Soft Selective Sweep on Chemosensory Genes Correlates with Ancestral Preference for Toxic Noni in a Specialist Drosophila Population
Abstract
1. Introduction
2. Materials and Methods
2.1. Population Genomics Analyses of Chemosensory Genes
2.2. Behavioral Feeding Preference Analyses
3. Results
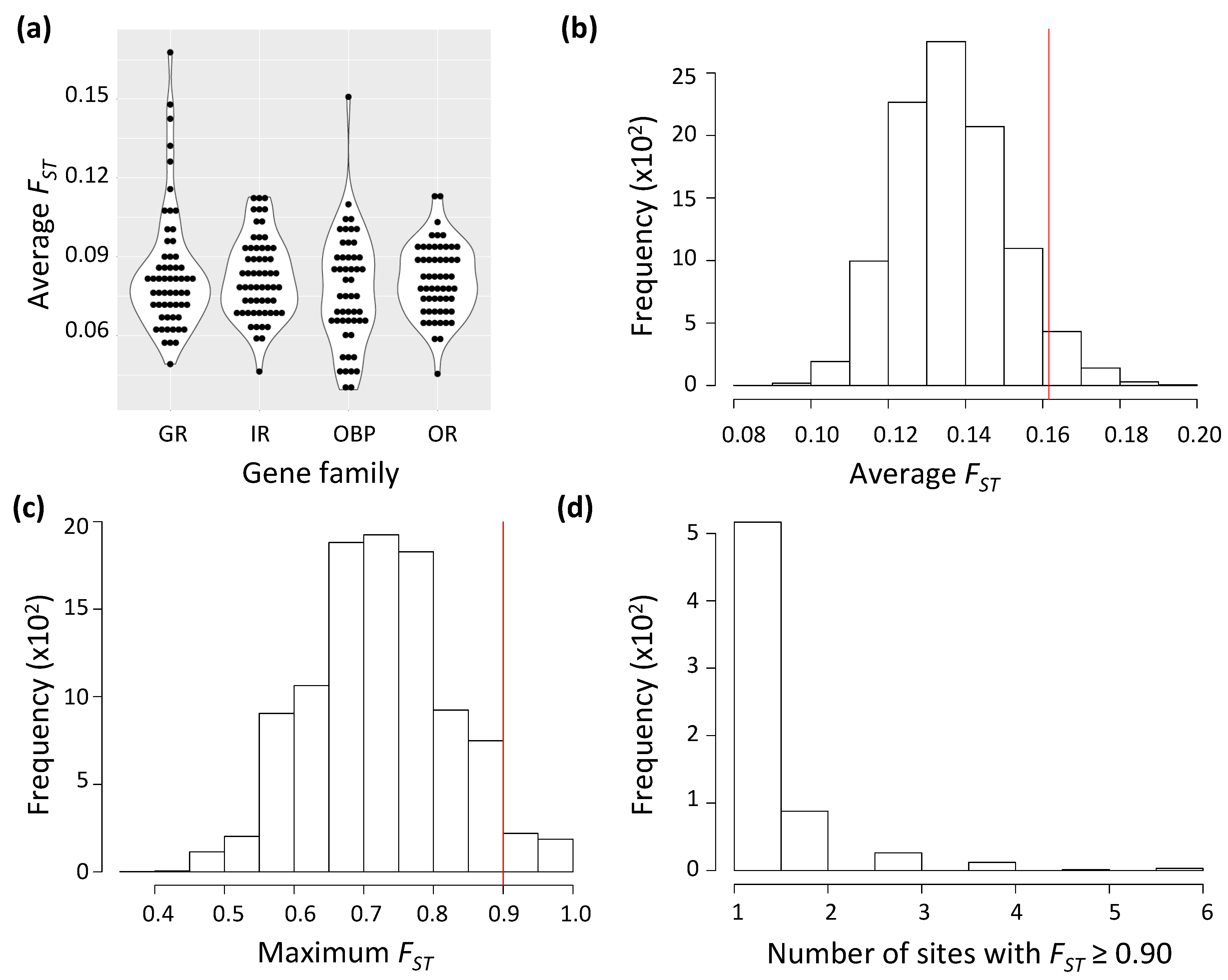
3.1. A Handful of Chemosensory Genes, Mostly Encoding Gustatory Receptors, Deviate from Neutral Expectations
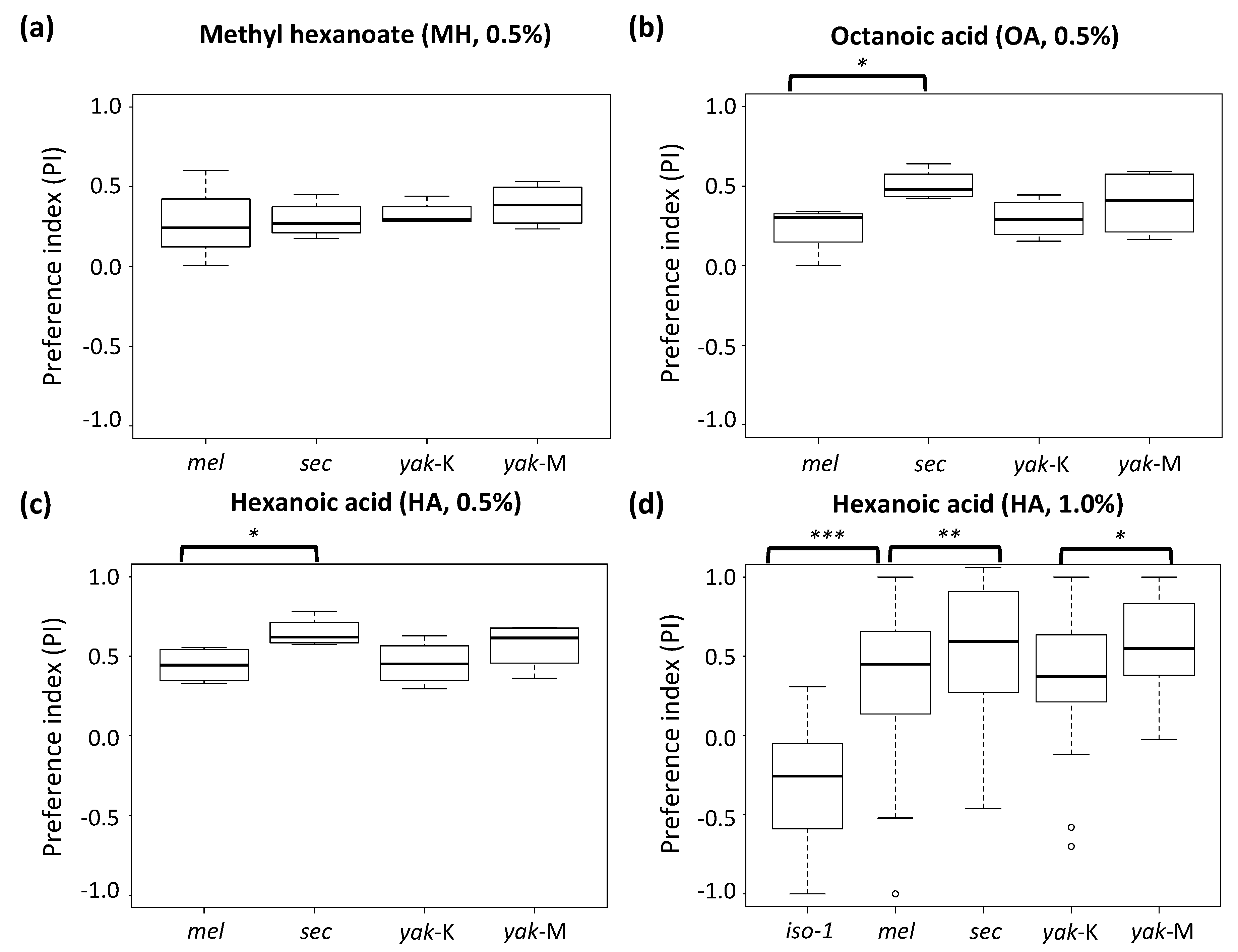
3.2. Generalist D. yakuba Has a Substantial Dose-Dependent Gustatory Preference for Noni Toxins
4. Discussion
4.1. Contrast between Olfactory and Gustatory Evolution in D. yakuba
4.2. Apparent Lack of Selection on Acid-Sensing Ionotropic Receptor Genes
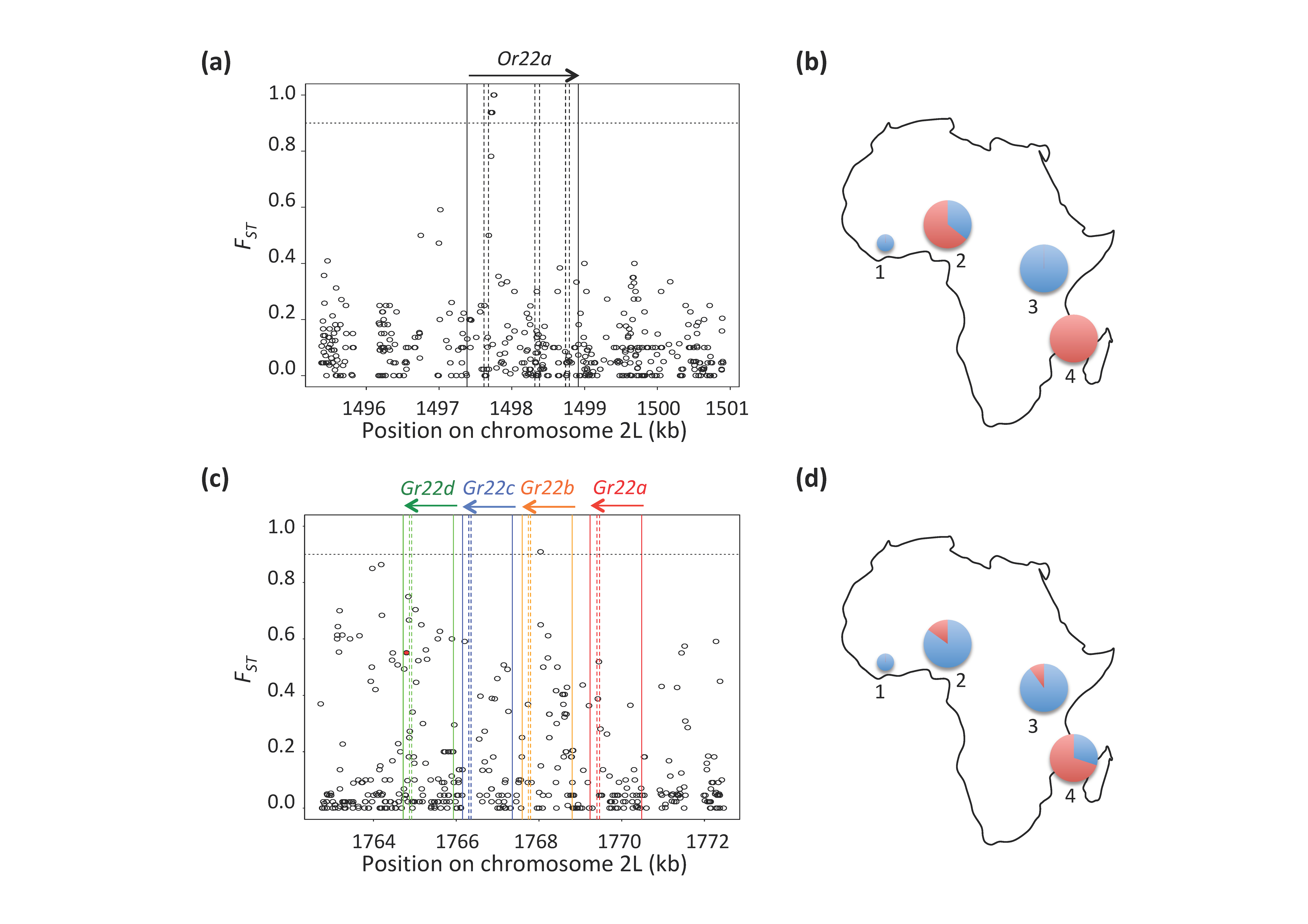
4.3. Variation in the Ester-Sensing Olfactory Receptor Or22a in Populations from the Ancestral Range
4.4. Selection on Bitter-Sensing Gustatory Receptors
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Futuyma, D.J.; Moreno, G. The Evolution of Ecological Specialization. Annu. Rev. Ecol. Syst. 1988, 19, 207–233. [Google Scholar] [CrossRef]
- Simon, J.-C.; d’Alençon, E.; Guy, E.; Jacquin-Joly, E.; Jacquiéry, J.; Nouhaud, P.; Peccoud, J.; Sugio, A.; Streiff, R. Genomics of Adaptation to Host-Plants in Herbivorous Insects. Brief. Funct. Genom. 2015, 14, 413–423. [Google Scholar] [CrossRef] [PubMed]
- Gloss, A.D.; Abbot, P.; Whiteman, N.K. How Interactions with Plant Chemicals Shape Insect Genomes. Curr. Opin. Insect Sci. 2019, 36, 149–156. [Google Scholar] [CrossRef] [PubMed]
- Vertacnik, K.L.; Linnen, C.R. Evolutionary Genetics of Host Shifts in Herbivorous Insects: Insights from the Age of Genomics. Ann. N. Y. Acad. Sci. 2017, 1389, 186–212. [Google Scholar] [CrossRef]
- Etges, W.J. Evolutionary Genomics of Host Plant Adaptation: Insights from Drosophila. Curr. Opin. Insect Sci. 2019, 36, 96–102. [Google Scholar] [CrossRef]
