Mitochondrial Genomes of the Genus Claassenia (Plecoptera: Perlidae) and Phylogenetic Assignment to Subfamily Perlinae
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Preparation and DNA Extraction
2.2. PCR Amplification and Sequencing
2.3. Mitogenome Assembly and Annotation
2.4. Phylogenetic Analysis
3. Results
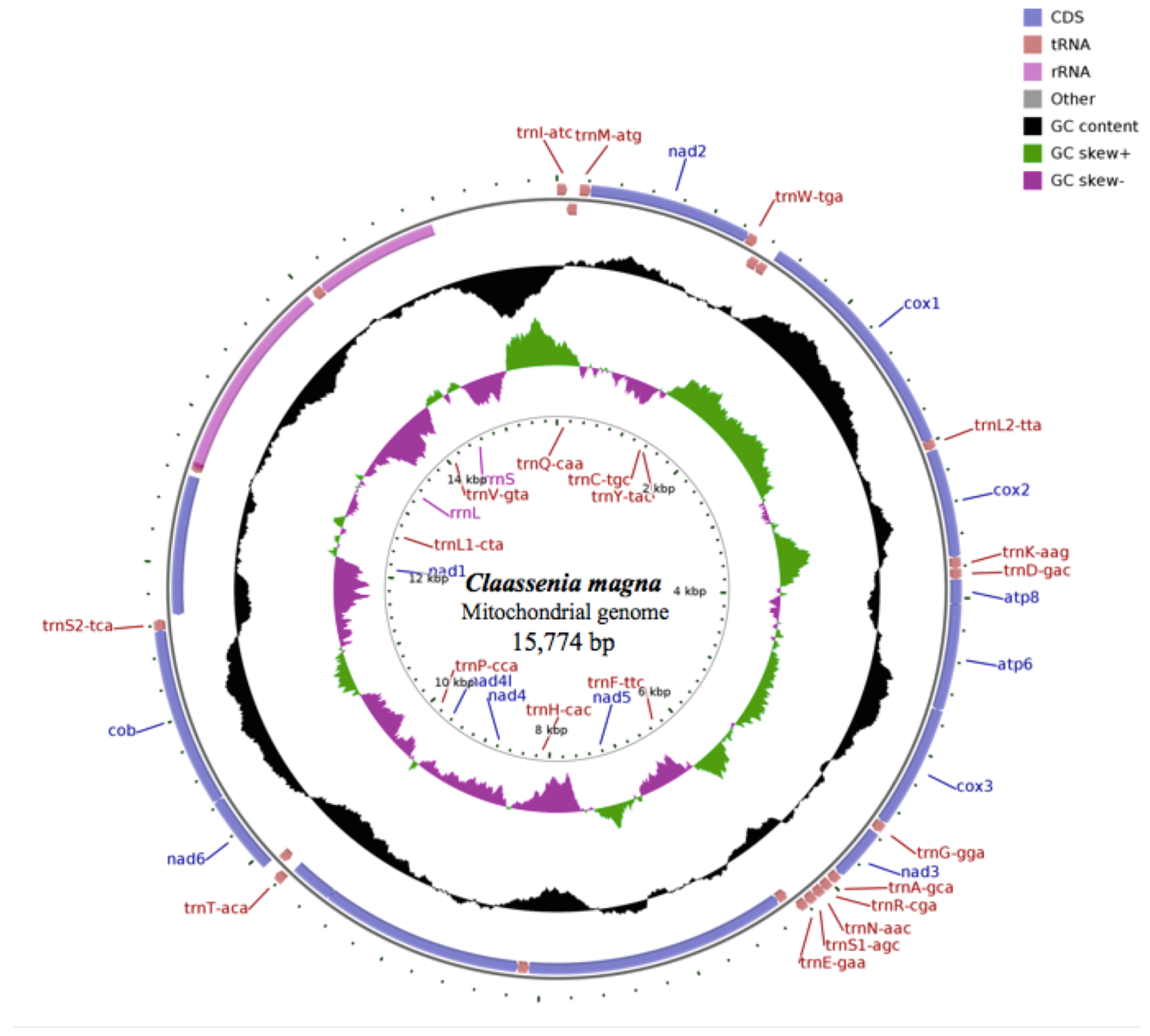
3.1. Mitogenome and Base Composition
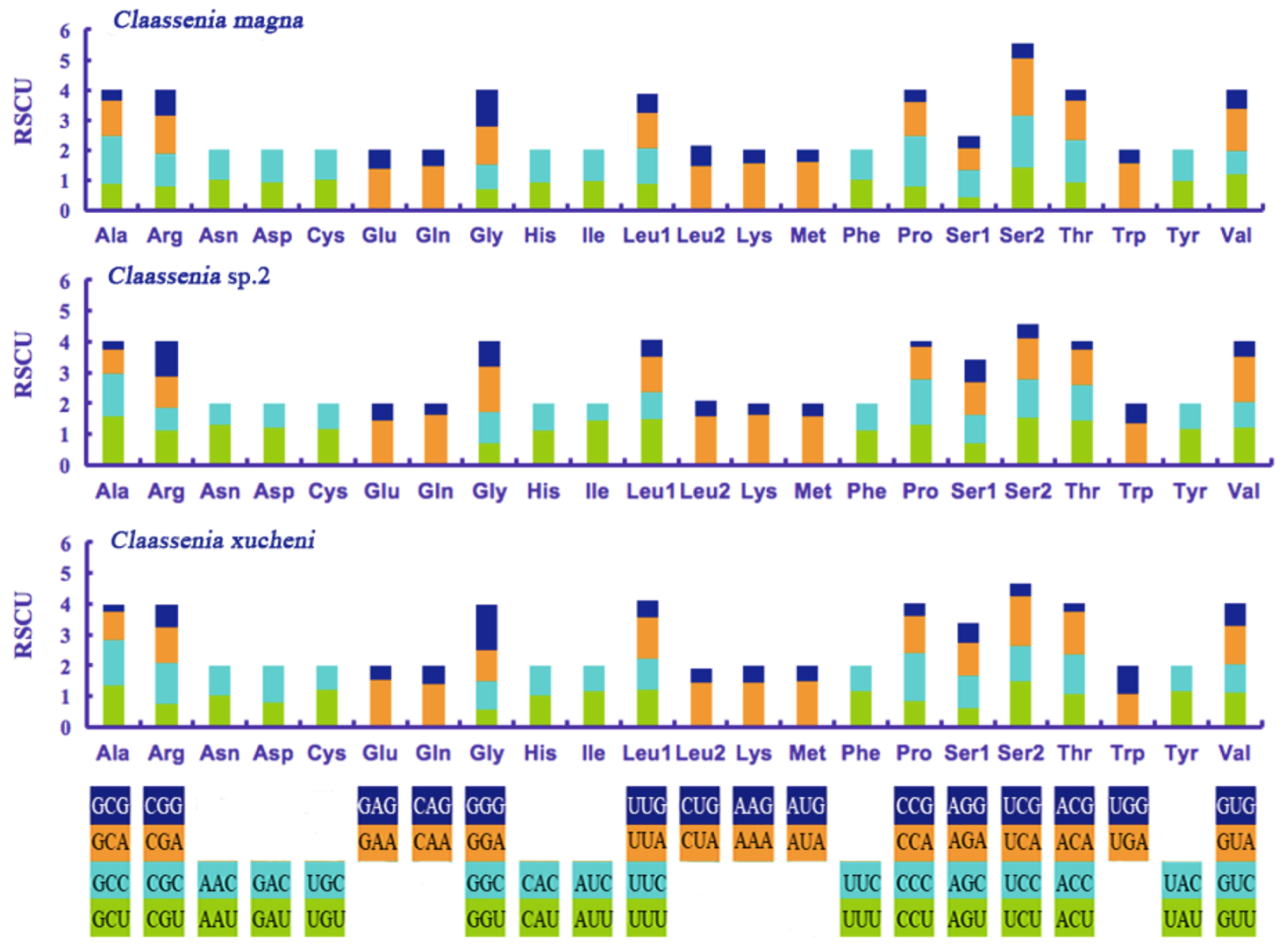
3.2. Protein-Coding Genes
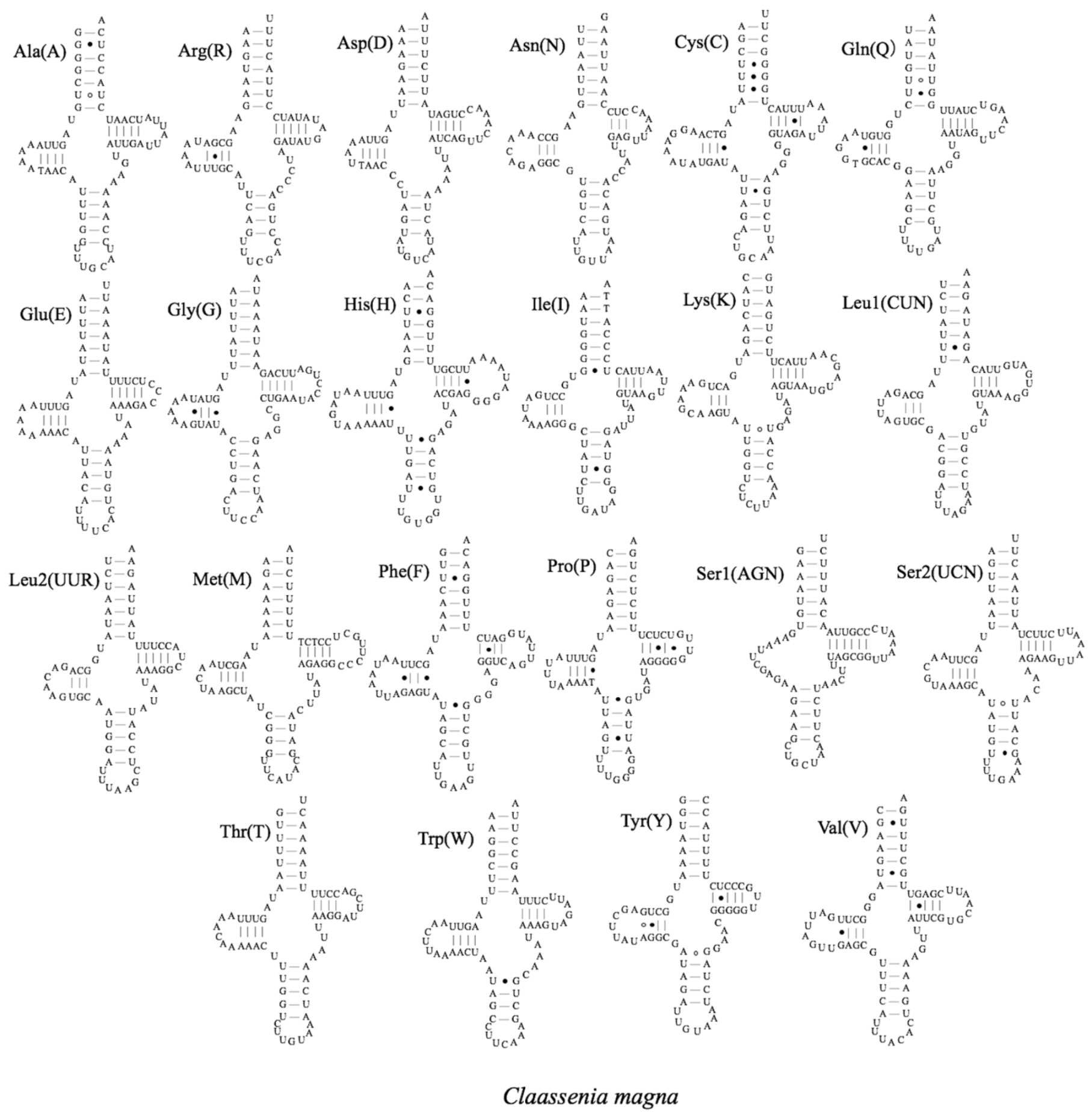
3.3. Transfer RNA Genes
3.4. Ribosomal RNA Genes
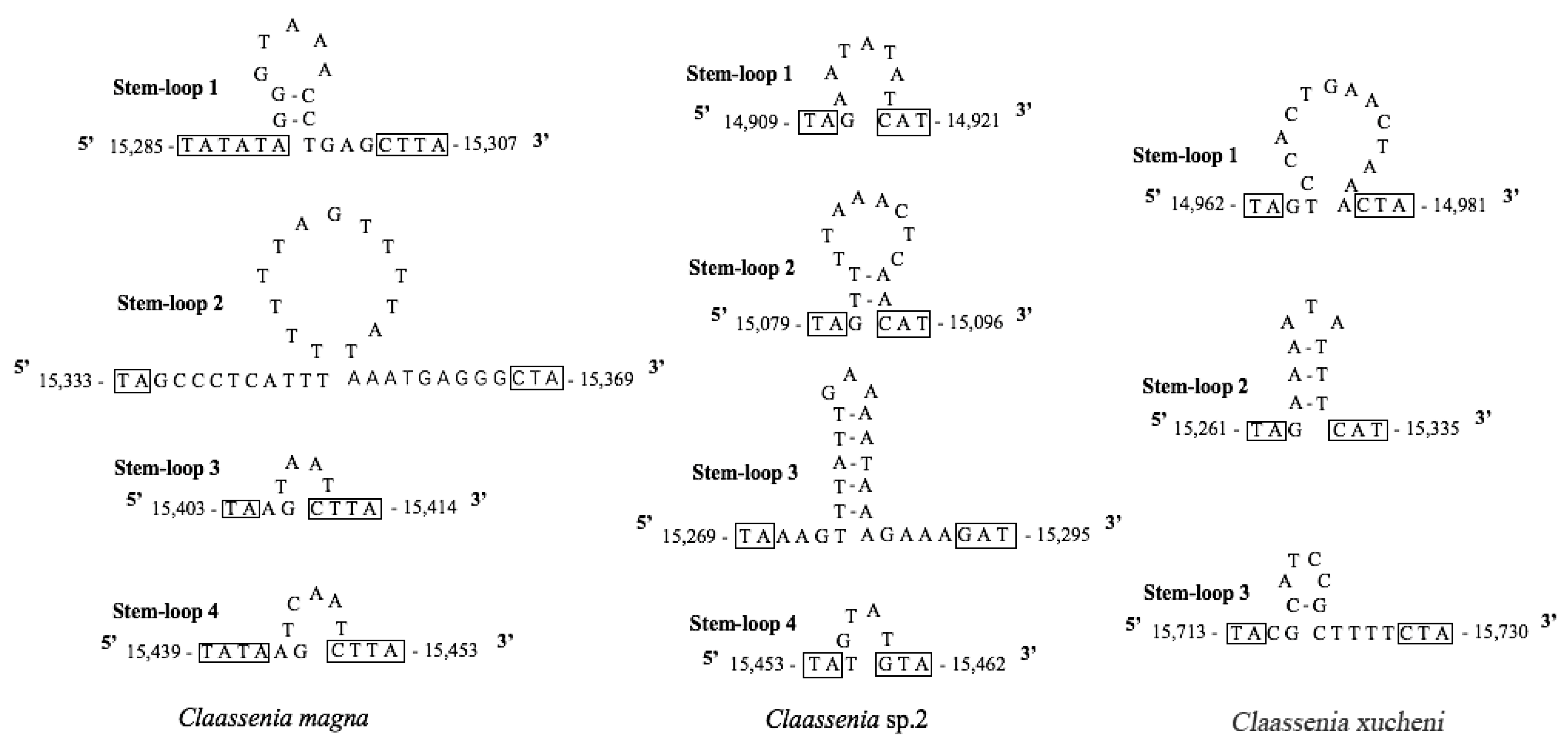
3.5. The Non-Coding Control Region
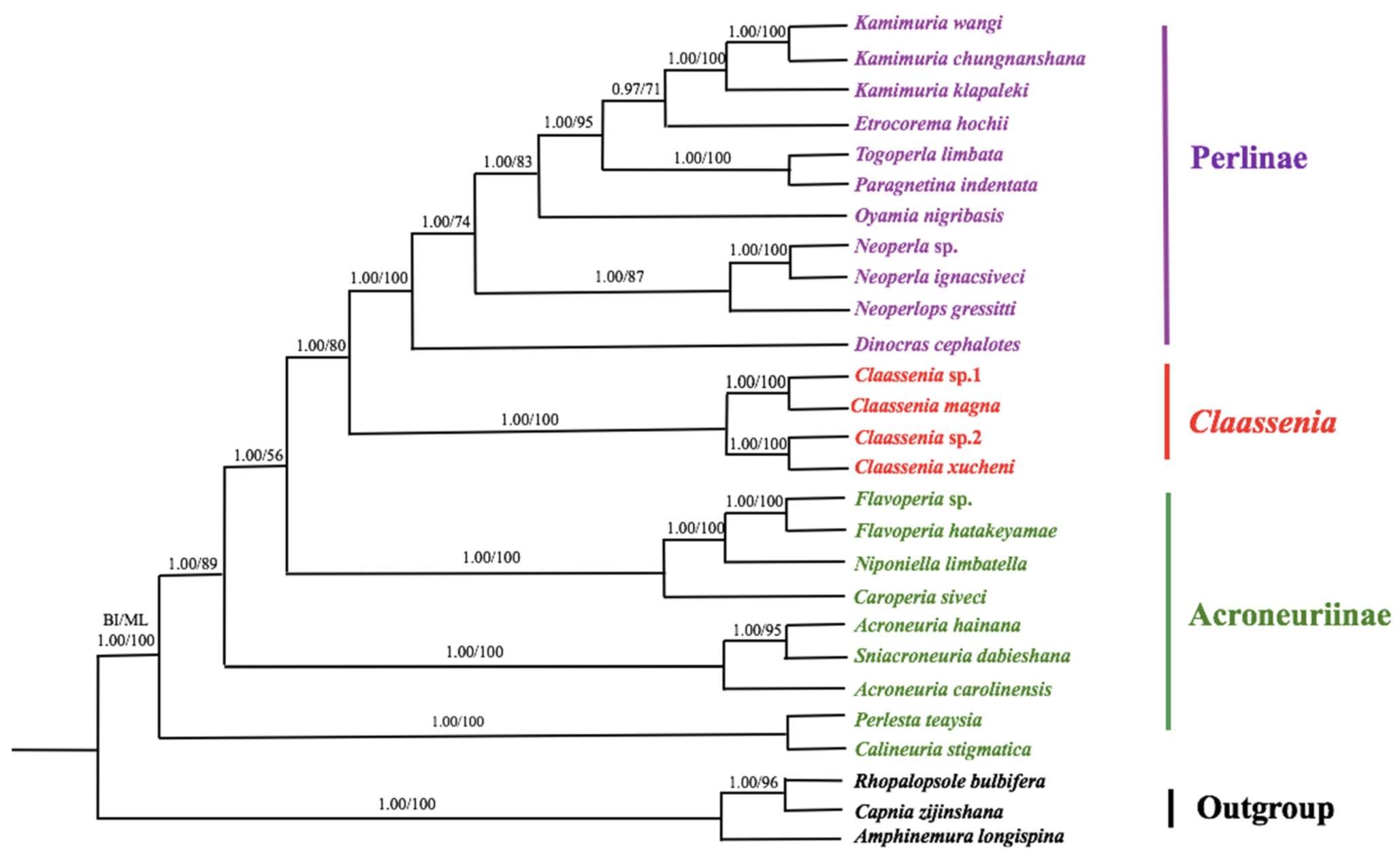
3.6. Phylogenetic Analyses
