A Comprehensive Integrated Genetic Map of the Complete Karyotype of Solea senegalensis (Kaup 1858)
Abstract
1. Introduction
2. Materials and Methods
2.1. BAC Sequencing and Annotation
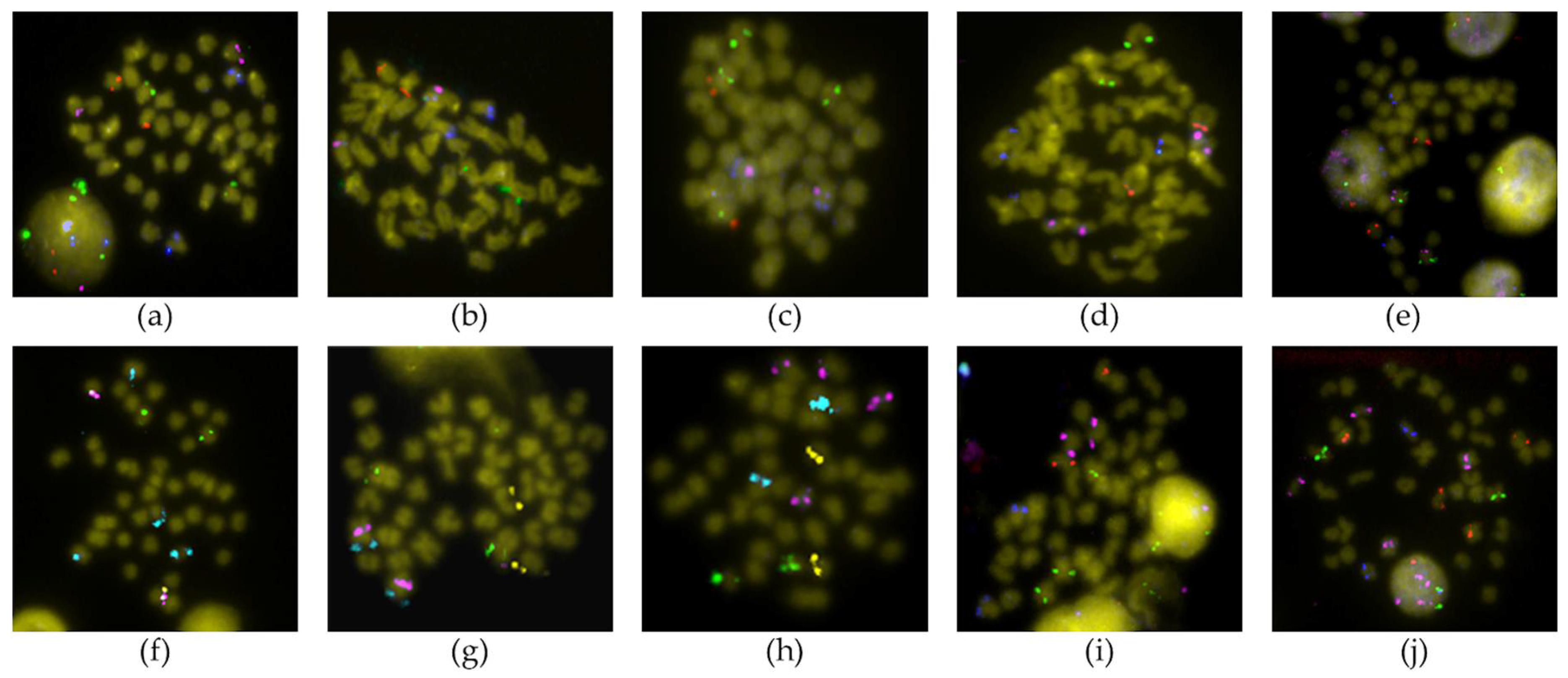
