4. Discussion
We presented a complex analysis of 22 differentiated single nucleotide polymorphisms in prediction of dental caries in primary dentition of children. This study is an extended analysis utilizing additional data from our previous studies [
14,
15]. Deep learning neural network models for caries prediction were applied in a homogeneous cohort of 2–3-year-old children living in an urban environment under similar cultural conditions.
The previous caries experience, independent of an individual’s age, has previously been reported as the strongest and the most universal risk factor for future caries development [
28]. However, it is challenging to assess the previous caries experience in infants and toddlers. Most of the previous studies and distinct caries prediction models apply to adults, school children or older preschoolers, while models for toddlers are scarce [
29]. Another obstacle in assessing well-performing caries prediction model are discrepancies among the studies, i.e., imprecise definition of caries phenotype and caries lesions or inconsistency in the terms used by the researchers [
9,
23,
30,
31]. Furthermore, the majority of acquired caries prediction models are based on demographical and environmental factors and the only biological feature that has been considered is the cariogenic bacteria colonization of oral cavity.
The first linkage studies in caries were performed in 2008 [
32] and the first genome-wide association studies for early childhood caries were published in 2011 [
33]. Abbasoğlu et al. [
10] were the first to correlate environmental and genetic factors in ECC in Turkish 2–5-year-olds, while Lewis et al. [
