [Google Scholar] [CrossRef]
- Dixon, G.R. The occurrence and economic impact of Plasmodiophora brassicae and clubroot disease. J. Plant Growth Regul. 2009, 28, 194–202. [Google Scholar] [CrossRef]
- Donald, C.; Porter, I. Integrated Control of Clubroot. J. Plant Growth Regul. 2009, 28, 289–303. [Google Scholar] [CrossRef]
- Suwabe, K.; Tsukazaki, H.; Iketani, H.; Hatakeyama, K.; Fujimura, M.; Nunome, T.; Fukuoka, H.; Matsumoto, S.; Hirai, M. Identification of two loci for resistance to clubroot (Plasmodiophora brassicae Woronin) in Brassica rapa L. Theor. Appl. Genet. 2003, 107, 997–1002. [Google Scholar] [CrossRef] [PubMed]
- Voorrips, R.E. Plasmodiophora brassicae: Aspects of pathogenesis and resistance in Brassica oleracea. Euphytica 1995, 83, 139–146. [Google Scholar] [CrossRef]
- Mehraj, H.; Akter, A.; Miyaji, N.; Miyazaki, J.; Shea, D.J.; Fujimoto, R.; Doullah, U. Genetics of clubroot and fusarium wilt disease resistance in Brassica Vegetables: The application of marker assisted breeding for disease resistance. Plants 2020, 9, 726. [Google Scholar] [CrossRef]
- Suwabe, K.; Suzuki, G.; Nunome, T.; Hatakeyama, K.; Mukai, Y.; Fukuoka, H.; Matsumoto, S. Microstructure of a Brassica rapa genome segment homoeologous to the resistance gene cluster on Arabidopsis chromosome 4. Breed. Sci. 2012, 62, 170–177. [Google Scholar] [CrossRef]
- Hirai, M.; Harada, T.; Kubo, N.; Tsukada, M.; Suwabe, K.; Matsumoto, S. A novel locus for CR in Brassica rapa and its linkage markers. Theor. Appl. Genet. 2004, 108, 639–643. [Google Scholar] [CrossRef]
- Suwabe, K.; Tsukazaki, H.; Iketani, H.; Hatakeyama, K.; Kondo, M.; Fujimura, M.; Nunome, T.; Fukuoka, H.; Hirai, M.; Matsumoto, S. Simple sequence repeat-based comparative genomics between Brassica rapa and Arabidopsis thaliana: The genetic origin of clubroot resistance. Genetics 2006, 173, 309–319. [Google Scholar] [CrossRef]
- Matsumoto, E.; Yasui, C.; Ohi, M.; Tsukada, M. Linkage analysis of RFLP markers for CR and pigmentation in Chinese cabbage (Brassica rapa ssp. pekinensis). Euphytica 1998, 104, 79–86. [Google Scholar] [CrossRef]
- Piao, Z.Y.; Deng, Y.Q.; Choi, S.R.; Park, Y.J.; Lim, Y.P. SCAR and CAPS mapping of CRb, a gene conferring resistance to Plasmodiophora brassicae in Chinese cabbage (Brassica rapa ssp. pekinensis). Theor. Appl. Genet. 2004, 108, 1458–1465. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Saito, A.; Hayashida, N.; Taguchi, G.; Matsumoto, E. Mapping of isolate-specific QTLs for clubroot resistance in Chinese cabbage (Brassica rapa ssp. pekinensis). Theor. Appl. Genet. 2008, 117, 759–767. [Google Scholar] [CrossRef] [PubMed]
- Pang, W.; Fu, P.; Li, X.; Zhan, Z.; Yu, S.; Piao, Z. Identification and mapping of the clubroot resistance gene CRd in Chinese cabbage (Brassica rapa ssp. pekinensis). Front. Plant Sci. 2018, 9, 653. [Google Scholar] [CrossRef]
- Chen, J.; Jing, J.; Zhan, Z.; Zhang, T.; Zhang, C.; Piao, Z. Identification of novel QTLs for isolate-specific partial resistance to Plasmodiophora brassicae in Brassica rapa. PLoS ONE 2013, 8, e85307. [Google Scholar] [CrossRef] [PubMed]
