Four Individuals with a Homozygous Mutation in Exon 1f of the PLEC Gene and Associated Myasthenic Features
Abstract
1. Introduction
2. Materials and Methods
2.1. Clinical Assessment
2.2. Standard Protocol Approvals, Registrations, and Patient Consents
2.3. Genetic Analysis
3. Results
3.1. Clinical Presentation
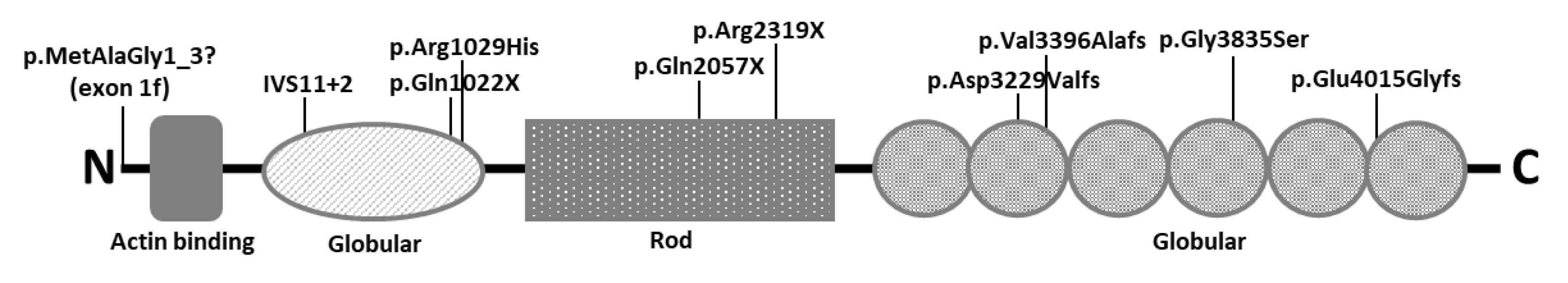
3.2. Variant Characterization
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Liu, C.G.; Maercker, C.; Castañon, M.J.; Hauptmann, R.; Wiche, G. Human plectin: Organization of the gene, sequence analysis, and chromosome localization (8q24). Proc. Natl. Acad. Sci. USA 1996, 93, 4278–4283. [Google Scholar] [CrossRef]
- Elliott, C.E.; Becker, B.; Oehler, S.; Castañón, M.J.; Hauptmann, R.; Wiche, G. Plectin transcript diversity: Identification and tissue distribution of variants with distinct first coding exons and rodless isoforms. Genomics 1997, 42, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Wiche, G.; Osmanagic-Myers, S.; Castanon, M.J. Networking and anchoring through plectin: A key to IF functionality and mechanotransduction. Curr. Opin. Cell Biol. 2014, 32, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Rezniczek, G.A.; Konieczny, P.; Nikolic, B.; Reipert, S.; Schneller, D.; Abrahamsberg, C.; Davies, K.E.; Winder, S.J.; Wiche, G. Plectin 1f scaffolding at the sarcolemma of dystrophic (mdx) muscle fibers through multiple interactions with beta-dystroglycan. J. Cell Biol. 2007, 176, 965–977. [Google Scholar] [CrossRef] [PubMed]
