Diverse Genetic Landscape of Suspected Retinitis Pigmentosa in a Large Korean Cohort
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Clinical Diagnosis of Retinitis Pigmentosa
2.3. Analysis of Genetic Variants
2.4. Statistical Analysis
3. Results
3.1. Demographics
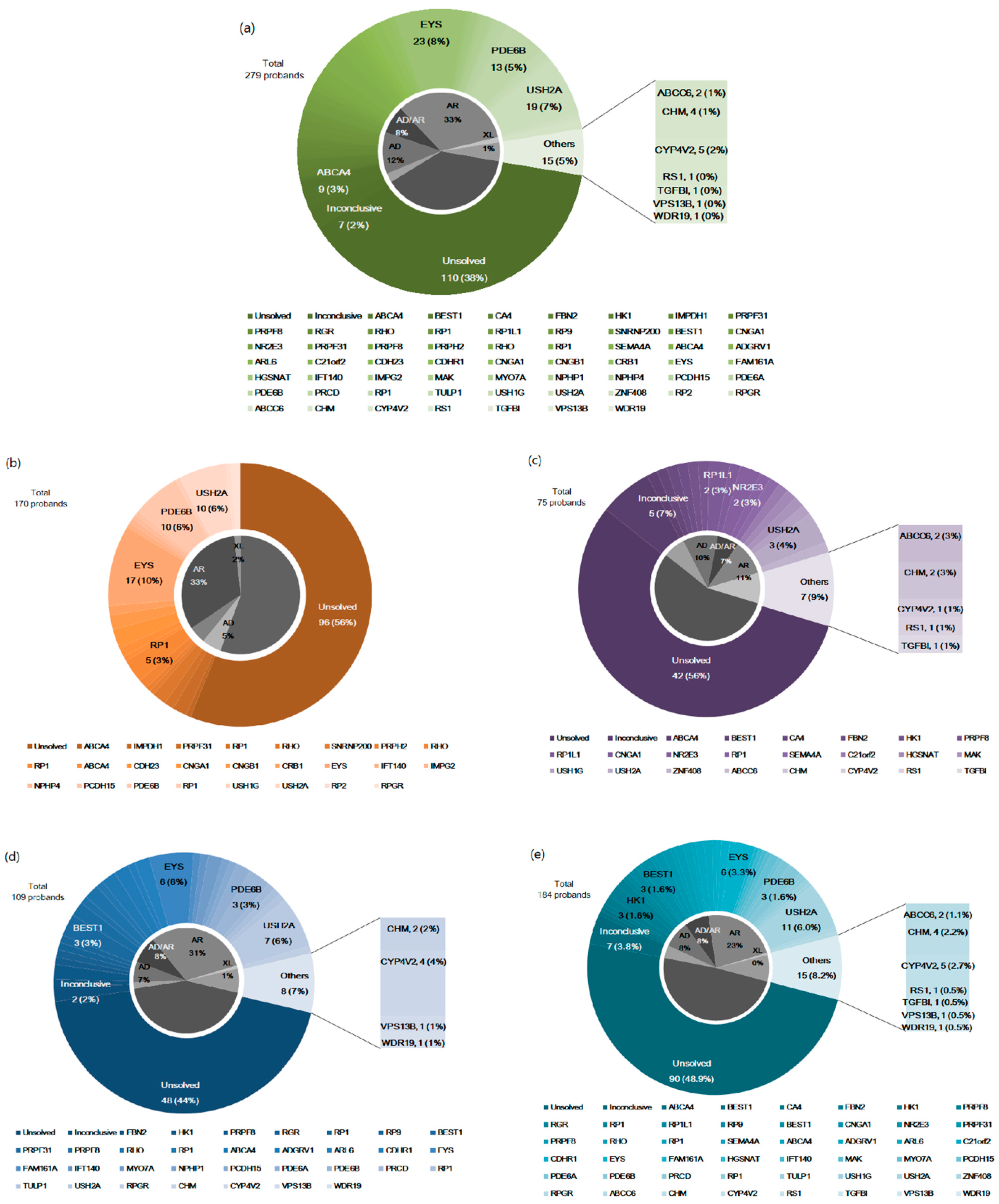
3.2. Genetic Distribution of RP in Korean Patients
3.3. Mutational Spectrum According to the Age at Initial Symptom
3.4. Mutational Spectrum According to Family History of RP
3.5. Patients with Unexpected Genotypes: Genes Causing IRDs Other than RP
- A 24-year-old woman had pseudoxanthoma elasticum (PXE) with retinal pigmentary dystrophy due to maternal-originated ABCC6, and the patient has had no other identified systemic disorders to date (Figure S1a).