34] considered several single nucleotide polymorphisms, however they were not included in the final caries prediction meta-analysis. Another technical obstacle in any caries prediction model implementation in practice is an almost total lack of replication and validation studies in independent populations [
13,
35]. Mejare et al. [
28] depicted 17 studies on caries prediction in preschoolers, defined as children of age <1 to 6 years old. None of them used genetic variables as prediction factors, most gave moderate prediction accuracy results and only one study by Holgerson et al. [
36] based on environmental factors and saliva sampling in 2-year-olds was validated. Persistent high dental caries rate in the general Polish population and nearly constant prevalence and severity index of the disease in children in the recent years [
37] should encourage both researchers and clinicians to develop better and validated prediction approaches that might be implemented in practice.
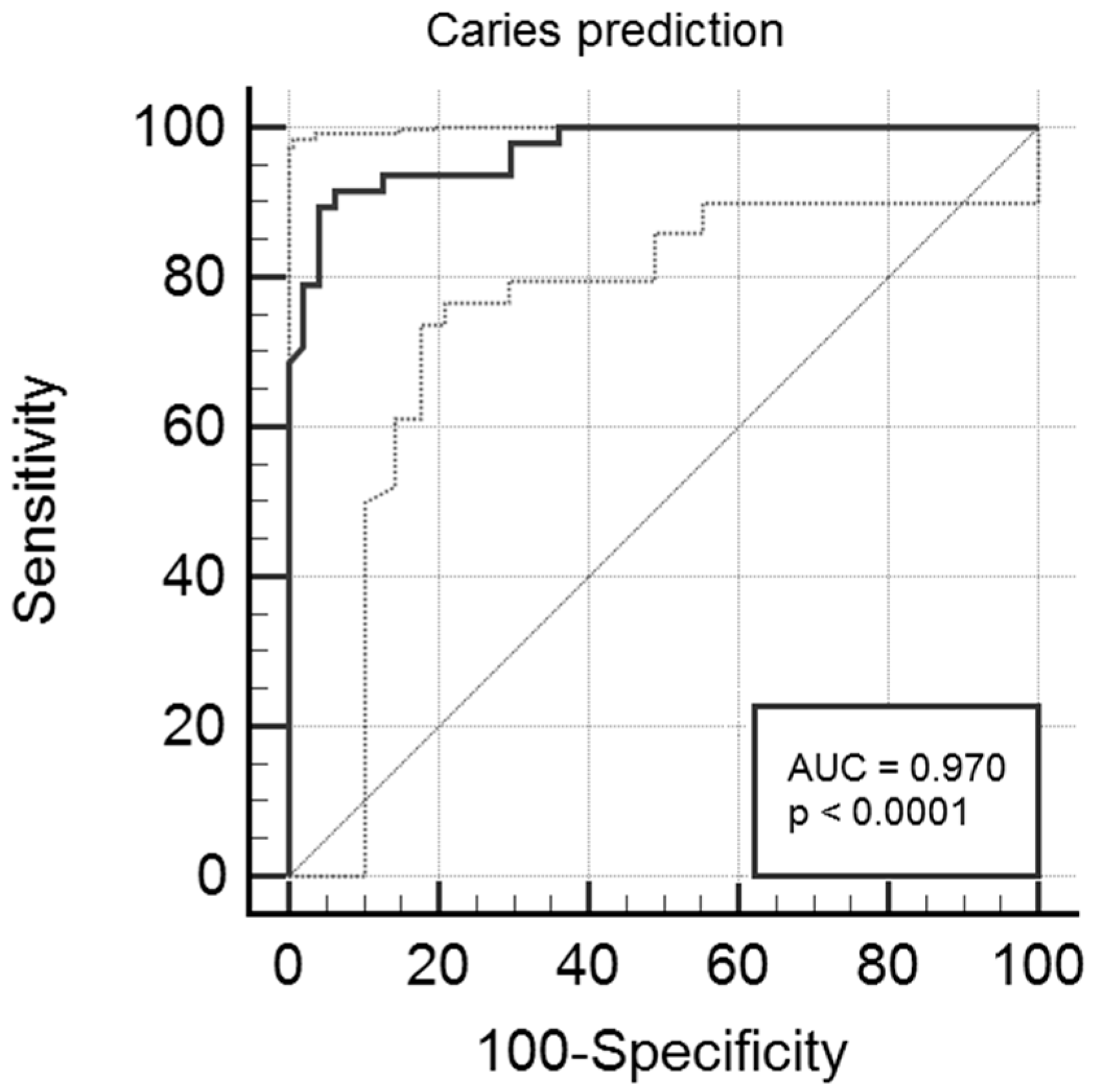
In this study, we obtained relatively high prediction accuracy ranging from 90.9% to 98.4%, depending on the prediction model, using a neural network approach. Additionally, multivariate logistic regression analysis showed accuracy of 93% with high sensitivity and specificity values, i.e., 89.6% and 95.7%, respectively. Results of both approaches were gender- and age-independent, as the two study subgroups were adjusted for both features. The most important predictors/indicators in both assays were
AMELX rs17878486 and
TUFT1 rs2337360 [
15]. In brief,
AMELX and
TUFT1 are the genes that play a crucial role in the enamel formation process and the single nucleotide polymorphisms have previously been attributed to high caries susceptibility, in primary as well as permanent dentition, although the risk allele varied depending on population and age [
12,
30,
