Prediction of Parkinson’s Disease Risk Based on Genetic Profile and Established Risk Factors
Abstract
:1. Introduction
2. Materials and Methods
2.1. Dataset
2.2. PRS Calculation
2.3. Selected Demographic Data
2.4. Imputation
2.5. Statistical Analysis
3. Results
3.1. Study Participants
3.2. Parkinson’s Disease Risk
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ibanez, L.; Dube, U.; Saef, B.; Budde, J.; Black, K.; Medvedeva, A.; Del-Aguila, J.L.; Davis, A.A.; Perlmutter, J.S.; Harari, O.; et al. Parkinson disease polygenic risk score is associated with Parkinson disease status and age at onset but not with α-synuclein cerebrospinal fluid levels. BMC Neurol. 2017, 17, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Moore, D.J.; West, A.; Dawson, V.; Dawson, T.M. Molecular Pathophysiology of Parkinson’s Disease. Annu. Rev. Neurosci. 2005, 28, 57–87. [Google Scholar] [CrossRef]
- Georgiou, A.; Demetriou, C.A.; Christou, Y.P.; Heraclides, A.; Leonidou, E.; Loukaides, P.; Yiasoumi, E.; Pantziaris, M.; Kleopa, K.A.; Papacostas, S.S.; et al. Genetic and Environmental Factors Contributing to Parkinson’s Disease: A Case-Control Study in the Cypriot Population. Front. Neurol. 2019, 10, 1047. [Google Scholar] [CrossRef] [PubMed]
- Jacobs, B.M.; Belete, D.; Bestwick, J.; Blauwendraat, C.; Bandres-Ciga, S.; Heilbron, K.; Dobson, R.; Nalls, M.A.; Singleton, A.; Hardy, J.; et al. Parkinson’s Disease Determinants, Prediction and Gene–Environment Interactions in the UK Biobank. J. Neurol. Neurosurg. Psychiatry 2020, 91, 1046–1054. [Google Scholar] [CrossRef] [PubMed]
- Grenn, F.P.; Kim, J.J.; Makarious, M.B.; Iwaki, H.; Illarionova, A.; Brolin, K.; Kluss, J.H.; Schumacher-Schuh, A.F.; Leonard, H.; Faghri, F.; et al. The Parkinson’s Disease Genome-Wide Association Study Locus Browser. Mov. Disord. 2020, 35, 2056–2067. [Google Scholar] [CrossRef]
- Erro, R.; Brigo, F.; Tamburin, S.; Zamboni, M.; Antonini, A.; Tinazzi, M. Nutritional Habits, Risk, and Progression of Parkinson Disease. J. Neurol. 2018, 265, 12–23. [Google Scholar] [CrossRef]
