Genetic Reconstruction and Forensic Analysis of Chinese Shandong and Yunnan Han Populations by Co-Analyzing Y Chromosomal STRs and SNPs
Abstract
1. Introduction
2. Methods and Materials
2.1. DNA Samples
2.2. Typing of 24 Y Haplogroup-Determining SNP Markers and 27 Y-STR Loci
2.3. Data Analysis
2.3.1. Forensic Parameters and Statistical Analysis
2.3.2. Y-Chromosomal Haplogroup-Based Network Analysis
2.3.3. Phylogenetic Reconstruction on the Population Level
2.3.4. Phylogenetic Reconstruction on the Individual Level
3. Result and Discussion
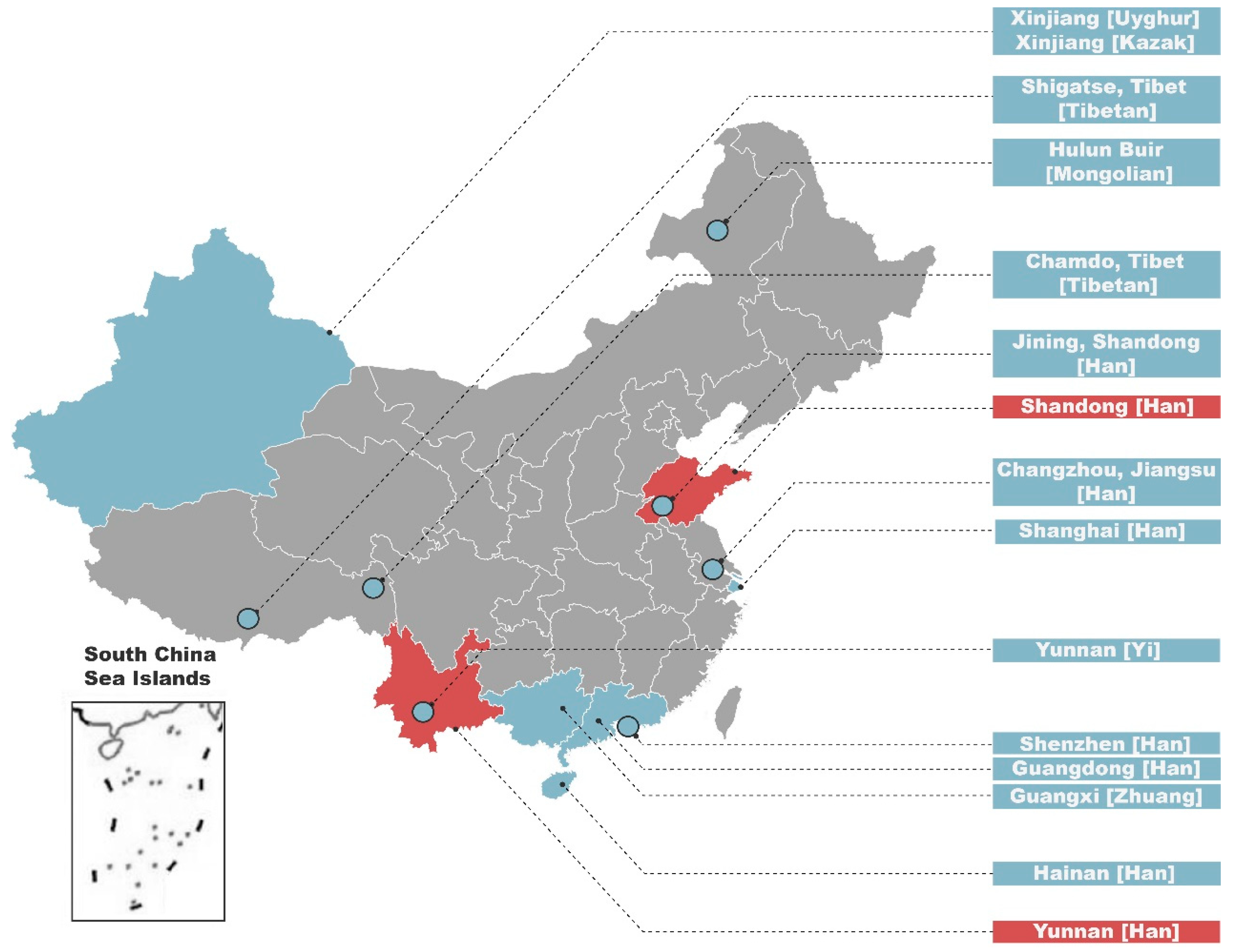
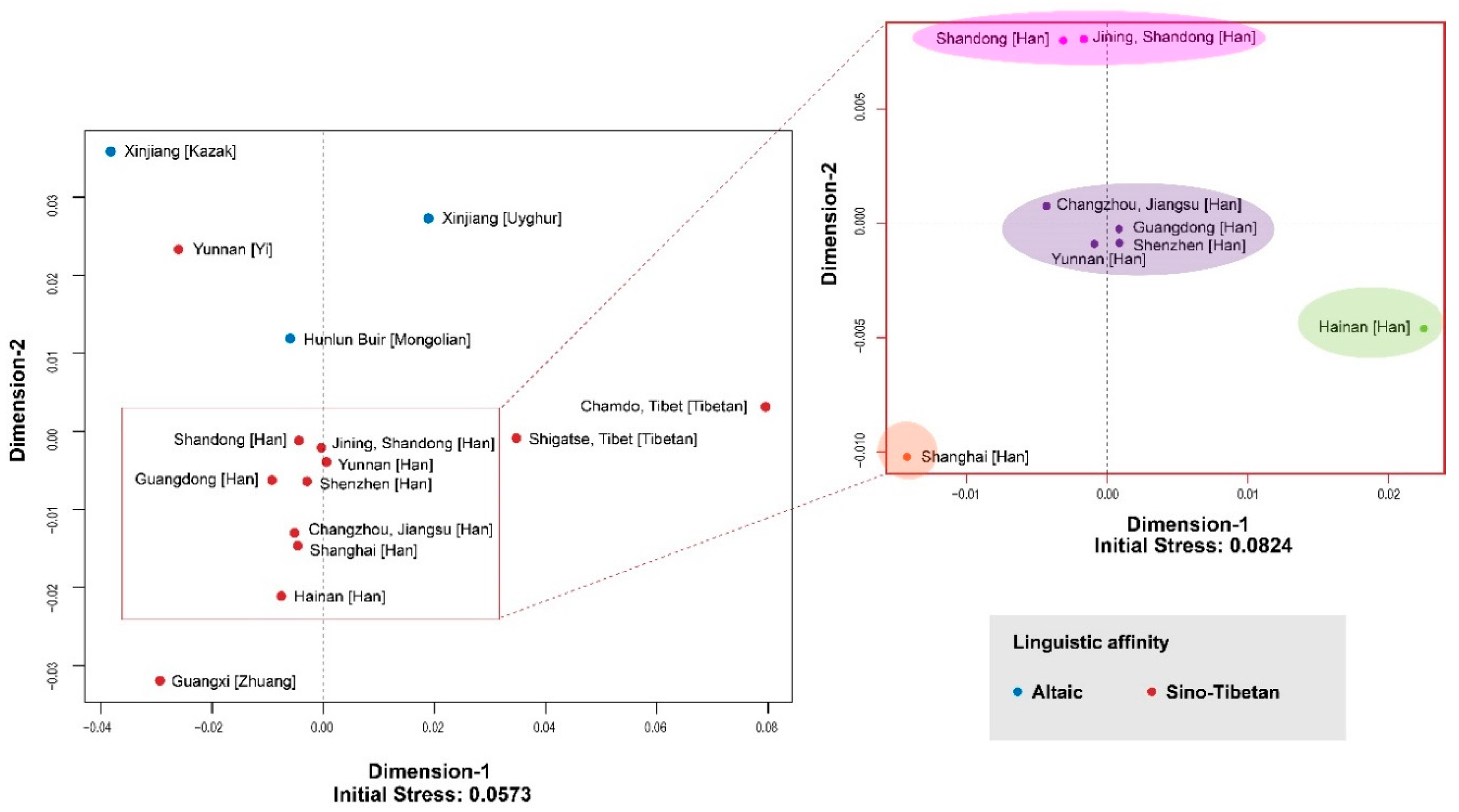
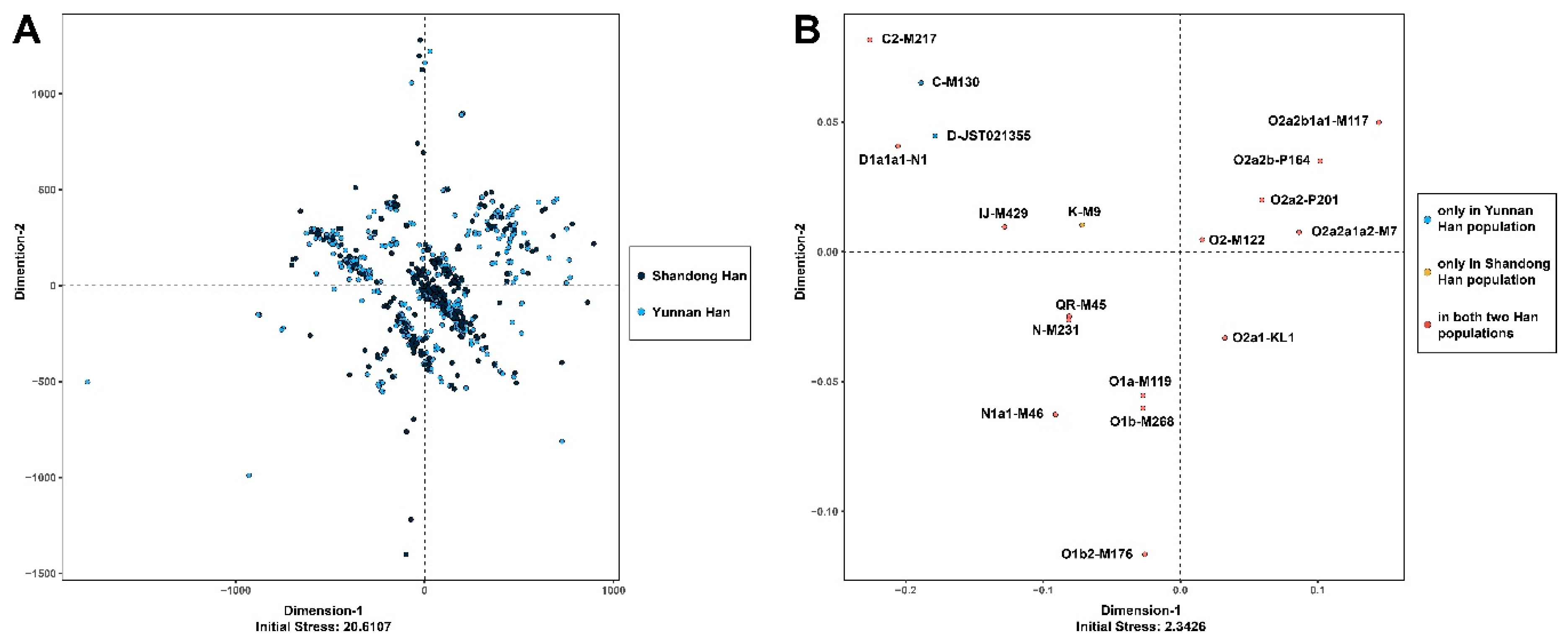
3.1. MDS and AMOVA
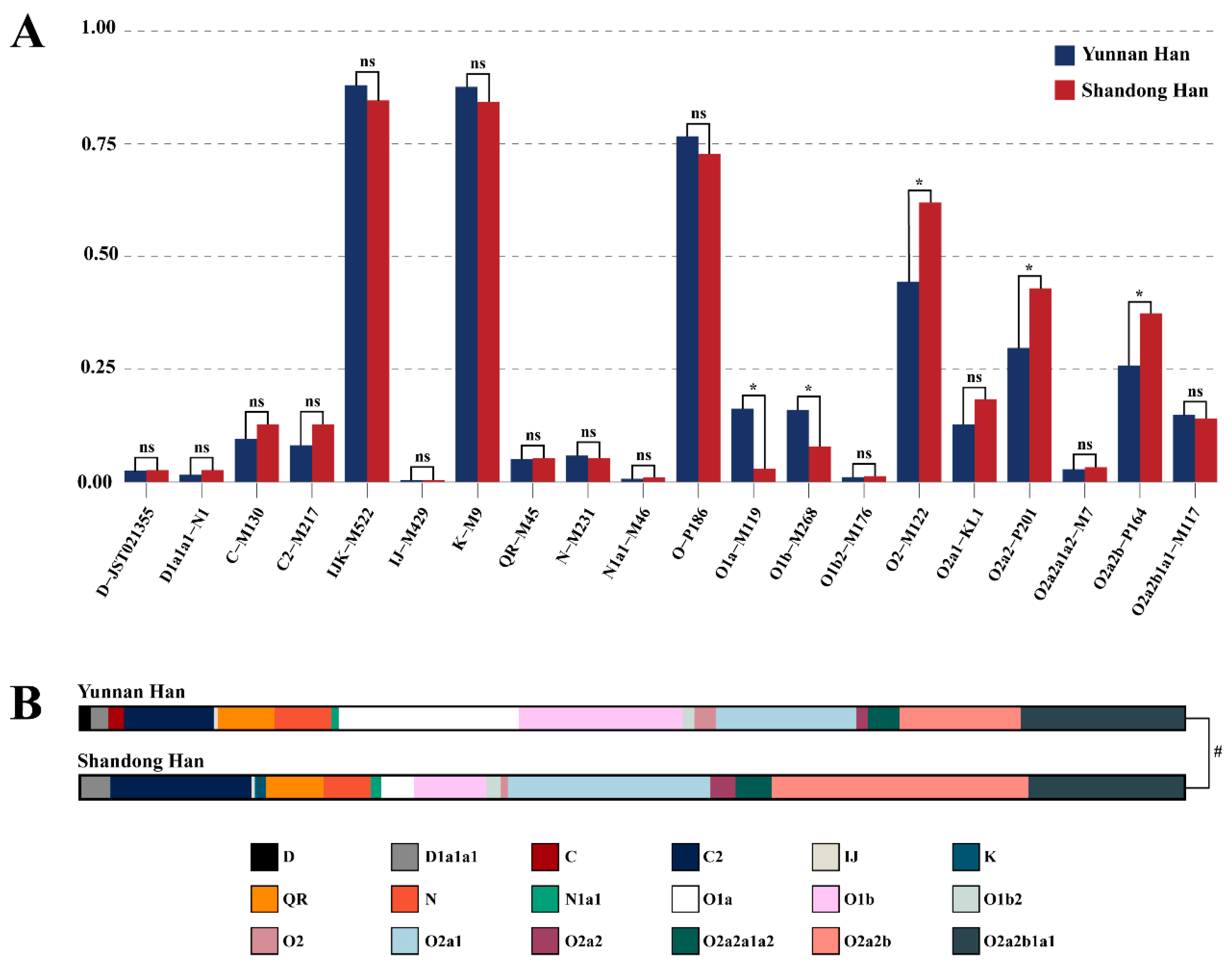
3.2. Y-SNP Allelic Frequencies and Haplogroup Distribution
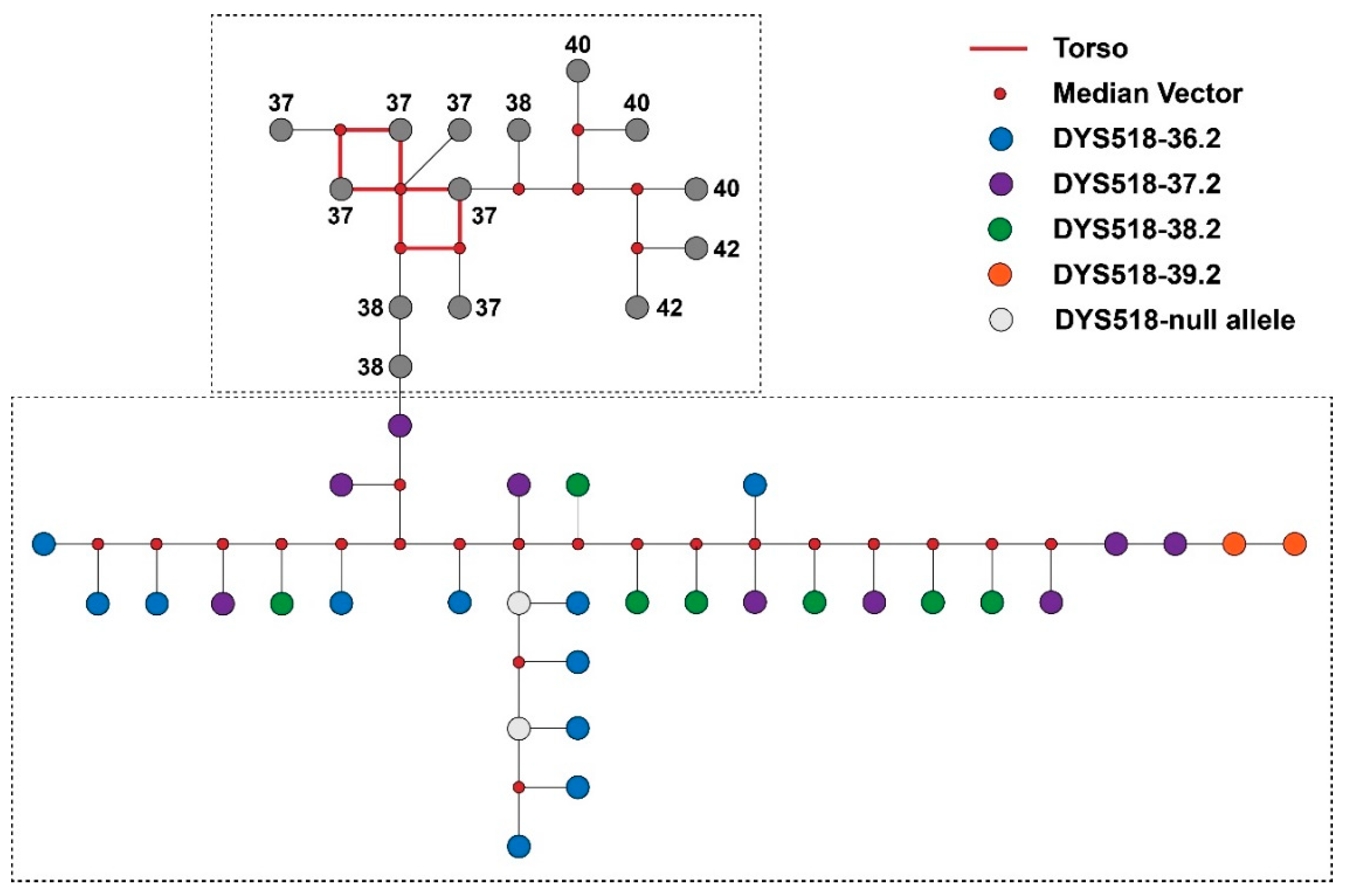
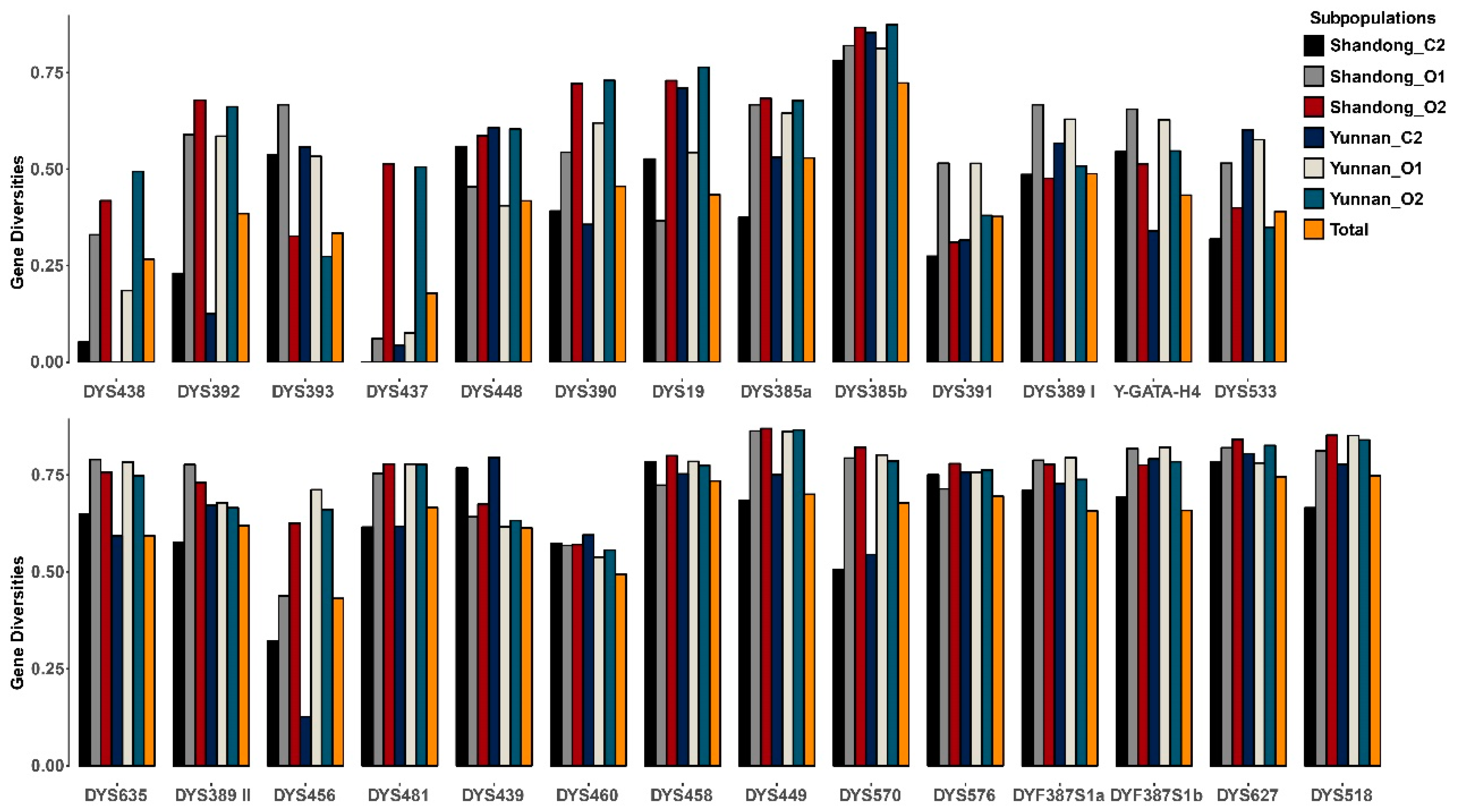
3.3. Y-STR Allele Variants
3.4. Distribution Spectrums of Y-STR Alleles within C2, O1, and O2 Haplogroups in Shandong and Yunnan Han Populations
3.5. Forensic Parameters
3.6. Phylogenetic Reconstruction on the Individual Level
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Santos, F.R.; Epplen, J.T.; Pena, S.D. Testing deficiency paternity cases with a Y-linked tetranucleotide repeat polymorphism. EXS 1993, 67, 261–265. [Google Scholar] [CrossRef]
- Kayser, M. Forensic use of Y-chromosome DNA: A general overview. Hum. Genet. 2017, 136, 621–635. [Google Scholar] [CrossRef]
- Corach, D.; Filgueira Risso, L.; Marino, M.; Penacino, G.; Sala, A. Routine Y-STR typing in forensic casework. Forensic Sci. Int. 2001, 118, 131–135. [Google Scholar] [CrossRef]
- Ballantyne, K.N.; Goedbloed, M.; Fang, R.; Schaap, O.; Lao, O.; Wollstein, A.; Choi, Y.; van Duijn, K.; Vermeulen, M.; Brauer, S.; et al. Mutability of Y-chromosomal microsatellites: Rates, characteristics, molecular bases, and forensic implications. Am. J. Hum. Genet. 2010, 87, 341–353. [Google Scholar] [CrossRef]
- Jobling, M.A.; Tyler-Smith, C. Human Y-chromosome variation in the genome-sequencing era. Nat. Rev. Genet. 2017, 18, 485–497. [Google Scholar] [CrossRef]
- Consortium, Y.C. A nomenclature system for the tree of human Y-chromosomal binary haplogroups. Genome Res. 2002, 12, 339–348. [Google Scholar] [CrossRef] [PubMed]