- McBride, C.S.; Arguello, J.R. Five Drosophila Genomes Reveal Nonneutral Evolution and the Signature of Host Specialization in the Chemoreceptor Superfamily. Genetics 2007, 177, 1395–1416. [Google Scholar] [CrossRef]
- McBride, C.S. Rapid Evolution of Smell and Taste Receptor Genes during Host Specialization in Drosophila Sechellia. Proc. Natl. Acad. Sci. USA 2007, 104, 4996–5001. [Google Scholar] [CrossRef]
- Almeida, F.C.; Sánchez-Gracia, A.; Campos, J.L.; Rozas, J. Family Size Evolution in Drosophila Chemosensory Gene Families: A Comparative Analysis with a Critical Appraisal of Methods. Genome Biol. Evol. 2014, 6, 1669–1682. [Google Scholar] [CrossRef]
- Kulmuni, J.; Wurm, Y.; Pamilo, P. Comparative Genomics of Chemosensory Protein Genes Reveals Rapid Evolution and Positive Selection in Ant-Specific Duplicates. Heredity 2013, 110, 538–547. [Google Scholar] [CrossRef]
- Sánchez-Gracia, A.; Vieira, F.G.; Almeida, F.C.; Rozas, J. Comparative Genomics of the Major Chemosensory Gene Families in Arthropods. Encycl. Life Sci. 2011. [Google Scholar] [CrossRef]
- Matzkin, L.M. Ecological Genomics of Host Shifts in Drosophila Mojavensis. Adv. Exp. Med. Biol. 2014, 781, 233–247. [Google Scholar] [CrossRef] [PubMed]
- Matzkin, L.M.; Watts, T.D.; Bitler, B.G.; Machado, C.A.; Markow, T.A. Functional Genomics of Cactus Host Shifts in Drosophila Mojavensis. Mol. Ecol. 2006, 15, 4635–4643. [Google Scholar] [CrossRef] [PubMed]
- R’Kha, S.; Capy, P.; David, J.R. Host-Plant Specialization in the Drosophila Melanogaster Species Complex: A Physiological, Behavioral, and Genetical Analysis. Proc. Natl. Acad. Sci. USA 1991, 88, 1835–1839. [Google Scholar] [CrossRef] [PubMed]
- Yassin, A.; Debat, V.; Bastide, H.; Gidaszewski, N.; David, J.R.; Pool, J.E. Recurrent Specialization on a Toxic Fruit in an Island Drosophila Population. Proc. Natl. Acad. Sci. USA 2016, 113, 4771–4776. [Google Scholar] [CrossRef]