38]. Another factor that was significantly associated with caries outcome in both uni- and multivariable logistic regression was
ENAM rs12640848.
ENAM encodes enamel-special protein enamelin that is a critical factor during enamel maturation. Our results were in agreement with other reports, supporting the presumable role of rs12640848_G as a protective factor in ECC [
10,
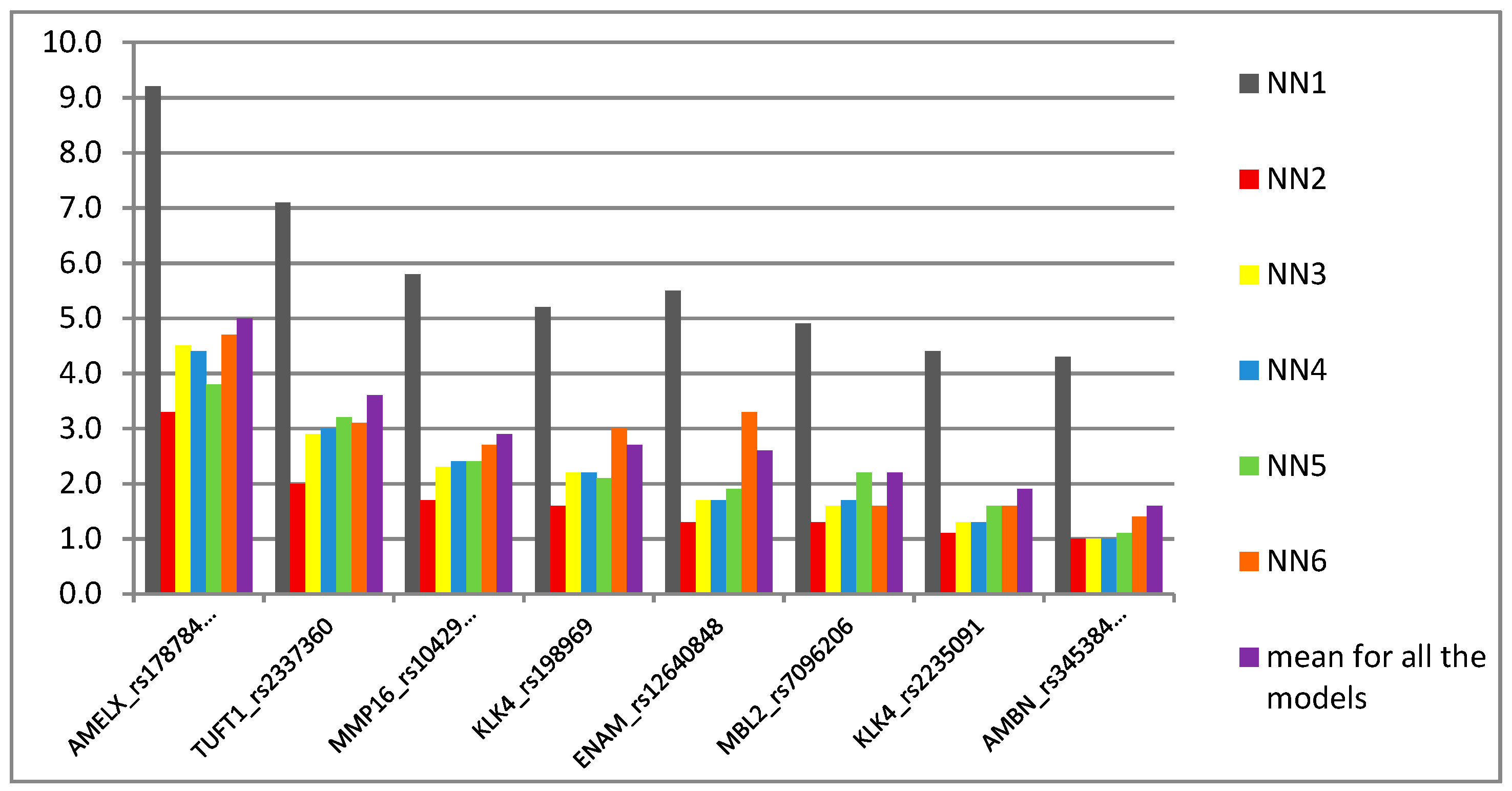
30]. Interestingly, when the SNP was used in this study as a predictor in neural network analysis, it turned out to be an intermediate marker in the scale of importance (
Figure 2). This might be explained by the fact that both wild AA and rare GG genotypes were associated with the opposite outcomes, i.e., caries and caries-free phenotype, respectively. Conversely, another SNP, i.e.,
MMP16 rs10429371, was the third most important predictor in NN models, although it did not hold the significance as an indicator in multivariate analysis. This is an interesting finding, since rs10429371 explained over 20% of the trait in single locus analysis and the TT variant was nearly 26 times more frequent in caries individuals when compared to controls, showing a strong recessive pattern of inheritance and suggesting TT as a risk variant for dental caries development in the studied individuals. Additionally, it was one of the variants for which alleles and genotypes were differently distributed in the cohort in this study in comparison to the CEU individuals in favor of wild C and CC variants (
Table 3). Matrix metalloproteinases (MMPs) play an important role in early tooth development by regulating ameloblast maturation and formation of enamel. Several types of MMPs have been described to be involved in dentin collagen degradation and dental caries lesion progression [
12,
34,
39,
40,
41,
42].