- Chu, M.; Song, T.; Falk, K.C.; Zhang, X.; Liu, X.; Chang, A.; Lahlali, R.; McGregor, L.; Gossen, B.D.; Yu, F.; et al. Fine mapping of Rcr1 and analyses of its effect on transcriptome patterns during infection by Plasmodiophora brassicae. BMC Genom. 2014, 15, 1166. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.Q.; Zhang, X.G.; Peng, G.; Falk, K.C.; Strelkov, S.E.; Gossen, B.D. Genotyping-by-sequencing reveals three QTL for clubroot resistance to six pathotypes of Plasmodiophora brassicae in Brassica rapa. Sci. Rep. 2017, 7, 4516. [Google Scholar] [CrossRef] [PubMed]
- Chukwu, S.C.; Rafii, M.Y.; Ramlee, S.I.; Ismail, S.I.; Oladosu, Y.O.; Okporie, E.O.; Onyishi, G.; Utobo, E.; Ekwu, L.G.; Swaray, S.; et al. Marker-assisted selection and gene pyramiding for resistance to bacterial leaf blight disease of rice (Oryza sativa L.). Biotechnol. Biotechnol. Equip. 2019, 33, 440–455. [Google Scholar] [CrossRef]
- Bharani, M.; Nagarajan, P.; Rabindran, R.; Saraswathi, R.; Balasubramanian, P.; Ramalingam, J. Bacterial leaf blight resistance genes (Xa21, xa13 and xa5) py-ramiding through molecular marker-assisted selection into rice cultivars. Arch. Phytopathol. Plant Prot. 2010, 43, 1032–1043. [Google Scholar] [CrossRef]
- Fu, C.; Wu, T.; Liu, W.; Wang, F.; Li, J.; Zhu, X.; Huang, H.; Liu, Z.; Liao, Y.; Zhu, M.; et al. Genetic improvement of resistance to blast and bacterial blight of the elite maintainer line Rongfeng B in hybrid rice (Oryza sativa L.) by using marker-assisted selection. Afr. J. Biotechnol. 2012, 11, 13104–13124. [Google Scholar] [CrossRef]
- Kloppers, F.J.; Pretorius, Z.A. Effects of combinations amongst genes Lr13, Lr34 and Lr37 on components of resistance in wheat to leaf rust. Plant Pathol. 1997, 46, 737–750. [Google Scholar] [CrossRef]
- Shanti, M.L.; George, M.L.; Cruz, C.M.; Bernardo, M.A.; Nelson, R.J.; Leung, H.; Reddy, J.N.; Sridhar, R. Identification of resistance genes effective against rice bacterial blight pathogen in eastern India. Plant Dis. 2001, 85, 506–512. [Google Scholar] [CrossRef][Green Version]
- Singh, V.K.; Singh, A.P.; Singh, S.P.; Ellur, R.K.; Choudhary, V.; Sarkel, S.; Singh, D.; Krishnan, S.G.; Nagarajan, M.; Vinod, K.K.; et al. Incorporation of blast resistance into “PRR78”, an elite Basmati rice restorer line, through marker-assisted backcross breeding. Field Crop. Res. 2012, 128, 8–16. [Google Scholar] [CrossRef]
- Prasanna, H.C.; Sinha, D.P.; Rai, G.K.; Krishna, R.; Kashyap, S.P.; Singh, N.K.; Singh, M.; Malathi, V.G. Pyramiding Ty-2 and Ty-3 genes for resistance to monopartite and bipartite tomato leaf curl viruses of India. Plant Pathol. 2014, 64, 256–264. [Google Scholar] [CrossRef]
- Matsumoto, E.; Ueno, H.; Aruga, D.; Sakamoto, K.; Hayashida, N. Accumulation of three CR genes through marker-assisted selection in Chinese cabbage (Brassica rapa ssp. pekinensis). J. Jpn. Soc. Hortic. Sci. 2012, 81, 184–190. [Google Scholar] [CrossRef]