- Farine, J.-P.; Legal, L.; Moreteau, B.; Le Quere, J.-L. Volatile Components of Ripe Fruits of Morinda Citrifolia and Their Effects on Drosophila. Phytochemistry 1996, 41, 433–438. [Google Scholar] [CrossRef]
- Higa, I.; Fuyama, Y. Genetics of Food Preference in Drosophila Sechellia. Genetica 1993, 88, 129–136. [Google Scholar] [CrossRef]
- Amlou, M.; Moreteau, B.; David, J.R. Genetic Analysis of Drosophila Sechellia Specialization: Oviposition Behavior Toward the Major Aliphatic Acids of Its Host Plant. Behav Genet 1998, 28, 455–464. [Google Scholar] [CrossRef]
- Dekker, T.; Ibba, I.; Siju, K.P.; Stensmyr, M.C.; Hansson, B.S. Olfactory Shifts Parallel Superspecialism for Toxic Fruit in Drosophila Melanogaster Sibling, D. Sechellia. Curr. Biol. 2006, 16, 101–109. [Google Scholar] [CrossRef]
- Prieto-Godino, L.L.; Rytz, R.; Cruchet, S.; Bargeton, B.; Abuin, L.; Silbering, A.F.; Ruta, V.; Peraro, M.D.; Benton, R. Evolution of Acid-Sensing Olfactory Circuits in Drosophilids. Neuron 2017, 93, 661–676.e6. [Google Scholar] [CrossRef]
- Auer, T.O.; Khallaf, M.A.; Silbering, A.F.; Zappia, G.; Ellis, K.; Álvarez-Ocaña, R.; Arguello, J.R.; Hansson, B.S.; Jefferis, G.S.X.E.; Caron, S.J.C.; et al. Olfactory Receptor and Circuit Evolution Promote Host Specialization. Nature 2020, 579, 402–408. [Google Scholar] [CrossRef]
- Matsuo, T.; Sugaya, S.; Yasukawa, J.; Aigaki, T.; Fuyama, Y. Odorant-Binding Proteins OBP57d and OBP57e Affect Taste Perception and Host-Plant Preference in Drosophila Sechellia. PLoS Biol. 2007, 5, e118. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.-E.; Chen, Y.; Amrein, H. Molecular Basis of Fatty Acid Taste in Drosophila. Elife Sci. 2017, 6, e30115. [Google Scholar] [CrossRef] [PubMed]
