Targeted Sequencing of Mitochondrial Genes Reveals Signatures of Molecular Adaptation in a Nearly Panmictic Small Pelagic Fish Species
Abstract
1. Introduction
2. Methods
2.1. Sample Collection and DNA Extraction
2.2. Mitochondrial DNA: Defining Target-Regions for Amplicon Sequencing
2.2.1. Sequencing, Filtering and Variant Calling at Targeted Mitochondrial Regions
2.2.2. Diversity and Distribution of Genetic Variation at Targeted Mitochondrial Regions
2.2.3. Relationship between Haplotype Composition and Environmental Variables at Each Location
2.2.4. Inferring Historical Events of Selection across Phylogenies with dN/dS Ratios-Based Tests
2.3. Microsatellite Amplification and Diversity Estimates
2.3.1. Estimates of Genetic Differentiation and Structure among Sampled Locations
2.3.2. Inferences on Deviations from Evolutionary Neutrality
3. Results
3.1. Mitochondrial DNA Sequencing and Variant Calling
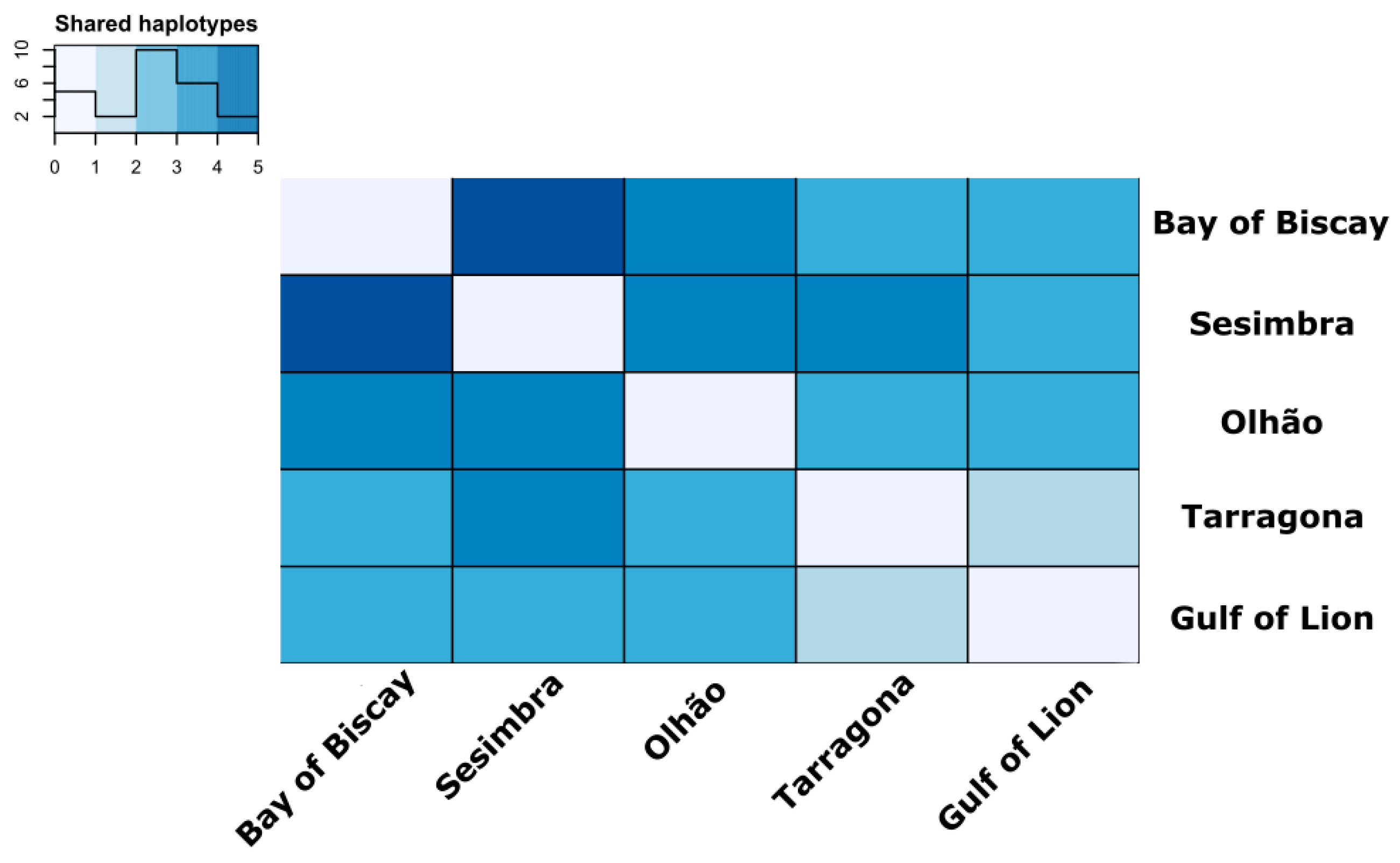
3.2. Estimates of Mitochondrial Genetic Diversity and Population Structure
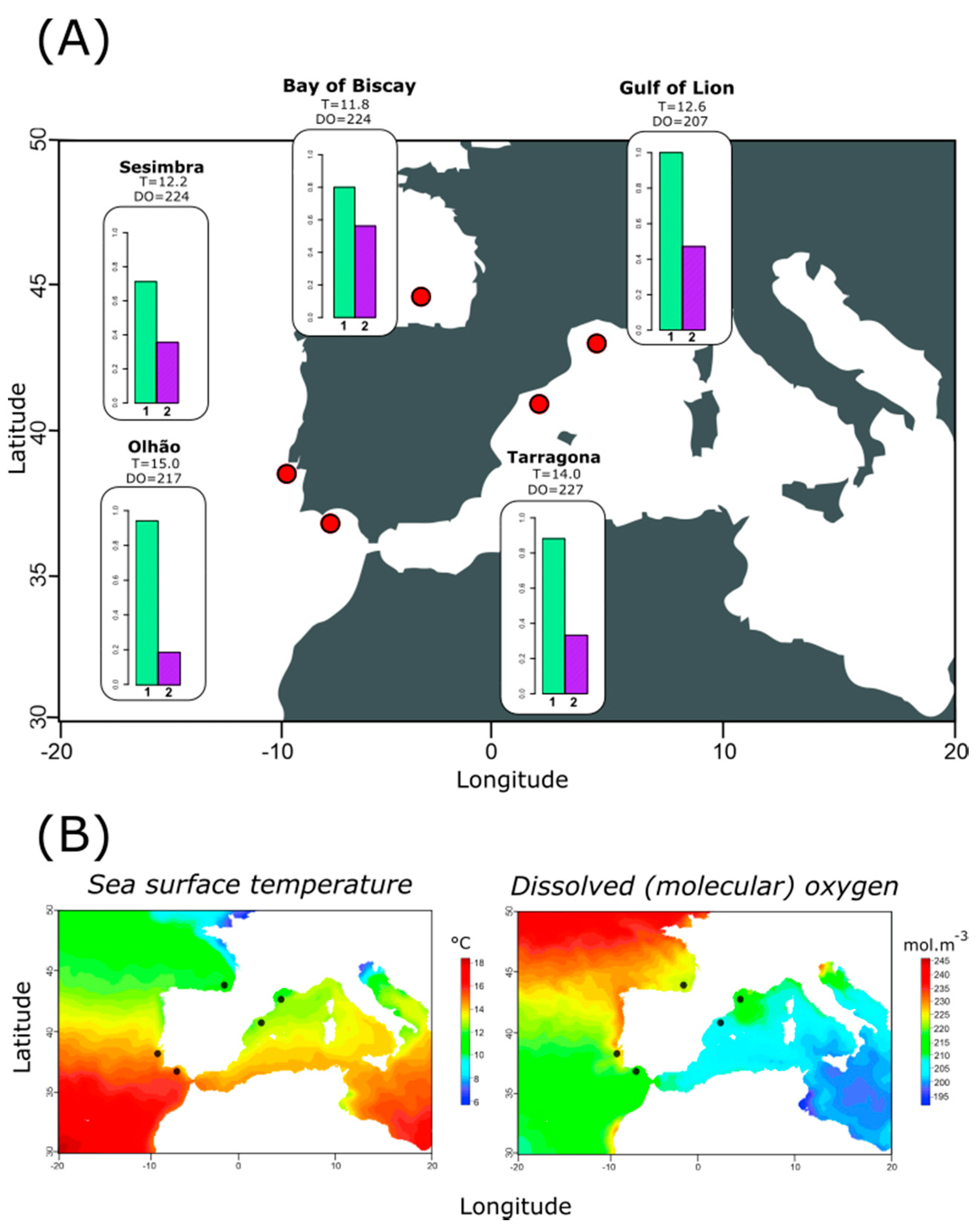
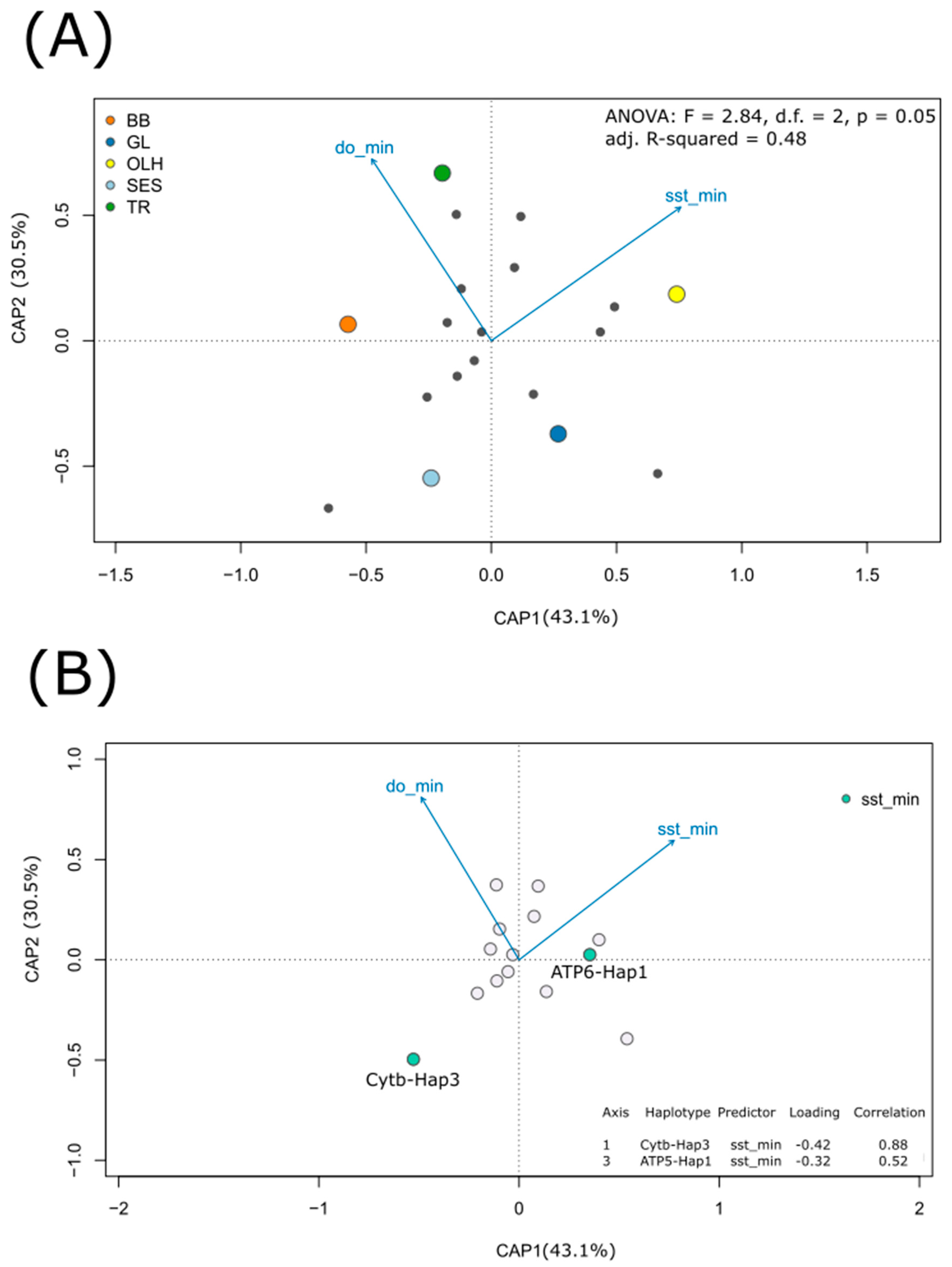
3.3. Relating Haplotype Frequencies and Environmental Variation
3.4. Inferences of Episodic Selection across the Phylogeny of Sampled Specimens
3.5. Microsatellite Amplification and Diversity Estimates
3.6. Estimates of Microsatellite Genetic Differentiation and Structure among Sampled Locations
4. Discussion
4.1. Population Structure Reveals High Connectivity around Iberian Peninsula
4.2. Temperature-Dependent Distribution of Most Frequently Observed Haplotypes
5. Conclusions and Future
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
Appendix A.1. Library Preparation and Amplicon Sequencing Protocol
Appendix A.2. Amplification and Genotyping of Microsatellite Loci
References
- Calosi, P.; De Wit, P.; Thor, P.; Dupont, S. Will life find a way? Evolution of marine species under global change. Evol. Appl. 2016, 9, 1035–1042. [Google Scholar] [CrossRef] [PubMed]
- Costa, B.H.E.; Assis, J.; Franco, G.; Erzini, K.; Henriques, M.; Gonçalves, E.; Caselle, J.; Gonçalves, E.J. Tropicalization of fish assemblages in temperate biogeographic transition zones. Mar. Ecol. Prog. Ser. 2014, 504, 241–252. [Google Scholar] [CrossRef]
- Vergés, A.; Steinberg, P.D.; Hay, M.E.; Poore, A.G.B.; Campbell, A.H.; Ballesteros, E.; Heck, K.L.; Booth, D.J.; Coleman, M.A.; Feary, D.A.; et al. The tropicalization of temperate marine ecosystems: Climate-mediated changes in herbivory and community phase shifts. Proc. R. Soc. B Biol. Sci. 2014, 281, 20140846. [Google Scholar] [CrossRef] [PubMed]
- Melzner, F.; Thomsen, J.; Koeve, W.; Oschlies, A.; Gutowska, M.A.; Bange, H.W.; Hansen, H.P.; Körtzinger, A. Future ocean acidification will be amplified by hypoxia in coastal habitats. Mar. Biol. 2012, 160, 1875–1888. [Google Scholar] [CrossRef]
