Genomic and Transcriptomic Analysis Identified Novel Putative Cassava lncRNAs Involved in Cold and Drought Stress
Abstract
1. Introduction
2. Materials and Methods
2.1. Genomes and Annotations
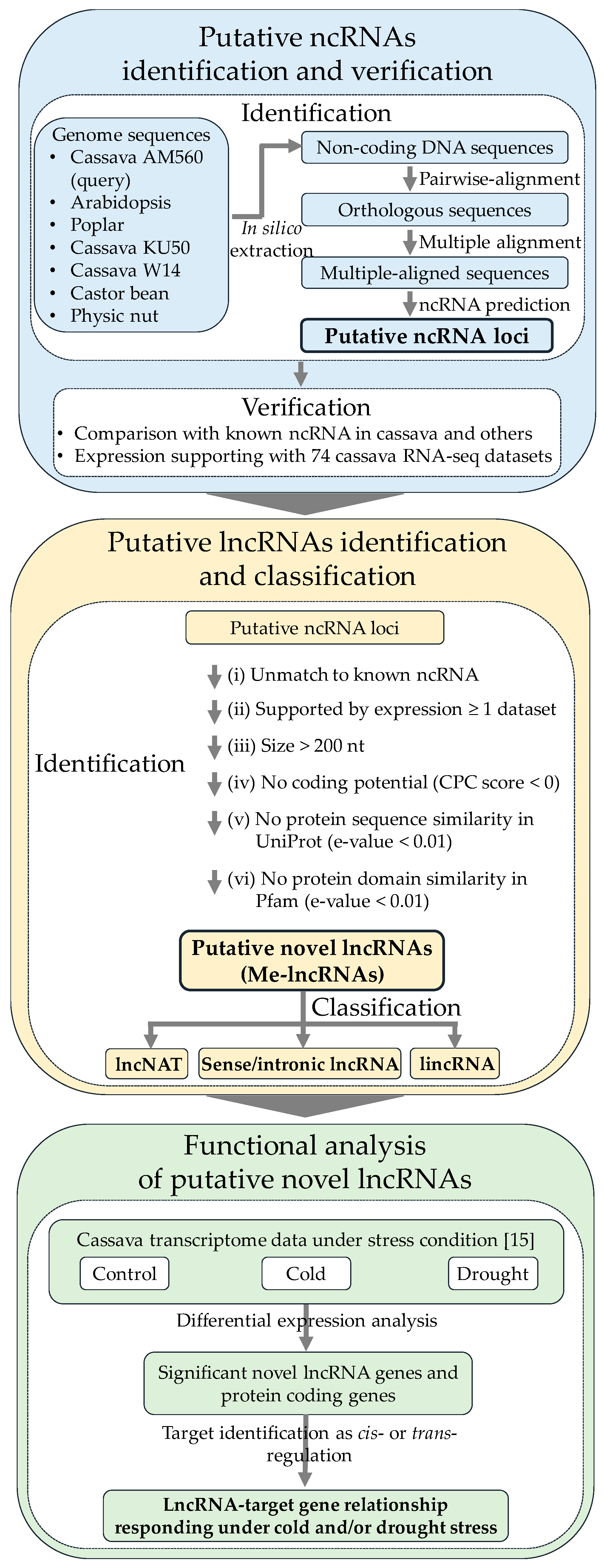
2.2. Identification of ncRNA Loci Based on Comparative Approach
2.3. Verification of Predicted ncRNA Loci
2.3.1. Comparison with Known ncRNAs based on Sequence or Structural Similarity
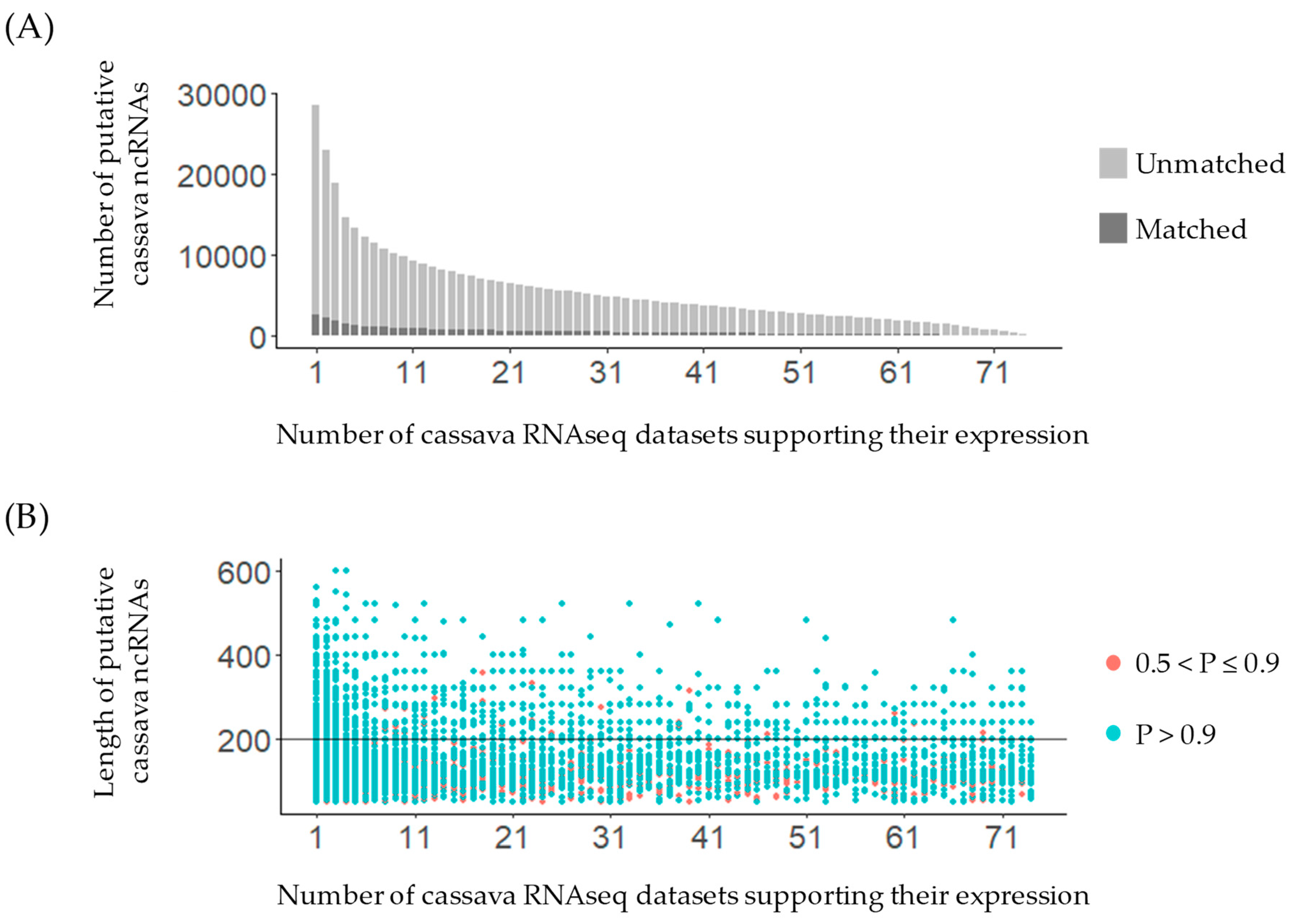
2.3.2. Expression Support Using Cassava RNA-Sequencing Datasets
2.4. Identification of Putative lncRNA Loci
2.5. Functional Analysis of Putative lncRNAs Involved in Cold or Drought Stress by RNA-seq Transcriptome Analysis
2.6. GO Enrichment Analysis and Visualization
3. Results and Discussion
3.1. Putative ncRNAs’ Identification and Verification
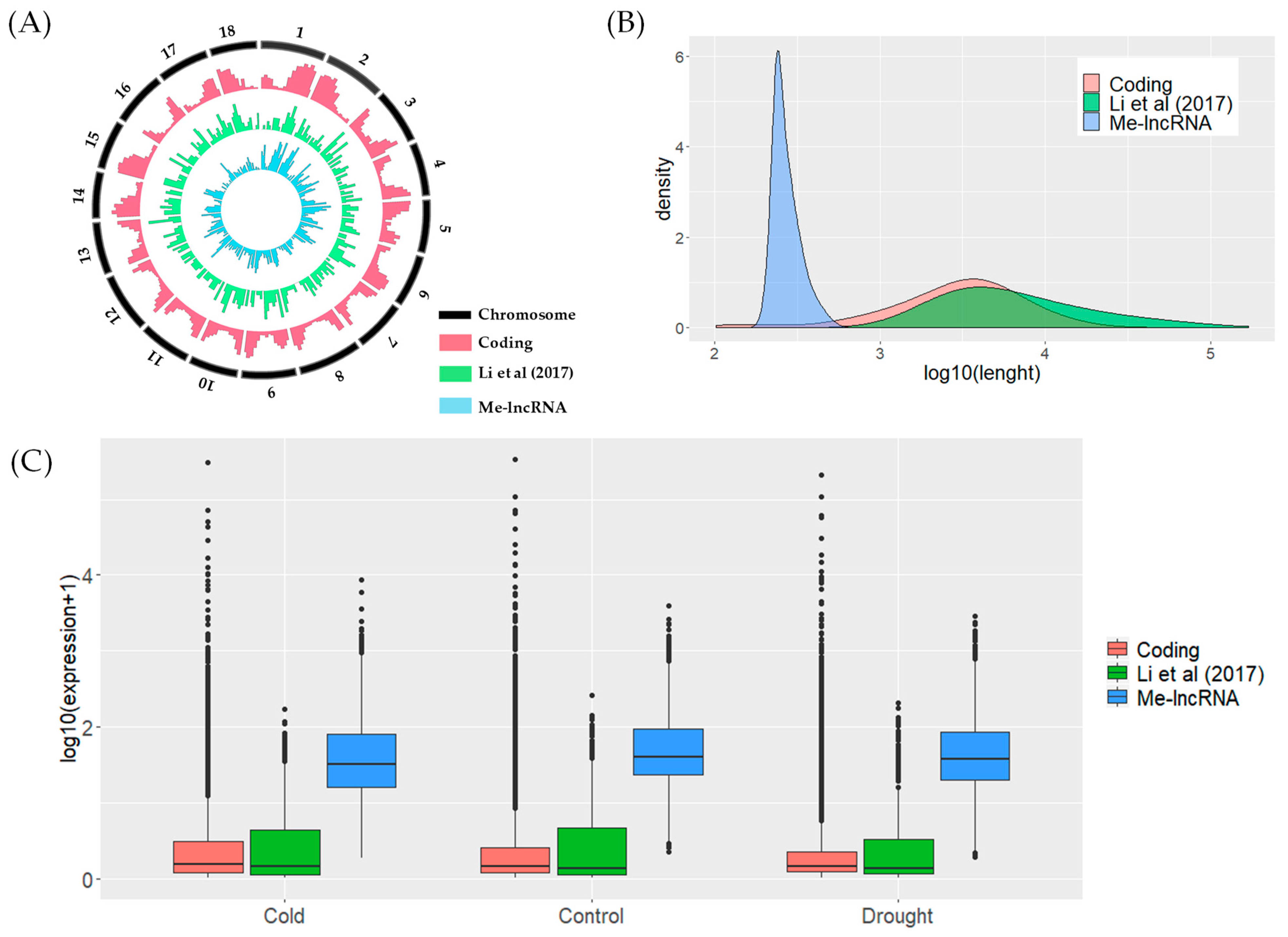
3.2. Potential Novel lncRNAs Classification and Characterization
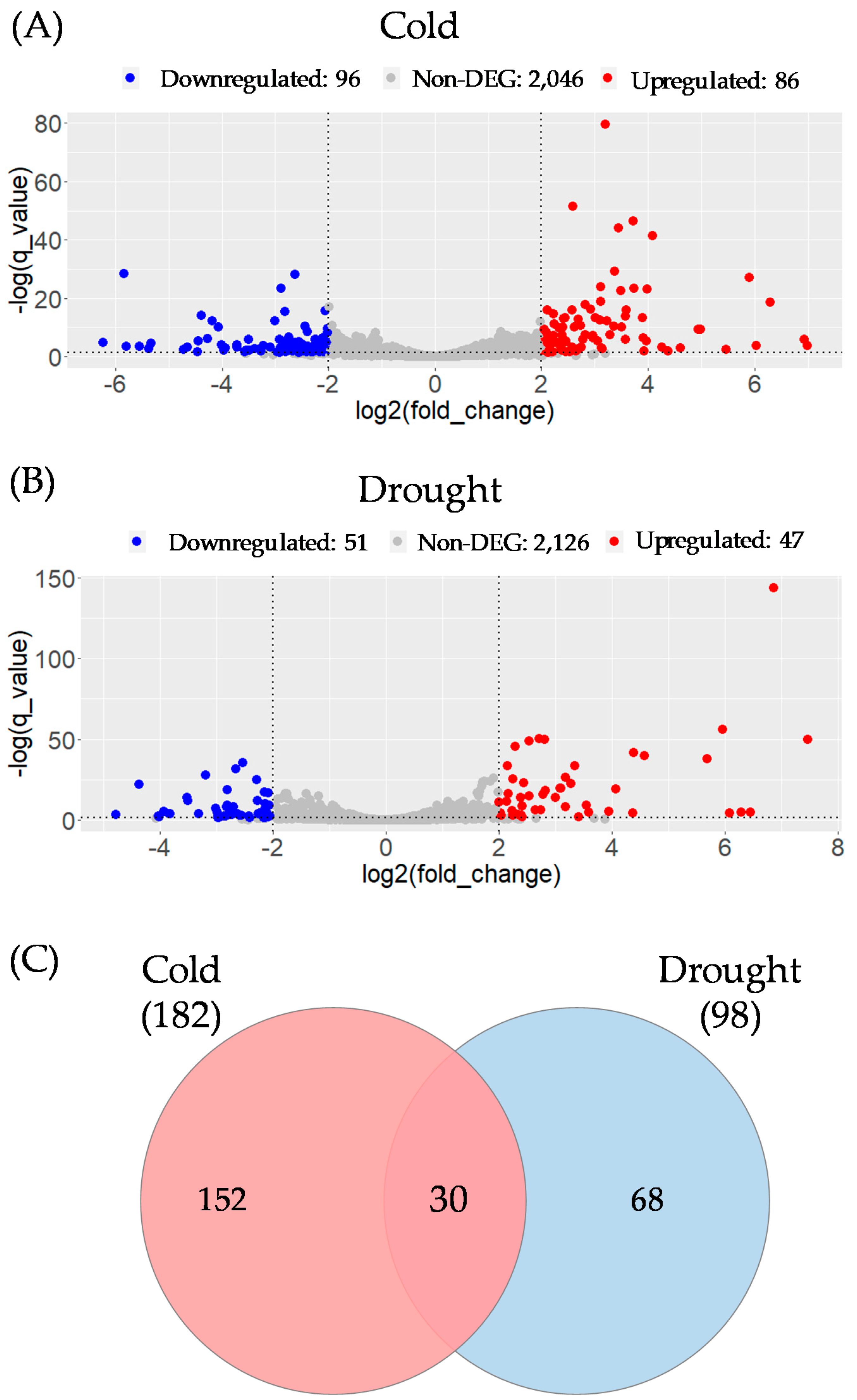
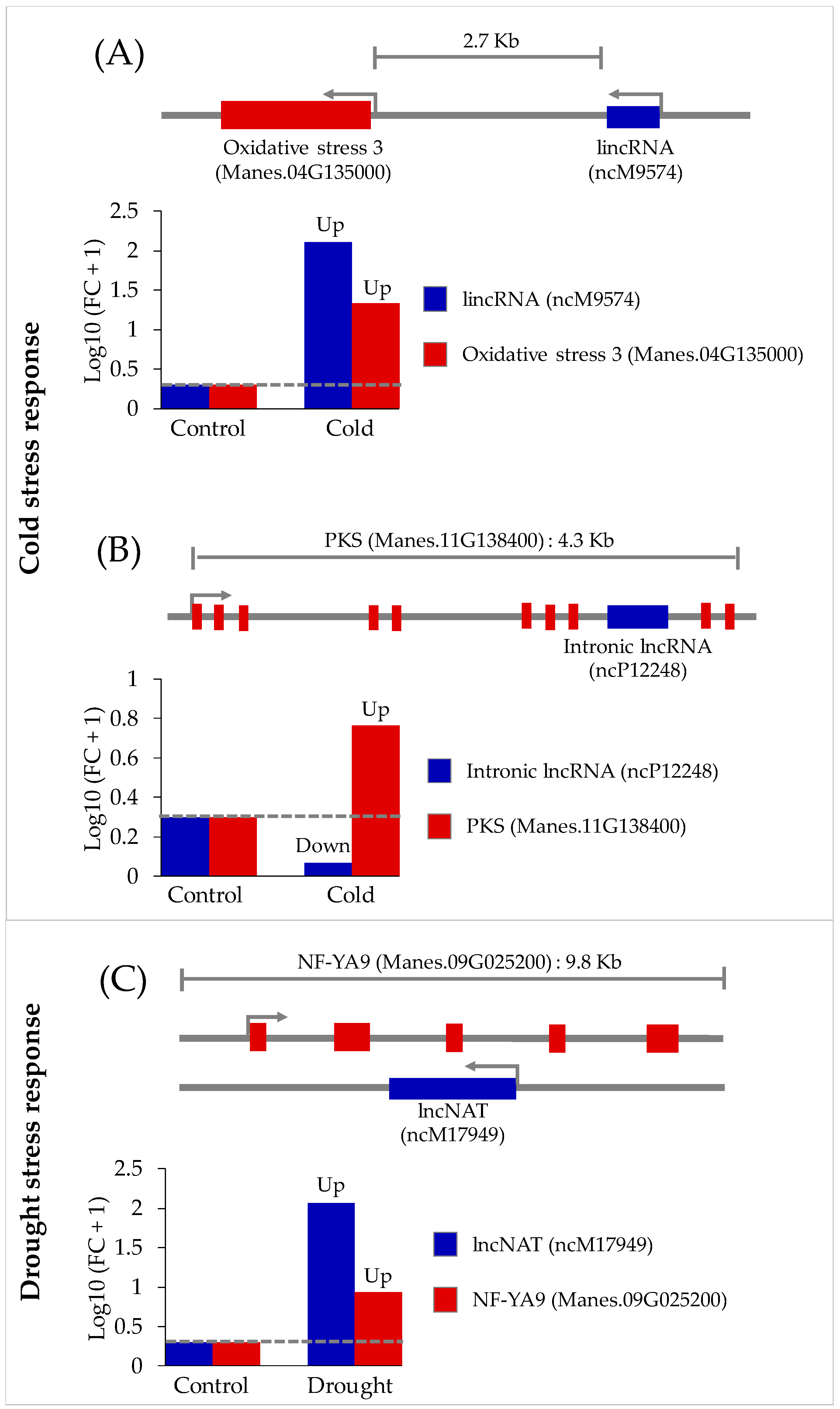
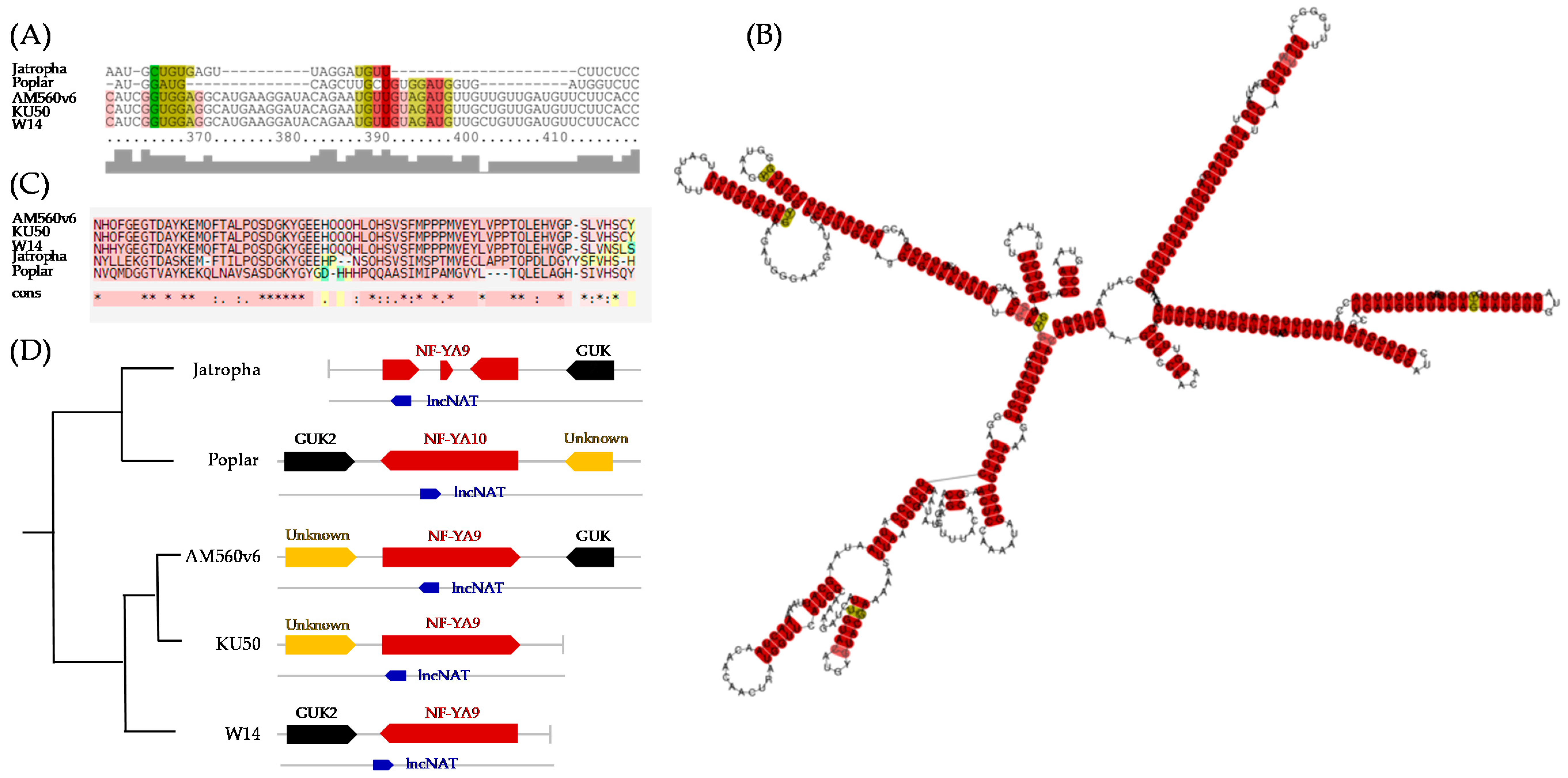
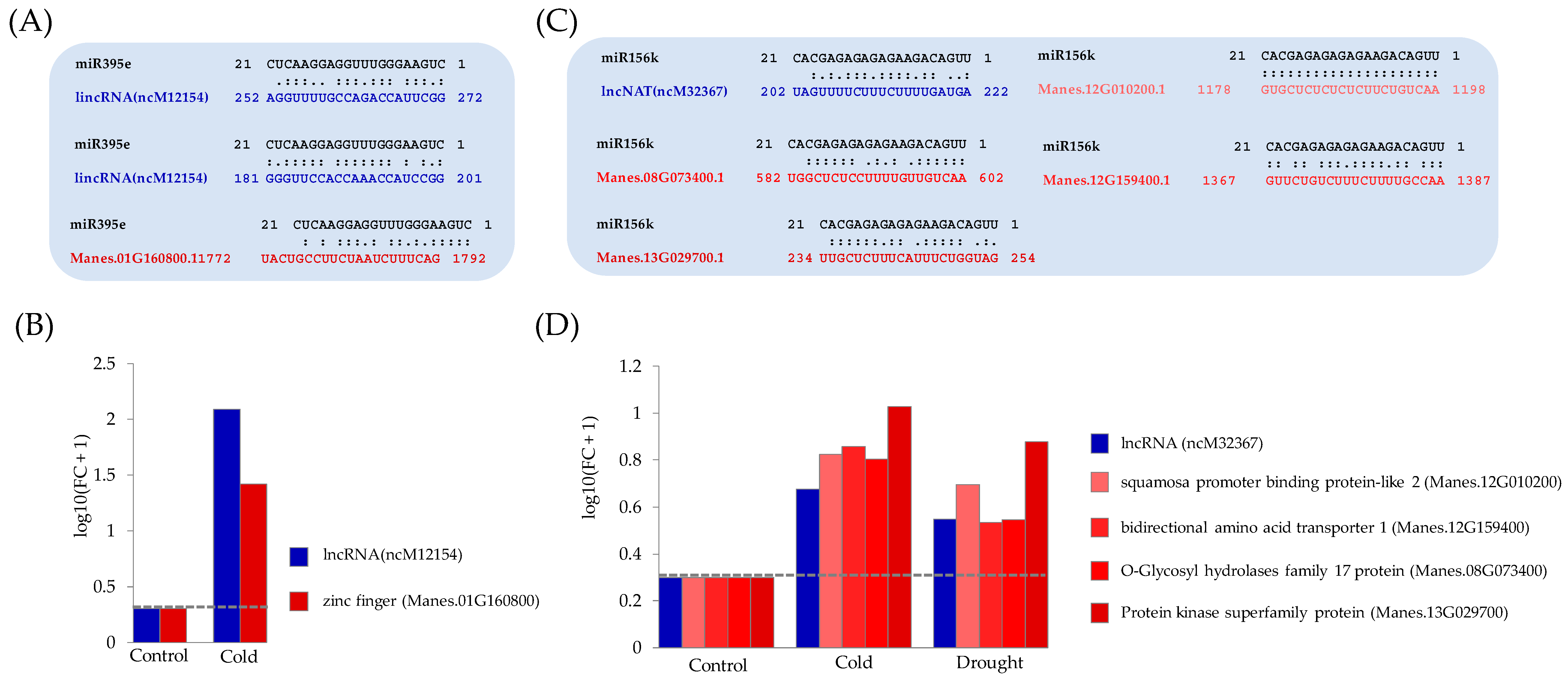
3.3. Functional Analysis of Potential Novel lncRNAs Involved in Cold and/or Drought Stress
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Glossary
References
- Yao, R.W.; Wang, Y.; Chen, L.L. Cellular functions of long noncoding RNAs. Nat. Cell Biol. 2019, 21, 542–551. [Google Scholar] [CrossRef] [PubMed]
- Prasanth, K.V.; Spector, D.L. Eukaryotic regulatory RNAs: An answer to the ‘genome complexity’ conundrum. Genes Dev. 2007, 21, 11–42. [Google Scholar] [CrossRef] [PubMed]
- Dinger, M.E.; Amaral, P.P.; Mercer, T.R.; Mattick, J.S. Pervasive transcription of the eukaryotic genome: Functional indices and conceptual implications. Brief. Funct. Genomic. Proteom. 2009, 8, 407–423. [Google Scholar] [CrossRef] [PubMed]