- Karafet, T.M.; Mendez, F.L.; Meilerman, M.B.; Underhill, P.A.; Zegura, S.L.; Hammer, M.F. New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res. 2008, 18, 830–838. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, C.; Yuan, L.; Ling, Y.; Wang, X.; Liu, C.; Pan, Y.; Zhang, X.; Ma, X.; Wang, Y.; et al. PGG.Han: The Han Chinese genome database and analysis platform. Nucleic Acids Res. 2020, 48, D971–D976. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Yin, X.; Li, S.; Jin, W.; Lou, H.; Yang, L.; Gong, X.; Wang, H.; Shen, Y.; Pan, X.; et al. Genomic dissection of population substructure of Han Chinese and its implication in association studies. Am. J. Hum. Genet. 2009, 85, 762–774. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, Y.; Kubo, S. Y-SNP and Y-STR analysis in a Japanese population. Leg. Med. (Tokyo) 2008, 10, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Watahiki, H.; Fujii, K.; Fukagawa, T.; Mita, Y.; Kitayama, T.; Mizuno, N. Polymorphisms and microvariant sequences in the Japanese population for 25 Y-STR markers and their relationships to Y-chromosome haplogroups. Forensic Sci. Int. Genet. 2019, 41, e1–e7. [Google Scholar] [CrossRef] [PubMed]
- Cortellini, V.; Verzeletti, A.; Cerri, N.; Marino, A.; De Ferrari, F. Y-chromosome polymorphisms and ethnic group—A combined STR and SNP approach in a population sample from northern Italy. Croat. Med. J. 2013, 54, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Erhart, D.; Berger, B.; Niederstatter, H.; Gassner, C.; Schennach, H.; Parson, W. Frequency data for 17 Y-chromosomal STRs and 19 Y-chromosomal SNPs in the Tyrolean district of Reutte, Austria. Int. J. Legal. Med. 2012, 126, 977–978. [Google Scholar] [CrossRef] [PubMed]
- Vullo, C.; Gomes, V.; Romanini, C.; Oliveira, A.M.; Rocabado, O.; Aquino, J.; Amorim, A.; Gusmao, L. Association between Y haplogroups and autosomal AIMs reveals intra-population substructure in Bolivian populations. Int. J. Legal. Med. 2015, 129, 673–680. [Google Scholar] [CrossRef]
- D’Atanasio, E.; Iacovacci, G.; Pistillo, R.; Bonito, M.; Dugoujon, J.M.; Moral, P.; El-Chennawi, F.; Melhaoui, M.; Baali, A.; Cherkaoui, M.; et al. Rapidly mutating Y-STRs in rapidly expanding populations: Discrimination power of the Yfiler Plus multiplex in northern Africa. Forensic Sci. Int. Genet. 2019, 38, 185–194. [Google Scholar] [CrossRef]
- Villaescusa, P.; Palencia-Madrid, L.; Campaner, M.A.; Jauregui-Rada, J.; Guerra-Rodriguez, M.; Rocandio, A.M.; de Pancorbo, M.M. Effective resolution of the Y chromosome sublineages of the Iberian haplogroup R1b-DF27 with forensic purposes. Int. J. Legal. Med. 2019, 133, 17–23. [Google Scholar] [CrossRef]