- Altieri, A.H.; Gedan, K.B. Climate change and dead zones. Glob. Chang. Biol. 2014, 21, 1395–1406. [Google Scholar] [CrossRef] [PubMed]
- Eizaguirre, C.; Baltazar-Soares, M. Evolutionary conservation—Evaluating the adaptive potential of species. Evol. Appl. 2014, 7, 963–967. [Google Scholar] [CrossRef]
- Nielsen, E.E.; Hemmer-Hansen, J.; Larsen, P.F.; Bekkevold, D. Population genomics of marine fishes: Identifying adaptive variation in space and time. Mol. Ecol. 2009, 18, 3128–3150. [Google Scholar] [CrossRef]
- Grummer, J.A.; Beheregaray, L.B.; Bernatchez, L.; Hand, B.K.; Luikart, G.; Narum, S.R.; Taylor, E.B. Aquatic Landscape Genomics and Environmental Effects on Genetic Variation. Trends Ecol. Evol. 2019, 34, 641–654. [Google Scholar] [CrossRef]
- Selkoe, K.A.; Henzler, C.M.; Gaines, S.D. Seascape genetics and the spatial ecology of marine populations. Fish Fish. 2008, 9, 363–377. [Google Scholar] [CrossRef]
- Hutchinson, W.F. The dangers of ignoring stock complexity in fishery management: The case of the North Sea cod. Biol. Lett. 2008, 4, 693–695. [Google Scholar] [CrossRef]
- Ernande, B.; Dieckmann, U.; Heino, M. Adaptive changes in harvested populations: Plasticity and evolution of age and size at maturation. Proc. R. Soc. B Biol. Sci. 2004, 271, 415–423. [Google Scholar] [CrossRef] [PubMed]
- Dunlop, E.S.; Eikeset, A.M.; Stenseth, N.C. From genes to populations: How fisheries-induced evolution alters stock productivity. Ecol. Appl. 2015, 25, 1860–1868. [Google Scholar] [CrossRef] [PubMed]
- Spielman, D.; Brook, B.W.; Frankham, R. Most species are not driven to extinction before genetic factors impact them. Proc. Natl. Acad. Sci. USA 2004, 101, 15261–15264. [Google Scholar] [CrossRef] [PubMed]
- Deyle, E.R.; Fogarty, M.; Hsieh, C.-H.; Kaufman, L.; MacCall, A.D.; Munch, S.B.; Perretti, C.T.; Ye, H.; Sugihara, G. Predicting climate effects on Pacific sardine. Proc. Natl. Acad. Sci. USA 2013, 110, 6430–6435. [Google Scholar] [CrossRef]
- Cury, P.; Bakun, A.; Crawford, R.J.M.; Jarre, A.; Quiñones, R.A.; Shannon, L.J.; Verheye, H.M. Small pelagics in upwelling systems: Patterns of interaction and structural changes in “wasp-waist” ecosystems. ICES J. Mar. Sci. 2000, 57, 603–618. [Google Scholar] [CrossRef]
- Rykaczewski, R.R.; Checkley, D.M. Influence of ocean winds on the pelagic ecosystem in upwelling regions. Proc. Natl. Acad. Sci. USA 2008, 105, 1965–1970. [Google Scholar] [CrossRef]
- Koslow, J.A.; Davison, P.; Lara-Lopez, A.; Ohman, M.D. Epipelagic and mesopelagic fishes in the southern California Current System: Ecological interactions and oceanographic influences on their abundance. J. Mar. Syst. 2014, 138, 20–28. [Google Scholar] [CrossRef]