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cameron, S.L. Insect mitochondrial genomics: Implications for evolution and phylogeny. Annu. Rev. Entomol. 2014, 59, 95–117. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Stewart, J.B.; Beckenbach, A.T. Insect mitochondrial genomics: The complete mitochondrial genome sequence of a giant stonefly, Pteronarcys princeps, asymmetric directional mutation bias, and conserved plecopteran A+T-region elements. Genome 2006, 49, 815–824. [Google Scholar] [CrossRef] [PubMed]
- Qian, Y.H.; Wu, H.Y.; Ji, X.Y.; Yu, W.W.; Du, Y.Z. Mitochondrial genome of the stonefly Kamimuria wangi (Plecoptera: Perlidae) and phylogenetic position of Plecoptera based on mitogenomes. PLoS ONE 2014, 9, e86328. [Google Scholar]
- Wu, H.Y.; Ji, X.Y.; Yu, W.W.; Du, Y.Z. Complete mitochondrial genome of the stonefly Cryptoperla stilifera Sivec (Plecoptera: Peltoperlidae) and the phylogeny of Polyneopteran insects. Gene 2014, 537, 177–183. [Google Scholar] [CrossRef]
- Huang, M.C.; Wang, Y.Y.; Liu, X.Y.; Li, W.; Kang, Z.; Wang, K.; Li, X.; Yang, D. The complete mitochondrial genome and its remarkable secondary structure for a stonefly Acroneuria hainana Wu (insecta: Plecoptera, perlidae). Gene 2015, 557, 52–60. [Google Scholar] [CrossRef]
- Chen, Z.T.; Wu, H.Y.; Du, Y.Z. The nearly complete mitochondrial genome of a stonefly species, Styloperla sp. (Plecoptera: Styloperlidae). Mitochondrial DNA Part A 2016, 27, 2728–2729. [Google Scholar] [CrossRef]
- Wang, K.; Ding, S.; Yang, D. The complete mitochondrial genome of a stonefly species, Kamimuria chungnanshana Wu, 1948 (Plecoptera: Perlidae). Mitochondrial DNA Part A 2016, 27, 3810–3811. [Google Scholar] [CrossRef]