MMP16 rs10429371 was previously associated with caries in white adults [
34]. Linhartova et al. [
41] observed higher incidence of rare T allele in caries children aged 13–15 years, although it did not reach the statistical significance level.
The second variant that showed significant association with caries outcome and was genotyped in this study was rs7096206 in
MBL2. Mannose binding protein (MBL2) is an acute phase protein that plays an important role in innate immunity. The study showed that
MBL2 polymorphisms are involved in several infectious diseases and the top ranked rs7096206 has been annotated as deleterious to the protein’s function and described as one of the most functionally important variants in the gene [
42]. Rs7096206_G was found to be a risk factor in Polish 5-year-olds in reference to higher vs. lower caries experience, while it had no effect in 12-year-olds [
1]. Interestingly, the same rare variant was significantly associated with no caries experience in 2–3-year-olds in our study, showing an additive model of inheritance. On the other hand, in the former study [
1], rs7096206_G was associated with higher caries experience in both age subgroups while in the haplotype with rs1800450_G, and the CG haplotype had the opposite effect. Alike, rs7096206 genotype distribution was insignificant, but in haplotype with rs7501477_T it correlated with caries experience in Saudi 5–13-year-old children [
43]. It might suggest that other SNPs could be associated with a more complex pattern of the disease and/or that distinct genetic variants could be involved at different ages. Such differences, i.e., in primary vs. permanent dentition, were also acknowledged by Wang et al. [
44]. Likewise, as the rs7096206_G variant is associated with lower MBL2 serum levels predisposing to infections [
45], other SNPs could show strong linkage disequilibrium accounting for possible protective mechanisms in the toddlers in our study. Nevertheless, the SNP was not significant in multivariate analysis, despite the drastic OR value, i.e., 3.36 × 10
−9.
We did not observe any association of other SNPs in MMP, i.e.,
MMP20 rs1711437 genotyped in this study and rs1784418 in the previous study [
15], with caries or caries-free phenotype. MMP20 (enamelysin) is the early protease secreted during enamel development and is involved in both dentin and enamel decomposition [
46]. Only Antunes et al. [
47] found both variants to be associated with early childhood caries. Yet, the results of other studies of
MMP20 SNPs were on the border of significance in 5-year-old Caucasian children with dental caries or have been associated more with poor oral hygiene and dietary habits than the disease itself in 5–14-year-old Caucasians [
16,
46]. Likewise, rs45447741 in
LTF, rs7217186 in
ALOX15 and rs1884302 in
SMAD6 presented no association with caries experience in this study in any of the statistical tests used.
The rest of significantly different distributed variants in the previous study [
15], i.e.,
AMBN rs34538475 and
KLK4 rs2235091and rs198969, were not significant in multivariate analysis and their importance in caries prediction models were of a moderate (rs2235091and rs198969) to low (rs34538475) degree (
Figure 2).
KLK4 plays an important role in the late stages of enamel development, while
AMBN, together with
AMELX, is crucial for enamel matrix formation and mineralization. The roles of the abovementioned SNPs in both genes as risk factors in caries development in children were partially in agreement with other authors, depending on studied population and children’s age [
10,
12,
46]. Additionally, the haplotype analysis showed that alleles of both variants in
KLK4 were necessary for the risk (i.e., rs2235091G_rs198969G haplotype) and protective (i.e., rs2235091A_rs198969C haplotype) effect.
It must be emphasized that the differences in association of genetic variants with caries experience and severity in the studies occurs not only due to the differences between the populations but also in one population itself, even from one individual to another [
34]. Single nucleotide polymorphisms are often highly variable between distinct ethnic cohorts and, at least partially, resemble divergence in human phenotypes, including different disease susceptibility or drug response. Dental caries is a highly complex trait, also in reference to environmental factors, i.e., socioeconomic and cultural factors, age, oral hygiene, eating habits, also the course of pregnancy and mother–child relations [
3,
6,
46,
48,
49,
50]. Age appears one of the key factors in caries analysis, as the disease experience is differentially defined in reference to the age of patients. Some authors emphasize that each age group should be characterized by other sets of temporal variables [
50], with the most important risk factors as follows: in 2-year-olds—allergies and infections before first tooth eruption and intake of drugs during the first 12 months of life, in 3-year-olds—mother’s age at the time of pregnancy and smoking during pregnancy [
29], consumption of sweetened food during first 12 months of life and nocturnal drinking of sweet drinks above 12th month [
3], in 4-year-olds and older children—frequency of tooth brushing, fluoride treatment and mother’s education [
51].