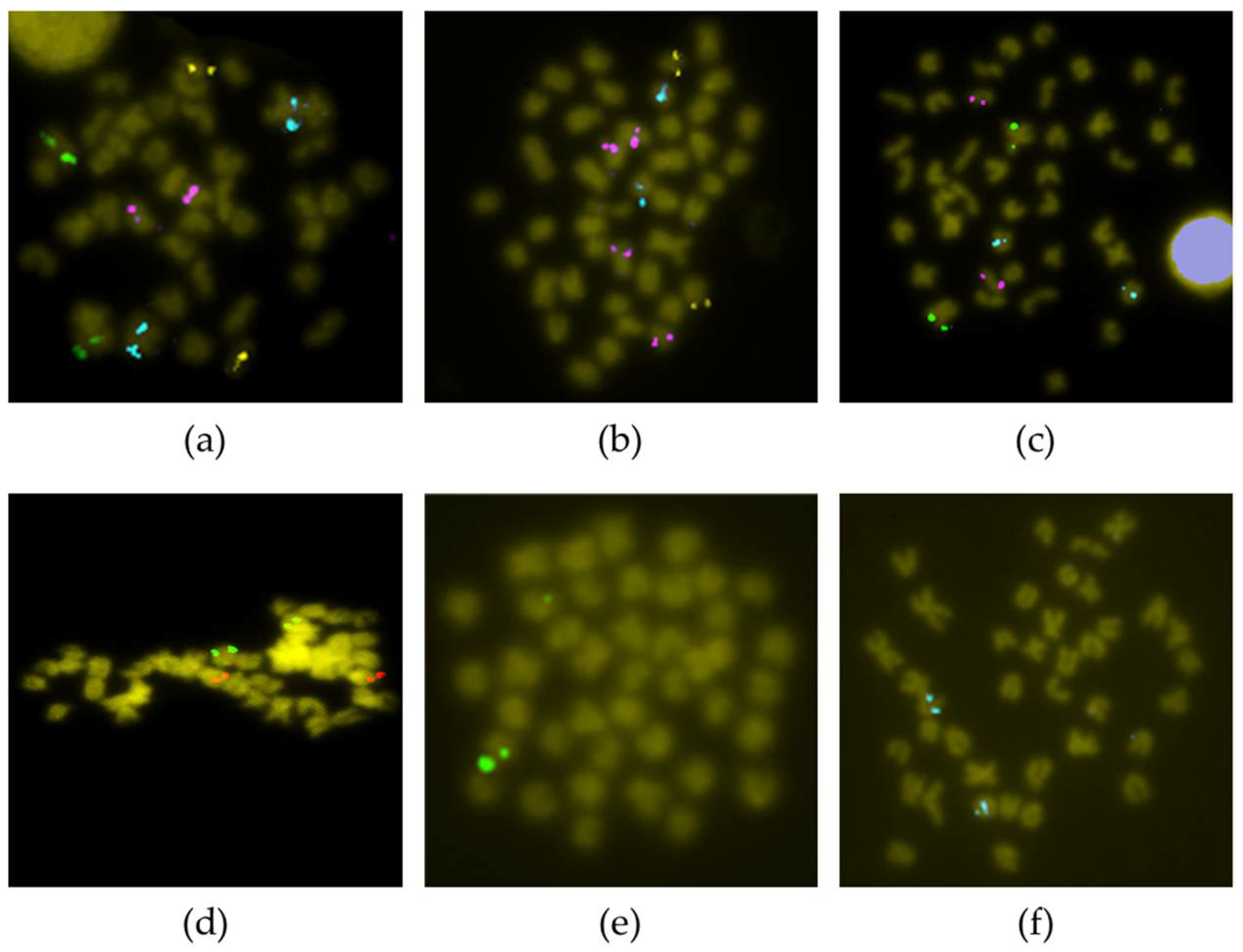
2.2. FISH Analysis
2.3. Synteny Analysis
3. Results
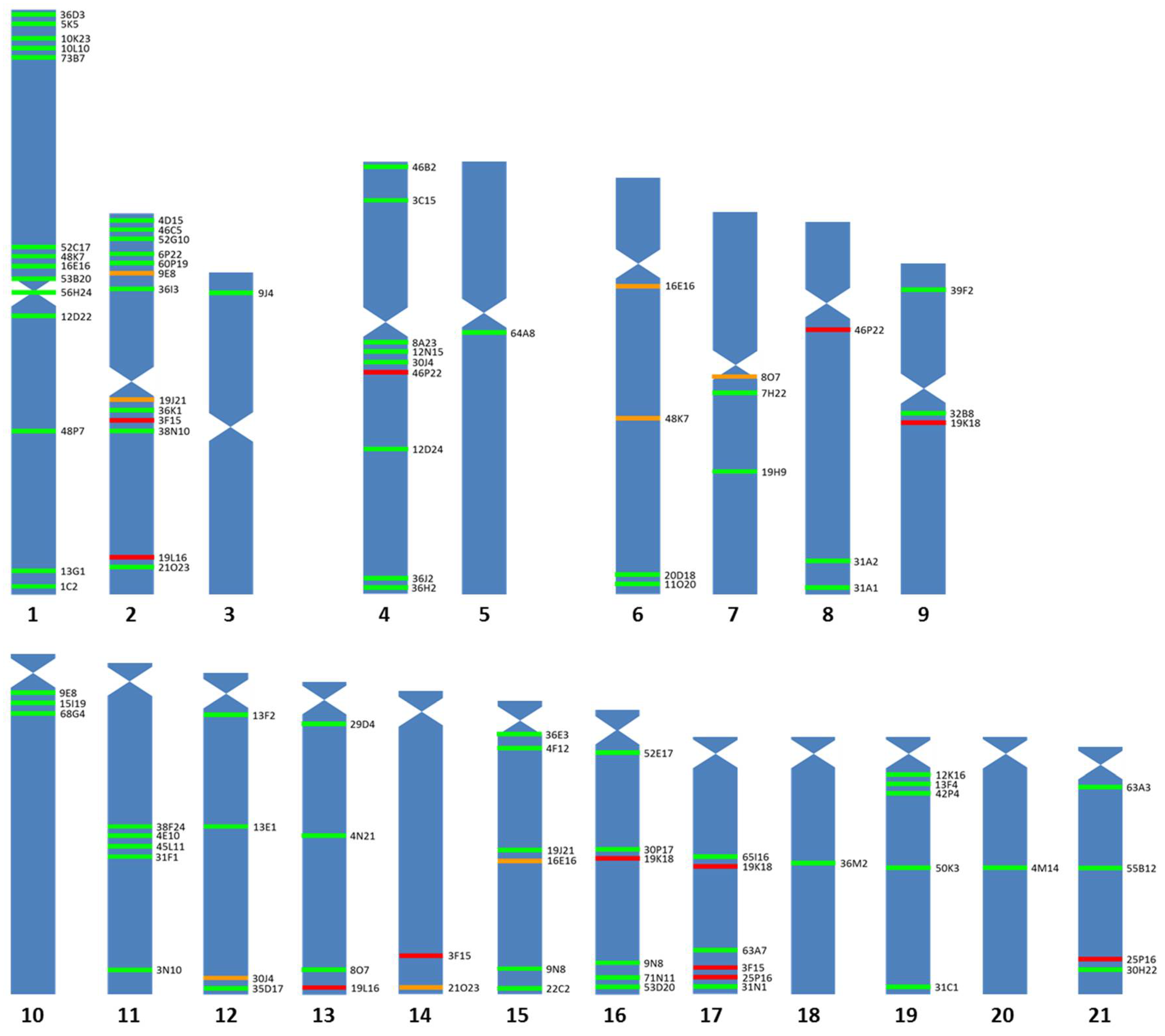
3.1. Chromosomal Map
3.2. BAC Sequence and Annotation Data