- Wang, K.; Wang, Y.; Yang, D. The complete mitochondrial genome of a stonefly species, Togoperla sp. (Plecoptera: Perlidae). Mitochondrial DNA Part A 2016, 27, 1703–1704. [Google Scholar]
- Yuan, M.L.; Zhang, Q.L.; Zhang, L.; Guo, Z.L.; Liu, Y.J.; Shen, Y.Y.; Shao, R. High-level phylogeny of the Coleoptera inferred with mitochondrial genome sequences. Mol. Phylogenet. Evol. 2016, 104, 99–111. [Google Scholar] [CrossRef]
- Li, H.; Leavengood, J.M.; Chapman, E.G.; Burkhardt, D.; Song, F.; Jiang, P.; Liu, J.; Zhou, X.; Cai, W. Mitochondrial phylogenomics of Hemiptera reveals adaptive innovations driving the diversification of true bugs. Proc. R. Soc. B. 2017, 284, 20171223. [Google Scholar] [CrossRef]
- Chen, Z.T.; Zhao, M.Y.; Xu, C.; Du, Y.Z. Molecular phylogeny of Systellognatha (Plecoptera: Arctoperlaria) inferred from mitochondrial genome sequences. Int. J. Biol. Macromol. 2018, 111, 542–547. [Google Scholar] [CrossRef]
- Shen, Y.; Du, Y.Z. The mitochondrial genome of Leuctra sp. (Plecoptera: Leuctridae) and its performance in phylogenetic analyses. Zootaxa 2019, 4671, 571–580. [Google Scholar] [CrossRef]
- Zhao, M.Y.; Huo, Q.B.; Du, Y.Z. Molecular phylogeny inferred from the mitochondrial genomes of Plecoptera with Oyamia nigribasis (Plecoptera: Perlidae). Sci. Rep. 2020, 10, 20955. [Google Scholar] [CrossRef]
- Simon, C.; Frati, F.; Beckenbach, A.; Crespi, B.; Liu, H.; Flook, P. Evolution, weighting, and phylogenetic utility of mitochondrial gene sequences and a compilation of conserved polymerase chain reaction primers. Ann. Entomol. Soc. Am. 1994, 87, 651–701. [Google Scholar] [CrossRef]
- Simon, C.; Buckley, T.R.; Frati, F.; Stewart, J.B.; Beckenbach, A.T. Incorporating molecular evolution into phylogenetic analysis, and a new compilation of conserved polymerase chain reaction primers for animal mitochondrial DNA. Annu. Rev. Ecol. Evol. Syst. 2006, 37, 545–579. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.T.; Du, Y.Z. The first two mitochondrial genomes from Taeniopterygidae (Insecta: Plecoptera): Structural features and phylogenetic implications. Int. J. Biol. Macromol. 2018, 111, 70–76. [Google Scholar] [CrossRef] [PubMed]
- DeWalt, R.E.; Maehr, M.D.; Neu-Becker, U.; Stueber, G. Plecoptera Species File Online. Available online: http://plecoptera.speciesfile.org/ (accessed on 7 November 2021).