- Mihailovska, E.; Raith, M.; Valencia, R.G.; Fischer, I.; Al Banchaabouchi, M.; Herbst, R.; Wiche, G. Neuromuscular synapse integrity requires linkage of acetylcholine receptors to postsynaptic intermediate filament networks via rapsyn-plectin 1f complexes. Mol. Biol. Cell 2014, 25, 4130–4149. [Google Scholar] [CrossRef]
- Rezniczek, G.A.; Winter, L.; Walko, G.; Wiche, G. Chapter thirteen—Functional and genetic analysis of plectin in skin and muscle. In Methods in Enzymology; Academic Press: Cambridge, MA, USA, 2016; Volume 569, pp. 235–259. [Google Scholar]
- Pulkkinen, L.; Smith, F.J.D.; Shimizu, H.; Murata, S.; Yaoita, H.; Hachisuka, H.; Nishikawa, T.; McLean, W.H.I.; Uitto, J. Homozygous deletion mutations in the plectin gene (PLEC1) in patients with epidermolysis bullosa simplex associated with late-onset muscular dystrophy. Hum. Mol. Genet. 1996, 5, 1539–1546. [Google Scholar] [CrossRef]
- Fine, J.D.; Stenn, J.; Johnson, L.; Wright, T.; Bock, H.G.O.; Horiguchi, Y. Autosomal recessive epidermolysis bullosa simplex: Generalized phenotypic features suggestive of junctional or dystrophic epidermolysis bullosa, and association with neuromuscular diseases. Arch. Dermatol. 1989, 125, 931–938. [Google Scholar] [CrossRef]
- Charlesworth, A.; Chiaverini, C.; Chevrant-Breton, J.; DelRio, M.; Diociaiuti, A.; Dupuis, R.P.; El Hachem, M.; Le Fiblec, B.; Sankari-Ho, A.M.; Valhquist, A.; et al. Epidermolysis bullosa simplex with PLEC mutations: New phenotypes and new mutations. Br. J. Dermatol. 2013, 168, 808–814. [Google Scholar] [CrossRef]
- Gundesli, H.; Talim, B.; Korkusuz, P.; Balci-Hayta, B.; Cirak, S.; Akarsu, N.A.; Topaloglu, H.; Dincer, P. Mutation in exon 1f of PLEC, leading to disruption of plectin isoform 1f, causes autosomal-recessive limb-girdle muscular dystrophy. Am. J. Hum. Genet. 2010, 87, 834–841. [Google Scholar] [CrossRef]