- Carninci, P.; Kasukawa, T.; Katayama, S.; Gough, J.; Frith, M.C.; Maeda, N.; Oyama, R.; Ravasi, T.; Lenhard, B.; Wells, C.; et al. The transcriptional landscape of the mammalian genome. Science 2005, 309, 1559–1563. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000, 403, 901–906. [Google Scholar] [CrossRef] [PubMed]
- McHugh, C.A.; Chen, C.K.; Chow, A.; Surka, C.F.; Tran, C.; McDonel, P.; Pandya-Jones, A.; Blanco, M.; Burghard, C.; Moradian, A.; et al. The Xist lncRNA directly interacts with SHARP to silence transcription through HDAC3. Nature 2015, 521, 232–236. [Google Scholar] [CrossRef]
- Heo, J.B.; Sung, S. Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 2011, 331, 76–79. [Google Scholar] [CrossRef]
- Liu, J.; Wang, H.; Chua, N.H. Long noncoding RNA transcriptome of plants. Plant Biotechnol. J. 2015, 13, 319–328. [Google Scholar] [CrossRef]
- Bhatia, G.; Goyal, N.; Sharma, S.; Upadhyay, S.K.; Singh, K. Present Scenario of Long Non-Coding RNAs in Plants. Non-Coding RNA. Noncoding RNA 2017, 3, 16. [Google Scholar] [CrossRef]
- Latgé, G.; Poulet, C.; Bours, V.; Josse, C.; Jerusalem, G. Natural Antisense Transcripts: Molecular Mechanisms and Implications in Breast Cancers. Int. J. Mol. Sci. 2018, 19, 123. [Google Scholar] [CrossRef]
- Bardou, F.; Ariel, F.; Simpson, C.G.; Romero-Barrios, N.; Laporte, P.; Balzergue, S.; Brown, J.W.; Crespi, M. Long noncoding RNA modulates alternative splicing regulators in Arabidopsis. Dev. Cell 2014, 30, 166–176. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.C.; Liao, J.Y.; Li, Z.Y.; Yu, Y.; Zhang, J.P.; Li, Q.F.; Qu, L.H.; Shu, W.S.; Chen, Y.Q. Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genome Biol. 2014, 15, 512. [Google Scholar] [CrossRef] [PubMed]