- Bellou, V.; Belbasis, L.; Tzoulaki, I.; Evangelou, E.; Ioannidis, J.P. Environmental risk factors and Parkinson’s disease: An umbrella review of meta-analyses. Park. Relat. Disord. 2015, 23, 1–9. [Google Scholar] [CrossRef]
- Tufail, M. Clinical Features and Risk Factors of Parkinson’s Disease in a Population of Khyber Pakhtunkhwa, Pakistan: A Case-Control Study. Neurodegener. Dis. 2019, 19, 211–217. [Google Scholar] [CrossRef]
- Escott-Price, V.; Nalls, M.A.; Morris, H.R.; Lubbe, S.; Brice, A.; Gasser, T.; Heutink, P.; Wood, N.W.; Hardy, J.; Singleton, A.B.; et al. Polygenic Risk of Parkinson Disease is Correlated with Disease Age at Onset. Ann. Neurol. 2015, 77, 582–591. [Google Scholar] [CrossRef]
- Escott-Price, V.; Shoai, M.; Pither, R.; Williams, J.; Hardy, J. Polygenic score prediction captures nearly all common genetic risk for Alzheimer’s disease. Neurobiol. Aging 2016, 49, 214.e7–214.e11. [Google Scholar] [CrossRef]
- Kara, E.; Xiromerisiou, G.; Spanaki, C.; Bozi, M.; Koutsis, G.; Panas, M.; Dardiotis, E.; Ralli, S.; Bras, J.; Letson, C.; et al. Assessment of Parkinson’s disease risk loci in Greece. Neurobiol. Aging 2013, 35, 442.e9–442.e16. [Google Scholar] [CrossRef]
- Do, C.B.; Tung, J.Y.; Dorfman, E.; Kiefer, A.K.; Drabant, E.M.; Francke, U.; Mountain, J.L.; Goldman, S.; Tanner, C.M.; Langston, J.W.; et al. Web-Based Genome-Wide Association Study Identifies Two Novel Loci and a Substantial Genetic Component for Parkinson’s Disease. PLoS Genet. 2011, 7, e1002141. [Google Scholar] [CrossRef] [PubMed]
- The UK Parkinson’s Disease Consortium and The Wellcome Trust Case Control Consortium 2; Spencer, C.C.; Plagnol, V.; Strange, A.; Gardner, M.; Paisan-Ruiz, C.; Band, G.; Barker, R.A.; Bellenguez, C.; Bhatia, K.; et al. Dissection of the genetics of Parkinson’s disease identifies an additional association 5′ of SNCA and multiple associated haplotypes at 17q21. Hum. Mol. Genet. 2010, 20, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, J.S.; Schulte, C.; Bras, J.; Sharma, M.; Gibbs, J.R.; Berg, D.; Paisan-Ruiz, C.; Lichtner, P.; Scholz, S.; Hernandez, D.G.; et al. Genome-wide association study reveals genetic risk underlying Parkinson’s disease. Nat. Genet. 2009, 41, 1308–1312. [Google Scholar] [CrossRef] [PubMed]