- Khubrani, Y.M.; Wetton, J.H.; Jobling, M.A. Extensive geographical and social structure in the paternal lineages of Saudi Arabia revealed by analysis of 27 Y-STRs. Forensic Sci. Int. Genet. 2018, 33, 98–105. [Google Scholar] [CrossRef]
- Yin, C.; Ren, Y.; Adnan, A.; Tian, J.; Guo, K.; Xia, M.; He, Z.; Zhai, D.; Chen, X.; Wang, L.; et al. Title: Developmental validation of Y-SNP pedigree tagging system: A panel via quick ARMS PCR. Forensic Sci. Int. Genet. 2020, 46, 102271. [Google Scholar] [CrossRef]
- van Oven, M.; Van Geystelen, A.; Kayser, M.; Decorte, R.; Larmuseau, M.H. Seeing the wood for the trees: A minimal reference phylogeny for the human Y chromosome. Hum. Mutat. 2014, 35, 187–191. [Google Scholar] [CrossRef]
- Willuweit, S.; Roewer, L.; International Forensic Y Chromosome User Group. Y chromosome haplotype reference database (YHRD): Update. Forensic Sci. Int. Genet. 2007, 1, 83–87. [Google Scholar] [CrossRef]
- Willuweit, S.; Roewer, L. The new Y Chromosome Haplotype Reference Database. Forensic Sci. Int. Genet. 2015, 15, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, S.; Zhong, C.; Nguyen, V.; Ge, J.; Lagace, R.E.; Short, M.L.; Mulero, J.J. Developmental validation of the Yfiler((R)) Plus PCR Amplification Kit: An enhanced Y-STR multiplex for casework and database applications. Forensic Sci. Int. Genet. 2016, 24, 164–175. [Google Scholar] [CrossRef]
- Nei, M.; Tajima, F. DNA polymorphism detectable by restriction endonucleases. Genetics 1981, 97, 145–163. [Google Scholar] [PubMed]
- Siegert, S.; Roewer, L.; Nothnagel, M. Shannon’s equivocation for forensic Y-STR marker selection. Forensic Sci. Int. Genet. 2015, 16, 216–225. [Google Scholar] [CrossRef] [PubMed]
- de Meeus, T.; Goudet, J. A step-by-step tutorial to use HierFstat to analyse populations hierarchically structured at multiple levels. Infect. Genet. Evol. 2007, 7, 731–735. [Google Scholar] [CrossRef] [PubMed]
- Bandelt, H.J.; Forster, P.; Rohl, A. Median-joining networks for inferring intraspecific phylogenies. Mol. Biol. Evol. 1999, 16, 37–48. [Google Scholar] [CrossRef]
- Nothnagel, M.; Fan, G.; Guo, F.; He, Y.; Hou, Y.; Hu, S.; Huang, J.; Jiang, X.; Kim, W.; Kim, K.; et al. Revisiting the male genetic landscape of China: A multi-center study of almost 38,000 Y-STR haplotypes. Hum. Genet. 2017, 136, 485–497. [Google Scholar] [CrossRef]