- Wallenhammar, A.; Arwidsson, O. Detection of plasmodiophora brassicae by PCR in naturally infested soils. Eur. J. Plant Pathol. 2001, 107, 313–321. [Google Scholar] [CrossRef]
- Watson, I.A.; Singh, D. The future for rust resistant wheat in Australia. J. Aust. Inst. Agric. Sci. 1952, 18, 190–197. [Google Scholar]
- Hittalmani, S.; Parco, A.; Mew, T.V.; Zeigler, R.S.; Huang, N. Fine mapping and DNA marker-assisted pyramiding of the three major genes for blast resistance in rice. Theor. Appl. Genet. 2000, 100, 1121–1128. [Google Scholar] [CrossRef]
- Pradhan, S.K.; Nayak, D.K.; Mohanty, S.; Behera, L.; Barik, S.R.; Pandit, E.; Lenka, S.; Anandan, A. Pyramiding of three bacterial blight resistance genes for broad-spectrum resistance in deepwater rice variety, Jalmagna. Rice 2015, 8, 51. [Google Scholar] [CrossRef] [PubMed]
- Shah, N.; Sun, J.; Yu, S.; Yang, Z.; Wang, Z.; Huang, F.; Dun, B.C.; Gong, J.; Liu, Y.; Li, Y.; et al. Genetic variation analysis of field isolates of clubroot and their responses to Brassica napus lines containing resistant genes CRb and PbBa8.1 and their combination in homozygous and heterozygous state. Mol. Breed. 2019, 39, 53. [Google Scholar] [CrossRef]
- Tomita, H.; Shimizu, M.; Asad-Ud Doullah, M.; Fujimoto, R.; Okazaki, K. Accumulation of quantitative trait loci conferring broad-spectrum clubroot resistance in Brassica oleracea. Mol. Breed. 2013, 32, 889–900. [Google Scholar] [CrossRef]
- Pang, W.; Liang, Y.; Zhan, Z.; Li, X.; Piao, Z. Development of a Sinitic clubroot differential set for the pathotype classification of Plasmodiophora brassicae. Front. Plant Sci. 2020, 11, 568771. [Google Scholar] [CrossRef]
- Li, X.; Ramchiary, N.; Choi, S.R.; Van Nguyen, D.; Hossain, M.J.; Yang, H.; Lim, Y.P. Development of a high density integrated reference genetic linkage map for the multinational Brassica rapa Genome Sequencing Project. Genome 2010, 53, 939–947. [Google Scholar] [CrossRef]
- Ge, Y.; Ramchiary, N.; Wang, T.; Liang, C.; Wang, N.; Wang, Z.; Choi, S.R.; Lim, Y.P.; Piao, Z.Y. Mapping quantitative trait loci for leaf and heading-related traits in Chinese cabbage (Brassica rapa L. ssp. pekinensis). Hortic. Environ. Biotechnol. 2011, 52, 494–501. [Google Scholar] [CrossRef]
- Singh, A.K.; Singh, V.K.; Singh, S.P.; Pandian, R.T.P.; Ellur, R.K.; Singh, D.; Prolay, K.; Bhowmick, S.; Gopala Krishnan, M.; Nagarajan, K.K.; et al. Molecular breeding for the development of multiple disease resistance in Basmati rice. AoB Plants 2012, 2012, pls029. [Google Scholar] [CrossRef]
- Das, G.; Rao, G.J.N. Molecular marker assisted gene stacking for biotic and abiotic stress resistance genes in an elite rice cultivar. Front. Plant Sci. 2015, 6, 698. [Google Scholar] [CrossRef]
- Ramalingam, J.; Raveendra, C.; Savitha, P.; Vidya, V.; Chaithra, T.L.; Velprabakaran, S.; Saraswathi, R.; Ramanathan, A.; Pillai, M.P.A.; Arumugachamy, S.; et al. Gene pyramiding for achieving enhanced resistance to bacterial blight, blast, and sheath blight diseases in rice. Front. Plant Sci. 2020, 11, 591457. [Google Scholar] [CrossRef]