- Edwards, T.L.; Scott, W.K.; Almonte, C.; Burt, A.; Powell, E.H.; Beecham, G.W.; Wang, L.; Züchner, S.; Konidari, I.; Wang, G.; et al. Genome-Wide Association Study Confirms SNPs in SNCA and the MAPT Region as Common Risk Factors for Parkinson Disease. Ann. Hum. Genet. 2010, 74, 97–109. [Google Scholar] [CrossRef]
- Nalls, M.A.; Pankratz, N.; Lill, C.M.; Do, C.B.; Hernandez, D.G.; Saad, M.; DeStefano, A.L.; Kara, E.; Bras, J.; Sharma, M.; et al. Large-Scale Meta-Analysis of Genome-Wide Association Data Identifies Six New Risk Loci for Parkinson’s Disease. Nat. Genet. 2014, 46, 989–993. [Google Scholar] [CrossRef] [PubMed]
- Mavaddat, N.; Michailidou, K.; Dennis, J.; Lush, M.; Fachal, L.; Lee, A.; Tyrer, J.P.; Chen, T.-H.; Wang, Q.; Bolla, M.K.; et al. Polygenic Risk Scores for Prediction of Breast Cancer and Breast Cancer Subtypes. Am. J. Hum. Genet. 2019, 104, 21–34. [Google Scholar] [CrossRef]
- Endelman, J.B. Ridge Regression and Other Kernels for Genomic Selection with R Package RrBLUP. Plant Genome 2011, 4, 250–255. [Google Scholar] [CrossRef]
- Van Buuren, S.; Groothuis-Oudshoorn, K. Mice: Multivariate Imputation by Chained Equations in R. J. Stat. Softw. 2011, 45, 1–67. [Google Scholar] [CrossRef]
- Pang, S.Y.-Y.; Ho, P.W.-L.; Liu, H.-F.; Leung, C.-T.; Li, L.; Chang, E.E.S.; Ramsden, D.B.; Ho, S.-L. The Interplay of Aging, Genetics and Environmental Factors in the Pathogenesis of Parkinson’s Disease. Transl. Neurodegener. 2019, 8, 23. [Google Scholar] [CrossRef]
- Jafari, S.; Etminan, M.; Aminzadeh, F.; Samii, A. Head injury and risk of Parkinson disease: A systematic review and meta-analysis. Mov. Disord. 2013, 28, 1222–1229. [Google Scholar] [CrossRef]
- Noyce, A.; Msc, J.P.B.; Silveira-Moriyama, L.; Hawkes, C.H.; Giovannoni, G.; Lees, A.J.; Schrag, A. Meta-analysis of early nonmotor features and risk factors for Parkinson disease. Ann. Neurol. 2012, 72, 893–901. [Google Scholar] [CrossRef]