- Mulero, J.J.; Chang, C.W.; Calandro, L.M.; Green, R.L.; Li, Y.; Johnson, C.L.; Hennessy, L.K. Development and validation of the AmpFlSTR Yfiler PCR amplification kit: A male specific, single amplification 17 Y-STR multiplex system. J. Forensic Sci. 2006, 51, 64–75. [Google Scholar] [CrossRef]
- Guo, F.; Li, J.; Chen, K.; Tang, R.; Zhou, L. Population genetic data for 27 Y-STR loci in the Zhuang ethnic minority from Guangxi Zhuang Autonomous Region in the south of China. Forensic Sci. Int. Genet. 2017, 27, 182–183. [Google Scholar] [CrossRef]
- Wang, C.Z.; Su, M.J.; Li, Y.; Chen, L.; Jin, X.; Wen, S.Q.; Tan, J.; Shi, M.S.; Li, H. Genetic polymorphisms of 27 Yfiler((R)) Plus loci in the Daur and Mongolian ethnic minorities from Hulunbuir of Inner Mongolia Autonomous Region, China. Forensic Sci. Int. Genet. 2019, 40, e252–e255. [Google Scholar] [CrossRef]
- Zhang, D.; Cao, G.; Xie, M.; Cui, X.; Xiao, L.; Tian, C.; Ye, Y. RETRACTED ARTICLE: Y Chromosomal STR haplotypes in Chinese Uyghur, Kazakh and Hui ethnic groups and genetic features of DYS448 null allele and DYS19 duplicated allele. Int. J. Legal. Med. 2019. [Google Scholar] [CrossRef]
- Fan, G.Y.; An, Y.R.; Peng, C.X.; Deng, J.L.; Pan, L.P.; Ye, Y. Forensic and phylogenetic analyses among three Yi populations in Southwest China with 27 Y chromosomal STR loci. Int. J. Legal. Med. 2019, 133, 795–797. [Google Scholar] [CrossRef] [PubMed]
- He, G.; Wang, Z.; Su, Y.; Zou, X.; Wang, M.; Chen, X.; Gao, B.; Liu, J.; Wang, S.; Hou, Y. Genetic structure and forensic characteristics of Tibeto-Burman-speaking U-Tsang and Kham Tibetan Highlanders revealed by 27 Y-chromosomal STRs. Sci. Rep. 2019, 9, 7739. [Google Scholar] [CrossRef] [PubMed]
- Du, W.; Feng, P.; Huang, H.; Wu, W.; Zhang, L.; Guo, Y.; Liu, C.; Liu, H.; Liu, C.; Chen, L. Technical note: Developmental validation of a novel 6-dye typing system with 36 Y-STR loci. Int. J. Legal. Med. 2019, 133, 1015–1027. [Google Scholar] [CrossRef]
- Wang, M.; Wang, Z.; Zhang, Y.; He, G.; Liu, J.; Hou, Y. Forensic characteristics and phylogenetic analysis of two Han populations from the southern coastal regions of China using 27 Y-STR loci. Forensic Sci. Int. Genet. 2017, 31, e17–e23. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.; Zhang, X.; Wang, X.; Ren, Z.; Li, W.; Long, R.; Liang, A.; Chen, J.; Song, T.; Qu, Y.; et al. Genetic analysis of 27 Y-STR loci in Han population from Hainan province, southernmost China. Forensic Sci. Int. Genet. 2018, 33, e9–e10. [Google Scholar] [CrossRef]