One of the hypotheses might be that distinct processes, and hence different genes and polymorphisms, play a significant role throughout the stages of a child’s growth and development. Since the genetic sequence does not change over the course of life, each stage might be sensitive to distinct variants, including those that shape susceptibility to caries development. Simultaneously, the sensitivity to the secondary environmental factors and the tertiary behavioral factors, responsible for the intermediate and distal effects, respectively, might also alter. Substantially, it can be explained by the order of teeth eruption, their type and number. Caries lesions appear first on incisors and molars, also maxillary teeth are more often affected than mandibular teeth. Caries formation predominantly affects teeth with deep and/or narrow pits and fissures on the occlusal surface, which is closely related to the morphology of primary dentition and molars [
4,
46,
50]. The deeper and narrower the pit, the easier it is for the bacterial plaque to penetrate and adhere to a tooth surface. The higher the number of teeth with deep and narrow pits, together with inappropriate brushing, the more frequent the predisposition to higher caries rate, which is especially relevant in preschoolers [
50,
52,
53]. The similar pattern of caries development has been observed in 3-year-old children in other studies [
53,
54,
55]. In this study, we did not observe differences in the total number of erupted teeth and the number of the teeth types was similar between affected and caries-free children and between boys and girls in both subgroups (not shown). Although we did not conduct a follow-up study, some authors pointed that caries susceptibility varies depending on the tooth morphology and rises sharply after 2–3 years after the tooth eruption, when posteruptive enamel maturation takes place [
4,
5]. Therefore, dental caries seems to be a remarkably divergent trait constantly changing in time and with its occurrence increasing with age. The prevalence of caries experience in our study was 18.3% and was comparable to children of the corresponding age of other ethnicities, i.e., Western Europe, Eastern-Southern Asia and Sub-Saharan Africa [
3,
29,
56]. Interestingly, it appeared to be much lower when compared to other studies concerning Polish children with active caries of distinct regions, i.e., 40.8% in the Podlasie region [
57], 53.8% in Lower Silesian, Malopolskie and Lubelskie voivodships [
4] or from 35% to 56.6% in the general 2–3-year-olds population [
3,
13,
57]. However, according to Werneck et al. [
13] caries rate can reach even 85% in preschoolers.
Another risk factor for caries lesions development, closely related to children’s age, is the presence and composition of bacterial plaque in the host. The level of colonization by cariogenic bacteria has been considered the strongest risk factor in 3-year-old Polish children [
49], however, it should be emphasized that it might be influenced by other features. Firstly, the activity of Lactobacilli and Mutans Streptococci is higher in preschoolers and in primary dentition in comparison with older children and permanent teeth [
10,
50,
58], therefore it might be a biological predisposition of the host. Secondly, the younger the child, the more attention and supervision over teeth brushing should be provided by the parents/guardians, therefore minimizing the bacterial film and setting a good hygiene habits. The mother not brushing a child’s teeth was found a major risk factor in Mutans Streptococci infections and ECC in Australian children aged 12–72 months but also in 9-month-old Thai children [
6,
59].