- Messer, P.W.; Petrov, D.A. Population Genomics of Rapid Adaptation by Soft Selective Sweeps. Trends Ecol. Evol. 2013, 28, 659–669. [Google Scholar] [CrossRef] [PubMed]
- Hermisson, J.; Pennings, P.S. Soft Sweeps and beyond: Understanding the Patterns and Probabilities of Selection Footprints under Rapid Adaptation. Methods Ecol. Evol. 2017, 8, 700–716. [Google Scholar] [CrossRef]
- Rogers, R.L.; Cridland, J.M.; Shao, L.; Hu, T.T.; Andolfatto, P.; Thornton, K.R. Landscape of Standing Variation for Tandem Duplications in Drosophila Yakuba and Drosophila Simulans. Mol. Biol. Evol. 2014, 31, 1750–1766. [Google Scholar] [CrossRef]
- Thurmond, J.; Goodman, J.L.; Strelets, V.B.; Attrill, H.; Gramates, L.S.; Marygold, S.J.; Matthews, B.B.; Millburn, G.; Antonazzo, G.; Trovisco, V.; et al. FlyBase 2.0: The next Generation. Nucleic Acids Res. 2019, 47, D759–D765. [Google Scholar] [CrossRef]
- Li, H. Minimap2: Pairwise Alignment for Nucleotide Sequences. Bioinformatics 2018, 34, 3094–3100. [Google Scholar] [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R. The Sequence Alignment/Map Format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef]
- Kofler, R.; Pandey, R.V.; Schlötterer, C. PoPoolation2: Identifying Differentiation between Populations Using Sequencing of Pooled DNA Samples (Pool-Seq). Bioinformatics 2011, 27, 3435–3436. [Google Scholar] [CrossRef]
- Hudson, R.R.; Slatkin, M.; Maddison, W.P. Estimation of Levels of Gene Flow from DNA Sequence Data. Genetics 1992, 132, 583–589. [Google Scholar]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A New Generation of Protein Database Search Programs. Nucl Acids Res 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Gutenkunst, R.N.; Hernandez, R.D.; Williamson, S.H.; Bustamante, C.D. Inferring the Joint Demographic History of Multiple Populations from Multidimensional SNP Frequency Data. PLoS Genet. 2009, 5. [Google Scholar] [CrossRef] [PubMed]