- Zhou, Y.; Shao, C.; Li, L.; Zhang, Y.; Liu, B.; Yang, Q.; Tang, Q.; Li, S.; Xie, J. Genetic analysis of 29 Y-STR loci in the Chinese Han population from Shanghai. Forensic Sci. Int. Genet. 2018, 32, e1–e4. [Google Scholar] [CrossRef]
- Tao, R.; Jin, M.; Ji, G.; Zhang, J.; Zhang, J.; Yang, Z.; Chen, C.; Zhang, S.; Li, C. Forensic characteristics of 36 Y-STR loci in a Changzhou Han population and genetic distance analysis among several Chinese populations. Forensic Sci. Int. Genet. 2019, 40, e268–e270. [Google Scholar] [CrossRef]
- Wang, Y.; Dang, Z.; Zhang, G.; Li, S.; Liu, Q.; Li, C.; Hou, X.; Li, H.; Chen, S.; Cui, W.; et al. Genetic diversity and haplotype structure of 27 Y-STR loci in a Han population from Jining, Shandong province, eastern China. Forensic Sci. Int. Genet. 2019, 42, e25–e26. [Google Scholar] [CrossRef]
- Excoffier, L.; Lischer, H.E. Arlequin suite ver 3.5: A new series of programs to perform population genetics analyses under Linux and Windows. Mol. Ecol. Resour. 2010, 10, 564–567. [Google Scholar] [CrossRef]
- Excoffier, L.; Smouse, P.E.; Quattro, J.M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 1992, 131, 479–491. [Google Scholar]
- Kruskal, J.B. Multidimensional-Scaling by Optimizing Goodness of Fit to a Nonmetric Hypothesis. Psychometrika 1964, 29, 1–27. [Google Scholar] [CrossRef]
- Nei, M. Phylogenetic analysis in molecular evolutionary genetics. Annu. Rev. Genet. 1996, 30, 371–403. [Google Scholar] [CrossRef] [PubMed]
- Nei, M. Genetic Distance, in Encyclopedia of Genetics; Brenner, S., Miller, J.H., Eds.; Academic Press: New York, NY, USA, 2001. [Google Scholar]
- Murtagh, F.; Contreras, P. Algorithms for hierarchical clustering: An overview, II. Wires Data Min. Knowl. 2017, 7. [Google Scholar] [CrossRef]
- Cavalli-Sforza, L.L.; Piazza, A.; Menozzi, P.; Mountain, J. Reconstruction of human evolution: Bringing together genetic, archaeological, and linguistic data. Proc. Natl. Acad. Sci. USA 1988, 85, 6002–6006. [Google Scholar] [CrossRef]
- Longobardi, G.; Ghirotto, S.; Guardiano, C.; Tassi, F.; Benazzo, A.; Ceolin, A.; Barbujani, G. Across language families: Genome diversity mirrors linguistic variation within Europe. Am. J. Phys. Anthropol. 2015, 157, 630–640. [Google Scholar] [CrossRef]
- Zhang, M.; Yan, S.; Pan, W.; Jin, L. Phylogenetic evidence for Sino-Tibetan origin in northern China in the Late Neolithic. Nature 2019, 569, 112–115. [Google Scholar] [CrossRef]