- In an 18-year-old man with reticular pigmentary dispersions with chorioretinal atrophy, choroideremia caused by a CHM mutation (Figure S1b) was diagnosed; we clearly confirmed maternal transmission from the segregation test. His mother showed a mild form of chorioretinal atrophy from an ophthalmologic study.
- In the case of a 52-year-old woman showing diffuse chorioretinal degeneration, we found an associated Bietti crystalline dystrophy (BCD) caused by CYP4V2 (Figure S1c). We retrospectively reviewed seven years of medical records and found yellow-white crystals on fundus photographs that correlated with BCD before severe retinal degeneration progressed.
- A 45-year-old man with diffuse atrophy in the right eye and sectoral retinal degeneration in the left eye had the RS1 mutation (Figure S1d). As he was blind from childhood due to chronic retinal detachment in the right eye according to medical records, fundus findings in the right eye showed diffuse atrophic changes indistinguishable from RP.
- We enrolled siblings aged 15 and 17 years who had one-year-old brothers with corneal dystrophy and concomitant retinal pigmentary degeneration and found a TGFBI mutation (Figure S1e). In their pedigree analysis, many maternal relatives had poor vision with corneal dystrophies in common.
- In a 41-year-old woman showing macula-dominant diffuse retinal dystrophy with mental retardation, Cohen syndrome caused by VPS13B (Figure S1f) was diagnosed.
- In a 22-year-old woman with retinal pigmentary changes, WDR19-related Senior-Løken syndrome (Figure S1g) was diagnosed.
3.6. Patients with Inconclusive Results
4. Discussion
4.1. Genetic Distribution in Korean Patients with RP
4.2. Mutational Spectrum According to the Age at First Symptom
4.3. Mutational Spectrum According to Family History
4.4. Genetic Tests and Correction of Clinical Diagnosis from Unexpected Causative Genes
- The ocular phenotypes of PXE caused by ABCC6 mutations are variable but may show a ‘peau d’orange’ fundus appearance in childhood with reticular pigmentary dystrophy and crystalline bodies underlying the lesion of retinal pigment epithelium (RPE) atrophy [31]. Our case showed yellowish mottled features initially considered as RPE pigmentation by the clinician.
- In choroideremia, the retina covered with pigmentary changes evolves into areas of atrophy, especially in the mid-peripheral retina [32]. Our patients carrying CHM variants showed pigmented clumps with RPE degeneration as RP phenotypes; however, they also developed petalloid pattern atrophic plaques, the characteristic findings of choroideremia.
- The CYP4V2 variants induce BCD characterized by multiple glistening intraretinal crystals scattered throughout the posterior poles of the eyes [33]. As noted in our case, since the crystals rarely become visible on the fundus examination in the advanced stages of retinal atrophy, en-face OCT images may be helpful for an accurate diagnosis in such cases.
- Retinoschisis is characterized by foveal retinal splitting and peripheral changes, with retinal pigmentations and vascular attenuation or sheathing, which can resemble RP [34]. Our patient demonstrated diffuse and sectoral RPE atrophy with a history of chronic retinal detachment, which was mis-interpreted as RP combined with cystoid macular edema.