- Hsu, Y.C.; Chiu, C.H.; Yap, R.; Tseng, Y.C.; Wu, Y.P. Pyramiding bacterial blight resistance genes in Tainung82 for broad spectrum resistance using marker-assisted selection. Int. J. Mol. Sci. 2020, 21, 1281. [Google Scholar] [CrossRef]
- Yamanaka, N.; Hossain, M. Pyramiding three rust-resistance genes confers a high level of resistance in soybean (Glycine max). Plant Breed. 2019, 138, 686–695. [Google Scholar] [CrossRef]
- Shi, A.; Chen, P.; Zheng, C.; Hou, A.; Zhu, S. Gene Pyramiding for Soybean Mosaic Virus resistance using microsatellite markers. Int. Meet. ASA-CSSA-SSSA 2006, 13. [Google Scholar] [CrossRef]
- Hanson, P.; Lu, S.F.; Wang, J.F.; Chen, W.; Kenyon, L.; Tan, C.W.; Tee, K.L.; Wang, Y.Y.; Hsu, Y.C.; Schafleitner, R.; et al. Conventional and molecular marker-assisted selection and pyramiding of genes for multiple disease resistance in tomato. Sci. Hortic. 2016, 201, 346–354. [Google Scholar] [CrossRef]
- Arunakumari, K.; Durgarani, C.V.; Satturu, V.; Sarikonda, K.R.; Chittoor, P.D.R.; Vutukuri, B.; Laha, G.S.; Nelli, A.P.K.; Gattu, S.; Jamal, M.; et al. Marker-assisted pyramiding of genes conferring resistance against bacterial blight and blast diseases into Indian rice variety MTU1010. Rice Sci. 2016, 23, 306–316. [Google Scholar] [CrossRef]
- Jamaloddin, M.; Durga Rani, C.V.; Swathi, G.; Anuradha, C.; Vanisri, S.; Rajan, C.P.C.; Krishnam Raju, S.; Bhuvaneshwari, V.; Jagadeeswar, R.; Laha, G.S.; et al. Marker assisted gene pyramiding (MAGP) for bacterial blight and blast resistance into mega rice variety “Tellahamsa”. PLoS ONE 2020, 15, e0234088. [Google Scholar] [CrossRef] [PubMed]
- Shah, N.; Li, Q.; Xu, Q.; Liu, J.; Huang, F.; Zhan, Z.; Qin, P.; Zhou, X.; Yu, W.; Zhu, L.; et al. CRb and PbBa8.1 synergically increases resistant genes expression upon infection of Plamodiophora brassicae in Brassica napus. Genes 2020, 11, 202. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Pang, W.; Chen, B.; Zhang, C.; Piao, Z. Transcriptome analysis of Brassica rapa near-isogenic lines carrying clubroot-resistant and -susceptible alleles in response to Plasmodiophora brassicae during early infection. Front. Plant Sci. 2016, 6, 1183. [Google Scholar] [CrossRef]
- Cobb, J.N.; Biswas, P.S.; Platten, J.D. Back to the future: Revisiting MAS as a tool for modern plant breeding. Theor. Appl. Genet. 2019, 132, 647–667. [Google Scholar] [CrossRef]
- Chen, S.; Xu, C.; Lin, X.; Zhang, Q. Improving bacterial blight resistance of ’6078’, an elite restorer line of hybrid rice, by molecular marker-assisted selection. Plant Breed. 2001, 120, 133–137. [Google Scholar] [CrossRef]
- Hasan, N.; Choudhary, S.; Naaz, N.; Sharma, N.; Laskar, R.A. Recent advancements in molecular marker-assisted selection and applications in plant breeding programmes. J. Genet. Eng. Biotechnol. 2021, 19, 128. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).