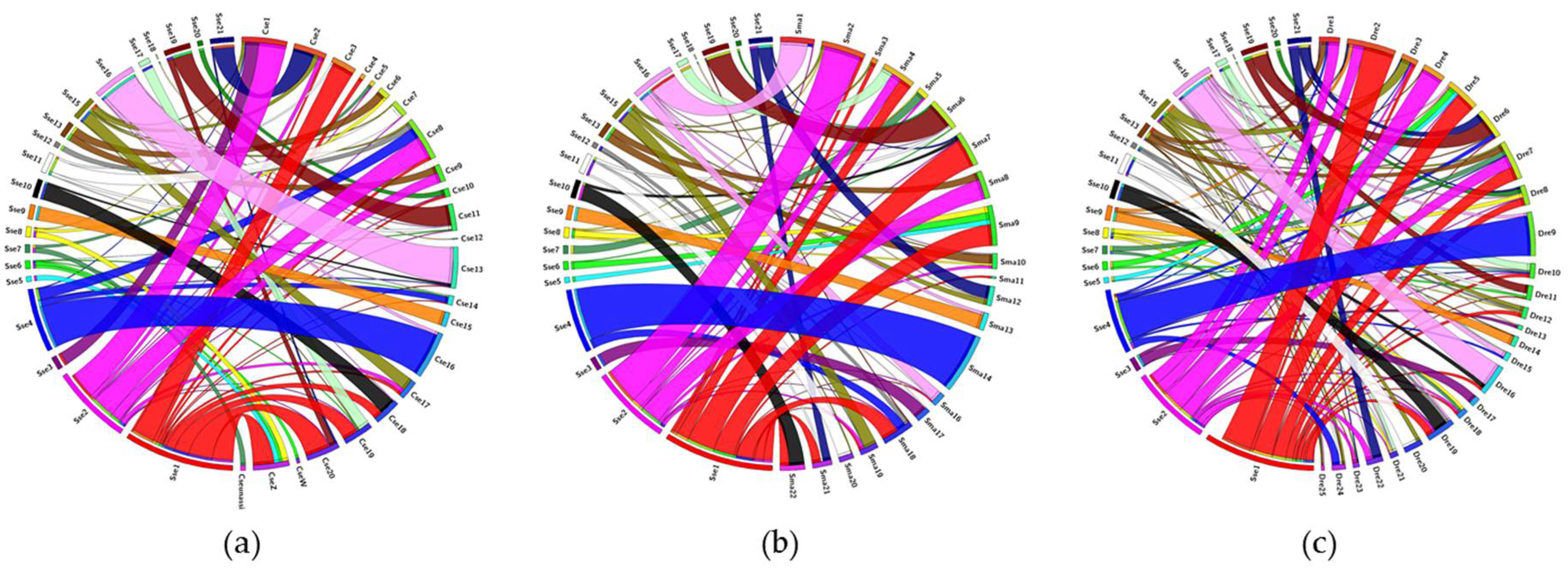
3.3. Comparative Genomic Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aquaculture in Spain 2020. Available online: www.apromar.es (accessed on 30 November 2020).