- Qin, T.; Zhao, H.; Cui, P.; Albesher, N.; Xiong, L. A Nucleus-Localized Long Non-Coding RNA Enhances Drought and Salt Stress Tolerance. Plant Physiol. 2017, 175, 1321–1336. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Wang, M.; Li, N.; Wang, H.; Qiu, P.; Pei, L.; Xu, Z.; Wang, T.; Gao, E.; Liu, J.; et al. Long noncoding RNAs involve in resistance to Verticillium dahliae, a fungal disease in cotton. Plant Biotechnol. J. 2018, 16, 1172–1185. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Yu, X.; Lei, N.; Cheng, Z.; Zhao, P.; He, Y.; Wang, W.; Peng, M. Genome-wide identification and functional prediction of cold and/or drought-responsive lncRNAs in cassava. Sci. Rep. 2017, 7, 45981. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.; Tie, W.; Fu, L.; Yan, Y.; Liu, G.; Yan, W.; Li, Y.; Wu, C.; Zhang, J.; Hu, W. Strand-specific RNA-seq based identification and functional prediction of drought-responsive lncRNAs in cassava. BMC Genom. 2019, 20, 214. [Google Scholar] [CrossRef]
- Ding, Z.H.; Wu, C.L.; Tie, W.W.; Yan, Y.; He, G.Y.; Hu, W. Strand-specific RNA-seq based identification and functional prediction of lncRNAs in response to melatonin and simulated drought stresses in cassava. Plant Physiol. Biochem. 2019, 140, 96–104. [Google Scholar] [CrossRef]
- Wu, C.; Ding, Z.; Chen, M.; Yang, G.; Tie, W.; Yan, Y.; Zeng, J.; Hea, G.; Hu, W. Identification and functional prediction of lncRNAs in response to PEG and ABA treatment in cassava. Environ. Exp. Bot. 2019, 166, 103809. [Google Scholar] [CrossRef]