- Cohen syndrome is an uncommon systemic disease caused by a VPS13B variant, presenting with mental impairment and retinal dystrophy [35]. Even though our patient had mental impairment, diagnosis of Cohen syndrome was possible after the confirmation of genetic analysis.
- Senior-Loken Syndrome, affecting the kidney and retina, leads to nephronophthisis, Calori disease, and RP [36]. Though a patient may show no clinical renal or hepatic diseases, a combination of clinical findings and genetic testing improves the accuracy of diagnoses in syndromic diseases.
4.5. Genetic Testing and Inconclusive Cases
4.6. Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Stephen, P.; Daiger, P.; Lori, S.; Sullivan, P.; Sara, J.; Bowne, P. RetNet: Retinal Information Network. Available online: https://sph.uth.edu/retnet/sum-dis.htm (accessed on 31 October 2020).
- Hanany, M.; Rivolta, C.; Sharon, D. Worldwide carrier frequency and genetic prevalence of autosomal recessive inherited retinal diseases. Proc. Natl. Acad. Sci. USA 2020, 117, 2710. [Google Scholar] [CrossRef] [PubMed]
- Hartong, D.T.; Berson, E.L.; Dryja, T.P. Retinitis pigmentosa. Lancet 2006, 368, 1795–1809. [Google Scholar] [CrossRef]
- Oishi, M.; Oishi, A.; Gotoh, N.; Ogino, K.; Higasa, K.; Iida, K.; Makiyama, Y.; Morooka, S.; Matsuda, F.; Yoshimura, N. Comprehensive Molecular Diagnosis of a Large Cohort of Japanese Retinitis Pigmentosa and Usher Syndrome Patients by Next-Generation Sequencing. Investig. Ophthalmol. Vis. Sci. 2014, 55, 7369–7375. [Google Scholar] [CrossRef]
- Bhatia, S.; Goyal, S.; Singh, I.R.; Singh, D.; Vanita, V. A novel mutation in the PRPF31 in a North Indian adRP family with incomplete penetrance. Doc. Ophthalmol. 2018, 137, 103–119. [Google Scholar] [CrossRef]
- Bhatia, S.; Kaur, N.; Singh, I.R.; Vanita, V. A novel mutation in MERTK for rod-cone dystrophy in a North Indian family. Can. J. Ophthalmol. 2019, 54, 40–50. [Google Scholar] [CrossRef]
- Gao, F.J.; Li, J.K.; Chen, H.; Hu, F.Y.; Zhang, S.H.; Qi, Y.H.; Xu, P.; Wang, D.D.; Wang, L.S.; Chang, Q.; et al. Genetic and Clinical Findings in a Large Cohort of Chinese Patients with Suspected Retinitis Pigmentosa. Ophthalmology 2019, 126, 1549–1556. [Google Scholar] [CrossRef]
- Martin-Merida, I.; Avila-Fernandez, A.; Del Pozo-Valero, M.; Blanco-Kelly, F.; Zurita, O.; Perez-Carro, R.; Aguilera-Garcia, D.; Riveiro-Alvarez, R.; Arteche, A.; Trujillo-Tiebas, M.J.; et al. Genomic Landscape of Sporadic Retinitis Pigmentosa: Findings from 877 Spanish Cases. Ophthalmology 2019, 126, 1181–1188. [Google Scholar] [CrossRef]