- Chairi, H.; Fernandez-Diaz, C.; Navas, J.I.; Manchado, M.; Rebordinos, L.; Blasco, J. In vivo genotoxicity and stress defenses in three flatfish species exposed to CuSO4. Ecotoxicol. Environ. Saf. 2010, 73, 1279–1285. [Google Scholar] [CrossRef] [PubMed]
- Anguis, V.; Cañavate, J.P. Spawning of captive Senegal sole (Solea senegalensis) under a naturally fluctuating temperature regime. Aquaculture 2005, 243, 133–145. [Google Scholar] [CrossRef]
- Flores, H.; Martinez, P. Ocular reversal in Paralichthys adspersus (Steindachner, 1867) small eyes flounder (Pleuronectiformes, Paralichthyidae): A case from the environment and analysis of cultured specimens. Lat. Am. J. Aquat. Res. 2013, 41, 57–67. [Google Scholar] [CrossRef]
- Vega, L.; Díaz, E.; Cross, I.; Rebordinos, L. Caracterizaciones citogenética e isoenzimática del lenguado Solea senegalensis Kaup, 1858. Bol. Inst. Esp. Oceanogr. 2002, 18, 245–250. [Google Scholar]
- García-Angulo, A.; Merlo, M.A.; Rodríguez, M.E.; Portela-Bens, S.; Liehr, T.; Rebordinos, L. Genome and Phylogenetic Analysis of Genes Involved in the Immune System of Solea senegalensis-Potential Applications in Aquaculture. Front. Genet. 2019, 11, 529. [Google Scholar] [CrossRef]
- Cross, I.; Garcia, E.; Rodríguez, M.E.; Arias-Perez, A.; Portela-Bens, S.; Merlo, M.A.; Rebordinos, L. The genomic structure of the highly-conserved dmrt1 gene in Solea senegalensis (Kaup, 1868) shows an unexpected intragenic duplication. PLoS ONE 2020, 15, e0241518. [Google Scholar] [CrossRef]
- Gjedrem, T. Genetic improvement of cold-water fish species. Aquac. Res. 2004, 31, 25–33. [Google Scholar] [CrossRef]
- Nguyen, N.H.; Rastas, P.M.A.; Premachandra, H.K.A.; Knibb, W. First high-density linkage map and single nucleotide polymorphisms significantly associated with traits of economic importance in yellowtail kingfish Seriola lalandi. Front. Genet. 2018, 9, 127. [Google Scholar] [CrossRef]
- Tsigenopoulos, C.S.; Louro, B.; Chatziplis, D.; Lagnel, J.; Vogiatzi, E.; Loukovitis, D.; Franch, R.; Sarropoulou, E.; Power, D.M.; Patarnello, T.; et al. Second generation genetic linkage map for the gilthead sea bream Sparus aurata L. Mar. Genom. 2014, 18, 77–82. [Google Scholar] [CrossRef]
- Siemon, H.S.; Chang, A.; Brown, G.D.; Koop, B.F.; Davidson, W.S. Type I microsatellite markers from Atlantic salmon (Salmo salar) expressed sequence tags. Mol. Ecol. Notes 2005, 5, 762–766. [Google Scholar] [CrossRef]
- Gonen, S.; Lowe, N.R.; Cezard, T.; Gharbi, K.; Bishop, S.P.C.; Houston, R.D. Linkage maps of the Atlantic salmon (Salmo salar) genome derived from RAD sequencing. BMC Genom. 2014, 15, 166. [Google Scholar] [CrossRef] [PubMed]
- Katagiri, T.; Kidd, C.; Tomasino, E.; Davis, J.T.; Wishon, C.; Stern, J.E.; Carleton, K.; Howe, A.E.; Kocher, T. A BAC-based physical map of the Nile tilapia genome. BMC Genom. 2005, 6, 89. [Google Scholar] [CrossRef] [PubMed]
- Rexroad, C.E.; Palti, Y.; Gahr, S.A.; Vallejo, R.L. A second generation genetic map for rainbow trout (Oncorhynchus mykiss). BMC Genet. 2008, 9, 74. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Kuang, Y.; Zhang, X.; Lu, C.; Cao, D.; Li, C.; Sun, X. A genetic linkage map and comparative genome analysis of common carp (Cyprinus carpio L.) using microsatellites and SNPs. Mol. Genet. Genom. 2011, 286, 261–277. [Google Scholar] [CrossRef]