- Yamaguchi, A.; Matsuno, K.; Abe, Y.; Arima, D.; Imai, I. Latitudinal variations in the abundance, biomass, taxonomic composition and estimated production of epipelagic mesozooplankton along the 155° E longitude in the western North Pacific during spring. Prog. Oceanogr. 2017, 150, 13–19. [Google Scholar] [CrossRef]
- Duarte, L.O.; Garcia, C. Trophic role of small pelagic fishes in a tropical upwelling ecosystem. Ecol. Model. 2004, 172, 323–338. [Google Scholar] [CrossRef]
- González-García, S.; Villanueva-Rey, P.; Belo, S.; Vázquez-Rowe, I.; Moreira, M.T.; Feijoo, G.; Arroja, L. Cross-vessel eco-efficiency analysis. A case study for purse seining fishing from North Portugal targeting Euro-pean pilchard. Int. J. Life Cycle Assess. 2015, 20, 1019–1032. [Google Scholar] [CrossRef]
- Vázquez-Rowe, I.; Villanueva-Rey, P.; Hospido, A.; Moreira, M.T.; Feijoo, G. Life cycle assessment of European pilchard (Sardina pilchardus) consumption. A case study for Galicia (NW Spain). Sci. Total Environ. 2014, 475, 48–60. [Google Scholar] [CrossRef] [PubMed]
- Jemaa, S.; Bacha, M.; Khalaf, G.; Dessailly, D.; Rabhi, K.; Amara, R. What can otolith shape analysis tell us about population structure of the European sardine, Sardina pilchardus, from Atlantic and Mediterranean waters? J. Sea Res. 2015, 96, 11–17. [Google Scholar] [CrossRef]
- Antonakakis, K.; Giannoulaki, M.; Machias, A.; Somarakis, S.; Sanchez, S.; Ibaibarriaga, L.; Uriarte, A. Assessment of the sardine (Sardina pilchardus Walbaum, 1792) fishery in the eastern Mediterranean basin (North Aegean Sea). Mediterr. Mar. Sci. 2011, 12, 333–357. [Google Scholar] [CrossRef]
- Letts, J.A.; Fiedorczuk, K.; Sazanov, J.A. The architecture of respiratory supercomplexes. Nat. Cell Biol. 2016, 537, 644–648. [Google Scholar] [CrossRef] [PubMed]
- Formosa, L.E.; Ryan, M.T. Mitochondrial OXPHOS complex assembly lines. Nat. Cell Biol. 2018, 20, 511–513. [Google Scholar] [CrossRef] [PubMed]
- Consuegra, S.; John, E.; Verspoor, E.; de Leaniz, C.G. Patterns of natural selection acting on the mitochondrial genome of a locally adapted fish species. Genet. Sel. Evol. 2015, 47, 58. [Google Scholar] [CrossRef]
- Silva, G.; Lima, F.P.; Martel, P.; Castilho, R. Thermal adaptation and clinal mitochondrial DNA variation of European anchovy. Proc. R. Soc. B Biol. Sci. 2014, 281, 20141093. [Google Scholar] [CrossRef]
- Sebastian, W.; Sukumaran, S.; Zacharia, P.; Muraleedharan, K.; Kumar, P.D.; Gopalakrishnan, A. Signals of selection in the mitogenome provide insights into adaptation mechanisms in heterogeneous habitats in a widely distributed pelagic fish. Sci. Rep. 2020, 10, 1–14. [Google Scholar]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. In Nucleic Acids Symposium Series; Information Retrieval Ltd.: London, UK, 1999; Volume 41, pp. 95–98. [Google Scholar]
- Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv 2013, arXiv:1303.3997. [Google Scholar]
- Li, H.; Durbin, R. Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 2010, 26, 589–595. [Google Scholar] [CrossRef]
- Catchen, J.; Hohenlohe, P.A.; Bassham, S.; Amores, A.; Cresko, W.A. Stacks: An analysis tool set for population genomics. Mol. Ecol. 2013, 22, 3124–3140. [Google Scholar] [CrossRef] [PubMed]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef]
- Excoffier, L.; Lischer, H.E.L. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef] [PubMed]
- Goslee, S.C.; Urban, D.L. The ecodist package for dissimilarity-based analysis of ecological data. J. Stat. Softw. 2007, 22, 1–19. [Google Scholar] [CrossRef]
- Leigh, J.W.; Bryant, D. popart: Full-feature software for haplotype network construction. Methods Ecol. Evol. 2015, 6, 1110–1116. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- Assis, J.; Tyberghein, L.; Bosch, S.; Verbruggen, H.; Serrão, E.A.; De Clerck, O. Bio-ORACLE v2.0: Extending marine data layers for bioclimatic modelling. Glob. Ecol. Biogeogr. 2018, 27, 277–284. [Google Scholar] [CrossRef]
- Rao, C.R. The use and interpretation of principal component analysis in applied research. Sankhyā Indian J. Stat. Ser. A 1964, 26, 329–358. [Google Scholar]
- Wickham, H.; Francois, R.; Henry, L.; Müller, K. Vegan: Community Ecology Package. R package version 1.17-4. 2010. Available online: https://cran.r-project.org/web/packages/vegan/index.html (accessed on 15 July 2020).
- Legendre, P.; Oksanen, J.; ter Braak, C.J.F. Testing the significance of canonical axes in redundancy analysis. Methods Ecol. Evol. 2011, 2, 269–277. [Google Scholar] [CrossRef]
- Weaver, S.; Shank, S.D.; Spielman, S.J.; Li, M.; Muse, S.V.; Kosakovsky Pond, S.L. Datamonkey 2.0: A Mod-ern Web Application for Characterizing Selective and Other Evolutionary Processes. Mol. Biol. Evol. 2018, 35, 773–777. [Google Scholar] [CrossRef]
- Pond, S.L.K.; Frost, S.D.W. Not So Different After All: A Comparison of Methods for Detecting Amino Acid Sites Under Selection. Mol. Biol. Evol. 2005, 22, 1208–1222. [Google Scholar] [CrossRef]
- Murrell, B.; Moola, S.; Mabona, A.; Weighill, T.; Sheward, D.; Pond, S.L.K.; Scheffler, K. FUBAR: A Fast, Unconstrained Bayesian AppRoximation for Inferring Selection. Mol. Biol. Evol. 2013, 30, 1196–1205. [Google Scholar] [CrossRef] [PubMed]
- Ruggeri, P.; Splendiani, A.; Bonanomi, S.; Arneri, E.; Cingolani, N.; Santojanni, A.; Belardinelli, A.; Giovannotti, M.; Barucchi, V.C. Temporal genetic variation as revealed by a microsatellite analysis of European sardine (Sardina pilchardus) archived samples. Can. J. Fish. Aquat. Sci. 2012, 69, 1698–1709. [Google Scholar] [CrossRef]