- Töpf, A.; Johnson, K.; Bates, A.; Phillips, L.; Chao, K.R.; England, E.M.; Laricchia, K.M.; Mullen, T.; Valkanas, E.; Xu, L.; et al. MYO-SEQ consortium, Straub, V. Sequential targeted exome sequencing of 1001 patients affected by unexplained limb-girdle weakness. Genet. Med. 2020. [Google Scholar] [CrossRef]
- Konieczny, P.; Fuchs, P.; Reipert, S.; Kunz, W.S.; Zeöld, A.; Fischer, I.; Paulin, D.; Schröder, R.; Wiche, G. Myofiber integrity depends on desmin network targeting to Z-disks and costameres via distinct plectin isoforms. J. Cell Biol. 2008, 181, 667–681. [Google Scholar] [CrossRef] [PubMed]
- Bolling, M.C.; Pas, H.H.; De Visser, M.; Aronica, E.; Pfendner, E.G.; Van den Berg, M.P.; Diercks, G.F.H.; Suurmeijer, A.J.H.; Jonkman, M.F. PLEC1 mutations underlie adult-onset dilated cardiomyopathy in epidermolysis bullosa simplex with muscular dystrophy. J. Investig. Dermatol. 2010, 130, 1178–1181. [Google Scholar] [CrossRef] [PubMed]
- Koss-Harnes, D.; Høyheim, B.; Jonkman, M.F.; De Groot, W.P.; De Weerdt, C.J.; Nikolic, B.; Wiche, G.; Gedde-Dahl, T., Jr. Life-long course and molecular characterization of the original Dutch family with epidermolysis bullosa simplex with muscular dystrophy due to a homozygous novel plectin point mutation. Acta Derm. Venereol. 2004, 84, 124–131. [Google Scholar] [PubMed]
- Kunz, M.; Rouan, F.; Pulkkinen, L.; Hamm, H.; Jeschke, R.; Bruckner-Tuderman, L.; Bröcker, E.B.; Wiche, G.; Uitto, J.; Zillikens, D. Mutation reports: Epidermolysis bullosa simplex associated with severe mucous membrane involvement and novel mutations in the plectin gene. J. Investig. Dermatol. 2000, 114, 376–380. [Google Scholar] [CrossRef]