- Wu, W.P. Migrant settlement and spatial distribution in metropolitan Shanghai. Prof. Geogr. 2008, 60, 101–120. [Google Scholar] [CrossRef]
- Deng, W.; Shi, B.; He, X.; Zhang, Z.; Xu, J.; Li, B.; Yang, J.; Ling, L.; Dai, C.; Qiang, B.; et al. Evolution and migration history of the Chinese population inferred from Chinese Y-chromosome evidence. J. Hum. Genet. 2004, 49, 339–348. [Google Scholar] [CrossRef]
- Zhong, H.; Shi, H.; Qi, X.B.; Duan, Z.Y.; Tan, P.P.; Jin, L.; Su, B.; Ma, R.Z. Extended Y chromosome investigation suggests postglacial migrations of modern humans into East Asia via the northern route. Mol Biol. Evol. 2011, 28, 717–727. [Google Scholar] [CrossRef]
- Wells, R.S.; Yuldasheva, N.; Ruzibakiev, R.; Underhill, P.A.; Evseeva, I.; Blue-Smith, J.; Jin, L.; Su, B.; Pitchappan, R.; Shanmugalakshmi, S.; et al. The Eurasian heartland: A continental perspective on Y-chromosome diversity. Proc. Natl. Acad. Sci. USA 2001, 98, 10244–10249. [Google Scholar] [CrossRef] [PubMed]
- Wen, B.; Li, H.; Lu, D.; Song, X.; Zhang, F.; He, Y.; Li, F.; Gao, Y.; Mao, X.; Zhang, L.; et al. Genetic evidence supports demic diffusion of Han culture. Nature 2004, 431, 302–305. [Google Scholar] [CrossRef]
- Yan, S.; Wang, C.C.; Li, H.; Li, S.L.; Jin, L.; Genographic, C. An updated tree of Y-chromosome Haplogroup O and revised phylogenetic positions of mutations P164 and PK4. Eur. J. Hum. Genet. 2011, 19, 1013–1015. [Google Scholar] [CrossRef] [PubMed]
- Fornarino, S.; Pala, M.; Battaglia, V.; Maranta, R.; Achilli, A.; Modiano, G.; Torroni, A.; Semino, O.; Santachiara-Benerecetti, S.A. Mitochondrial and Y-chromosome diversity of the Tharus (Nepal): A reservoir of genetic variation. BMC Evol. Biol. 2009, 9, 154. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Mo, X.; Shang, L.; Jin, X.; Chen, D.; Zhu, H.; Zhang, Y.; Kang, B.; Li, W.; Ye, J. Genetic analysis of 29 Y-STR loci in Han population from Dongfang, Southern China. Int. J. Legal. Med. 2019, 133, 1033–1035. [Google Scholar] [CrossRef]
- Wang, H.; Ba, H.; Yang, C.; Zhang, J.; Tai, Y. Inner and inter population structure construction of Chinese Jiangsu Han population based on Y23 STR system. PLoS ONE 2017, 12, e0180921. [Google Scholar] [CrossRef]
- Chen, P.; Han, Y.; He, G.; Luo, H.; Gao, T.; Song, F.; Wan, D.; Yu, J.; Hou, Y. Genetic diversity and phylogenetic study of the Chinese Gelao ethnic minority via 23 Y-STR loci. Int. J. Legal. Med. 2018, 132, 1093–1096. [Google Scholar] [CrossRef]