- Chen, J.; Guan, Z.; Wang, L.; Song, G.; Ma, B.; Wang, Y. Meta-Analysis: Overweight, Obesity, and Parkinson’s Disease. Int. J. Endocrinol. 2014, 2014, 203930. [Google Scholar]
- Noyce, A.; Kia, D.A.; Hemani, G.; Nicolas, A.; Price, T.R.; De Pablo-Fernández, E.; Haycock, P.C.; Lewis, P.A.; Foltynie, T.; Smith, G.D.; et al. Estimating the causal influence of body mass index on risk of Parkinson disease: A Mendelian randomisation study. PLoS Med. 2017, 14, e1002314. [Google Scholar] [CrossRef]
- Butcher, N.J.; Merico, D.; Zarrei, M.; Ogura, L.; Marshall, C.R.; Chow, E.W.C.; Lang, A.; Scherer, S.W.; Bassett, A.S. Whole-genome sequencing suggests mechanisms for 22q11.2 deletion-associated Parkinson’s disease. PLoS ONE 2017, 12, e0173944. [Google Scholar] [CrossRef]
- Nalls, M.A.; Blauwendraat, C.; Vallerga, C.L.; Heilbron, K.; Bandres-Ciga, S.; Chang, D.; Tan, M.; Kia, D.A.; Noyce, A.J.; Xue, A.; et al. Expanding Parkinson’s Disease Genetics: Novel Risk Loci, Genomic Context, Causal Insights and Heritable Risk. BioRxiv 2019, 388165. [Google Scholar] [CrossRef]
- Paul, K.; Schulz, J.; Bronstein, J.M.; Lill, C.M.; Ritz, B.R. Association of Polygenic Risk Score With Cognitive Decline and Motor Progression in Parkinson Disease. JAMA Neurol. 2018, 75, 360–366. [Google Scholar] [CrossRef]
- Iwaki, H.; Blauwendraat, C.; Makarious, M.B.; Bandrés-Ciga, S.; Leonard, H.L.; Gibbs, J.R.; Hernandez, D.G.; Scholz, S.W.; Faghri, F.; Nalls, M.A.; et al. Penetrance of Parkinson’s Disease in LRRK2 p.G2019S Carriers is Modified by a Polygenic Risk Score. Mov. Disord. 2020, 35, 774–780. [Google Scholar] [CrossRef]
- Han, Y.; Teeple, E.; Shankara, S.; Sadeghi, M.; Zhu, C.; Liu, D.; Wang, C.; Frau, F.; Klinger, K.W.; Madden, S.L.; et al. Genome-Wide Polygenic Risk Score Identifies Individuals at Elevated Parkinson’s Disease Risk. MedRxiv 2020, 96. [Google Scholar] [CrossRef]

| Georgiou et al. [3] | Previous Studies | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| # | SNP | Nearest Gene | Alleles | Minor Allele | MAF | OR (95% CI) * | p-Value | European MAF * | Minor Allele | OR (95% CI) ** | p-Value | References |