- Doriguzzi, C.; Palmucci, L.; Mongini, T.; Bertolotto, A.; Maniscalco, M.; Chiadò-Piat, L.; Zina, A.M.; Bundino, S. Congenital muscular dystrophy associated with familial junctional epidermolysis bullosa letalis. Eur. Neurol. 1993, 33, 454–460. [Google Scholar] [CrossRef]
- Schröder, R.; Fürst, D.O.; Klasen, C.; Reimann, J.; Herrmann, H.; Van der Ven, P.F. Association of plectin with Z-discs is a prerequisite for the formation of the intermyofibrillar desmin cytoskeleton. Lab. Investig. 80 2000, 80, 455–464. [Google Scholar] [CrossRef][Green Version]
- Cowton, J.A.; Beattie, T.J.; Gibson, A.A.; Mackie, R.; Skerrow, C.J.; Cockburn, F. Epidermolysis bullosa in association with aplasia cutis congenita and pyloric atresia. Acta Paediatr. Scand. 1982, 71, 155–160. [Google Scholar] [CrossRef]
- Gedde-Dahl, T., Jr. Epidermolysis Bullosa: A Clinical, Genetic and Epidemiological Study; Johns Hopkins Press (Pub.): Baltimore, MD, USA, 1971. [Google Scholar]
- Deev, R.V.; Bardakov, S.N.; Mavlikeev, M.O.; Yakovlev, I.A.; Umakhanova, Z.R.U.; Akhmedova, P.G.; Magomedova, R.M.; Chekmaryeva, I.A.; Dalgatov, G.D.; Isaev, A.A. Glu20Ter variant in PLEC 1f isoform causes limb-girdle muscle dystrophy with lung injury. Front. Neurol. 2017, 8, 367. [Google Scholar] [CrossRef]
- Selcen, D.; Juel, V.C.; Hobson-Webb, L.D.; Smith, E.C.; Stickler, D.E.; Bite, A.V.; Ohno, K.; Engel, A.G. Myasthenic syndrome caused by plectinopathy. Neurology 2011, 76, 327–336. [Google Scholar] [CrossRef]