- Sivec, I.; Stark, B.P.; Uchida, S. Synopsis of the world genera of Perlinae (Plecoptera: Perlidae). Scopolia 1988, 16, 1–66. [Google Scholar]
- Du, Y.Z. A preliminary study on the generic level phylogeny of the subfamily Perlinae (Plecoptera: Perlidae). J. Zhejiang Agric. Univ. 1999, 25, 170–174. [Google Scholar]
- Klapálek, F. Analytická tabulka fam. Perlidae a jeji dvou subfam., Perlinae a Acroneuriinae (Plecoptera). Časopis České Společnosti Entomol. 1914, 11, 53–69. [Google Scholar]
- Wu, C.F. A homonym of a plecopterous genus. Ann. Entomol. Soc. Am. 1934, 27, 256. [Google Scholar] [CrossRef]
- Stark, B.P.; Gaufin, A.R. The Nearctic Genera of Perlidae (Plecoptera); Entomological Society of America: Annapolis, MD, USA, 1976; Volume 10, pp. 1–80.
- Chen, M.D.; Cao, J.J.; Li, W.H.; Wang, Y. The mitochondrial genome from the stonefly: Claassenia sp. (Plecoptera: Perlidae). Mitochondrial DNA Part B 2019, 4, 3790–3791. [Google Scholar] [CrossRef] [Green Version]
- Du, Y.Z. Advances in Plecoptera systematics in China. Chin. J. Appl. Entomol. 2020, 57, 284–297. [Google Scholar]
- Wang, Y.; Cao, J.J.; Liu, Z.Z. The complete mitochondrial genome analysis of a stonefly, Acroneuria carolinensis (Plecoptera: Perlidae). Mitochondrial DNA Part B 2020, 5, 2777–2778. [Google Scholar] [CrossRef]
- Wang, Y.; Cao, J.J.; Chen, J.J.; Li, W.H. The mitochondrial genome of the stonefly Togoperla limbata Pictet (Plecoptera: Perlidae). Mitochondrial DNA Part B 2020, 5, 2812–2813. [Google Scholar] [CrossRef]
- Grant, J.R.; Stothard, P. The CGView server: A comparative genomics tool for circular genomes. Nucl. Acids Res. 2008, 36, 181–184. [Google Scholar] [CrossRef] [PubMed]
- Bernt, M.; Donath, A.; Jühling, F.; Externbrinka, F.; Florentz, C.; Fritzsch, G.; Pützh, J.; Middendorf, M.; Stadler, P.F. MITOS: Improved de novo metazoan mitochondrial genome annotation. Mol. Phylogenet. Evol. 2013, 69, 313–319. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [Green Version]
- Perna, N.T.; Kocher, T.D. Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J. Mol. Evol. 1995, 41, 353–358. [Google Scholar] [CrossRef] [PubMed]
- Vaidya, G.; Lohman, D.J.; Meier, R. SequenceMatrix: Concatenation software for the fast assembly of multi-gene datasets with character set and codon information. Cladistics 2010, 27, 171–180. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.T.; Schmidt, H.A.; Haeseler, A.V.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Gao, F.L.; Shen, J.G.; Liao, F.R.; Cai, W.; Lin, S.Q.; Yang, H.K.; Chen, S.L. The first complete genome sequence of narcissus latent virus from Narcissus. Arch. Virol. 2018, 163, 1383–1386. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.T.; Du, Y.Z. First mitochondrial genome from Nemouridae (Plecoptera) reveals novel features of the elongated control region and phylogenetic implications. Int. J. Mol. Sci. 2017, 18, 996. [Google Scholar] [CrossRef]
- Clary, D.O.; Wolstenholme, D.R. The ribosomal RNA genes of Drosophila mitochondrial DNA. Nucl. Acids. Res. 1985, 13, 4029–4045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ojala, D.; Montoya, J.; Attardi, G. tRNA punctuation model of RNA processing in human mitochondria. Nature 1981, 290, 470–474. [Google Scholar] [CrossRef] [PubMed]