- Liu, C.; Han, X.; Min, Y.; Liu, H.; Xu, Q.; Yang, X.; Huang, S.; Chen, Z.; Liu, C. Genetic polymorphism analysis of 40 Y-chromosomal STR loci in seven populations from South China. Forensic Sci. Int. 2018, 291, 109–114. [Google Scholar] [CrossRef]
- Du, W.; Wu, W.; Wu, Z.; Guo, L.; Wang, B.; Chen, L. Genetic polymorphisms of 32 Y-STR loci in Meizhou Hakka population. Int. J. Legal. Med. 2019, 133, 465–466. [Google Scholar] [CrossRef]
- Zhang, J.; Mo, X.; Zhang, Y.; Ding, G.; Wang, X.; Li, W.; Zhao, X.; Ye, J. Genetic analysis of 26 Y-STR loci in Han population from Leshan, Southwest China. Forensic Sci. Int. Genet. 2018, 37, e15–e16. [Google Scholar] [CrossRef]
- Ye, Y.; Gao, J.; Fan, G.; Liao, L.; Hou, Y. Population genetics for 23 Y-STR loci in Tibetan in China and confirmation of DYS448 null allele. Forensic Sci. Int. Genet. 2015, 16, e7–e10. [Google Scholar] [CrossRef] [PubMed]
- Hu, L.; Gu, T.; Fan, X.; Yuan, X.; Rao, M.; Pang, J.B.; Nie, A.; Du, L.; Zhang, X.; Nie, S. Genetic polymorphisms of 24 Y-STR loci in Hani ethnic minority from Yunnan Province, Southwest China. Int. J. Legal. Med. 2017, 131, 1235–1237. [Google Scholar] [CrossRef] [PubMed]
- Cao, S.; Bai, P.; Zhu, W.; Chen, D.; Wang, H.; Jin, B.; Zhang, L.; Liang, W. Genetic portrait of 27 Y-STR loci in the Tibetan ethnic population of the Qinghai province of China. Forensic Sci. Int. Genet. 2018, 34, e18–e19. [Google Scholar] [CrossRef] [PubMed]
- Grugni, V.; Raveane, A.; Ongaro, L.; Battaglia, V.; Trombetta, B.; Colombo, G.; Capodiferro, M.R.; Olivieri, A.; Achilli, A.; Perego, U.A.; et al. Analysis of the human Y-chromosome haplogroup Q characterizes ancient population movements in Eurasia and the Americas. BMC Biol. 2019, 17, 3. [Google Scholar] [CrossRef] [PubMed]
- Lang, M.; Liu, H.; Song, F.; Qiao, X.; Ye, Y.; Ren, H.; Li, J.; Huang, J.; Xie, M.; Chen, S.; et al. Forensic characteristics and genetic analysis of both 27 Y-STRs and 143 Y-SNPs in Eastern Han Chinese population. Forensic Sci. Int. Genet. 2019, 42, e13–e20. [Google Scholar] [CrossRef]
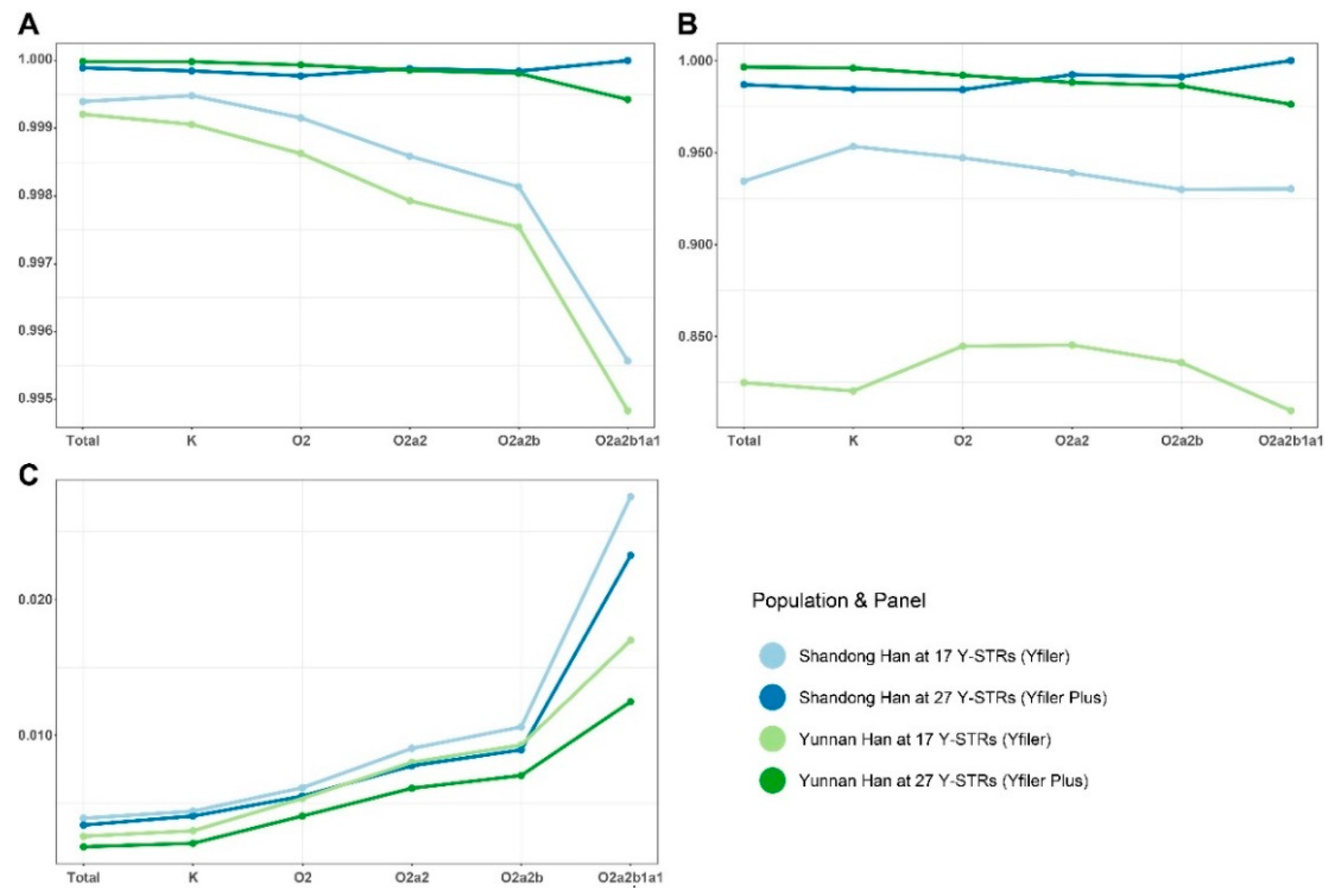

| Population | Panel | HD | DC | MP | Unique Haplotypes | Sample Size |
|---|---|---|---|---|---|---|
| Yunnan Han | ||||||
| Total | Yfiler-17 | 0.99921 | 0.82478 | 0.00256 | 466 | 565 |
| YfilerPlus-27 | 0.99999 | 0.99646 | 0.00178 | 563 | 565 | |
| K | Yfiler-17 | 0.99906 | 0.82020 | 0.00296 | 406 | 495 |
| YfilerPlus-27 | 0.99998 | 0.99596 | 0.00204 | 493 | 495 | |
