Antibiotic Resistance Is Associated with Integrative and Conjugative Elements and Genomic Islands in Naturally Circulating Streptococcus pneumoniae Isolates from Adults in Liverpool, UK
Abstract
1. Introduction
2. Materials and Methods
2.1. EHPC Clinical Studies
2.1.1. Study Design
2.1.2. Detection of Pneumococcal Carriage—Baseline Samples
2.2. Susceptibility Testing
2.3. Filter-Mating Assays
2.3.1. Streptococcus pneumoniae Isolates
2.3.2. Filter-Mating Procedure
2.4. Whole Genome Sequencing and Bioinformatic Analysis
3. Results and Discussion
3.1. Determination of the Genetic Basis for Resistance
3.2. Characterisation of Resistance Associated ICEs
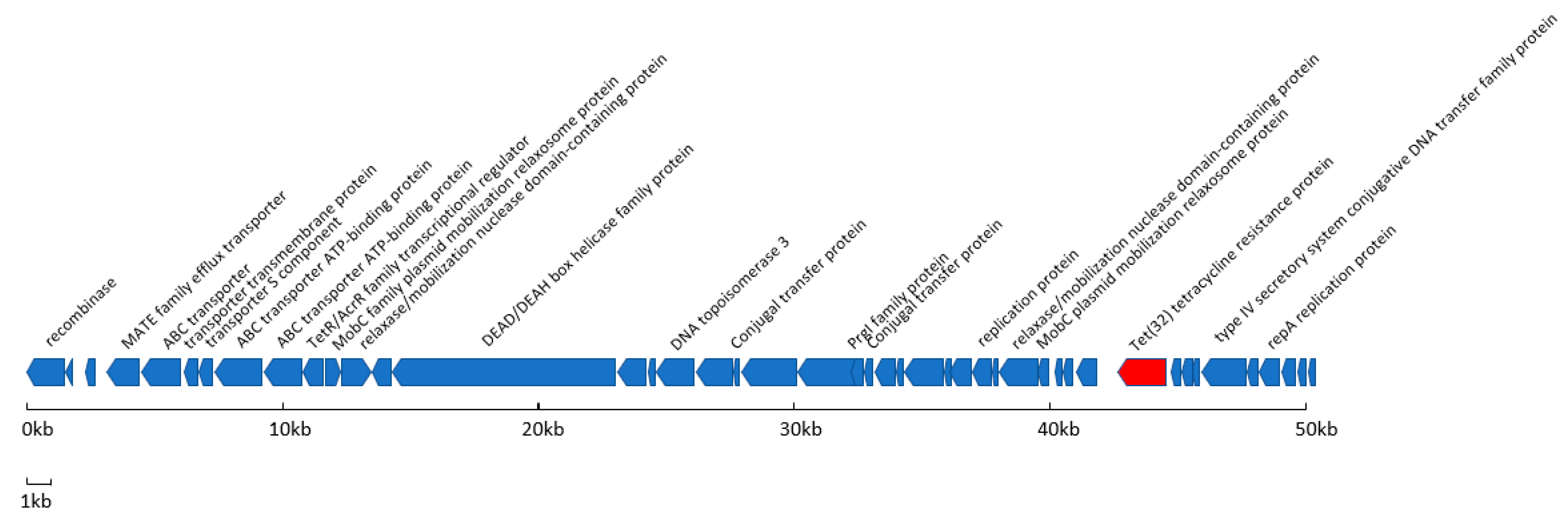
3.3. Characterisation of a Novel, Resistance Associated Genomic Island
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- O’Brien, K.L.; Wolfson, L.J.; Watt, J.P.; Henkle, E.; Deloria-Knoll, M.; McCall, N.; Lee, E.; Mulholland, K.; Levine, O.S.; Cherian, T.; et al. Burden of disease caused by Streptococcus pneumoniae in children younger than 5 years: Global estimates. Lancet 2009, 374, 893–902. [Google Scholar] [CrossRef]
- Welte, T.; Torres, A.; Nathwani, D. Clinical and economic burden of community-acquired pneumonia among adults in Europe. Thorax 2012, 67, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Goldblatt, D.; Hussain, M.; Andrews, N.; Ashton, L.; Virta, C.; Melegrano, A.; Pedoby, R.; George, R.; Soininen, A.; Edmunds, J.; et al. Antibody responses to nasopharyngeal carriage of S. pneumoniae in adults: A longitudinal household study. J. Infect. Dis. 2005, 192, 387–393. [Google Scholar] [CrossRef]
- Numminen, E.; Chewapreecha, C.; Turner, C.; Goldblatt, D.; Nosten, F.; Bentley, S.D.; Turner, P.; Corander, J. Climate induces seasonality in pneumococcal transmission. Sci. Rep. 2015, 12, e11344. [Google Scholar] [CrossRef] [PubMed]
- Heinsbroek, E.; Tafatatha, T.; Phiri, A.; Ngwira, B.; Crampin, A.M.; Read, J.M.; French, N. Persisting high prevalence of pneumococcal carriage among HIV-infected adults receiving antiretroviral therapy in Malawi: A cohort study. AIDS 2015, 29, 1837–1844. [Google Scholar] [CrossRef] [PubMed]
- Dawid, S.; Roche, A.M.; Weiser, J.N. The blp bacteriocins of Streptococcus pneumoniae mediate intraspecies competition both in vitro and in vivo. Infect. Immun. 2007, 75, 443–451. [Google Scholar] [CrossRef]
- Brugger, S.D.; Hathaway, L.J.; Mühlemann, K. Detection of Streptococcus pneumoniae strain cocolonization in the nasopharynx. J. Clin. Microbiol. 2009, 47, 1750–1756. [Google Scholar] [CrossRef][Green Version]
- Rodrigues, F.; Danon, L.; Moralas-Aza, B.; Sikora, P.; Thors, V.; Ferreira, M.; Gould, K.; Hinds, J.; Finn, A. Pneumococcal serotypes colonise the nasopharynx in children at different densities. PLoS ONE 2016, 11, e0163435. [Google Scholar] [CrossRef]