- Zhao, L.; Zhang, Y.; Ji, P.; Zhang, X.; Zhao, Z.; Hou, G.; Huo, L.; Liu, G.; Li, C.; Xu, P.; et al. A dense genetic linkage map for common carp and its integration with a BAC-based physical map. PLoS ONE 2013, 8, e63928. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Karsi, A.; Li, P.; Cao, D.; Dunham, R. An AFLP-based genetic linkage map of channel catfish (Ictalurus punctatus) constructed by using an interspecific hybrid resource family. Genetics 2003, 165, 687–694. [Google Scholar]
- Poompuang, S.; Na-Nakorn, U. A preliminary genetic map of walking catfish (Clarias macrocephalus). Aquaculture 2004, 232, 195–203. [Google Scholar] [CrossRef]
- Zhang, J.; Shao, C.; Zhang, L.; Liu, K.; Gao, F.; Dong, Z.; Xu, P.; Chen, S. A first generation BAC-based physical map of the half-smooth tongue sole (Cynoglossus semilaevis) genome. BMC Genom. 2014, 15, 215. [Google Scholar] [CrossRef]
- Taboada, X.; Pansonato-Alves, J.C.; Foresti, F.; Martinez, P.; Viñas, A.; Pardo, B.; Bouza, C. Consolidation of the genetic and cytogenetic maps of turbot (Scophthalmus maximus) using FISH with BAC clones. Chromosoma 2014, 123, 281–291. [Google Scholar] [CrossRef]
- Shao, C.; Li, C.; Wang, N.; Qin, Y.; Xu, W.; Liu, Q.; Zhou, Q.; Zhao, Y.; Li, X.; Liu, S.; et al. Chromosome-level genome assembly of the spotted sea bass, Lateolabrax Maculatus. Gigascience 2018, 7, giy114. [Google Scholar] [CrossRef]
- Phillips, R.B.; Keatley, K.A.; Morash, M.R.; Ventura, A.B.; Lubieniecki, K.P.; Koop, B.F.; Dnazmann, R.G.; Davidson, W.S. Assignment of Atlantic salmon (Salmo salar) linkage groups to specific chromosomes: Conservation of large syntenic blocks corresponding to whole chromosome arms in rainbow trout (Oncorhynchus mykiss). BMC Genet. 2009, 10, 46. [Google Scholar] [CrossRef]
- Phillips, R.B.; Nichols, K.M.; Dekoning, J.J.; Morasch, M.R.; Keatley, K.A.; Rexroad, C.; Gahr, S.A.; Danzmann, R.G.; Drew, R.E.; Thorgaard, G.H. Assignment of rainbow trout linkage groups to specific chromosomes. Genetics 2006, 174, 1661–1670. [Google Scholar] [CrossRef]
- García-Cegarra, A.; Merlo, M.A.; Ponce, M.; Portela-Bens, S.; Cross, I.; Manchado, M.; Rebordinos, L. A preliminary genetic map in Solea senegalensis (Pleuronectiformes, Soleidae) using BAC-FISH and next-generation sequencing. Cytogenet. Genome Res. 2013, 141, 227–240. [Google Scholar] [CrossRef]
- Manchado, M.; Planas, J.V.; Cousin, X.; Rebordinos, L.; Claros, M.G. Current status in other finfish species: Description of current genomic resources for the gilthead seabream (Sparus aurata) and soles (Solea senegalensis and Solea solea). In Genomics in Aquaculture; MacKencie, S., Jentoft, S., Eds.; Academic Press: San Diego, CA, USA, 2016; pp. 195–221. [Google Scholar]
- Diopere, E.; Maes, G.E.; Komen, H.; Volckaert, F.; Groenen, M. A Genetic Linkage Map of Sole (Solea solea): A Tool for Evolutionary and Comparative Analyses of Exploited (Flat) Fishes. PLoS ONE 2014, 9, e115040. [Google Scholar] [CrossRef] [PubMed]
- Molina-Luzón, M.J.; Hermida, M.; Navajas, R.; Robles, F.; Navas, J.I.; Bouza, C.; Martinez, P.; Herran, R. First haploid genetic map based on microsatellite markers in Senegalese sole (Solea senegalensis, Kaup 1858). Mar. Biotechnol. 2015, 17, 8–22. [Google Scholar] [CrossRef] [PubMed]