| O2 | Yfiler-17 | 0.99863 | 0.84462 | 0.00535 | 212 | 251 |
| YfilerPlus-27 | 0.99994 | 0.99203 | 0.00405 | 249 | 251 | |
| O2a2 | Yfiler-17 | 0.99793 | 0.84524 | 0.00801 | 142 | 168 |
| YfilerPlus-27 | 0.99986 | 0.98810 | 0.00609 | 166 | 168 | |
| O2a2b | Yfiler-17 | 0.99754 | 0.83562 | 0.00929 | 122 | 146 |
| YfilerPlus-27 | 0.99981 | 0.98630 | 0.00704 | 144 | 146 | |
| O2a2b1a1 | Yfiler-17 | 0.99484 | 0.80952 | 0.01701 | 68 | 84 |
| YfilerPlus-27 | 0.99943 | 0.97619 | 0.01247 | 82 | 84 | |
| Shandong Han | ||||||
| Total | Yfiler-17 | 0.99940 | 0.93443 | 0.00388 | 285 | 305 |
| YfilerPlus-27 | 0.99989 | 0.98689 | 0.00339 | 301 | 305 | |
| K | Yfiler-17 | 0.99948 | 0.95331 | 0.00441 | 245 | 257 |
| YfilerPlus-27 | 0.99985 | 0.98444 | 0.00404 | 253 | 257 | |
| O2 | Yfiler-17 | 0.99916 | 0.94709 | 0.00613 | 179 | 189 |
| YfilerPlus-27 | 0.99977 | 0.98413 | 0.00551 | 186 | 189 | |
| O2a2 | Yfiler-17 | 0.99859 | 0.93893 | 0.00903 | 123 | 131 |
| YfilerPlus-27 | 0.99988 | 0.99237 | 0.00775 | 130 | 131 | |
| O2a2b | Yfiler-17 | 0.99814 | 0.92982 | 0.01062 | 106 | 114 |
| YfilerPlus-27 | 0.99984 | 0.99123 | 0.00893 | 113 | 114 | |
| O2a2b1a1 | Yfiler-17 | 0.99557 | 0.93023 | 0.02758 | 40 | 43 |
| YfilerPlus-27 | 1.00000 | 1.00000 | 0.02326 | 43 | 43 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yin, C.; Su, K.; He, Z.; Zhai, D.; Guo, K.; Chen, X.; Jin, L.; Li, S. Genetic Reconstruction and Forensic Analysis of Chinese Shandong and Yunnan Han Populations by Co-Analyzing Y Chromosomal STRs and SNPs. Genes 2020, 11, 743. https://doi.org/10.3390/genes11070743
Yin C, Su K, He Z, Zhai D, Guo K, Chen X, Jin L, Li S. Genetic Reconstruction and Forensic Analysis of Chinese Shandong and Yunnan Han Populations by Co-Analyzing Y Chromosomal STRs and SNPs. Genes. 2020; 11(7):743. https://doi.org/10.3390/genes11070743
Chicago/Turabian StyleYin, Caiyong, Kaiyuan Su, Ziwei He, Dian Zhai, Kejian Guo, Xueyun Chen, Li Jin, and Shilin Li. 2020. "Genetic Reconstruction and Forensic Analysis of Chinese Shandong and Yunnan Han Populations by Co-Analyzing Y Chromosomal STRs and SNPs" Genes 11, no. 7: 743. https://doi.org/10.3390/genes11070743
APA StyleYin, C., Su, K., He, Z., Zhai, D., Guo, K., Chen, X., Jin, L., & Li, S. (2020). Genetic Reconstruction and Forensic Analysis of Chinese Shandong and Yunnan Han Populations by Co-Analyzing Y Chromosomal STRs and SNPs. Genes, 11(7), 743. https://doi.org/10.3390/genes11070743