- Durbin, R.M.; Altshuler, D.; Durbin, R.M.; Abecasis, G.R.; Bentley, D.R.; Chakravarti, A.; Clark, A.G.; Collins, F.S.; De La Vega, F.M.; Donnelly, P.; et al. A map of human genome variation from population-scale sequencing. Nature 2010, 467, 1061–1073. [Google Scholar] [CrossRef]
- Seo, G.H.; Kim, T.; Choi, I.H.; Park, J.Y.; Lee, J.; Kim, S.; Won, D.G.; Oh, A.; Lee, Y.; Choi, J.; et al. Diagnostic yield and clinical utility of whole exome sequencing using an automated variant prioritization system, EVIDENCE. Clin. Genet. 2020, 98, 562–570. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Aziz, N.; Bale, S.; Bick, D.; Das, S.; Gastier-Foster, J.; Grody, W.W.; Hegde, M.; Lyon, E.; Spector, E.; et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 2015, 17, 405–424. [Google Scholar] [CrossRef]
- Tavtigian, S.V.; Greenblatt, M.S.; Harrison, S.M.; Nussbaum, R.L.; Prabhu, S.A.; Boucher, K.M.; Biesecker, L.G.; ClinGen Sequence Variant Interpretation Working, G. Modeling the ACMG/AMP variant classification guidelines as a Bayesian classification framework. Genet. Med. 2018, 20, 1054–1060. [Google Scholar] [CrossRef]
- Huang, X.F.; Huang, F.; Wu, K.C.; Wu, J.; Chen, J.; Pang, C.P.; Lu, F.; Qu, J.; Jin, Z.B. Genotype-phenotype correlation and mutation spectrum in a large cohort of patients with inherited retinal dystrophy revealed by next-generation sequencing. Genet Med. 2015, 17, 271–278. [Google Scholar] [CrossRef]
- Carss, K.J.; Arno, G.; Erwood, M.; Stephens, J.; Sanchis-Juan, A.; Hull, S.; Megy, K.; Grozeva, D.; Dewhurst, E.; Malka, S.; et al. Comprehensive Rare Variant Analysis via Whole-Genome Sequencing to Determine the Molecular Pathology of Inherited Retinal Disease. Am. J. Hum. Genet. 2017, 100, 75–90. [Google Scholar] [CrossRef] [PubMed]
- Birtel, J.; Gliem, M.; Mangold, E.; Müller, P.L.; Holz, F.G.; Neuhaus, C.; Lenzner, S.; Zahnleiter, D.; Betz, C.; Eisenberger, T.; et al. Next-generation sequencing identifies unexpected genotype-phenotype correlations in patients with retinitis pigmentosa. PLoS ONE 2018, 13, e0207958. [Google Scholar] [CrossRef] [PubMed]
- Numa, S.; Oishi, A.; Higasa, K.; Oishi, M.; Miyata, M.; Hasegawa, T.; Ikeda, H.O.; Otsuka, Y.; Matsuda, F.; Tsujikawa, A. EYS is a major gene involved in retinitis pigmentosa in Japan: Genetic landscapes revealed by stepwise genetic screening. Sci. Rep. 2020, 10, 20770. [Google Scholar] [CrossRef] [PubMed]
- Arai, Y.; Maeda, A.; Hirami, Y.; Ishigami, C.; Kosugi, S.; Mandai, M.; Kurimoto, Y.; Takahashi, M. Retinitis Pigmentosa with EYS Mutations Is the Most Prevalent Inherited Retinal Dystrophy in Japanese Populations. J. Ophthalmol. 2015, 2015, 819760. [Google Scholar] [CrossRef]
