Spatial and Temporal Expression of High-Mobility-Group Nucleosome-Binding (HMGN) Genes in Brain Areas Associated with Cognition in Individuals with Down Syndrome
Abstract
:1. Introduction
2. Methodology
2.1. Data Mining
2.2. Data Preprocessing
2.3. Quantification of the Differential HMGN Genes Expression
2.4. Gene-Dosage Imbalanced Quantification
2.5. Construction of HMGN Genes Network Using GeneMania
2.6. Protein–Protein Interaction Analysis
2.7. Statistical Analysis
3. Results
3.1. Expression of HMGN Genes in Brain Areas from Individuals with DS
3.2. Age Dependent Expression of HMGN Genes in the Brain of Individuals with DS
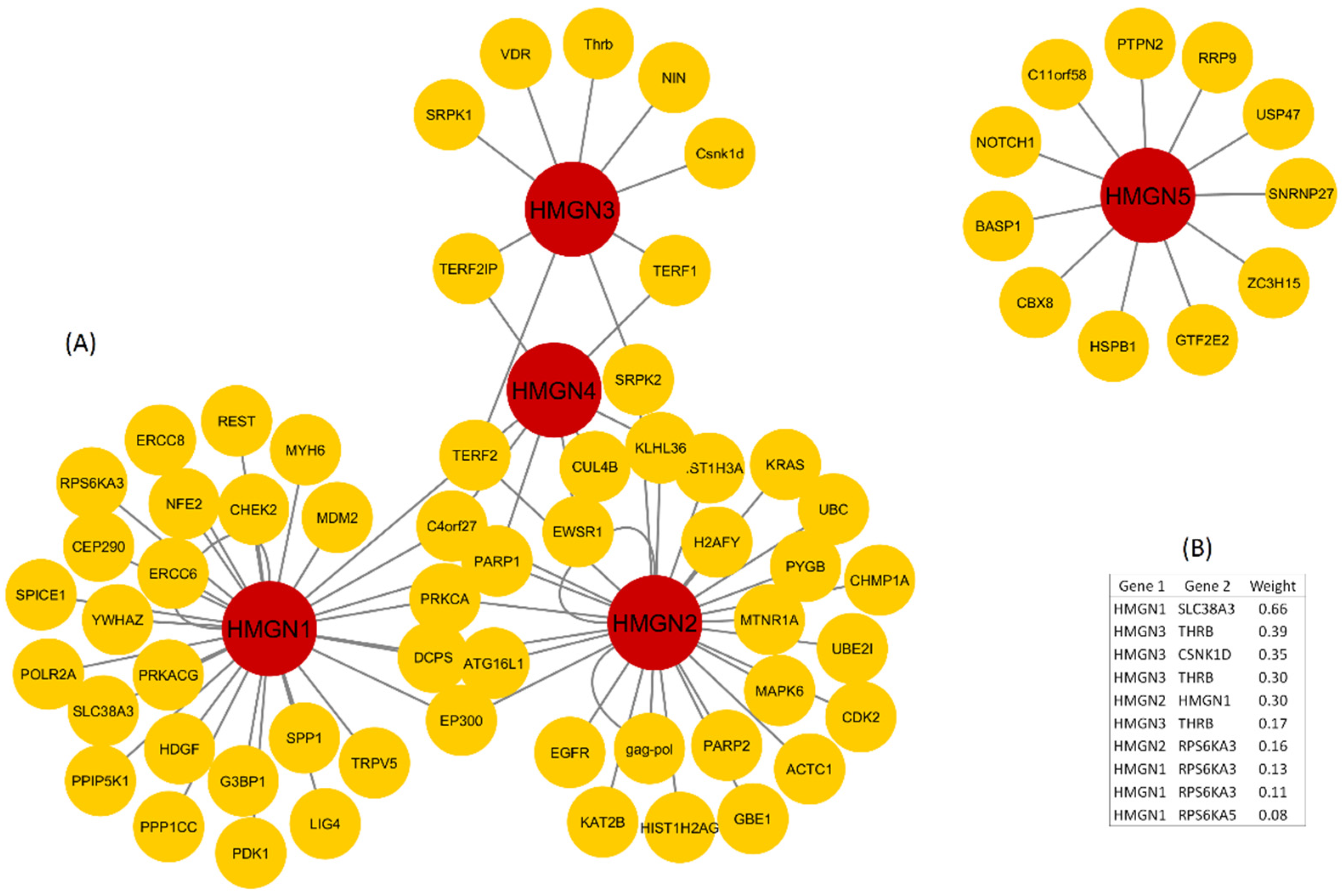
3.3. Protein–Protein Interaction Network and GO Categories
3.4. HMGN Protein Interaction with Histones of Nucleosome Core and Linker H1
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zhu, F.; Farnung, L.; Kaasinen, E.; Sahu, B.; Yin, Y.; Wei, B.; Dodonova, S.O.; Nitta, K.R.; Morgunova, E.; Taipale, M.; et al. The interaction landscape between transcription factors and the nucleosome. Nature 2018, 562, 76–81. [Google Scholar] [CrossRef]
- Hochedlinger, K.; Jaenisch, R. Induced pluripotency and epigenetic reprogramming. Cold Spring Harb. Perspect. Biol. 2015, 7, a019448. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kugler, J.E.; Deng, T.; Bustin, M. The HMGN family of chromatin-binding proteins: Dynamic modulators of epigenetic processes. Biochimica et Biophysica acta 2012, 1819, 652–656. [Google Scholar] [CrossRef] [Green Version]
- Patterson, D. Genetic mechanisms involved in the phenotype of Down Syndrome. Ment. Retard Dev. Disabil. Res. Rev. 2007, 13, 199–206. [Google Scholar] [CrossRef] [PubMed]
- Carothers, A.D.; Hecht, C.A.; Hook, E.B. International variation in reported live birth prevalence rates of Down Syndrome, adjusted for maternal age. J. Med. Genet. 1999, 36, 386–393. [Google Scholar]
- Canfield, M.A.; Honein, M.A.; Yuskiv, N.; Xing, J.; Mai, C.T.; Collins, J.S.; Devine, O.; Petrini, J.; Ramadhani, T.A.; Hobbs, C.A.; et al. National estimates and race/ethnic-specific variation of selected birth defects in the United States, 1999–2001. Birth Defects Res. A Clin. Mol. Teratol. 2006, 76, 747–756. [Google Scholar] [CrossRef]
- Glasson, E.J.; Sullivan, S.G.; Hussain, R.; Petterson, B.A.; Montgomery, P.D.; Bittles, A.H. The changing survival profile of people with Down’s syndrome: Implications for genetic counselling. Clin. Genet. 2002, 62, 390–393. [Google Scholar] [CrossRef]
- Antonarakis, S.E. Down Syndrome and the complexity of genome dosage imbalance. Nat. Rev. Genet. 2016, 18, 147–163. [Google Scholar] [CrossRef]
- Do, C.; Xing, Z.; Yu, Y.E.; Tycko, B. Trans-acting epigenetic effects of chromosomal aneuploidies: Lessons from Down Syndrome and mouse models. Epigenomics 2017, 9, 189–207. [Google Scholar] [CrossRef] [Green Version]