- Kamng’ona, A.W.; Hinds, J.; Bar-Zeev, N.; Gould, K.A.; Chaguza, C.; Msefula, C.; Cornick, J.E.; Kulohoma, B.W.; Gray, K.; Bently, S.D.; et al. High multiple colonization and emergence of Streptococcus pneumoniae vaccine serotype variants in Malawian children. BMC Infect. Dis. 2015, 15, 234. [Google Scholar] [CrossRef]
- Htar, M.T.T.; Stuurman, A.L.; Ferreira, G.; Alicino, C.; Bollaerts, K.; Paganino, C.; Reinert, R.R.; Schmitt, H.J.; Trucchi, C.; Vestraeten, T.; et al. Effectiveness of pneumococcal vaccines in preventing pneumonia in adults, a systematic review and meta-analyses of observational studies. PLoS ONE 2017, 12, e0177985. [Google Scholar] [CrossRef]
- Weinberger, D.M.; Trzcinski, K.; Lu, Y.J.; Bogaert, D.; Brandes, A.; Galagan, J.; Anderson, P.W.; Malley, R.; Lipsitch, M. Pneumococcal capsular polysaccharide structure predicts serotype prevalence. PLoS Pathog. 2009, 5, e1000476. [Google Scholar] [CrossRef] [PubMed]
- Gladstone, R.A.; Jefferies, J.M.; Faust, S.N.; Clarke, S.C. Continued control of pneumococcal disease in the UK-the impact of vaccination. J. Med. Microbiol. 2011, 60, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Isea-Peña, M.C.; Sanz-Moreno, J.C.; Esteban, J.; Fernández-Roblas, R.; Fernández-Guerrero, M.L. Risk factors and clinical significance of invasive infections caused by levofloxacin-resistant Streptococcus pneumoniae. Infection 2013, 41, 935–939. [Google Scholar] [CrossRef] [PubMed]
- Song, J.H.; Dagan, R.; Klugman, K.P.; Fritzell, B. The relationship between pneumococcal serotypes and antibiotic resistance. Vaccine 2012, 30, 2728–2737. [Google Scholar] [CrossRef] [PubMed]
- Gritzfeld, J.F.; Wright, A.D.; Collins, A.M.; Pennington, S.H.; Wright, A.K.; Kadioglu, A.; Ferreira, D.M.; Gordon, S.B. Experimental Human Pneumococcal Colonization. J. Vis. Exp. 2013, 72, e50115. [Google Scholar] [CrossRef]
- Adler, H.; Nikolaou, E.; Gould, K.; Hinds, J.; Collins, A.M.; Connor, V.; Hales, C.; Hill, H.; Hyder-Wright, A.D.; Zaidi, S.R.; et al. Pneumococcal colonization in healthy adult research participants in the conjugate vaccine era, United Kingdom, 2010–2017. J. Infect. Dis. 2019, 219, 1989–1993. [Google Scholar] [CrossRef]
- Ferreira, D.M.; Neill, D.R.; Bangert, M.; Gritzfeld, J.F.; Green, N.; Wright, A.K.; Pennington, S.H.; Bricio-Moreno, L.; Moreno, A.T.; Miyaji, E.N.; et al. Controlled human infection and rechallenge with Streptococcus pneumoniae reveals the protective efficacy of carriage in healthy adults. Am. J. Respir. Crit. Care Med. 2013, 187, 855–864. [Google Scholar] [CrossRef]
- The European Committee on Antimicrobial Susceptibility Testing. Breakpoint tables for interpretation of MICs and zone diameters. Version 10.0. 2020. Available online: http://www.eucast.org (accessed on 5 June 2020).