- Kim, M.S.; Joo, K.; Seong, M.W.; Kim, M.J.; Park, K.H.; Park, S.S.; Woo, S.J. Genetic Mutation Profiles in Korean Patients with Inherited Retinal Diseases. J. Korean Med. Sci. 2019, 34, e161. [Google Scholar] [CrossRef] [PubMed]
- Dias, M.F.; Joo, K.; Kemp, J.A.; Fialho, S.L.; da Silva Cunha, A., Jr.; Woo, S.J.; Kwon, Y.J. Molecular genetics and emerging therapies for retinitis pigmentosa: Basic research and clinical perspectives. Prog. Retin. Eye. Res. 2018, 63, 107–131. [Google Scholar] [CrossRef]
- Karali, M.; Testa, F.; Brunetti-Pierri, R.; Di Iorio, V.; Pizzo, M.; Melillo, P.; Barillari, M.R.; Torella, A.; Musacchia, F.; D’Angelo, L.; et al. Clinical and Genetic Analysis of a European Cohort with Pericentral Retinitis Pigmentosa. Int. J. Mol. Sci. 2019, 21, 86. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.J.; Lin, K.H.; Lee, S.H.; Shen, R.J.; Feng, Z.K.; Wang, X.F.; Huang, X.F.; Huang, Z.Q.; Jin, Z.B. Mutation spectrum and genotype-phenotype correlation of inherited retinal dystrophy in Taiwan. Clin. Exp. Ophthalmol. 2020, 48, 486–499. [Google Scholar] [CrossRef]
- Na, K.-H.; Kim, H.J.; Kim, K.H.; Han, S.; Kim, P.; Hann, H.J.; Ahn, H.S. Prevalence, Age at Diagnosis, Mortality, and Cause of Death in Retinitis Pigmentosa in Korea—A Nationwide Population-based Study. Am. J. Ophthalmol. 2017, 176, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Pierrache, L.H.; Hartel, B.P.; van Wijk, E.; Meester-Smoor, M.A.; Cremers, F.P.; de Baere, E.; de Zaeytijd, J.; van Schooneveld, M.J.; Cremers, C.W.; Dagnelie, G.; et al. Visual Prognosis in USH2A-Associated Retinitis Pigmentosa Is Worse for Patients with Usher Syndrome Type IIa Than for Those with Nonsyndromic Retinitis Pigmentosa. Ophthalmology 2016, 123, 1151–1160. [Google Scholar] [CrossRef] [PubMed]
- Iftikhar, M.; Lemus, M.; Usmani, B.; Campochiaro, P.A.; Sahel, J.A.; Scholl, H.P.N.; Shah, S.M.A. Classification of disease severity in retinitis pigmentosa. Br. J. Ophthalmol. 2019, 103, 1595–1599. [Google Scholar] [CrossRef] [PubMed]
- Verbakel, S.K.; van Huet, R.A.C.; Boon, C.J.F.; den Hollander, A.I.; Collin, R.W.J.; Klaver, C.C.W.; Hoyng, C.B.; Roepman, R.; Klevering, B.J. Non-syndromic retinitis pigmentosa. Prog. Retin. Eye. Res. 2018, 66, 157–186. [Google Scholar] [CrossRef] [PubMed]
- Ormerod, L.D.; Puklin, J.E.; Sobel, J.D. Syphilitic Posterior Uveitis: Correlative Findings and Significance. Clin. Infect. Dis. 2001, 32, 1661–1673. [Google Scholar] [CrossRef] [PubMed]
- Farrell, D.F. Retinal toxicity to antimalarial drugs: Chloroquine and hydroxychloroquine: A neurophysiologic study. Clin. Ophthalmol. 2012, 6, 377–383. [Google Scholar] [CrossRef]
- Grange, L.; Dalal, M.; Nussenblatt, R.B.; Sen, H.N. Autoimmune retinopathy. Am. J. Ophthalmol. 2014, 157, 266–272.e261. [Google Scholar] [CrossRef]