- Garey, J.R.; Wolstenholme, D.R. Platyhelminth mitochondrial DNA: Evidence for early evolutionary origin of a tRNAserAGN that contains a dihydrouridine arm replacement loop, and of serine-specifying AGA and AGG codons. J. Mol. Evol. 1989, 28, 374–387. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cao, J.J.; Chen, J.J.; Li, W.H. The mitochondrial genome of the stonefly Togoperla limbata Pictet (Plecoptera: Perlidae). Mitochondrial DNA Part B 2020, 5, 2812–2813. [Google Scholar] [CrossRef]
- Duran, D.P.; Laroche, R.A.; Gough, H.M.; Gwiazdowski, R.A.; Knisley, C.B.; Herrmann, D.P.; Roman, S.J.; Egan, S.P. Geographic life history differences predict genomic divergence better than mitochondrial barcodes or phenotype. Genes 2020, 11, 265. [Google Scholar] [CrossRef] [Green Version]
| Order | Subfamily | Species | GenBank Accession No. |
|---|---|---|---|
| Plecoptera | Perlinae | Dinocras cephalotes | KF484757 |
| Neoperla sp. FS-2017 | KX091859 | ||
| Neoperla ignacsiveci | KX091858 | ||
| Neoperlops gressitti | MN400756 | ||
| Oyamia nigribasis | MN548290 | ||
| Kamimuria wangi | KC894944 | ||
| Kamimuria chungnanshana | KT186102 | ||
| Kamimuria klapaleki | MN400755 | ||
| Paragnetina indentata | MN627431 | ||
| Togoperla limbata | MN969990 | ||
| Etrocorema hochii | MK905888 | ||
| Claassenia sp. YW-2019 | MN419914 | ||
| Claassenia magna | OK012602 | ||
| Claassenia sp. 2 | OK021652 | ||
| Claassenia xucheni | OK021653 | ||
| Acroneuriinae | Sniacroneuria dabieshana | MK492253 | |
| Acroneuria hainana | KM199685 | ||
| Acroneuria carolinensis | MN969989 | ||
| Perlesta teaysia | MN627432 | ||
| Calineuria stigmatica | MG677941 | ||
| Flavoperla sp. YZD-2020 | MK905206 | ||
| Flavoperia hatakeyamae | MN821010 | ||
| Niponiella limbatella | MK686067 | ||
| Caroperia siveci | MG677942 | ||
| Leuctrinae | Rhopalopsole bulbifera | MK111419 | |
| Nemouroidea | Capnia zijinshana | KX094942 | |
| Amphinemurinae | Amphinemura longispina | MH085446 |
| Species | Whole Genome | PCGs | tRNAs | rRNAs | Control Region | |||||
|---|---|---|---|---|---|---|---|---|---|---|
| Size(bp) | A+T (%) | Size (bp) | A+T (%) | Size (bp) | A+T (%) | Size (bp) | A+T (%) | Size (bp) | A+T (%) | |
| Claassenia magna | 15,774 | 61.46 | 11,177 | 59.40 | 1493 | 66.98 | 2201 | 64.15 | 832 | 71.51 |
| Claassenia sp. 2 | 15,777 | 65.81 | 11,232 | 64.12 | 1492 | 68.70 | 2222 | 69.04 | 869 | 74.45 |
| Claasseniaxucheni | 15,746 | 62.89 | 11,139 | 60.62 | 1491 | 67.40 | 2204 | 67.06 | 832 | 73.95 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xiang, Y.; Zhao, M.; Huo, Q.; Du, Y. Mitochondrial Genomes of the Genus Claassenia (Plecoptera: Perlidae) and Phylogenetic Assignment to Subfamily Perlinae. Genes 2021, 12, 1986. https://doi.org/10.3390/genes12121986
Xiang Y, Zhao M, Huo Q, Du Y. Mitochondrial Genomes of the Genus Claassenia (Plecoptera: Perlidae) and Phylogenetic Assignment to Subfamily Perlinae. Genes. 2021; 12(12):1986. https://doi.org/10.3390/genes12121986
Chicago/Turabian StyleXiang, Yanan, Mengyuan Zhao, Qingbo Huo, and Yuzhou Du. 2021. "Mitochondrial Genomes of the Genus Claassenia (Plecoptera: Perlidae) and Phylogenetic Assignment to Subfamily Perlinae" Genes 12, no. 12: 1986. https://doi.org/10.3390/genes12121986
APA StyleXiang, Y., Zhao, M., Huo, Q., & Du, Y. (2021). Mitochondrial Genomes of the Genus Claassenia (Plecoptera: Perlidae) and Phylogenetic Assignment to Subfamily Perlinae. Genes, 12(12), 1986. https://doi.org/10.3390/genes12121986