- Ewing, G.; Hermisson, J. MSMS: A Coalescent Simulation Program Including Recombination, Demographic Structure and Selection at a Single Locus. Bioinformatics 2010, 26, 2064–2065. [Google Scholar] [CrossRef] [PubMed]
- Comeron, J.M.; Ratnappan, R.; Bailin, S. The Many Landscapes of Recombination in Drosophila Melanogaster. PLoS Genet. 2012, 8, 33–35. [Google Scholar] [CrossRef] [PubMed]
- Bastide, H.; Lange, J.D.; Lack, J.B.; Yassin, A.; Pool, J.E. A Variable Genetic Architecture of Melanic Evolution in Drosophila Melanogaster. Genetics 2016, 204, 1307–1319. [Google Scholar] [CrossRef]
- Sellier, M.-J.; Reeb, P.; Marion-Poll, F. Consumption of Bitter Alkaloids in Drosophila Melanogaster in Multiple-Choice Test Conditions. Chem. Senses 2011, 36, 323–334. [Google Scholar] [CrossRef]
- De Chaumont, F.; Dallongeville, S.; Chenouard, N.; Hervé, N.; Pop, S.; Provoost, T.; Meas-Yedid, V.; Pankajakshan, P.; Lecomte, T.; Le Montagner, Y.; et al. Icy: An Open Bioimage Informatics Platform for Extended Reproducible Research. Nat. Methods 2012, 9, 690–696. [Google Scholar] [CrossRef]
- R Core Team R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2016.
- Delventhal, R.; Carlson, J.R. Bitter Taste Receptors Confer Diverse Functions to Neurons. eLife 2016, 5, 1–23. [Google Scholar] [CrossRef]
- Lachaise, D.; Harry, M.; Solignac, M.; Lemeunier, F.; Bénassi, V.; Cariou, M.-L. Evolutionary Novelties in Islands: Drosophila Santomea, a New Melanogaster Sister Species from São Tomé. Proc. R. Soc. Lond. B Biol. Sci. 2000, 267, 1487–1495. [Google Scholar] [CrossRef]
- Masek, P.; Keene, A.C. Drosophila Fatty Acid Taste Signals through the PLC Pathway in Sugar-Sensing Neurons. PLoS Genet. 2013, 9, e1003710. [Google Scholar] [CrossRef]
- Gerber, B.; Stocker, R.F.; Tanimura, T.; Thum, A.S. Smelling, Tasting, Learning: Drosophila as a Study Case. Results Probl. Cell Differ. 2009, 47, 139–185. [Google Scholar] [CrossRef] [PubMed]