- Xu, Y.; Guan, L.; Xiao, X.; Zhang, J.; Li, S.; Jiang, H.; Jia, X.; Yang, J.; Guo, X.; Yin, Y.; et al. Mutation analysis in 129 genes associated with other forms of retinal dystrophy in 157 families with retinitis pigmentosa based on exome sequencing. Mol. Vis. 2015, 21, 477–486. [Google Scholar] [PubMed]
- Wright, A.F.; Chakarova, C.F.; Abd El-Aziz, M.M.; Bhattacharya, S.S. Photoreceptor degeneration: Genetic and mechanistic dissection of a complex trait. Nat. Rev. Genet. 2010, 11, 273–284. [Google Scholar] [CrossRef]
- Audo, I.; Vanakker, O.M.; Smith, A.; Leroy, B.P.; Robson, A.G.; Jenkins, S.A.; Coucke, P.J.; Bird, A.C.; De Paepe, A.; Holder, G.E.; et al. Pseudoxanthoma Elasticum with Generalized Retinal Dysfunction, a Common Finding? Investig. Ophthalmol. Vis. Sci. 2007, 48, 4250–4256. [Google Scholar] [CrossRef] [PubMed]
- Khan, K.N.; Islam, F.; Moore, A.T.; Michaelides, M. Clinical and Genetic Features of Choroideremia in Childhood. Ophthalmology 2016, 123, 2158–2165. [Google Scholar] [CrossRef]
- Gocho, K.; Kameya, S.; Akeo, K.; Kikuchi, S.; Usui, A.; Yamaki, K.; Hayashi, T.; Tsuneoka, H.; Mizota, A.; Takahashi, H. High-Resolution Imaging of Patients with Bietti Crystalline Dystrophy with CYP4V2 Mutation. J. Ophthalmol. 2014, 2014, 283603. [Google Scholar] [CrossRef] [PubMed]
- Sikkink, S.K.; Biswas, S.; Parry, N.R.A.; Stanga, P.E.; Trump, D. X-linked retinoschisis: An update. J. Med. Genet. 2007, 44, 225–232. [Google Scholar] [CrossRef]
- Chandler, K.E.; Biswas, S.; Lloyd, I.C.; Parry, N.; Clayton-Smith, J.; Black, G.C.M. The ophthalmic findings in Cohen syndrome. Br. J. Ophthalmol. 2002, 86, 1395–1398. [Google Scholar] [CrossRef]
- Coussa, R.G.; Otto, E.A.; Gee, H.Y.; Arthurs, P.; Ren, H.; Lopez, I.; Keser, V.; Fu, Q.; Faingold, R.; Khan, A.; et al. WDR19: An ancient, retrograde, intraflagellar ciliary protein is mutated in autosomal recessive retinitis pigmentosa and in Senior-Loken syndrome. Clin. Genet. 2013, 84, 150–159. [Google Scholar] [CrossRef]
- Kannabiran, C.; Klintworth, G.K. TGFBI gene mutations in corneal dystrophies. Hum. Mutat. 2006, 27, 615–625. [Google Scholar] [CrossRef]
- Wang, S.K.; Xue, Y.; Cepko, C.L. Microglia modulation by TGF-β1 protects cones in mouse models of retinal degeneration. J. Clin. Investig. 2020, 130, 4360–4369. [Google Scholar] [CrossRef] [PubMed]
- Ma, W.; Silverman, S.M.; Zhao, L.; Villasmil, R.; Campos, M.M.; Amaral, J.; Wong, W.T. Absence of TGFβ signaling in retinal microglia induces retinal degeneration and exacerbates choroidal neovascularization. eLife 2019, 8. [Google Scholar] [CrossRef]