- García, E.; Cross, I.; Portela-Bens, S.; Rodriguez, M.E.; García-Angulo, A.; Molina, B.; Cuadrado, A.; Liehr, T.; Rebordinos, L. Integrative genetic map of repetitive DNA in the sole Solea senegalensis genome shows a Rex transposon located in a proto-sex chromosome. Sci. Rep. 2019, 9, 17146. [Google Scholar] [CrossRef] [PubMed]
- Manchado, M.; Planas, J.V.; Cousin, X.; Rebordinos, L.; Claros, M.G. Chap. Genetic and Genomic Characterization of Soles. In The Biology of Sole; Muñoz-Cueto, J.A., Mañanos, E., Sanchez, F.J., Eds.; CRC Press: Boca Raton, FL, USA, 2019; pp. 375–394. [Google Scholar]
- Benzekri, H.; Armesto, P.; Cousin, X.; Rovira, M.; Crespo, D.; Merlo, M.A.; Mazurais, D.; Bautista, R.; Guerrero-Fernández, D.; Fernández-Pozo, N.; et al. De novo assembly, characterization and functional annotation of Senegalese sole (Solea senegalensis) and common sole (Solea solea) transcriptomes: Integration in a database and design of a microarray. BMC Genom. 2014, 15, 952. [Google Scholar] [CrossRef] [PubMed]
- Cross, I.; Merlo, M.A.; Manchado, M.; Infante, C.; Cañavate, J.P.; Rebordinos, L. Cytogenetic Characterization of the sole Solea senegalensis (Teleostei: Pleuronectiformes: Soleidae): Ag-NOR, (GATA)n, (TTAGGG)n and ribosomal genes by one-color and two-color FISH. Genetica 2006, 128, 253–259. [Google Scholar] [CrossRef]
- Manchado, M.; Zuasti, E.; Cross, I.; Merlo, M.A.; Infante, C.; Rebordinos, L. Molecular characterization and chromosomal mapping of the 5S rRNA gene in Solea senegalensis: A new linkage to the U1, U2, and U5 small nuclear RNA genes. Genome 2006, 49, 79–86. [Google Scholar] [CrossRef]
- Garcia-Angulo, A.; Merlo, M.A.; Portela-Bens, S.; Rodriguez, M.E.; Garcia, E.; Al-Rikabi, A.; Liehr, T.; Rebordinos, L. Evidence for a Robertsonian fusion in Solea senegalensis (Kaup, 1858) revealed by zoo-FISH and comparative genome analysis. BMC Genom. 2018, 19, 818. [Google Scholar] [CrossRef]
- Arias-Perez, A.; Ramirez, D.; Rodriguez, M.E.; Portela-Bens, S.; Garcia, E.; Merlo, M.A.; Garcia-Angulo, A.; Cross, I.; Liehr, T.; Rebordinos, L. In Silico Detection and FISH Analysis to Determine Location of miRNAs in Solea senegalensis Chromosomes Using BACs. OBM Genet. 2018, 2, 4. [Google Scholar] [CrossRef]
- Portela-Bens, S.; Merlo, M.A.; Rodríguez, M.E.; Cross, I.; Manchado, M.; Kosyakova, N.; Liehr, T.; Rebordinos, L. Integrated gene mapping and synteny studies give insights into the evolution of a sex proto-chromosome in Solea senegalensis. Chromosoma 2017, 126, 261–277. [Google Scholar] [CrossRef] [PubMed]
- Merlo, M.A.; Iziga, R.; Portela-Bens, S.; Cross, I.; Kosyakova, N.; Liehr, T.; Manchado, M.; Rebordinos, L. Analysis of the histone cluster in Senegalese sole (Solea senegalensis): Evidence for a divergent evolution of two canonical histone clusters. Genome 2017, 60, 441–453. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, M.E.; Molina, B.; Merlo, M.A.; Arias-Perez, A.; Garcia-Angulo, A.; Cross, I.; Liehr, T.; Rebordinos, L. Evolution of the Proto Sex-Chromosome in Solea senegalensis. Int. J. Mol. Sci. 2019, 20, 5111. [Google Scholar] [CrossRef] [PubMed]
- Lewis, S.E.; Searle, S.M.J.; Harris, N.; Gibson, M.; Iyer, V.; Richter, J.; Wiel, C.; Bayraktaroglu, L.; Birney, E.; Crosby, M.A.; et al. Apollo: A sequence annotation editor. Genom. Biol. 2002, 3, research0082.1. [Google Scholar] [CrossRef]