- Xiao, L.; Shang, X.H.; Cao, S.; Xie, X.Y.; Zeng, W.D.; Lu, L.Y.; Chen, S.B.; Yan, H.B. Comparative physiology and transcriptome analysis allows for identification of lncRNAs imparting tolerance to drought stress in autotetraploid cassava. BMC Genom. 2019, 20, 514. [Google Scholar] [CrossRef]
- Pérez-Quintero, Á.L.; Quintero, A.; Urrego, O.; Vanegas, P.; López, C. Bioinformatic identification of cassava miRNAs differentially expressed in response to infection by Xanthomonas axonopodis pv. manihotis. BMC Plant Biol. 2012, 12, 29. [Google Scholar] [CrossRef]
- Washietl, S.; Hofacker, I.L.; Stadler, P.F. Fast and reliable prediction of noncoding RNAs. Proc. Natl. Acad. Sci. USA 2005, 102, 2454–2459. [Google Scholar] [CrossRef] [PubMed]
- Srisuk, T.; Pornputtapong, N.; Cheevadhanarak, S.; Thammarongtham, C. Prediction of Non-coding RNA and Their Targets in Spirulina platensis Genome. In Proceedings of the 1st Computational Systems-Biology and Bioinformatics Conference (CSBio 2010), Bangkok, Thailand, 3–5 November 2010; pp. 106–117. [Google Scholar]
- Mourier, T.; Carret, C.; Kyes, S.; Christodoulou, Z.; Gardner, P.P.; Jeffares, D.C.; Pinches, R.; Barrell, B.; Berriman, M.; Griffiths-Jones, S.; et al. Genome-wide discovery and verification of novel structured RNAs in Plasmodium falciparum. Genome Res. 2008, 18, 281–292. [Google Scholar] [CrossRef] [PubMed]
- Rose, D.; Hackermüller, J.; Washietl, S.; Reiche, K.; Hertel, J.; Findeiss, S.; Stadler, P.F.; Prohaska, S.J. Computational RNomics of Drosophilids. BMC Genom. 2007, 8, 406. [Google Scholar] [CrossRef] [PubMed]
- Washietl, S.; Hofacker, I.L.; Lukasser, M.; Hüttenhofer, A.; Stadler, P.F. Mapping of conserved RNA secondary structures predicts thousands of functional noncoding RNAs in the human genome. Nat. Biotechnol. 2005, 23, 1383–1390. [Google Scholar] [CrossRef]