Both genetic and host factors fluctuate with age, and with a broad spectrum of environmental factors, they contribute to the development of caries lesion differently in every individual. Moreover, even children of the same ethnicity that are exposed to the same levels of risk factors might present distinct caries severity index and patterns of the disease [
13], which strongly supports the existence of other, intrinsic, i.e., genetic components. Still, depending on the population and its habits, prediction models using the same risk factors might be characterized by distinct final prediction parameters, as well as comparable prediction features of different models might be determined by distinct input variables [
8]. To mention a few, Kalhan et al. reached a high AUC value of 0.81–0.91 of caries prediction in 2-year-olds and 0.79 in 3-year-olds [
29], while Fontana et al. [
60] assessed AUC of 0.73 in 4-year-old children. Tamaki et al. [
61] came across 33 studies on caries prediction, however, the majority of them used logistic regression analyses and only some authors used additional, advanced machine learning algorithms to compare the results, namely artificial neural network, decision analysis or classification and regression trees [
27,
50,
61,
62,
63]. The benefit of logistic regression analysis is that it can be conducted using non-complicated software; however, the procedure is descriptive in its nature and should be applied to assess risk indicators, not predictors. On the other hand, advanced machine learning methods consider the unequal strength of potential markers, so that a weak factor is not hidden by the strong one and predictive power is more reliable [
9,
51]. In fact, we observed some differences in the importance level of studied variants between the two approaches, although it is not entirely adequate to compare the results of different algorithms, even describing the same features. Still, one of NN models—NN5, in which logistic function was applied as an activation algorithm in both hidden and output layers, gave the most outstanding importance values of all tested predictors, while in the remaining 5 models based on distinct algorithms were lower but comparable with each other within one model. The NN5 model presented the second-best prediction accuracy in this study; however, using logistic methods only might suggest over-fitting of the model, even in advanced machine learning approaches. Likewise, So and Sham [
64] stated that ROC curve analysis is not directly correlated with the disease risk and even high significance, i.e., low
p-values, is not equal to a good predictive power. Javed et al. [
27] obtained 99% accuracy in caries prediction in 6–14-year-old Indian children using neural network modeling, and it was the method of choice in other studies with medical prediction [
65,
66,
67,
68].
Another issue worth mentioning is the nature of variables themselves. When studying some environmental, behavioral and biological factors, a collinearity might occur, e.g., the presence of dental plaque might be the result of a poor oral hygiene, which in turn might be correlated with a lack of guardians’ control over child’s brushing. [
63]. While it is not a problem in advanced modeling approaches, it should be avoided in logistic regression analyses, since it tends to cause over-fitting of the data and spurious results [
63,
69]. The strength of this study is a homogeneous study group, that is sex- and age-adjusted, since both factors are found to be potential confounders in case–control studies [
24]. Secondly, the assessment and comparison of the results and performance of two different statistical approaches, adjusted for the data and therefore more reliable, appears to be crucial when implementing the method into the clinics. The obvious limitation of this study and a necessary future direction is a larger study group and validation of the model.
The polymorphisms analyzed in this study and a high caries prediction rates indicate a strong genetic component in the course of the disease. Nevertheless, one has to remember about the multifactorial nature of caries and that even the best prediction model cannot fully describe the real life scenarios and neither solely genetic nor solely environmental factors can completely explain the disease cause. Hence, analyses exploring environmental and genetic predictors need to be conducted very carefully. Firstly, some genetic variants might influence behavioral habits, e.g., SNPs in taste genes [
11] as well as non-genetic factors themselves might be correlated with one another, yielding spurious results. Secondly, compared to the environmental impact, if some genetic effect on a disease trait exists, it remains the main and unchangeable element in the disease incidence, especially early in life [
70,
71]. Even being under the influence of the environment, genetic factors combined with non-genetic findings have been replicated only partially and with the lack of statistical power [
70]. Although many rare variants or the ones with smaller effect might be missed in purely genetic case–control studies, the subgroup analyses and high homogeneity of the study subgroups greatly improves the prediction [
72,
73]. Some authors [
70,
74] have studied and described interesting gene–environment association tests, that successively and carefully analyze both components in multiple testing approaches. McAllister et al. [
75] have developed novel statistical approaches to detect genetic–non-genetic interactions with regard to different durations of exposure to environmental factors. Therefore, a fully developed caries prediction model undoubtedly requires a complex analysis of multiple factors as well as replication studies before implementation in practice. Still, the high prediction value of genetic polymorphisms presented in this study comprises valuable findings on early caries development that sets the direction of the future research.