- Liehr, T. (Ed.) Fluorescence In Situ Hybridization (FISH) Application Guide; Springer: Berlin, Germany, 2009. [Google Scholar]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genom. Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Sader, M.A.; Dias, Y.; Costa, Z.P.; Munhoz, C.; Penha, H.; Berges, H.; Vieira, M.L.C.; Pedrosa-Harand, A. Identification of passion fruit (Passiflora edulis) chromosomes using BAC-FISH. Chromosom. Res. 2019, 27, 299–311. [Google Scholar] [CrossRef]
- Dong, G.; Shen, J.; Zhang, Q.; Wang, J.; Yu, Q.; Ming, R.; Wang, K.; Zhang, J. Development and Applications of Chromosome-Specific Cytogenetic BAC-FISH Probes in S. spontaneum. Front. Plant. Sci. 2018, 9, 218. [Google Scholar] [CrossRef]
- Mazzuchelli, J.; Kocher, T.D.; Yang, F.; Martins, C. Integrating cytogenetics and genomics in comparative evolutionary studies of cichlid fish. BMC Genom. 2012, 13, 463. [Google Scholar] [CrossRef]
- Yang, Z.; Li, X.; Liao, H.; Hu, L.; Peng, C.; Wang, S.; Huang, X.; Bao, Z. A Molecular Cytogenetic Map of Scallop (Patinopecten yessoensis). Mar. Biotechnol. 2019, 21, 731–742. [Google Scholar] [CrossRef]
- Glasauer, S.; Neuhauss, S. Whole-genome duplication in teleost fishes and its evolutionary consequences. Mol. Genet. Genom. 2014, 289, 1045–1060. [Google Scholar] [CrossRef] [PubMed]
- Kaitetzidou, E.; Katsiadaki, I.; Lagnei, J.; Antonopoulou, E.; Sarropoulou, E. Unravelling paralogous gene expression dynamics during three-spined stickleback embryogenesis. Sci. Rep. 2019, 9, 3752. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Hu, H.; Panserat, S.; Marandel, L. Evolutionary history of DNA methylation related genes in chordates: New insights from multiple whole genome duplications. Sci. Rep. 2020, 10, 970. [Google Scholar] [CrossRef]
- Firmino, J.P.; Carballo, C.; Armesto, P.; Campinho, M.C.; Power, D.W.; Manchado, M. Phylogeny, expression patterns and regulation of DNA methyltransferases in early development of the flatfish, Solea senegalensis. BMC Dev. Biol. 2017, 17, 1. [Google Scholar] [CrossRef]
- Jabalameli, M.; Horscroft, C.; Vergara-Lope, A.; Pengelly, R.; Collins, A. Gene-dense autosomal chromosomes show evidence for increased selection. Heredity 2019, 123, 774–783. [Google Scholar] [CrossRef]
- Yu, L.; Ma, X.; Deng, B.; Yue, J.; Ming, R. Construction of high-density genetic maps de ned sex determination region of the Y chromosome in spinach. Mol. Genet. Genom. 2020. [Google Scholar] [CrossRef]
- Kobayashi, T.; Matsuda, M.; Kajiura-Kobayashi, H.; Suzuki, A.; Saito, N.; Nakamoto, M.; Shibata, N.; Gagahama, Y. Two DM domain genes, DMY and DMRT1, involved in testicular differentiation and development in the medaka, Oryzias latipes. Dev. Dyn. 2004, 231, 518–526. [Google Scholar] [CrossRef]
- Pan, Q.; Guiguen, Y.; Herpin, A. Chap. Evolution of Sex Determining Genes in Fish. In Encyclopedia of Reproduction, 2nd ed.; Skinner, M.K., Ed.; Washington State University: Pullman, WA, USA, 2018; pp. 168–275. [Google Scholar]
- Charlesworth, D.; Charlesworth, B.; Marais, G. Steps in the evolution of heteromorphic sex chromosomes. Heredity 2005, 95, 118–128. [Google Scholar] [CrossRef]