- Song, D.; Yang, Y.; Yu, B.; Zheng, B.; Deng, Z.; Lu, B.L.; Chen, X.; Jiang, T. Computational prediction of novel non-coding RNAs in Arabidopsis thaliana. BMC Bioinform. 2009, 10, S36. [Google Scholar] [CrossRef]
- JGI Phytozome 12 The Plant Genomics Resource Home Page. Available online: https://phytozome.jgi.doe.gov/pz/portal.html (accessed on 27 August 2014).
- Jatropha Genome Database Hame Page. Available online: www.kazusa.or.jp/jatropha/ (accessed on 27 August 2014).
- Cassava China Database Home Page. Available online: http://www.cassava-genome.cn/ (accessed on 27 August 2014).
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef]
- Gruber, A.R.; Findeiß, S.; Washietl, S.; Hofacker, I.L.; Stadler, P.F. RNAz 2.0: Improved noncoding RNA detection. In Proceedings of the Pacific Symposium, Kamuela, HI, USA, 4–8 January 2010; pp. 69–79. [Google Scholar]
- Wang, W.; Feng, B.; Xiao, J.; Xia, Z.; Zhou, X.; Li, P.; Zhang, W.; Wang, Y.; Møller, B.L.; Zhang, P.; et al. Cassava genome from a wild ancestor to cultivated varieties. Nat. Commun. 2014, 5, 5110. [Google Scholar] [CrossRef]
- Quintero, A.; Pérez-Quintero, A.L.; López, C. Identification of ta-siRNAs and Cis-nat-siRNAs in Cassava and Their Roles in Response to Cassava Bacterial Blight. Genom. Proteom. Bioinform. 2013, 11, 172–181. [Google Scholar] [CrossRef]
- Ballén-Taborda, C.; Plata, G.; Ayling, S.; Rodríguez-Zapata, F.; Becerra Lopez-Lavalle, L.A.; Duitama, J.; Tohme, J. Identification of Cassava MicroRNAs under Abiotic Stress. Int. J. Genomics. 2013, 2013, 857986. [Google Scholar] [CrossRef]
- Xia, J.; Zeng, C.; Chen, Z.; Zhang, K.; Chen, X.; Zhou, Y.; Song, S.; Lu, C.; Yang, R.; Yang, Z.; et al. Endogenous small-noncoding RNAs and their roles in chilling response and stress acclimation in Cassava. BMC Genom. 2014, 15, 634. [Google Scholar] [CrossRef]