- Mao, R.; Zielke, C.L.; Zielke, H.R.; Pevsner, J. Global up-regulation of chromosome 21 gene expression in the developing Down Syndrome brain. Genomics 2003, 81, 457–467. [Google Scholar] [CrossRef]
- Antonarakis, S.E.; Lyle, R.; Dermitzakis, E.T.; Reymond, A.; Deutsch, S. Chromosome 21 and down Syndrome: From genomics to pathophysiology. Nat Rev Genet 2004, 5, 725–738. [Google Scholar] [CrossRef]
- Gensous, N.; Franceschi, C.; Salvioli, S.; Garagnani, P.; Bacalini, M.G. Down Syndrome, Ageing and Epigenetics. Subcell Biochem. 2019, 91, 161–193. [Google Scholar] [CrossRef] [PubMed]
- Sailani, M.R.; Santoni, F.A.; Letourneau, A.; Borel, C.; Makrythanasis, P.; Hibaoui, Y.; Antonarakis, S.E. DNA-methylation patterns in trisomy 21 using cells from monozygotic twins. PLoS ONE 2015, 10, e013555. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rochman, M.; Postnikov, Y.; Correll, S.; Malicet, C.; Wincovitch, S.; Karpova, T.S.; McNally, J.G.; Wu, X.; Bubunenko, N.A.; Grigoryev, S.; et al. The Interaction of NSBP1 with Nucleosomes in Euchromatin Counteracts Linker Histone-Mediated Chromatin Compaction and Modulates The Fidelity Of The Cellular Transcription Profile. Mol. Cell. 2009, 35, 642–656. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shirakawa, H.; Landsman, D.; Postnikov, Y.V.; Bustin, M. NBP-45, a novel nucleosomal binding protein with a tissue-specific and developmentally regulated expression. J. Biol. Chem. 2000, 275, 6368–6374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ueda, T.; Catez, F.; Gerlitz, G.; Bustin, M. Delineation of the protein module that anchors HMGN proteins to nucleosomes in the chromatin of living cells. Mol. Cell Biol. 2008, 28, 2872–2883. [Google Scholar] [CrossRef] [Green Version]
- Mardian, J.K.; Paton, A.E.; Bunick, G.J.; Olins, D.E. Nucleosome cores have two specific binding sites for nonhistone chromosomal proteins HMG 14 and HMG 17. Science 1980, 209, 1534–1536. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sandeen, G.; Wood, W.I.; Felsenfeld, G. The interaction of high mobility proteins HMG14 and 17 with nucleosomes. Nucleic Acids Res. 1980, 8, 3757–3778. [Google Scholar] [CrossRef] [Green Version]
- Rice, J.C.; Allis, C.D. Histone methylation versus histone acetylation: New insights into epigenetic regulation. Curr. Opin Cell Biol. 2001, 13, 263–273. [Google Scholar] [CrossRef]
- Belova, G.I.; Postnikov, Y.V.; Furusawa, T.; Birger, Y.; Bustin, M. Chromosomal protein HMGN1 enhances the heat shock-induced remodeling of Hsp70 chromatin. J. Biol. Chem. 2008, 283, 8080–8088. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tao, D.; Yuri, P.; Zhang, S.; Lillian, G.; Lore, B.; Ildikó, R.; Sabine, M.; Hölter, W.W.; Helmut, F.; Valerie, G.-D.; et al. Interplay between H1 and HMGN epigenetically regulates OLIG1&2 expression and oligodendrocyte differentiation. Nucleic Acids Res. 2017, 45, 3031–3045. [Google Scholar] [CrossRef]
- Singh, G.P.; Ganapathi, M.; Dash, D. Role of intrinsic disorder in transient interactions of hub proteins. Proteins 2007, 66, 761–765. [Google Scholar] [CrossRef]