- Fattahi, Z.; Kahrizi, K.; Nafissi, S.; Fadaee, M.; Abedini, S.S.; Kariminejad, A.; Akbari, M.R.; Najmabadi, H. Report of a patient with limb-girdle muscular dystrophy, ptosis and ophthalmoparesis caused by plectinopathy. Arch. Iran. Med. 2015, 18, 60–64. [Google Scholar]
- Gonzalez Garcia, A.; Tutmaher, M.; Upadhyayula, S.; Sanchez Russo, R.; Verma, S. Novel PLEC gene variants causing congenital myasthenic syndrome. Muscle Nerve 2019, 60, E40–E43. [Google Scholar] [CrossRef] [PubMed]
- Forrest, K.; Mellerio, J.E.; Robb, S.; Dopping-Hepenstal, P.J.; McGrath, J.A.; Liu, L.; Buk, S.J.; Al-Sarraj, S.; Wraige, E.; Jungbluth, H. Congenital muscular dystrophy, myasthenic symptoms and epidermolysis bullosa simplex (EBS) associated with mutations in the PLEC1 gene encoding plectin. Neuromuscul. Disord. 2010, 20, 709–711. [Google Scholar] [CrossRef]
- Banwell, B.L.; Russel, J.; Fukudome, T.; Shen, X.M.; Stilling, G.; Engel, A.G. Myopathy, myasthenic syndrome, and epidermolysis bullosa simplex due to plectin deficiency. J. Neuropathol. Exp. Neurol. 1999, 58, 832–846. [Google Scholar] [CrossRef]
- Gache, Y.; Chavanas, S.; Lacour, J.P.; Wiche, G.; Owaribe, K.; Meneguzzi, G.; Ortonne, J.P. Defective expression of plectin/HD1 in epidermolysis bullosa simplex with muscular dystrophy. J. Clin. Investig. 1996, 97, 2289–2298. [Google Scholar] [CrossRef] [PubMed]
- Winter, L.; Wiche, G. The many faces of plectin and plectinopathies: Pathology and mechanisms. Acta Neuropathol. 2013, 125, 77–93. [Google Scholar] [CrossRef] [PubMed]
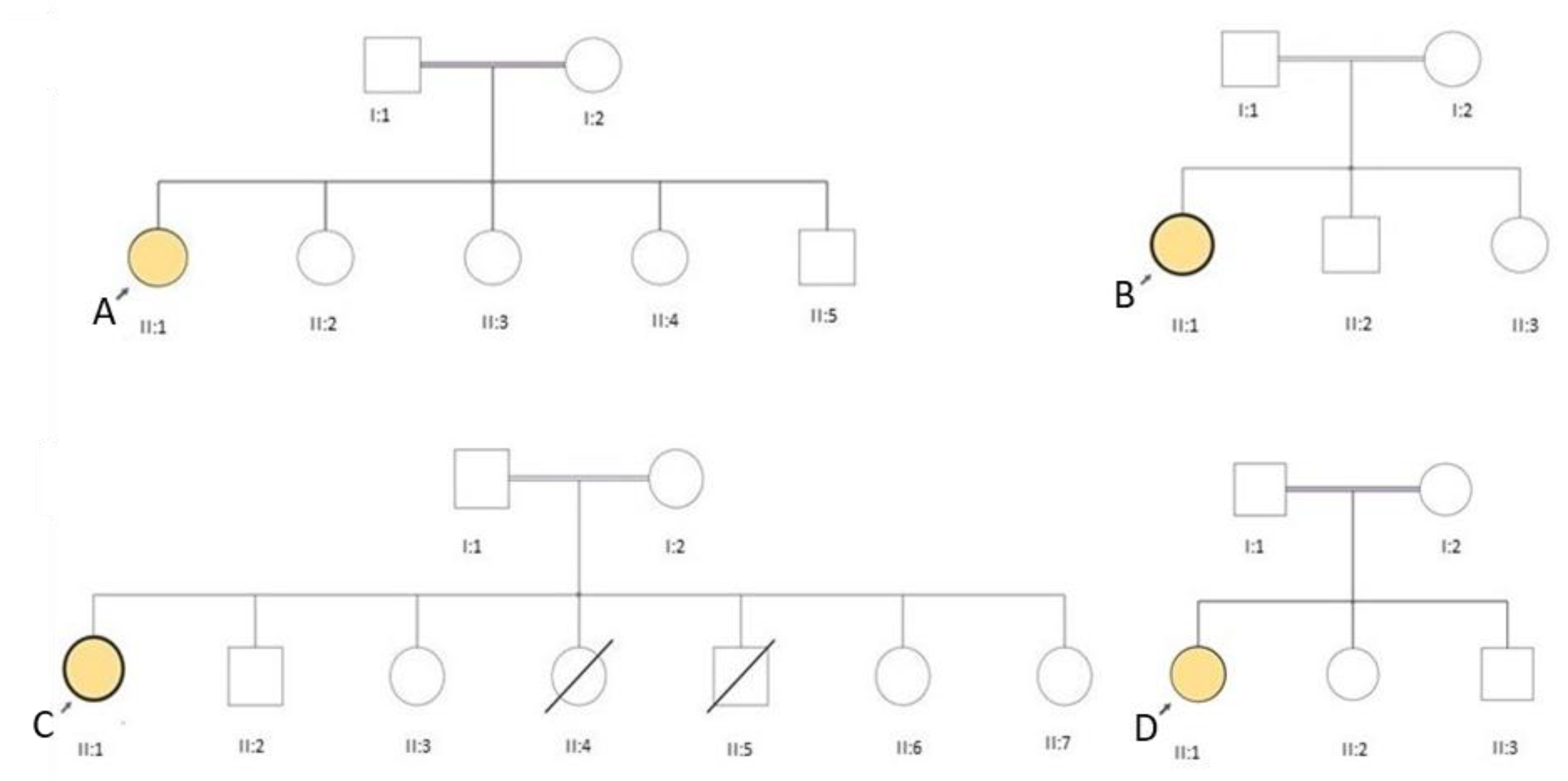
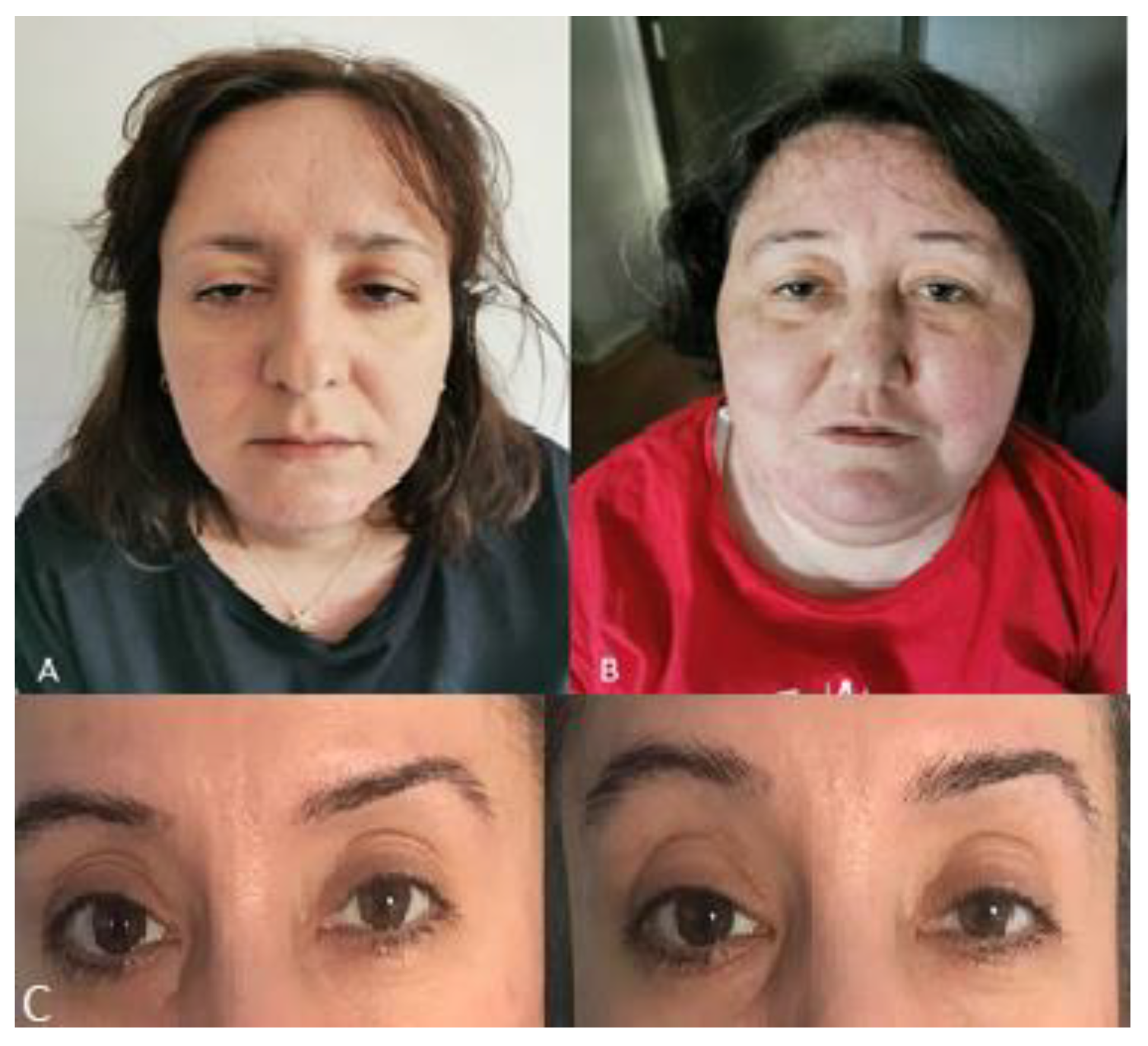
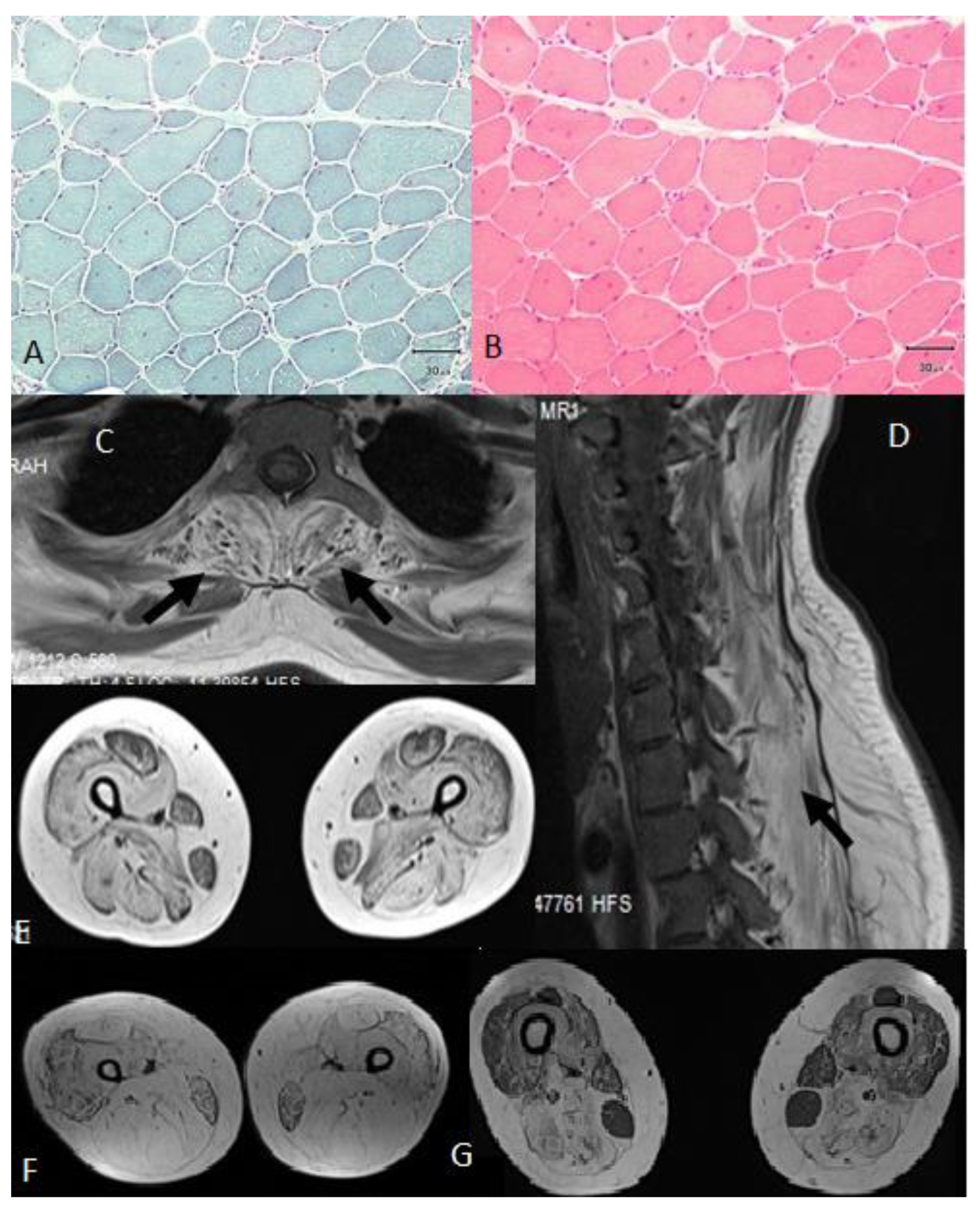
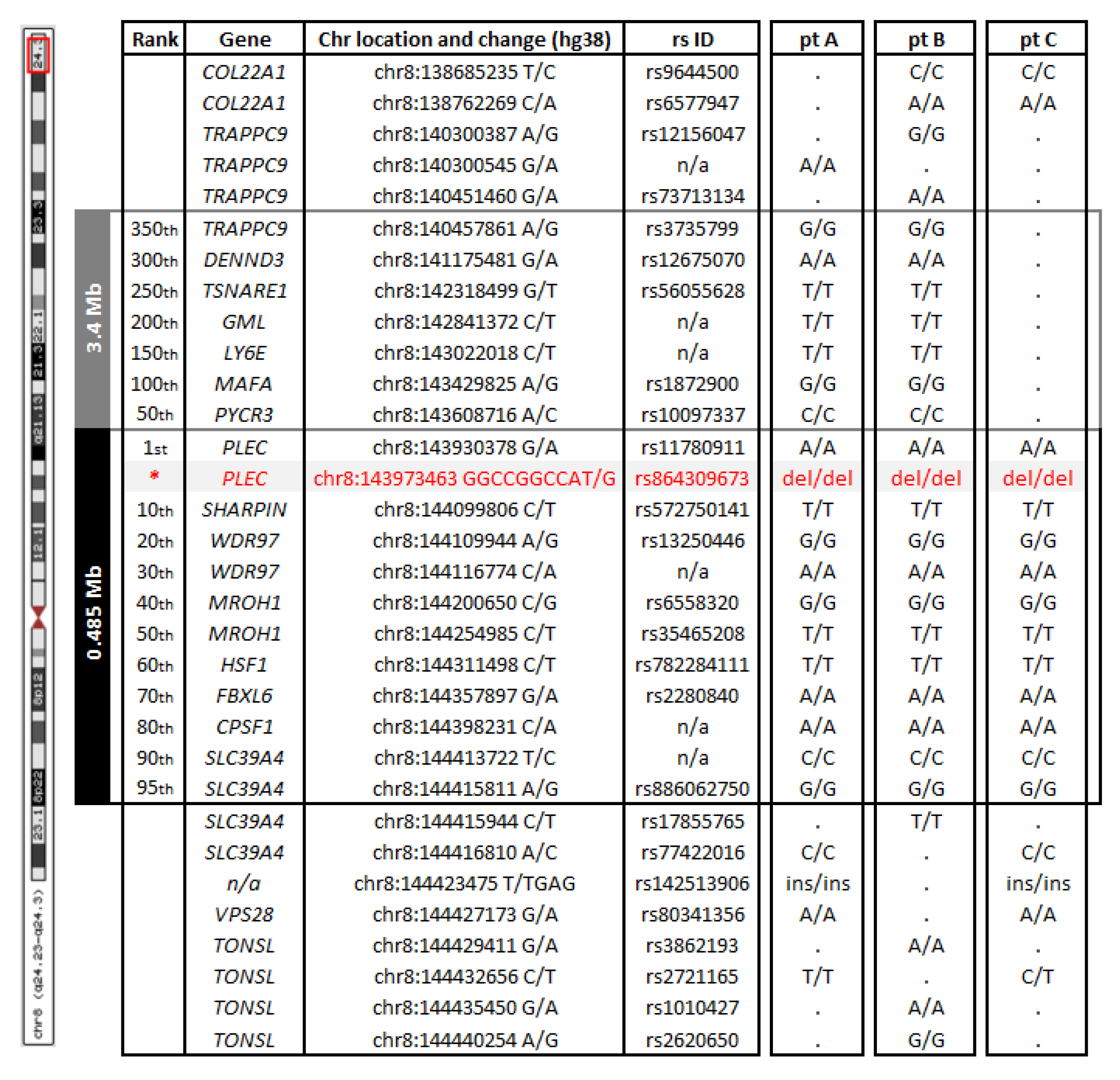