- Kasapidis, P.; Silva, A.; Zampicinini, G.; Magoulas, A. Evidence for microsatellite hitchhiking selection in European sardine (Sardina pilchardus) and implications in inferring stock structure. Sci. Mar. 2011, 76, 123–132. [Google Scholar] [CrossRef]
- Kalinowski, S.T. hp-rare 1.0: A computer program for performing rarefaction on measures of allelic richness. Mol. Ecol. Notes 2005, 5, 187–189. [Google Scholar] [CrossRef]
- Van Oosterhout, C.; Hutchinson, W.F.; Wills, D.P.M.; Shipley, P. micro-checker: Software for identifying and correcting genotyping errors in microsatellite data. Mol. Ecol. Notes 2004, 4, 535–538. [Google Scholar] [CrossRef]
- Jombart, T. adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- Frankham, R.; Briscoe, D.A.; Ballou, J.D. Introduction to Conservation Genetics; Cambridge University Press: Cambridge, UK, 2002. [Google Scholar]
- Baibai, T.; Oukhattar, L.; Quinteiro, J.V.; Mesfioui, A.; Rey-Mendez, M. First global approach: Morphological and biological variability in a genetically homogeneous population of the European pilchard, Sardina pilchardus (Walbaum, 1792) in the North Atlantic coast. Rev. Fish Biol. Fish. 2012, 22, 63–80. [Google Scholar] [CrossRef]
- Silva, A.; Carrera, P.; Masse, J.; Uriarte, A.; Santos, M.; Oliveira, P.B.; Soares, E.; Porteiro, C.; Stratoudakis, Y. Geographic variability of sardine growth across the northeastern Atlantic and the Mediterranean Sea. Fish. Res. 2008, 90, 56–69. [Google Scholar] [CrossRef]
- Van Beveren, E.; Fromentin, J.-M.; Rouyer, T.; Bonhommeau, S.; Brosset, P.; Saraux, C. The fisheries history of small pelagics in the Northern Mediterranean. ICES J. Mar. Sci. 2016, 73, 1474–1484. [Google Scholar] [CrossRef]
- Pinsky, M.L.; Palumbi, S.R. Meta-analysis reveals lower genetic diversity in overfished populations. Mol. Ecol. 2014, 23, 29–39. [Google Scholar] [CrossRef]
- Ramon, M.; Castro, J. Genetic variation in natural stocks of Sardina pilchardus (sardines) from the western Mediterranean Sea. Heredity 1997, 78, 520–528. [Google Scholar] [CrossRef]
- Correia, A.T.; Hamer, P.; Carocinho, B.; Silva, A. Evidence for meta-population structure of Sardina pilchardus in the Atlantic Iberian waters from otolith elemental signatures of a strong cohort. Fish. Res. 2014, 149, 76–85. [Google Scholar] [CrossRef]
- Bakun, A.; Black, B.A.; Bograd, S.J.; Garcia-Reyes, M.; Miller, A.J.; Rykaczewski, R.R.; Sydeman, W.J. Anticipated effects of climate change on coastal upwelling ecosystems. Curr. Clim. Chang. Rep. 2015, 1, 85–93. [Google Scholar] [CrossRef]
- Miranda, P.; Alves, J.; Serra, N. Climate change and upwelling: Response of Iberian upwelling to atmospheric forcing in a regional climate scenario. Clim. Dyn. 2013, 40, 2813–2824. [Google Scholar] [CrossRef]