- Uversky, V.N.; Oldfield, C.J.; Midic, U.; Xie, H.; Xue, B.; Vucetic, S.; Iakoucheva, L.M.; Obradovic, Z.; Dunker, A.K. Unfoldomics of human diseases: Linking protein intrinsic disorder with diseases. BMC Genom. 2009, 10, S7. [Google Scholar] [CrossRef] [Green Version]
- Zhang, S.; Deng, T.; Tang, W.; He, B.; Furusawa, T.; Ambs, S.; Bustin, M. Epigenetic regulation of REX1 expression and chromatin binding specificity by HMGNs. Nucleic Acids Res. 2019, 47, 4449–4461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olmos-Serrano, J.L.; Kang, H.J.; Tyler, W.A.; Silbereis, J.C.; Cheng, F.; Zhu, Y.; Pletikos, M.; Jankovic-Rapan, L.; Cramer, N.P.; Galdzicki, Z.; et al. Down Syndrome Developmental Brain Transcriptome Reveals Defective Oligodendrocyte Differentiation and Myelination. Neuron 2016, 89, 1208–1222. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Irizarry, R.A.; Hobbs, B.; Collin, F.; Beazer-Barclay, Y.D.; Antonellis, K.J.; Scherf, U.; Speed, T.P. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 2003, 4, 249–264. [Google Scholar] [CrossRef] [Green Version]
- Therneau, T.M.; Ballman, K.V. What does PLIER really do? Cancer Inform. 2008, 6, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Cheadle, C.; Vawter, M.P.; Freed, W.J.; Becker, K.G. Analysis of microarray data using Z score transformation. J. Mol. Diagn. JMD 2003, 5, 73–81. [Google Scholar] [CrossRef] [Green Version]
- Aït Yahya-Graison, E.; Aubert, J.; Dauphinot, L.; Rivals, I.; Prieur, M.; Golfier, G.; Rossier, J.; Personnaz, L.; Creau, N.; Bléhaut, H.; et al. Classification of human chromosome 21 gene-expression variations in Down Syndrome: Impact on disease phenotypes. Am. J. Hum. Genet. 2007, 81, 475–491. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mostafavi, S.; Ray, D.; Warde-Farley, D.; Grouios, C.; Morris, Q. GeneMANIA: A real-time multiple association network integration algorithm for predicting gene function. Genome Biol. 2008, 9 (Suppl. 1), S4. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Franz, M.; Rodriguez, H.; Lopes, C.; Zuberi, K.; Montojo, J.; Bader, G.D.; Morris, Q. GeneMANIA update 2018. Nucleic Acids Res 2018, 46, W60–W64. [Google Scholar] [CrossRef] [Green Version]
- Breitkreutz, B.J.; Stark, C.; Tyers, M. The GRID: The General Repository for Interaction Datasets. Genome Biol. 2003, 4, R23. [Google Scholar] [CrossRef] [PubMed]
- Oughtred, R.; Stark, C.; Breitkreutz, B.J.; Rust, J.; Boucher, L.; Chang, C.; Kolas, N.; O’Donnell, L.; Leung, G.; McAdam, R.; et al. The BioGRID interaction database: 2019 update. Nucleic Acids Res. 2019, 47, D529–D541. [Google Scholar] [CrossRef] [Green Version]
- Costa, V.; Angelini, C.; D’Apice, L.; Mutarelli, M.; Casamassimi, A.; Sommese, L.; Gallo, M.A.; Aprile, M.; Esposito, R.; Leone, L.; et al. Massive-scale RNA-Seq analysis of non ribosomal transcriptome in human trisomy 21. PLoS ONE 2011, 6, e18493. [Google Scholar] [CrossRef]