- Yawichai, A.; Kalapanulak, S.; Thammarongtham, C.; Saithong, T. Genome-Wide Identification of Putative MicroRNAs in Cassava (Manihot esculenta Crantz) and Their Functional Landscape in Cellular Regulation. Biomed. Res. Int. 2019, 2019, 2019846. [Google Scholar] [CrossRef] [PubMed]
- Quinlan, A.R.; Hall, I.M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 2010, 26, 841–842. [Google Scholar] [CrossRef] [PubMed]
- Kalvari, I.; Argasinska, J.; Quinones-Olvera, N.; Nawrocki, E.P.; Rivas, E.; Eddy, S.R.; Bateman, A.; Finn, R.D.; Petrov, A.I. Rfam 13.0: Shifting to a genome-centric resource for non-coding RNA families. Nucleic Acids Res. 2017, 46, D335–D342. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.C.; Mutka, A.M.; Hummel, A.W.; Berry, J.; Chauhan, R.D.; Vijayaraghavan, A.; Taylor, N.J.; Voytas, D.F.; Chitwood, D.H.; Bart, R.S. Gene expression atlas for the food security crop cassava. New Phytol. 2017, 213, 1632–1641. [Google Scholar] [CrossRef] [PubMed]
- Amuge, T.; Berger, D.K.; Katari, M.S.; Myburg, A.A.; Goldman, S.L.; Ferguson, M.E. A time series transcriptome analysis of cassava (Manihot esculenta Crantz) varieties challenged with Ugandan cassava brown streak virus. Sci. Rep. 2017, 7, 9747. [Google Scholar] [CrossRef] [PubMed]
- Pootakham, W.; Shearman, J.R.; Ruang-Areerate, P.; Sonthirod, C.; Sangsrakru, D.; Jomchai, N.; Yoocha, T.; Triwitayakorn, K.; Tragoonrung, S.; Tangphatsornruang, S. Large-scale SNP discovery through RNA sequencing and SNP genotyping by targeted enrichment sequencing in cassava (Manihot esculenta Crantz). PLoS ONE 2014, 9, e116028. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (accessed on 27 August 2014).