- Moreteau, B.; R’Kha, S.; David, J.R. Genetics of a Nonoptimal Behavior: Oviposition Preference ofDrosophila Mauritiana for a Toxic Resource. Behav. Genet. 1994, 24, 433–441. [Google Scholar] [CrossRef]
- Legal, L.; Chappe, B.; Jallon, J.M. Molecular Basis ofMorinda Citrifolia (L.): Toxicity on Drosophila. J. Chem. Ecol. 1994, 20, 1931–1943. [Google Scholar] [CrossRef] [PubMed]
- Mansourian, S.; Stensmyr, M.C. The Chemical Ecology of the Fly. Curr. Opin. Neurobiol. 2015, 34, 95–102. [Google Scholar] [CrossRef]
- Prieto-Godino, L.L.; Rytz, R.; Bargeton, B.; Abuin, L.; Arguello, J.R.; Peraro, M.D.; Benton, R. Olfactory Receptor Pseudo-Pseudogenes. Nature 2016, 539, 93–97. [Google Scholar] [CrossRef]
- Sung, H.Y.; Jeong, Y.T.; Lim, J.Y.; Kim, H.; Oh, S.M.; Hwang, S.W.; Kwon, J.Y.; Moon, S.J. Heterogeneity in the Drosophila Gustatory Receptor Complexes That Detect Aversive Compounds. Nat. Commun. 2017, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Gloss, A.D.; Groen, S.C.; Whiteman, N.K. A Genomic Perspective on the Generation and Maintenance of Genetic Diversity in Herbivorous Insects. Annu. Rev. Ecol. Evol. Syst. 2016, 47, 165–187. [Google Scholar] [CrossRef]
| Position | Alleles (ref./alt.) | Gene | Effect | Allele Frequencies | FST | ||
|---|---|---|---|---|---|---|---|
| Cameroon | Kenya | Mayotte | |||||
| X:10060167 | G/A | Or13a | Downstream | 8/4 | 1/11 | 22/0 | 0.917 |
| X:13654476 | T/A | Ir7a | Downstream | 5/9 | 10/0 | 4/40 | 0.909 |
| X:13654477 | C/A | Ir7a | Downstream | 5/9 | 10/0 | 4/40 | 0.909 |
| 2L:1497713 | G/T | Or22a | Synonymous | 6/10 | 15/1 | 0/44 | 0.938 |
| 2L:1497721 | C/A | Or22a | Synonymous | 6/10 | 15/1 | 0/44 | 0.938 |
| 2L:1497723 | T/G | Or22a | Nonsynonymous (V/G) | 6/10 | 15/1 | 0/44 | 0.938 |