| No. of Probands, n (%) | Sex, M:F (%) | Family History, n (Y:N, %) | Age at Genetic Examination, Years | Age at Symptom Onset, Years | Age at Diagnosis, Years | BCVA, LogMAR | ||
|---|---|---|---|---|---|---|---|---|
| RE | LE | |||||||
| Total subjects | 279 | 131:148 (47.0:53.0) | 92:187 (33.0:67.0) | 47.6 ± 15.7 | 25.6 ± 16.9 | 41.1 ± 15.2 | 0.8 ± 1.0 | 0.8 ± 1.0 |
| Subgroup analysis according to age at symptom onset | ||||||||
| ≤10 years | 69 (24.7) | 41:28 (59.4:40.6) | 25:44 (36.2:63.8) | 40.6 ± 16.7 | 7.7 ± 2.1 | 30.7 ± 14.7 | 0.9 ± 1.1 | 0.8 ± 1.0 |
| 11–20 years | 82 (29.4) | 43:39 (52.4:47.6) | 28:54 (34.1:65.9) | 41.6 ± 14.8 | 16.0 ± 3.1 | 36.7 ± 13.8 | 0.9 ± 1.1 | 0.9 ± 1.1 |
| 21–40 years | 61 (21.9) | 21:40 (34.4:65.6) | 19:42 (31.1:68.9) | 52.2 ± 12.8 | 33.2 ± 5.5 | 44.6 ± 10.4 | 0.6 ± 0.9 | 0.7 ± 1.0 |
| ≥41 years | 61 (21.9) | 23:38 (37.7:62.3) | 19:42 (31.1:68.9) | 59.9 ± 7.9 | 51.0 ± 6.2 | 55.9 ± 7.9 | 0.7 ± 1.0 | 0.7 ± 0.9 |
| asymptomatic | 6 (2.1) | 3:3 (50.0:50.0) | 1:5 (16.7:83.3) | 39.2 ± 15.4 | N/A | 36.2 ± 13.0 | 0.1 ± 0.1 | 0.0 ± 0.2 |
| No. of Probands, n | No. of Segregation Analyses, n | No. of Probands with Detected Variants, n (Detection Rate, %) | No. of Probands with Inconclusive Results, n (%) | ||
|---|---|---|---|---|---|
| Total subjects | 279 | 60 | 161 (57.7) | 10 (3.6) | |
| Genetic analysis | TGS | 170 | 29 | 74 (43.5) | |
| WES from unsolved TGS | 75 | 23 | 28 (37.3) | 8 (10.7) | |
| WES only | 109 | 31 | 59 (54.1) | 2 (1.8) | |
| Age at symptom onset | ≤10 years | 69 | 19 | 52 (75.4) | 2 (2.9) |
| 11–20 years | 82 | 19 | 49 (59.8) | 4 (4.9) | |
| 21–40 years | 61 | 11 | 27 (44.3) | 4 (6.6) | |
| ≥41 years | 61 | 10 | 30 (49.2) | 0 (0.0) | |
| asymptomatic | 6 | 1 | 3 (50.0) | 0 (0.0) | |
| Family history | + | 91 | 28 | 65 (71.4) | 2 (2.2) |
| - | 188 | 32 | 96 (51.1) | 8 (4.3) | |
| Subject No. | Causative Gene | NM Number | Chromosome | HGVS DNA | HGVS Protein Change | Zygosity | Inheritance | Origin | Chromosome | ACMG Criteria | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 6-6 | RS1 | NM_000330.3 | X | c.78+1G>A | Hemi | XL | Unknown | LPV | PVS1, PM2 | Novel | |
| 7-7 | CYP4V2 | NM_207352.3 | 4 | c.809_810C | Hetero | AR | Unknown | PV | PVS1, PM2, PP5 | ||
| CYP4V2 | NM_207352.3 | 4 | c.992A>C | p.His331Pro | Hetero | AR | Unknown | LPV | PS1, PM2, PP2, PP3 | ||
| 14-17 | ABCC6 | NM_001171.5 | 16 | c.3703C>T | p.Arg1235Trp | Hetero | AD | Maternal | LPV | PS1, PM2, PP3 | |
| 21-28 | TGFBI | NM_000358.2 | 5 | c.371G>A | p.Arg124His | Hetero | AD | Unknown | PV | PS1, PS3, PM1, PM5, PP2, PP3 | |
| 21-29 | TGFBI | NM_000358.2 | 5 | c.371G>A | p.Arg124His | Hetero | AD | Unknown | PV | PS1, PS3, PM1, PM5, PP2, PP3 | |