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Smid, M.; Coebergh van den Braak, R.R.J.; van de Werken, H.J.G.; van Riet, J.; van Galen, A.; de Weerd, V.; van der Vlugt-Daane, M.; Bril, S.I.; Lalmahomed, Z.S.; Kloosterman, W.P.; et al. Gene length corrected trimmed mean of M-values (GeTMM) processing of RNA-seq data performs similarly in intersample analyses while improving intrasample comparisons. BMC Bioinform. 2018, 19, 236. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- Li, J.; Ma, W.; Zeng, P.; Wang, J.; Geng, B.; Yang, J.; Cui, Q. LncTar: A tool for predicting the RNA targets of long noncoding RNAs. Brief. Bioinform. 2015, 16, 806–812. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Zhuang, Z.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef] [PubMed]
- Klopfenstein, D.V.; Zhang, L.; Pedersen, B.S.; Ramírez, F.; Warwick Vesztrocy, A.; Naldi, A.; Mungall, C.J.; Yunes, J.M.; Botvinnik, O.; Weigel, M.; et al. GOATOOLS: A Python library for Gene Ontology analyses. Sci. Rep. 2018, 8, 10872. [Google Scholar] [CrossRef] [PubMed]
- Data Visualization with ggplot2 Cheat Sheet. Available online: https://rstudio.com/wp-content/uploads/2016/11/ggplot2-cheatsheet-2.1.pdf (accessed on 27 August 2018).
- Will, S.; Joshi, T.; Hofacker, I.L.; Stadler, P.F.; Backofen, R. LocARNA-P: Accurate boundary prediction and improved detection of structural RNAs. RNA 2012, 18, 900–914. [Google Scholar] [CrossRef] [PubMed]
- Di Tommaso, P.; Moretti, S.; Xenarios, I.; Orobitg, M.; Montanyola, A.; Chang, J.M.; Taly, J.F.; Notredame, C. T-Coffee: A web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 2011, 39, W13–W17. [Google Scholar] [CrossRef]
- Thorvaldsdóttir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-performance genomics data visualization and exploration. Brief. Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef]
- Pedersen, J.S.; Bejerano, G.; Siepel, A.; Rosenbloom, K.; Lindblad-Toh, K.; Lander, E.S.; Kent, J.; Miller, W.; Haussler, D. Identification and Classification of Conserved RNA Secondary Structures in the Human Genome. PLoS Comput. Biol. 2006, 2, e33. [Google Scholar] [CrossRef]
- Harrow, J.; Frankish, A.; Gonzalez, J.M.; Tapanari, E.; Diekhans, M.; Kokocinski, F.; Aken, B.L.; Barrell, D.; Zadissa, A.; Searle, S.; et al. GENCODE: The reference human genome annotation for The ENCODE Project. Genome Res. 2012, 22, 1760–1774. [Google Scholar] [CrossRef]
- Golicz, A.A.; Singh, M.B.; Bhalla, P.L. The long intergenic noncoding RNA (lincRNA) landscape of the soybean genome. Plant Physiol. 2018, 176, 2133–2147. [Google Scholar] [CrossRef]
- Wang, M.; Zhao, W.; Gao, L.; Zhao, L. Genome-wide profiling of long non-coding RNAs from tomato and a comparison with mRNAs associated with the regulation of fruit ripening. BMC Plant Biol. 2018, 18, 75. [Google Scholar] [CrossRef]
- Zhang, Y.; Huang, H.; Zhang, D.; Qiu, J.; Yang, J.; Wang, K.; Zhu, L.; Fan, J.; Yang, J. A Review on Recent Computational Methods for Predicting Noncoding RNAs. Biomed. Res. Int. 2017, 2017, 9139504. [Google Scholar] [CrossRef] [PubMed]
- Eom, S.H.; Lee, H.J.; Lee, J.H.; Wi, S.H.; Kim, S.K.; Hyun, T.K. Identification and functional prediction of drought-responsive long non-coding RNA in tomato. Agronomy 2019, 9, 629. [Google Scholar] [CrossRef]
- Kader, D.Z.A.; Saleh, A.A.H.; Elmeleigy, S.A.; Dosoky, N.S. Chilling-induced oxidative stress and polyamines regulatory role in two wheat varieties. J. Taibah Univ. Sci. 2011, 5, 14–24. [Google Scholar] [CrossRef]
- Wang, T.Z.; Liu, M.; Zhao, M.G.; Chen, R.; Zhang, W.H. Identification and characterization of long non-coding RNAs involved in osmotic and salt stress in Medicago truncatula using genome-wide high-throughput sequencing. BMC Plant Biol. 2015, 15, 131. [Google Scholar] [CrossRef]
- Li, X.; Xing, X.; Xu, S.; Zhang, M.; Wang, Y.; Wu, H.; Sun, Z.; Huo, Z.; Chen, F.; Yang, T. Genome-wide identification and functional prediction of tobacco lncRNAs responsive to root-knot nematode stress. PLoS ONE 2018, 13, e0204506. [Google Scholar] [CrossRef]
- Ma, J.; Bai, X.; Luo, W.; Feng, Y.; Shao, X.; Bai, Q.; Sun, S.; Long, Q.; Wan, D. Genome-Wide Identification of Long Noncoding RNAs and Their Responses to Salt Stress in Two Closely Related Poplars. Front. Genet. 2019, 10, 777. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, Z.; Dong, X.; Feng, Y.; Liu, X.; Gao, B.; Wang, J.; Zhang, L.; Wang, J.; Shi, S.; et al. Identification and functional characterization of three type III polyketide synthases from Aquilaria sinensis calli. Biochem. Biophys. Res. Commun. 2017, 486, 1040–1047. [Google Scholar] [CrossRef]