| ID | Age (Years) | Age of Onset (Years) | Motor Milestones | First Symptom | Fatigability | Weakness (MRC) | Calf Pseudohypertrophy | Wheelchair (Age) | CK (N < 220 U/L) | EMG | Biopsy | Refs |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A | 37 | 6 | Normal | Difficulty in walking | Yes | Ptosis, neck flexor 2/5, upper proximal (3/5), distal (4/5), lower proximal (2/5), distal (4/5), paraspinal and abdominal muscle weakness | Yes | Yes (38) | 1379 | Myopathic, high jitter in SF-EMG | Dystrophic, increased number of internal nuclei, a few angular atrophic fibers | This study |
| B | 38 | 26 | Normal | Difficulty in climbing stairs | Yes | Bilateral ptosis, neck flexors 3/5, upper proximal (4/5), upper distal 4/5 lower proximal (3/5) and distal (4/5), paraspinal and abdominal muscle weakness | Yes | No | 3290 | Myopathic and high jitter in SF-EMG. 10% decremental response in Trapezius muscle on RNS | Dystrophic increased number of internal nuclei, a few angular atrophic fibers | This study |
| C | 40 | Early childhood | Delayed—Age of onset for walking 2.5 years | Delayed walking, slower than her peers | Yes | Mild ptosis, nasal speech, tongue weakness, upper proximal 2/5, distal 3/5; lower proximal 1-2/5, distal 2/5, paraspinal and abdominal muscle weakness | Yes | Yes (35) | 4980 | Myopathic, spontaneous pathogenic activity, high jitter in SF-EMG. 8.7% decremental response in trapezius on RNS | Dystrophic, increased number of internal nuclei, a few angular atrophic fibers | This study |