| 22-30 | CHM | NM_000390.3 | X | c.688delinsTG | Hemi | XL | Maternal | LPV | PVS1, PM2 | Novel | |
| 22-31 | CHM | NM_000390.3 | X | c.688delinsTG | Hemi | XL | Unknown | LPV | PVS1, PM2 | Novel | |
| 61-96 | ABCC6 | NM_001171.5 | 16 | c.3698T>C | p.Val1233Ala | Hetero | AD | Unknown | VUS | PM1, PM2, PP3 | |
| 72-112 | CHM | NM_000390.3 | X | c.1718_1719del | p.Tyr573CysfsTer12 | Hemi | XL | Unknown | PV | PVS1, PM2, PP5 | |
| 72-113 | CHM | NM_000390.3 | X | c.1718_1719del | p.Tyr573CysfsTer12 | Hemi | XL | Unknown | PV | PVS1, PM2, PP5 | |
| 172-223 | WDR19 | NM_025132.3 | 4 | c.2645+1G>T | Hetero | AR | Unknown | LPV | PVS1, PM2 | ||
| WDR19 | NM_025132.3 | 4 | c.1613G>T | p.Gly538Val | Hetero | AR | Unknown | VUS | PM2, PP3 | Novel | |
| 188-252 | CYP4V2 | NM_207352.3 | 4 | c.1072G>T | p.Glu358Ter | Homo | AR | Unknown | LPV | PVS1, PM2 | |
| 193-261 | CHM | NM_000390.3 | X | c.2T>A | p.Met1Lys | Hemi | XL | Unknown | VUS | PVS1_M, PM2 | Novel |
| 207-280 | VPS13B | NM_017890.4 | 8 | c.7220_7221A | Hetero | AR | Maternal | PV | PM2, PP3, BP1 | ||
| VPS13B | NM_017890.4 | c.11468G>C | p.Gly3823Arg | Hetero | AR | Unknown | VUS | PVS1, PM2, PP5 | Novel | ||
| 225-311 | CYP4V2 | NM_207352.3 | 4 | c.809_810C | Homo | AR | Unknown | PV | PVS1, PM2, PP5 | ||
| 240-337 | CYP4V2 | NM_207352.3 | 4 | c.802_807A | Hetero | AR | Unknown | LPV | PVS1, PM2 | ||
| CYP4V2 | NM_207352.3 | c.219T>A | p.Phe73Leu | Hetero | AR | Unknown | VUS | PM2, PP2, PP3 | |||
| 248-349 | CYP4V2 | NM_207352.3 | 4 | c.675-1G>A | Hetero | AR | Unknown | LPV | PVS1, PM2 | Novel | |
| CYP4V2 | NM_207352.3 | 4 | c.802-8_807del | Hetero | AR | Unknown | LPV | PVS1, PM2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, Y.-J.; Kim, Y.-N.; Yoon, Y.-H.; Seo, E.-J.; Seo, G.-H.; Keum, C.; Lee, B.-H.; Lee, J.-Y. Diverse Genetic Landscape of Suspected Retinitis Pigmentosa in a Large Korean Cohort. Genes 2021, 12, 675. https://doi.org/10.3390/genes12050675
Kim Y-J, Kim Y-N, Yoon Y-H, Seo E-J, Seo G-H, Keum C, Lee B-H, Lee J-Y. Diverse Genetic Landscape of Suspected Retinitis Pigmentosa in a Large Korean Cohort. Genes. 2021; 12(5):675. https://doi.org/10.3390/genes12050675
Chicago/Turabian StyleKim, Yoon-Jeon, You-Na Kim, Young-Hee Yoon, Eul-Ju Seo, Go-Hun Seo, Changwon Keum, Beom-Hee Lee, and Joo-Yong Lee. 2021. "Diverse Genetic Landscape of Suspected Retinitis Pigmentosa in a Large Korean Cohort" Genes 12, no. 5: 675. https://doi.org/10.3390/genes12050675
APA StyleKim, Y.-J., Kim, Y.-N., Yoon, Y.-H., Seo, E.-J., Seo, G.-H., Keum, C., Lee, B.-H., & Lee, J.-Y. (2021). Diverse Genetic Landscape of Suspected Retinitis Pigmentosa in a Large Korean Cohort. Genes, 12(5), 675. https://doi.org/10.3390/genes12050675