- Lane, A.A.; Chapuy, B.; Lin, C.Y.; Tivey, T.; Li, H.; Townsend, E.C.; van Bodegom, D.; Day, T.A.; Wu, S.C.; Liu, H.; et al. Triplication of a 21q22 region contributes to B cell transformation through HMGN1 overexpression and loss of histone H3 Lys27 trimethylation. Nat. Genet. 2014, 46, 618–623. [Google Scholar] [CrossRef] [Green Version]
- Kahmann, N.H.; Rake, A.V. Altered nucleosome spacing associated with Down Syndrome. Biochem. Genet. 1993, 31, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Hock, R.; Furusawa, T.; Ueda, T.; Bustin, M. HMG chromosomal proteins in development and disease. Trends Cell Biol. 2007, 17, 72–79. [Google Scholar] [CrossRef] [Green Version]
- Nanduri, R.; Furusawa, T.; Bustin, M. Biological Functions of HMGN Chromosomal Proteins. Int. J. Mol. Sci. 2020, 21, 449. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rubin, R.D.; Watson, P.D.; Duff, M.C.; Cohen, N.J. The role of the hippocampus in flexible cognition and social behavior. Front. Hum. Neurosci. 2014, 8, 742. [Google Scholar] [CrossRef] [Green Version]
- Abuhatzira, L.; Shamir, A.; Schones, D.E.; Schäffer, A.A.; Bustin, M. The chromatin-binding protein HMGN1 regulates the expression of methyl CpG-binding protein 2 (MECP2) and affects the behavior of mice. J. Biol. Chem. 2011, 286, 42051–44262. [Google Scholar] [CrossRef] [Green Version]
- Kugler, J.E.; Horsch, M.; Huang, D.; Furusawa, T.; Rochman, M.; Garrett, L.; Becker, L.; Bohla, A.; Hölter, S.M.; Prehn, C.; et al. High mobility group N proteins modulate the fidelity of the cellular transcriptional profile in a tissue- and variant-specific manner. J. Biol. Chem. 2013, 288, 16690–16703. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ito, Y.; Bustin, M. Immunohistochemical localization of the nucleosome-binding protein HMGN3 in mouse brain. J. Histochem. Cytochem. 2002, 50, 1273–1275. [Google Scholar] [CrossRef] [Green Version]
- Mohamed, O.A.; Bustin, M.; Clarke, H.J. High-mobility group proteins 14 and 17 maintain the timing of early embryonic development in the mouse. Dev. Biol. 2001, 229, 237–249. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Korner, U.; Bustin, M.; Scheer, U.; Hock, R. Developmental role of HMGN proteins in Xenopus laevis. Mech. Dev. 2003, 120, 1177–1192. [Google Scholar] [CrossRef] [PubMed]
- Garza-Manero, S.; Sindi, A.A.A.; Mohan, G.; Rehbini, O.; Jeantet, V.H.M.; Bailo, M.; Latif, F.A.; West, M.P.; Gurden, R.; Finlayson, L.; et al. Maintenance of active chromatin states by HMGN2 is required for stem cell identity in a pluripotent stem cell model. Epigenetics Chromatin. 2019, 12, 73. [Google Scholar] [CrossRef]
| Gene | ID * | Locus | Gene Name |
|---|---|---|---|
| HMGN1 | 3150 | 21q22.2 | High mobility group nucleosome binding domain 1 |
| HMGN2 | 3151 | 1p36.11 | High mobility group nucleosome binding domain 2 |
| HMGN3 | 9324 | 6q14.1 | High mobility group nucleosome binding domain 3 |