| 1 | rs12185268 & | SPPL2C | A/G | G | 0.26 | 0.69 (0.52–0.90) | 0.006 | 0.22 | G | 0.77 (0.72–0.82) | 2.72 × 10−14 | [12] |
| 2 | rs10513789 | MCCC1 | T/G | G | 0.18 | 1.09 (0.82–1.45) | 0.57 | 0.20 | G | 0.80 (0.75–0.86) | 2.67 × 10−10 | [12] |
| 3 | rs6599389 & | TMEM175 | G/A | A | 0.09 | 1.50 (1.04–2.16) | 0.03 | 0.07 | A | 1.31 (1.19–1.44) | 3.87 × 10−8 | [12] |
| 4 | rs356220 & | SNCA | C/T | T | 0.37 | 1.33 (1.05–1.67) | 0.02 | 0.37 | T | 1.29 (1.22–1.36) | 2.29 × 10−19 | [12] |
| 5 | rs7617877 & | LINC00693 | A/G | A | 0.26 | 1.03 (0.80–1.34) | 0.8 | 0.35 | A | 1.23 (1.13–1.33) | 4.49 × 10−7 | [12,13] |
| 6 | rs17115100 | CYP17A1 | G/T | T | 0.09 | 1.06 (0.74–1.53) | 0.75 | 0.1 | T | 0.80 (NR) | 7.44 × 10−8 | [12,14] |
| 7 | rs13312 | USP24 | C/G | G | 0.14 | 1.68 (1.23–2.28) | 0.001 | 0.24 | G | 0.76 (0.66–0.86) | Not reported | [12] |
| 8 | rs1801582 | PARK2 | C/G | G | 0.16 | 1.08 (0.80–1.46) | 0.63 | 0.16 | G | 0.79 (0.64–0.97) | Not reported | [12] |
| 9 | rs4837628 & | BRINP1 | C/T | C | 0.34 | 0.89 (0.69–1.14) | 0.36 | 0.42 | C | 0.79 (0.72–0.87) | 1.07 × 10−6 | [12,15] |
| 10 | rs823118 & | NUCKS1 | C/T | C | 0.37 | 0.79 (0.62–1.01) | 0.056 | 0.44 | T *** | 1.12 (NR) | 1.66 × 10−16 | [16] |
| 11 | rs356182 & | SNCA | A/G | G | 0.33 | 1.24 (0.98–1.57) | 0.076 | 0.36 | A *** | 0.76 (NR) | 4.16 × 10−73 | [16] |
| 12 | rs17649553 & | MAPT | C/T | T | 0.26 | 0.71 (0.54–0.93) | 0.013 | 0.21 | T *** | 0.77 (0.74–0.80) | 2.37 × 10−48 | [16] |
| Controls | Cases | |||
|---|---|---|---|---|
| Age (Years, Mean ± SD) | 65 ± 10.7 | 66.5 ± 10.5 | ||
| Age of Onset (Mean ± SD) | 60.4 ± 11.4 | |||
| BMI | 28.28 ± 5.01 | 26.29 ± 4.15 | ||
| Variable | Total n (%) | Controls n (%) | Cases n (%) | |
| Gender | Male | 359 (51.36) | 231 (49.78) | 128 (54.47) |
| Female | 340 (48.64) | 233 (50.22) | 107 (45.53) | |
| NA | 0 (0) | 0 (0) | 0 (0) | |
| Outdoor Work | No | 512 (73.25) | 348 (75) | 164 (69.79) |
| Yes | 168 (24.03) | 103 (22.2) | 65 (27.66) | |
| NA | 19 (2.72) | 13 (2.80) | 6 (2.55) | |