- Santoro, F.; Oggioni, M.R.; Pozzi, G.; Iannelli, F. Nucleotide sequence and functional analysis of the tet(M)-carrying conjugative transposon Tn5251 of Streptococcus pneumoniae. FEMS Microbiol. Lett. 2010, 308, 150–158. [Google Scholar] [CrossRef]
- Roberts, A.P.; Mullany, P.; Wilson, M. Gene transfer in bacterial biofilms. Methods Enzymol. 2001, 336, 60–65. [Google Scholar] [CrossRef]
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Zankari, E.; Hasman, H.; Cosentino, S.; Vestergaard, M.; Rasmussen, S.; Lund, O.; Aarestrup, F.M.; Larsen, M.V. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 2012, 67, 2640–2644. [Google Scholar] [CrossRef] [PubMed]
- Carattoli, A.; Zankari, E.; Garcia-Fernandez, A.; Larsen, M.V.; Lund, O.; Vila, L.; Aarestrup, F.M.; Hasman, H. In silico detection and typing of plasmids using PlasmidFinder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Li, X.B.; Xie, Y.Z.; Bi, D.X.; Sun, J.Y.; Li, J.; Tai, C.; Deng, Z.X.; Ou, H.Y. ICEberg 2.0: An updated database of bacterial integrative and conjugative elements. Nucleic Acids Res. 2019, 47, D660–D665. [Google Scholar] [CrossRef] [PubMed]
- Guglielmini, J.; Néron, B.; Abby, S.S.; Garcillán-Barcia, M.P.; de la Cruz, F.; Rocha, E.P. Key components of the eight classes of type IV secretion systems involved in bacterial conjugation or protein secretion. Nucleic Acids Res. 2014, 42, 5715–5727. [Google Scholar] [CrossRef]
- Abby, S.; Cury, J.; Guglielmini, J.; Neron, B.; Touchon, M.; Rocha, E.P.C. Identification of protein secretion systems in bacterial genomes. Sci. Rep. 2016, 6, 23080. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef]
- KBase. Prokka (v1.12). Available online: https://github.com/tseemann/prokka (accessed on 5 June 2020).
- GSL Biotech. SnapGene® software. Available online: https://www.snapgene.com/ (accessed on 5 June 2020).
- Chopra, I.; Roberts, M. Tetracycline antibiotics: Tetracycline antibiotics: Mode of action, applications, molecular biology, and epidemiology of bacterial resistance. Microbiol. Mol. Biol. Rev. 2001, 65, 232–260. [Google Scholar] [CrossRef]
- Warburton, P.J.; Amodeo, N.; Roberts, A.P. Mosaic tetracycline resistance genes encoding ribosomal protection proteins. J. Antimicrob. Chemoth. 2016, 71, 3333–3339. [Google Scholar] [CrossRef]
- Brodersen, D.E.; Clemons, W.M., Jr.; Carter, A.P. The structural basis for the action of the antibiotics tetracyclin, pectamycin and hygromycin B on the 30S ribosomal subunit. Cell 2000, 103, 1143–1154. [Google Scholar] [CrossRef]
- Santoro, S.; Vianna, M.E.; Roberts, A.P. Variation on a theme; an overview of the Tn916/Tn1545 family of mobile genetic elements in the oral and nasopharyngeal streptococci. Front. Microbiol. 2014, 535. [Google Scholar] [CrossRef]
- Melville, C.M.; Scott, K.P.; Mercer, D.K.; Flint, H.J. Novel tetracycline resistance gene, tet(32), in the Clostridium-related human colonic anaerobe K10 and its transmission in vitro to the rumen anaerobe Butyrivibrio fibrisolvens. Antimicrob Agents Chemother. 2001, 45, 3246–3249. [Google Scholar] [CrossRef] [PubMed]
- Stanton, T.B.; Humphrey, S.B.; Scott, K.P.; Flint, H.J. Hybrid tet genes and tet gene nomenclature: Request for opinion. Antimicrob. Agents Chemother. 2005, 49, 1265–1266. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Patterson, A.J.; Rincon, M.T.; Flint, H.J.; Scott, K.P. Mosaic tetracycline resistance genes are widespread in human and animal fecal samples. Antimicrob. Agents Chemother. 2007, 51, 1115–1118. [Google Scholar] [CrossRef][Green Version]
- Weisblum, B. Erythromycin resistance by ribosome modification. Antimicrob. Agents Chemother. 1995, 39, 577–585. [Google Scholar] [CrossRef] [PubMed]