| D | 27 | Early childhood | Delayed | Difficulty in climbing up stairs | Yes | Bilateral mild ptosis, upper proximal (4/5), distal (5/5), lower proximal (3/5), distal (4/5), paraspinal, and abdominal muscle weakness | Mild | No | 6200 | Myopathic, spontaneous pathogenic activity, high jitter in SF-EMG. 12% decremental response in trapezius on RNS | Dystrophic, increased number of internal nuclei, a few angular atrophic fibers | This study |
| VI:1 (*) | 19 | Early childhood | Delayed walking at 3 years | Delayed milestones, difficulty climbing stairs | No | No facial involvement, proximal muscle strength of 3+/5 | No | No | 5584 | Myopathic | Dystrophic | Gundesli et al. [10] |
| Family II patient 20 | 4 | 2 | Normal | Muscular weakness | No | NI | No | No | 4159 | NI | Dystrophic | Gundesli et al. [10] |
| Family III patient 47 | 5 | Early childhood | Delayed walking at 2.5 years | Difficulty climbing stairs, slower in running than peers | No | NI | No | No | 3704 | NI | Dystrophic | Gundesli et al. [10] |
| Type | Mutation | Consang. | Clinical Features | Onset | RNS/SF-EMG | Response to Treatment | Refs. |
|---|---|---|---|---|---|---|---|
| EBS-MD-MyS | c.6169 C>T; p.Gln2057X c.12043dupG; p.Glu4015GlyfsX69 | no | AW, B, EBS, EO, F, FW, LW, P, PW, WD | Childhood | RNS 11–25% decrement | Positive to 3,4-DAP, negative to AchE inhibitors | [21] |
| EBS-MD-MyS | c.6955 C>T; p.Arg2319X c.12043dupG; p.Glu4015GlyfsX69 | no | AW, B, EBS, EO, F, FW, LW, P, R, W, WD | Early Childhood | RNS 67% decrement | Negative | [21] |
| EBS-MD-MyS | c.3064 C>T; p.Gln1022X c.11503G>A; Gly3835Ser | no | AW, B, F, FD, FW, LW, P, PW | Childhood | SF-EMG negative | Negative | [22] |
| EBS-MD-MyS | c.3086 G>A; p.Arg1029His c.9679_9766del88; p.Asp3229ValfsX21 | no | AW, BW, DMM, EBS, F, FW, LW, P, PW, RFM | Congenital | RNS 12% decrement, SF-EMG positive 1 | Positive to pyridostigmine | [23] |