| 2L:1497724 | A/G | Or22a | Nonsynonymous (V/G) | 6/10 | 15/1 | 0/44 | 0.938 |
| 2L:1497730 | C/T | Or22a | Synonymous | 6/14 | 15/1 | 0/44 | 0.938 |
| 2L:1497751 | A/G | Or22a | Nonsynonymous (I/M) | 9/5 | 16/0 | 0/44 | 1.000 |
| 2L:1497754 | T/C | Or22a | Synonymous | 9/5 | 16/0 | 0/44 | 1.000 |
| 2L:1497757 | T/C | Or22a | Synonymous | 5/13 | 16/0 | 0/44 | 1.000 |
| 2L:1764188 | A/C | Gr22d | Downstream | 4/14 | 0/20 | 19/3 | 0.864 |
| 2L:1768039 | T/A | Gr22b | Synonymous | 20/0 | 20/0 | 2/20 | 0.909 |
| 2R:18929879 | G/A | Gr59a | Upstream | 16/4 | 20/0 | 5/39 | 0.886 |
| 3R:18490921 | G/A | Gr93b | Downstream | 16/2 | 20/0 | 3/19 | 0.864 |
| 3R:18494463 | A/G | Gr93c | Nonsynonymous (F/S) | 12/8 | 18/0 | 4/40 | 0.909 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ferreira, E.A.; Lambert, S.; Verrier, T.; Marion-Poll, F.; Yassin, A. Soft Selective Sweep on Chemosensory Genes Correlates with Ancestral Preference for Toxic Noni in a Specialist Drosophila Population. Genes 2021, 12, 32. https://doi.org/10.3390/genes12010032
Ferreira EA, Lambert S, Verrier T, Marion-Poll F, Yassin A. Soft Selective Sweep on Chemosensory Genes Correlates with Ancestral Preference for Toxic Noni in a Specialist Drosophila Population. Genes. 2021; 12(1):32. https://doi.org/10.3390/genes12010032
Chicago/Turabian StyleFerreira, Erina A., Sophia Lambert, Thibault Verrier, Frédéric Marion-Poll, and Amir Yassin. 2021. "Soft Selective Sweep on Chemosensory Genes Correlates with Ancestral Preference for Toxic Noni in a Specialist Drosophila Population" Genes 12, no. 1: 32. https://doi.org/10.3390/genes12010032
APA StyleFerreira, E. A., Lambert, S., Verrier, T., Marion-Poll, F., & Yassin, A. (2021). Soft Selective Sweep on Chemosensory Genes Correlates with Ancestral Preference for Toxic Noni in a Specialist Drosophila Population. Genes, 12(1), 32. https://doi.org/10.3390/genes12010032