- Avery, P.J. The population genetics of haplo-diploids and X-linked genes. Genet. Res. 1984, 44, 321–341. [Google Scholar] [CrossRef]
- Levin, L.; Zhidkov, I.; Gurman, Y.; Hawlena, H.; Mishmar, D. Functional Recurrent Mutations in the Human Mitochondrial Phylogeny: Dual Roles in Evolution and Disease. Genome Biol. Evol. 2013, 5, 876–890. [Google Scholar] [CrossRef] [PubMed]
- Chamary, J.V.; Hurst, L.D. Evidence for selection on synonymous mutations affecting stability of mRNA secondary structure in mammals. Genome Biol. 2005, 6, R75. [Google Scholar] [CrossRef]
- Drummond, D.A.; Wilke, C.O. Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution. Cell 2008, 134, 341–352. [Google Scholar] [CrossRef]
- Castellana, S.; Vicario, S.; Saccone, C. Evolutionary Patterns of the Mitochondrial Genome in Metazoa: Exploring the Role of Mutation and Selection in Mitochondrial Protein–Coding Genes. Genome Biol. Evol. 2011, 3, 1067–1079. [Google Scholar] [CrossRef]
- Uddin, A.; Chakraborty, S. Synonymous codon usage pattern in mitochondrial CYB gene in pisces, aves, and mammals. Mitochondrial DNA Part A 2015, 28, 187–196. [Google Scholar] [CrossRef]
- Cvijović, I.; Good, B.H.; Desai, M.M. The Effect of Strong Purifying Selection on Genetic Diversity. Genetics 2018, 209, 1235–1278. [Google Scholar] [CrossRef] [PubMed]
- Barreto, F.S.; Watson, E.T.; Lima, T.G.; Willett, C.S.; Edmands, S.; Li, W.; Burton, R.S. Genomic signatures of mitonuclear coevolution across populations of Tigriopus californicus. Nat. Ecol. Evol. 2018, 2, 1250–1257. [Google Scholar] [CrossRef] [PubMed]
- Hill, G.E. Mitonuclear Compensatory Coevolution. Trends Genet. 2020, 36, 403–414. [Google Scholar] [CrossRef] [PubMed]
- Alheit, J.; Pohlmann, T.; Casini, M.; Greve, W.; Hinrichs, R.; Mathis, M.; O’Driscoll, K.; Vorberg, R.; Wagner, C. Climate variability drives anchovies and sardines into the North and Baltic Seas. Prog. Oceanogr. 2012, 96, 128–139. [Google Scholar] [CrossRef]
- Schickele, A.; Goberville, E.; Leroy, B.; Beaugrand, G.; Hattab, T.; Francour, P.; Raybaud, V. European small pelagic fish distribution under global change scenarios. Fish Fish. 2021, 22, 212–225. [Google Scholar] [CrossRef]
- Schulte, P.M.; Healy, T.M.; Fangue, N.A. Thermal Performance Curves, Phenotypic Plasticity, and the Time Scales of Temperature Exposure. Integr. Comp. Biol. 2011, 51, 691–702. [Google Scholar] [CrossRef]
- Yampolsky, L.Y.; Schaer, T.M.M.; Ebert, D. Adaptive phenotypic plasticity and local adaptation for temperature tolerance in freshwater zooplankton. Proc. R. Soc. B Biol. Sci. 2014, 281, 20132744. [Google Scholar] [CrossRef]
- McCaw, B.A.; Stevenson, T.J.; Lancaster, L.T. Epigenetic Responses to Temperature and Climate. Integr. Comp. Biol. 2020, 60, 1469–1480. [Google Scholar] [CrossRef]