| EBS-MD-Mys | c.10187_10190delTGTC, p.Val3396AlafsX11 IVS11+2 T>G | no | AW, DMM, EBS, EO, F, FD, FW, H, IGH, LW, P, PW, RFM, W | Congenital | RNS 16% decrement | Positive to pyridostygmine | [24] |
| EBS-MD with ptosis | ? | no | AW, DMM, EBS, FW, LW, P, PW, R, WD | Congenital | ? | ? | [16] |
| EBS-MD with congenital myasthenia gravis | ? | yes | EBS, F; congenital myasthenic syndrome treated with thymectomy | Congenital | ? | ? | [8] |
| EBS-MD with ocular signs | ? | no | EBS, LW, PW, WD, slightly restricted eye movement, and reduced strength in eyelid closure | Congenital | ? | ? | [25] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mroczek, M.; Durmus, H.; Töpf, A.; Parman, Y.; Straub, V. Four Individuals with a Homozygous Mutation in Exon 1f of the PLEC Gene and Associated Myasthenic Features. Genes 2020, 11, 716. https://doi.org/10.3390/genes11070716
Mroczek M, Durmus H, Töpf A, Parman Y, Straub V. Four Individuals with a Homozygous Mutation in Exon 1f of the PLEC Gene and Associated Myasthenic Features. Genes. 2020; 11(7):716. https://doi.org/10.3390/genes11070716
Chicago/Turabian StyleMroczek, Magdalena, Hacer Durmus, Ana Töpf, Yesim Parman, and Volker Straub. 2020. "Four Individuals with a Homozygous Mutation in Exon 1f of the PLEC Gene and Associated Myasthenic Features" Genes 11, no. 7: 716. https://doi.org/10.3390/genes11070716
APA StyleMroczek, M., Durmus, H., Töpf, A., Parman, Y., & Straub, V. (2020). Four Individuals with a Homozygous Mutation in Exon 1f of the PLEC Gene and Associated Myasthenic Features. Genes, 11(7), 716. https://doi.org/10.3390/genes11070716