- Schroeder, M.R.; Lohsen, S.; Chancey, S.T.; Stephens, D.S. High-level macrolide resistance due to the mega element [mef(E)/mel] in Streptococcus pneumoniae. Front. Microbiol. 2019, 10, 868. [Google Scholar] [CrossRef]
- Schroeder, M.R.; Stephens, D.S. Macrolide resistance in Streptococcus pneumoniae. Front. Cell Infect. Microbiol. 2016, 6, 98. [Google Scholar] [CrossRef] [PubMed]
- Tait-Kamradt, A.; Clancy, J.; Cronan, M.; Dib-Hajj, F.; Wondrack, L.; Yuan, W.; Sutcliffe, J. mefE is necessary for the erythromycin-resistant M phenotype in Streptococcus pneumoniae. Antimicrob. Agents Chemother. 1997, 41, 2251–2255. [Google Scholar] [CrossRef]
- Warburton, P.J.; Palmer, R.M.; Munson, M.A.; Wade, W.G. Demonstration of in vivo transfer of doxycycline resistance mediated by a novel transposon. J. Antimicrob. Chemother. 2007, 60, 973–980. [Google Scholar] [CrossRef]
- Li, Y.; Tomita, H.; Lv, Y.; Liu, J.; Xue, F.; Zheng, B.; Ike, Y. Molecular characterization of erm(B)-and mef(E)-mediated erythromycin-resistant Streptococcus pneumoniae in China and complete DNA sequence of Tn2010. J. Appl. Microbiol. 2011, 110, 254–265. [Google Scholar] [CrossRef]
- Roberts, A.P.; Mullany, P. A modular master on the move: The Tn916 family of mobile genetic elements. Trends Microbiol. 2009, 17, 251–258. [Google Scholar] [CrossRef]
- Kittana, F.N.A.; Mustak, I.B.; Hascelik, G.; Saricam, S.; Gurler, N.; Diker, K.S. Erythromycin-resistant Streptococcus pneumoniae: Phenotypes, genotypes, transposons and pneumococcal vaccine coverage rates. J. Med. Microbiol. 2019, 68, 874–881. [Google Scholar] [CrossRef] [PubMed]
- Tansirichaiya, S.; Rahman, M.A.; Roberts, A.P. The Transposon Registry. Mob DNA. 2019, 10, 40. [Google Scholar] [CrossRef] [PubMed]
| Bacterial Isolate | Date Isolated | Antibiotic Resistance Genes | GenBank Accession Numbers |
|---|---|---|---|
| S. pneumoniae 291015 | 29 October 2015 | erm(B), tet(M) | JABAHF000000000 |
| S. pneumoniae 210415 | 21 April 2015 | erm(B), tet(M) | JABAHE000000000 |
| S. pneumoniae 080217 | 8 February 2017 | mef(A), msr(D), erm(B), tet(M) | JABAHD000000000 |
| S. pneumoniae 131016 | 13 October 2016 | tet(32) | JABAHC000000000 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nikolaou, E.; Hubbard, A.T.M.; Botelho, J.; Marschall, T.A.M.; Ferreira, D.M.; Roberts, A.P. Antibiotic Resistance Is Associated with Integrative and Conjugative Elements and Genomic Islands in Naturally Circulating Streptococcus pneumoniae Isolates from Adults in Liverpool, UK. Genes 2020, 11, 625. https://doi.org/10.3390/genes11060625
Nikolaou E, Hubbard ATM, Botelho J, Marschall TAM, Ferreira DM, Roberts AP. Antibiotic Resistance Is Associated with Integrative and Conjugative Elements and Genomic Islands in Naturally Circulating Streptococcus pneumoniae Isolates from Adults in Liverpool, UK. Genes. 2020; 11(6):625. https://doi.org/10.3390/genes11060625
Chicago/Turabian StyleNikolaou, Elissavet, Alasdair T. M. Hubbard, João Botelho, Taylor A. M. Marschall, Daniela M. Ferreira, and Adam P. Roberts. 2020. "Antibiotic Resistance Is Associated with Integrative and Conjugative Elements and Genomic Islands in Naturally Circulating Streptococcus pneumoniae Isolates from Adults in Liverpool, UK" Genes 11, no. 6: 625. https://doi.org/10.3390/genes11060625
APA StyleNikolaou, E., Hubbard, A. T. M., Botelho, J., Marschall, T. A. M., Ferreira, D. M., & Roberts, A. P. (2020). Antibiotic Resistance Is Associated with Integrative and Conjugative Elements and Genomic Islands in Naturally Circulating Streptococcus pneumoniae Isolates from Adults in Liverpool, UK. Genes, 11(6), 625. https://doi.org/10.3390/genes11060625