- Best, C.; Ikert, H.; Kostyniuk, D.J.; Craig, P.M.; Navarro-Martin, L.; Marandel, L.; Mennigen, J.A. Epigenetics in teleost fish: From molecular mechanisms to physiological phenotypes. Comp. Biochem. Physiol. Part B Biochem. Mol. Biol. 2018, 224, 210–244. [Google Scholar] [CrossRef]
- Heckwolf, M.J.; Meyer, B.S.; Häsler, R.; Höppner, M.P.; Eizaguirre, C.; Reusch, T.B. Two different epigenetic information channels in wild three-spined sticklebacks are involved in salinity adaptation. Sci. Adv. 2020, 6, eaaz1138. [Google Scholar] [CrossRef]
- Binladen, J.; Gilbert, M.T.P.; Bollback, J.P.; Panitz, F.; Bendixen, C.; Nielsen, R.; Willerslev, E. The Use of Coded PCR Primers Enables High-Throughput Sequencing of Multiple Homolog Amplification Products by 454 Parallel Sequencing. PLoS ONE 2007, 2, e197. [Google Scholar] [CrossRef] [PubMed]
- McInnes, J.C.; Jarman, S.N.; Lea, M.-A.; Raymond, B.; Deagle, B.E.; Phillips, R.A.; Catry, P.; Stanworth, A.; Weimerskirch, H.; Kusch, A. DNA metabarcoding as a marine conservation and management tool: A circumpolar examination of fishery discards in the diet of threatened albatrosses. Front. Mar. Sci. 2017, 4, 277. [Google Scholar] [CrossRef]
| Gene | Population | n | S | nHap | Hd | π | Tajima’s-D |
|---|---|---|---|---|---|---|---|
| ATP6 Respiratory Complex V | Bay of Biscay | 15 | 3 | 4 | 0.371 | 0.00135 | −1.31654 |
| Gulf of Lion | 15 | 0 | 1 | 0 | 0 | na | |
| Olhão | 18 | 2 | 2 | 0.11 | 0.00058 | −1.50776 | |
| Sesimbra | 14 | 3 | 4 | 0.396 | 0.0012 | −1.67053 | |
| Tarragona | 17 | 0 | 1 | 0 | 0 | na | |
| COI Respiratory Complex IV | Bay of Biscay | 17 | 2 | 3 | 0.228 | 0.00081 | −1.50358 |
| Gulf of Lion | 20 | 1 | 2 | 0.1 | 0.00034 | −1.16439 | |
| Olhão | 15 | 1 | 2 | 0.133 | 0.00046 | −1.15945 | |
| Sesimbra | 14 | 1 | 2 | 0.143 | 0.00049 | −1.15524 | |
| Tarragona | 18 | 2 | 3 | 0.216 | 0.00076 | −1.50776 | |
| CYTB Respiratory Complex III | Bay of Biscay | 16 | 3 | 4 | 0.642 | 0.0051 | 0.38767 |
| Gulf of Lion | 17 | 3 | 4 | 0.676 | 0.00563 | 0.78185 | |
| Olhão | 16 | 4 | 5 | 0.717 | 0.00589 | −0.05743 | |
| Sesimbra | 14 | 4 | 5 | 0.769 | 0.00716 | 0.476 | |
| Tarragona | 15 | 4 | 5 | 0.781 | 0.00625 | 0.07027 | |
| ND5 Respiratory Complex I | Bay of Biscay | 19 | 4 | 5 | 0.696 | 0.00284 | −0.53717 |
| Gulf of Lion | 20 | 1 | 2 | 0.521 | 0.00158 | 1.53133 | |
| Olhão | 19 | 2 | 3 | 0.526 | 0.0017 | −0.04521 | |
| Sesimbra | 13 | 2 | 3 | 0.615 | 0.0021 | 0.2084 | |
| Tarragona | 18 | 4 | 5 | 0.66 | 0.00273 | −0.67309 |
| Maximum Temperature and Dissolved Molecular Oxygen | ||||
| df | SS | F | p (>F) | |
| model | 2 | 0.366 | 1.858 | 0.117 |
| Residuals | 2 | 0.197 | - | - |
| Minimum Temperature and Dissolved Molecular Oxygen | ||||
| df | SS | F | p (>F) | |
| model | 2 | 0.416 | 2.841 | 0.050 |