| Pesticides or Toxic Substances | No | 384 (54.94) | 261 (56.25) | 123 (52.34) |
| Yes | 315 (45.06) | 203 (43.75) | 112 (47.66) | |
| NA | 0 (0) | 0 (0) | 0 (0) | |
| Pesticides | No | 462 (66.09) | 309 (66.59) | 153 (65.11) |
| Yes | 237 (33.91) | 155 (33.41) | 82 (34.89) | |
| NA | 0 (0) | 0 (0) | 0 (0) | |
| Well Water Drinking | No | 296 (42.35) | 201 (43.32) | 95 (40.43) |
| Yes | 389 (55.65) | 258 (55.6) | 131 (55.74) | |
| NA | 14 (2.00) | 5 (1.08) | 9 (3.83) | |
| Head Injury | No | 488 (69.81) | 344 (74.14) | 144 (61.28) |
| Yes | 199 (28.47) | 117 (25.22) | 82 (34.89) | |
| NA | 12 (1.72) | 3 (0.65) | 9 (3.83) | |
| Family history of PD | No | 582 (83.26) | 425 (91.59) | 157 (66.81) |
| Yes | 112 (16.02) | 38 (8.19) | 74 (31.49) | |
| NA | 5 (0.72) | 1 (0.22) | 4 (1.70) | |
| Hypertension | No | 404 (57.8) | 268 (57.76) | 136 (57.87) |
| Yes | 284 (40.63) | 191 (41.16) | 93 (39.57) | |
| NA | 11 (1.57) | 5 (1.08) | 6 (2.55) | |
| Statin Use | No | 463 (66.24) | 313 (67.46) | 150 (63.83) |
| Yes | 222 (31.76) | 143 (30.82) | 79 (33.62) | |
| NA | 14 (2.00) | 8 (1.72) | 6 (2.55) | |
| Depression | No | 504 (72.1) | 393 (84.7) | 111 (47.23) |
| Yes | 140 (20.03) | 45 (9.7) | 95 (40.43) | |
| NA | 55 (7.87) | 26 (5.60) | 29 (12.34) | |
| Smoking (Current or ever) | No | 350 (50.07) | 222 (47.84) | 128 (54.47) |
| Yes | 332 (47.5) | 233 (50.22) | 99 (42.13) | |
| NA | 17 (2.43) | 9 (1.95) | 8 (3.40) | |
| Physical Activity | No | 481 (68.81) | 331 (71.34) | 150 (63.83) |
| Yes | 180 (25.75) | 128 (27.59) | 52 (22.13) | |
| NA | 38 (5.44) | 5 (1.08) | 33 (14.04) | |
| Coffee Consumption | No | 32 (4.58) | 19 (4.09) | 13 (5.53) |
| Yes | 654 (93.56) | 439 (94.61) | 215 (91.49) | |
| NA | 13 (1.86) | 6 (1.29) | 7 (2.98) |
| p-Value | OR (95% CI) | AUC | |
|---|---|---|---|
| PRS | 1.87 × 10−2 ** | 1.39 (1.06–1.84) | 0.55 |
| Age | 1.25 × 10−5 ** | 1.04 (1.02–1.05) | 0.62 |
| Gender (female) | 2.42 × 10−1 | 0.83 (0.6–1.13) | 0.52 |
| Outdoor work | 1.14 × 10−1 | 1.34 (0.93–1.92) | 0.53 |
| Pesticides or toxic substances | 3.27 × 10−1 | 1.17 (0.85–1.6) | 0.52 |
| Pesticides | 6.95 × 10−1 | 1.07 (0.77–1.48) | 0.51 |