| HMGN4 | 10473 | 6p22.2 | High mobility group nucleosome binding domain 4 |
| HMGN5 | 79366 | Xq21.1 | High mobility group nucleosome binding domain 5 |
| Gene | Gene ID | Brain | HIP | CBC | DFC | OFC | V1C | VFC | ITC |
|---|---|---|---|---|---|---|---|---|---|
| HMGN1 | 3150 | 2.81 | 3.00 | 2.35 | 2.63 | 2.07 | 3.00 | 1.86 | 2.45 |
| HMGN2 | 3151 | 0.06 | 0.23 | 0.13 | 0.54 | 0.05 | −0.62 | 0.68 | 0.99 |
| HMGN3 | 9324 | 2.48 | 1.29 | 1.60 | 1.78 | 1.57 | 2.82 | 2.31 | 1.86 |
| HMGN4 | 10473 | 0.37 | 0.85 | 0.77 | 0.27 | 0.84 | 0.41 | 0.19 | −0.27 |
| HMGN5 | 79366 | 2.65 | 2.18 | 1.60 | 2.49 | 1.13 | 1.49 | 2.53 | 2.88 |
| Gene | 16–22 WG | 0–12 M | 2–10 Y | 12–22 Y | 30–39 Y | 40–42 Y |
|---|---|---|---|---|---|---|
| HMGN1 | 0.66 | 1.45 | 1.94 | 1.97 | 2.18 | 1.55 |
| HMGN2 | 2.13 | 0.22 | 0.54 | 0.54 | 0.31 | 0.49 |
| HMGN3 | 1.08 | 3.61 | 2.71 | 2.55 | 2.59 | 2.94 |
| HMGN4 | 4.71 | 1.12 | 1.51 | 1.65 | 1.84 | 1.67 |
| HMGN5 | 0.16 | 1.89 | 0.78 | 1.15 | 0.76 | 0.82 |
| Interactor | Interaction | Experimental Evidence | Throughput | Score * |
|---|---|---|---|---|
| HMGN1 | HIST1H4A | Affinity Capture-MS (§) | High | >0.75 |
| HMGN2 | HIST1H2BA | Affinity Capture-MS | High | 0.99 |
| HIST1H3A | Affinity Capture-MS | High | 0.90 | |
| HIST1H2AG | Affinity Capture-MS | High | 0.77 | |
| HIST1H1A | Proximity Label-MS (§§) | High | >0.75 | |
| HMGN3 | HIST1H4A | Affinity Capture-MS | High | >0.75 |
| HMGN4 | HIST1H2AG | Affinity Capture-MS | High | 0.92 |
| HIST1H3A | Proximity Label-MS | High | >0.75 | |
| HIST2H2AB | Affinity Capture-MS | High | 0.88 | |
| HMGN5 | HIST1H4A | Affinity Capture-MS | High | >0.75 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodríguez-Ortiz, A.; Montoya-Villegas, J.C.; García-Vallejo, F.; Mina-Paz, Y. Spatial and Temporal Expression of High-Mobility-Group Nucleosome-Binding (HMGN) Genes in Brain Areas Associated with Cognition in Individuals with Down Syndrome. Genes 2021, 12, 2000. https://doi.org/10.3390/genes12122000
Rodríguez-Ortiz A, Montoya-Villegas JC, García-Vallejo F, Mina-Paz Y. Spatial and Temporal Expression of High-Mobility-Group Nucleosome-Binding (HMGN) Genes in Brain Areas Associated with Cognition in Individuals with Down Syndrome. Genes. 2021; 12(12):2000. https://doi.org/10.3390/genes12122000
Chicago/Turabian StyleRodríguez-Ortiz, Alejandra, Julio César Montoya-Villegas, Felipe García-Vallejo, and Yecid Mina-Paz. 2021. "Spatial and Temporal Expression of High-Mobility-Group Nucleosome-Binding (HMGN) Genes in Brain Areas Associated with Cognition in Individuals with Down Syndrome" Genes 12, no. 12: 2000. https://doi.org/10.3390/genes12122000
APA StyleRodríguez-Ortiz, A., Montoya-Villegas, J. C., García-Vallejo, F., & Mina-Paz, Y. (2021). Spatial and Temporal Expression of High-Mobility-Group Nucleosome-Binding (HMGN) Genes in Brain Areas Associated with Cognition in Individuals with Down Syndrome. Genes, 12(12), 2000. https://doi.org/10.3390/genes12122000