| Residuals | 2 | 0.146 | - | - |
| Range of Temperature and Dissolved Molecular Oxygen | ||||
| df | SS | F | p (>F) | |
| model | 2 | 0.150 | 1.087 | 0.483 |
| Residuals | 2 | 0.413 | - | - |
| Mean of Temperature and Dissolved Molecular Oxygen | ||||
| df | SS | F | p (>F) | |
| model | 2 | 0.202 | 0.560 | 0.867 |
| Residuals | 2 | 0.361 | - | - |
| df | SS | F | p (>F) | |
| CAP1 | 1 | 0.244 | 3.343 | 0.050 |
| CAP2 | 1 | 0.171 | 2.339 | 0.150 |
| Residual | 2 | 0.146 | ||
| df | SS | F | p (>F) | |
| sst_min | 1 | 0.230 | 3.109 | 0.033 |
| do_min | 1 | 0.202 | 2.756 | 0.058 |
| Residual | 2 | 0.146 |
| Method | Gene | Codon | Site | p Value | pp [α > β] |
|---|---|---|---|---|---|
| SLAC | CYTB | 14 | 42 | 0.1 | n/a |
| 15 | 45 | 0.1 | n/a | ||
| 16 | 48 | 0.1 | n/a | ||
| 26 | 78 | 0.1 | n/a | ||
| 40 | 120 | 0.1 | n/a | ||
| 48 | 144 | 0.1 | n/a | ||
| 53 | 159 | 0.1 | n/a | ||
| 57 | 171 | 0.1 | n/a | ||
| FEL | ATP6 | 53 | 159 | 0.1 | n/a |
| ND5 | 58 | 174 | 0.1 | n/a | |
| FUBAR | ATP6 | 24 | 72 | n/a | 0.9 |
| 49 | 147 | n/a | 0.9 | ||
| 36 | 108 | n/a | 0.9 | ||
| 52 | 156 | n/a | 0.9 | ||
| 53 | 159 | n/a | 0.9 | ||
| ND5 | 58 | 174 | n/a | 0.9 | |
| 64 | 192 | n/a | 0.9 | ||
| 90 | 270 | n/a | 0.9 | ||
| COI | 8 | 24 | n/a | 0.9 | |
| 44 | 132 | n/a | 0.9 | ||
| 53 | 159 | n/a | 0.9 | ||
| 57 | 171 | n/a | 0.9 |
| Locations | n | rAR | pA | Ho | He |
|---|---|---|---|---|---|
| Bay of Biscay | 20 | 13.44 | 2.24 | 0.71 | 0.86 |
| Gulf of Lion | 20 | 12.41 | 1.80 | 0.73 | 0.83 |
| Olhão | 20 | 12.89 | 2.10 | 0.74 | 0.86 |
| Sesimbra | 16 | 12.00 | 1.78 | 0.65 | 0.84 |
| Tarragona | 20 | 12.18 | 1.18 | 0.63 | 0.80 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Baltazar-Soares, M.; de Araújo Lima, A.R.; Silva, G. Targeted Sequencing of Mitochondrial Genes Reveals Signatures of Molecular Adaptation in a Nearly Panmictic Small Pelagic Fish Species. Genes 2021, 12, 91. https://doi.org/10.3390/genes12010091
Baltazar-Soares M, de Araújo Lima AR, Silva G. Targeted Sequencing of Mitochondrial Genes Reveals Signatures of Molecular Adaptation in a Nearly Panmictic Small Pelagic Fish Species. Genes. 2021; 12(1):91. https://doi.org/10.3390/genes12010091
Chicago/Turabian StyleBaltazar-Soares, Miguel, André Ricardo de Araújo Lima, and Gonçalo Silva. 2021. "Targeted Sequencing of Mitochondrial Genes Reveals Signatures of Molecular Adaptation in a Nearly Panmictic Small Pelagic Fish Species" Genes 12, no. 1: 91. https://doi.org/10.3390/genes12010091
APA StyleBaltazar-Soares, M., de Araújo Lima, A. R., & Silva, G. (2021). Targeted Sequencing of Mitochondrial Genes Reveals Signatures of Molecular Adaptation in a Nearly Panmictic Small Pelagic Fish Species. Genes, 12(1), 91. https://doi.org/10.3390/genes12010091