| Well water drinking | 6.63 × 10−1 | 1.07 (0.78–1.48) | 0.51 |
| Head injury | 3.21 × 10−3 ** | 1.67 (1.19–2.36) | 0.55 |
| Family history of PD | 4.55 × 10−14 ** | 5.27 (3.44–8.19) | 0.62 |
| Hypertension | 8.02 × 10−1 | 0.96 (0.69–1.32) | 0.51 |
| Statin use | 4.08 × 10−1 | 1.15 (0.82–1.61) | 0.52 |
| Depression | <2.00 × 10−16 ** | 7.47 (4.98–11.37) | 0.68 |
| Smoking (current or ever) | 6.18 × 10−2 | 0.74 (0.53–1.01) | 0.54 |
| Physical activity | 5.68 × 10−1 | 0.9 (0.61–1.3) | 0.51 |
| BMI | 4.03 × 10−6 ** | 0.91 (0.87–0.94) | 0.62 |
| Coffee consumption | 3.65 × 10−1 | 0.72 (0.35–1.51) | 0.51 |
| PRS Score Quartiles | Controls n (%) | Cases n (%) | p-Value | OR (95% CI) |
| 0–25 | 110 (24.2) | 42 (18.3) | 2.21 × 10−1 | 0.74 (0.46–1.19) |
| * 25–50 | 117 (25.7) | 60 (26.2) | ||
| 50–75 | 112 (24.6) | 59 (25.8) | 9.05 × 10−1 | 1.03 (0.66–1.60) |
| 75–100 | 116 (25.5) | 68 (29.7) | 5.44 × 10−1 | 1.14 (0.74–1.76) |
| <NA> | 9 (1.9) | 6 (2.6) |
| BMI Categories | Controls n (%) | Cases n (%) | p-Value | OR (95% CI) |
| Underweight (<20) | 8 (1.7) | 14 (6) | 2.07 × 10−2 ** | 2.99 (1.21–7.88) |
| * Normal (20–24.94) | 99 (21.3) | 58 (24.7) | ||
| Obesity (24.95–29.94) | 133 (28.7) | 33 (14) | 7.64 × 10−4 ** | 0.42 (0.25–0.69) |
| Overweight (>29.95) | 174 (37.5) | 83 (35.3) | 3.33 × 10−1 | 0.81 (0.54–1.24) |
| <NA> | 50 (10.8) | 47 (20) |
| p-Value | OR (95% CI) | |
|---|---|---|
| (Intercept) | 3.49 × 10−1 | 0.36 (0.04–3.06) |
| PRS score | 6.19 × 10−2 | 1.45 (0.99–2.15) |
| Current age | 1.62 × 10−3 | 1.04 (1.01–1.06) |
| Gender | 1.41 × 10−2 | 0.51 (0.3–0.87) |
| Head injury | 3.86 × 10−3 | 1.99 (1.25–3.18) |
| Family history of PD | 1.44 × 10−5 | 3.48 (1.99–6.14) |
| Depression | 1.13 × 10−13 | 7.01 (4.23–11.86) |
| Smoking (current or ever) | 1.33 × 10−1 | 0.67 (0.4–1.13) |
| BMI | 1.16 × 10−3 | 0.92 (0.87–0.97) |
| AUC (95% CI) | 0.79 (0.75–0.83) | |
| GOF | 0.09 | |
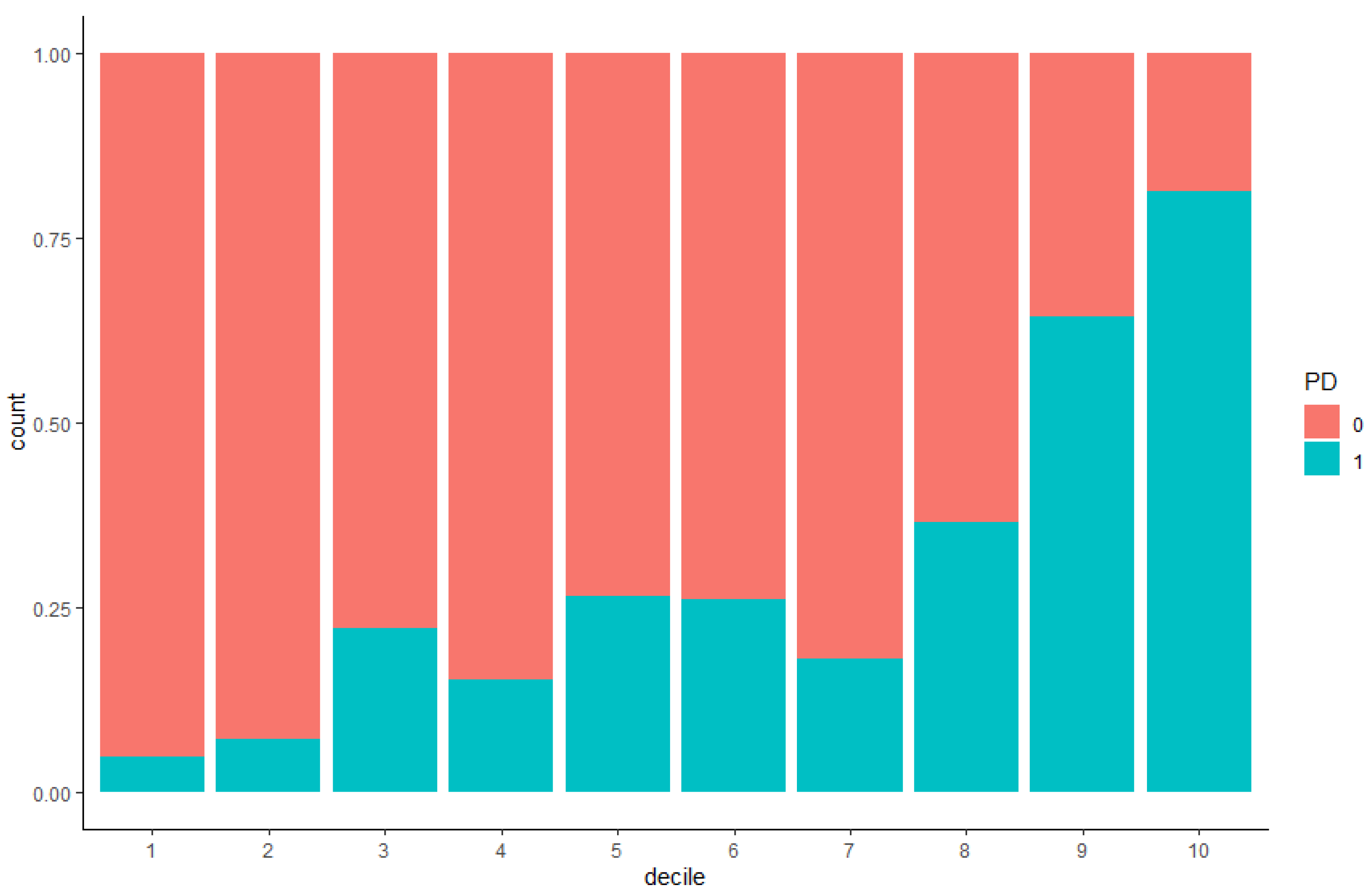
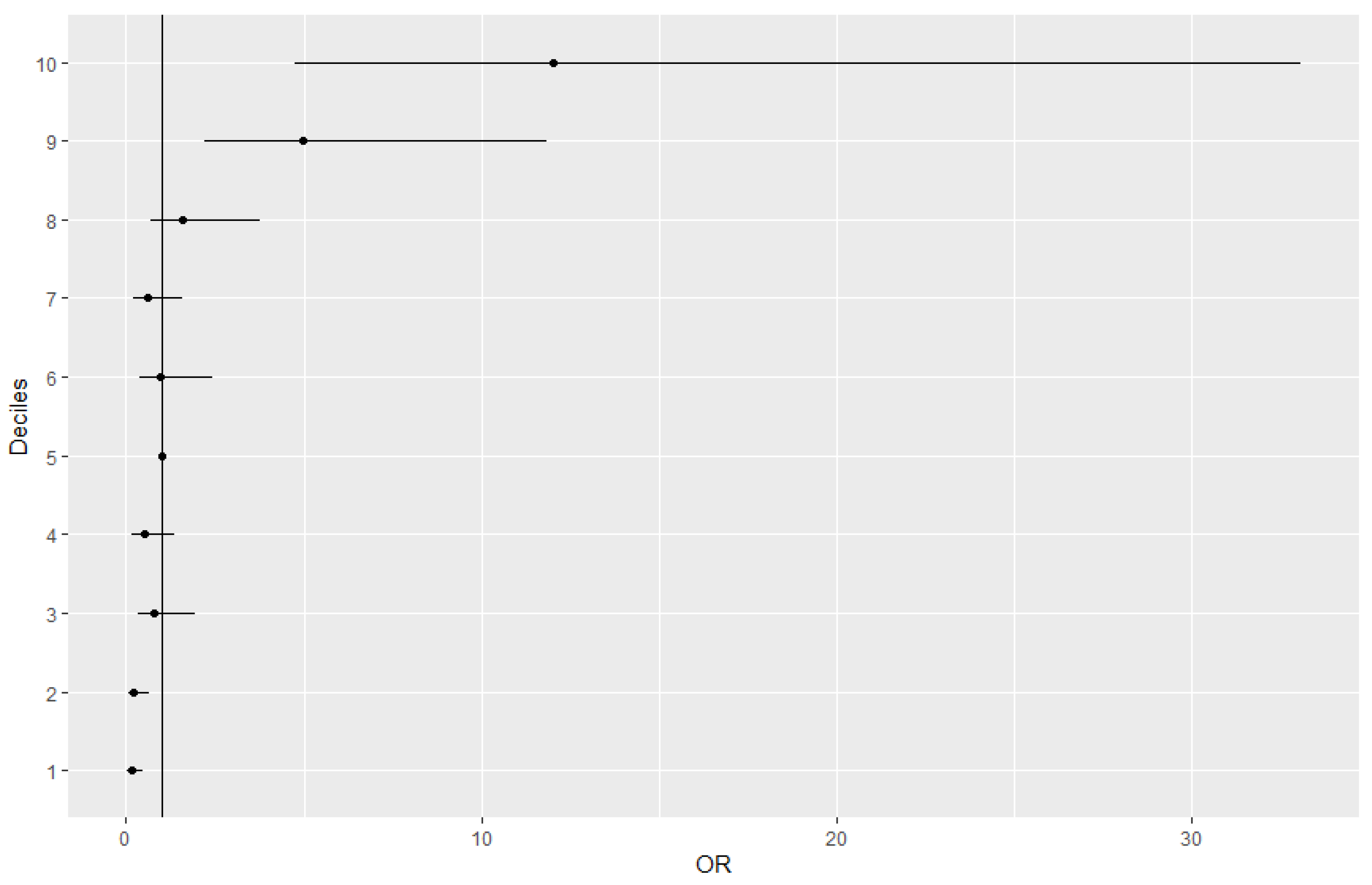
| Deciles | OR (95% CI) | p-Value | Controls | Controls (%) | Cases | Cases (%) |