- Schemberger, M.; do Nascimento, V.D.; Coan, R.; Ramos, E.; Nogaroto, V.; Ziemniczak, K.; Valente, G.; Moreira-Filho, O. DNA transposon invasion and microsatellite accumulation guide W chromosome differentiation in a Neotropical fish genome. Chromosoma 2019, 128, 547–560. [Google Scholar] [CrossRef]
- Al-Shahrour, F.; Minguez, P.; Marques-Bonet, T.; Gazave, E.; Navarro, A.; Dopazo, J. Selection upon genome architecture: Conservation of functional neighborhoods with changing genes. PLoS Comput. Biol. 2010, 6, e1000953. [Google Scholar] [CrossRef]
- Lee, J.; Sonnhammer, E. Genomic gene clustering analysis of pathways in eukaryotes. Genom. Res. 2003, 13, 875–882. [Google Scholar] [CrossRef]
- Batada, N.; Hurst, L. Evolution of chromosome organization driven by selection for reduced gene expression noise. Nat. Genet. 2007, 39, 945–949. [Google Scholar] [CrossRef]
- Singer, G.; Lloyde, A.; Huminiecki, L.; Wolfe, K. Clusters of co-expressed genes in mammalian genomes are conserved by natural selection. Mol. Biol. Evol. 2005, 22, 767–775. [Google Scholar] [CrossRef]
- Garcia-Angulo, A.; Merlo, M.A.; Iziga, R.; Rodriguez, M.E.; Portela-Bens, S.; Al-Rikabi, A.; Liehr, T.; Rebordinos, L. Gene clusters related to metamorphosis in Solea senegalensis are highly conserved. Comp. Biochem. Physiol. Part D Genom. Proteom. 2020, 35, 100706. [Google Scholar] [CrossRef]


| Chromosome | Number of BAC | Number of Genes a | Size (bp) a | Gene Density (Genes/Mb) |
|---|---|---|---|---|
| 1 | 14 | 113 | 2,741,505 | 41.22 |
| 2 | 9 | 80 | 991,406 | 80.69 |
| 3 | 1 | 12 | 140,126 | 85.64 |
| 4 | 8 | 69 | 1,200,692 | 57.47 |
| 5 | 1 | 5 | 211,201 | 23.67 |
| 6 | 2 | 8 | 256,445 | 31.20 |
| 7 | 2 | 8 | 185,169 | 43.20 |
| 8 | 2 | 9 | 146,485 | 61.44 |
| 9 | 2 | 14 | 273,741 | 51.14 |
| 10 | 2 | 15 | 203,915 | 73.56 |
| 11 | 5 | 18 | 555,500 | 32.40 |
| 12 | 2 | 1 | 137,654 | 7.27 |
| 13 | 3 | 15 | 432,796 | 34.66 |
| 14 | ||||
| 15 | 3 | 21 | 431,021 | 48.72 |
| 16 | 5 | 40 | 583,480 | 58.56 |
| 17 | 3 | 11 | 448,777 | 24.51 |
| 18 | 1 | 1 | 81,656 | 12.25 |
| 19 | 5 | 25 | 758,436 | 32.96 |
| 20 | 1 | 5 | 283,462 | 17.64 |
| 21 | 3 | 24 | 413,134 | 58.09 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Merlo, M.A.; Portela-Bens, S.; Rodríguez, M.E.; García-Angulo, A.; Cross, I.; Arias-Pérez, A.; García, E.; Rebordinos, L. A Comprehensive Integrated Genetic Map of the Complete Karyotype of Solea senegalensis (Kaup 1858). Genes 2021, 12, 49. https://doi.org/10.3390/genes12010049
Merlo MA, Portela-Bens S, Rodríguez ME, García-Angulo A, Cross I, Arias-Pérez A, García E, Rebordinos L. A Comprehensive Integrated Genetic Map of the Complete Karyotype of Solea senegalensis (Kaup 1858). Genes. 2021; 12(1):49. https://doi.org/10.3390/genes12010049
Chicago/Turabian StyleMerlo, Manuel A., Silvia Portela-Bens, María E. Rodríguez, Aglaya García-Angulo, Ismael Cross, Alberto Arias-Pérez, Emilio García, and Laureana Rebordinos. 2021. "A Comprehensive Integrated Genetic Map of the Complete Karyotype of Solea senegalensis (Kaup 1858)" Genes 12, no. 1: 49. https://doi.org/10.3390/genes12010049
APA StyleMerlo, M. A., Portela-Bens, S., Rodríguez, M. E., García-Angulo, A., Cross, I., Arias-Pérez, A., García, E., & Rebordinos, L. (2021). A Comprehensive Integrated Genetic Map of the Complete Karyotype of Solea senegalensis (Kaup 1858). Genes, 12(1), 49. https://doi.org/10.3390/genes12010049