|---|---|---|---|---|---|---|
| 0–10% | 0.14(0.03–0.47) | 3.66 × 10–3 ** | 59 | 16.2 | 3 | 1.9 |
| 10–20% | 0.21(0.06–0.66) | 1.14 × 10–2 ** | 52 | 14.2 | 4 | 2.6 |
| 20–30% | 0.79(0.32–1.96) | 6.11 × 10–1 | 42 | 11.5 | 12 | 7.8 |
| 30–40% | 0.50(0.17–1.35) | 1.81 × 10–1 | 39 | 10.7 | 7 | 4.5 |
| 40–50% * | 36 | 9.9 | 13 | 8.4 | ||
| 50–60% | 0.98(0.39–2.45) | 9.61 × 10–1 | 34 | 9.3 | 12 | 7.8 |
| 60–70% | 0.61(0.23–1.57) | 3.10 × 10–1 | 41 | 11.2 | 9 | 5.8 |
| 70–80% | 1.59(0.69–3.79) | 2.82 × 10–1 | 33 | 9 | 19 | 12.3 |
| 80–90% | 4.98(2.20–11.84) | 1.70 × 10–4 ** | 20 | 5.5 | 36 | 23.4 |
| 90–100% | 12.00(4.76–33.08) | 4.26 × 10–7 ** | 9 | 2.5 | 39 | 25.3 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chairta, P.P.; Hadjisavvas, A.; Georgiou, A.N.; Loizidou, M.A.; Yiangou, K.; Demetriou, C.A.; Christou, Y.P.; Pantziaris, M.; Michailidou, K.; Zamba-Papanicolaou, E. Prediction of Parkinson’s Disease Risk Based on Genetic Profile and Established Risk Factors. Genes 2021, 12, 1278. https://doi.org/10.3390/genes12081278
Chairta PP, Hadjisavvas A, Georgiou AN, Loizidou MA, Yiangou K, Demetriou CA, Christou YP, Pantziaris M, Michailidou K, Zamba-Papanicolaou E. Prediction of Parkinson’s Disease Risk Based on Genetic Profile and Established Risk Factors. Genes. 2021; 12(8):1278. https://doi.org/10.3390/genes12081278
Chicago/Turabian StyleChairta, Paraskevi P., Andreas Hadjisavvas, Andrea N. Georgiou, Maria A. Loizidou, Kristia Yiangou, Christiana A. Demetriou, Yiolanda P. Christou, Marios Pantziaris, Kyriaki Michailidou, and Eleni Zamba-Papanicolaou. 2021. "Prediction of Parkinson’s Disease Risk Based on Genetic Profile and Established Risk Factors" Genes 12, no. 8: 1278. https://doi.org/10.3390/genes12081278
APA StyleChairta, P. P., Hadjisavvas, A., Georgiou, A. N., Loizidou, M. A., Yiangou, K., Demetriou, C. A., Christou, Y. P., Pantziaris, M., Michailidou, K., & Zamba-Papanicolaou, E. (2021). Prediction of Parkinson’s Disease Risk Based on Genetic Profile and Established Risk Factors. Genes, 12(8), 1278. https://doi.org/10.3390/genes12081278