- Nelson, D.E.; Repetti, P.P.; Adams, T.R.; Creelman, R.A.; Wu, J.; Warner, D.C.; Anstrom, D.C.; Bensen, R.J.; Castiglioni, P.P.; Donnarummo, M.G.; et al. Plant nuclear factor Y (NF-Y) B subunits confer drought tolerance and lead to improved corn yields on water-limited acres. Proc. Natl. Acad. Sci. USA 2007, 104, 16450–16455. [Google Scholar] [CrossRef]
- Han, X.; Tang, S.; An, Y.; Zheng, D.C.; Xia, X.L.; Yin, W.L. Overexpression of the poplar NF-YB7 transcription factor confers drought tolerance and improves water-use efficiency in Arabidopsis. J. Exp. Bot. 2013, 64, 4589–4601. [Google Scholar] [CrossRef]
- Chen, M.; Zhao, Y.; Zhuo, C.; Lu, S.; Guo, Z. Overexpression of a NF-YC transcription factor from Bermuda grass confers tolerance to drought and salinity in transgenic rice. Plant Biotechnol. J. 2015, 13, 482–491. [Google Scholar] [CrossRef]
- Pegueroles, C.; Iraola-Guzmán, S.; Chorostecki, U.; Ksiezopolska, E.; Saus, E.; Gabaldón, T. Transcriptomic analyses reveal groups of co-expressed, syntenic lncRNAs in four species of the genus Caenorhabditis. RNA Biol. 2019, 16, 320–329. [Google Scholar] [CrossRef] [PubMed]
- Quinn, J.J.; Zhang, Q.C.; Georgiev, P.; Ilik, I.A.; Akhtar, A.; Chang, H.Y. Rapid evolutionary turnover underlies conserved lncRNA-genome interactions. Genes Dev. 2016, 30, 191–207. [Google Scholar] [CrossRef] [PubMed]
- Mohammadin, S.; Edger, P.P.; Pires, J.C.; Schranz, M.E. Positionally-conserved but sequence-diverged: Identification of long non-coding RNAs in the Brassicaceae and Cleomaceae. BMC Plant Biol. 2015, 15, 217. [Google Scholar] [CrossRef] [PubMed]
- Deng, P.; Liu, S.; Nie, X.; Weining, S.; Wu, L. Conservation analysis of long non-coding RNAs in plants. Sci. China Life Sci. 2018, 61, 190–198. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhu, W.; Fang, L.; Sun, X.; Su, L.; Liang, Z.; Wang, N.; Londo, J.P.; Li, S.; Xin, H. Genome-wide identification of WRKY family genes and their response to cold stress in Vitis vinifera. BMC Plant Biol. 2014, 14, 103. [Google Scholar] [CrossRef]
- Finatto, T.; Viana, V.E.; Woyann, L.G.; Busanello, C.; Maia, L.; Oliveira, A.C. Can WRKY transcription factors help plants to overcome environmental challenges? Genet. Mol. Biol. 2018, 41, 533–544. [Google Scholar] [CrossRef]
- Jiang, J.; Ma, S.; Ye, N.; Jiang, M.; Cao, J.; Zhang, J. WRKY transcription factors in plant responses to stresses. J. Integr. Plant Biol. 2017, 59, 86–101. [Google Scholar] [CrossRef]
- Lv, Y.; Hu, F.; Zhou, Y.; Wu, F.; Gaut, B.S. Maize transposable elements contribute to long non-coding RNAs that are regulatory hubs for abiotic stress response. BMC Genom. 2019, 20, 864. [Google Scholar] [CrossRef]
- Daszkowska-Golec, A.; Szarejko, I. Open or close the gate—Stomata action under the control of phytohormones in drought stress conditions. Front. Plant Sci. 2013, 4, 138. [Google Scholar] [CrossRef]
- Geiger, D.; Scherzer, S.; Mumm, P.; Stange, A.; Marten, I.; Bauer, H.; Ache, P.; Matschi, S.; Liese, A.; Al-Rasheid, K.A. Activity of guard cell anion channel SLAC1 is controlled by drought-stress signaling kinase-phosphatase pair. Proc. Natl. Acad. Sci. USA 2009, 106, 21425–21430. [Google Scholar] [CrossRef]
- Sah, S.K.; Reddy, K.R.; Li, J. Abscisic Acid and Abiotic Stress Tolerance in Crop Plants. Front. Plant Sci. 2016, 7, 571. [Google Scholar] [CrossRef] [PubMed]
- Deforges, J.; Reis, R.S.; Jacquet, P.; Sheppard, S.; Gadekar, V.P.; Hart-Smith, G.; Tanzer, A.; Hofacker, I.L.; Iseli, C.; Xenarios, I.; et al. Control of Cognate Sense mRNA Translation by cis-Natural Antisense RNAs. Plant Physiol. 2019, 180, 305–322. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B. MicroRNA: A new target for improving plant tolerance to abiotic stress. J. Exp. Bot. 2015, 66, 1749–1761. [Google Scholar] [CrossRef] [PubMed]
- Klein, J.; Saedler, H.; Huijser, P. A new family of DNA binding proteins includes putative transcriptional regulators of the Antirrhinum majus floral meristem identity gene SQUAMOSA. Mol. Gen. Genet. 1996, 250, 7–16. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Zhang, Z.; Liu, D.; Zhang, K.; Li, A.; Mao, L. SQUAMOSA promoter-binding protein-like transcription factors: Star players for plant growth and development. J. Integr. Plant Biol. 2010, 52, 946–951. [Google Scholar] [CrossRef]
- Chen, X.; Xia, J.; Xia, Z.; Zhang, H.; Zeng, C.; Lu, C.; Zhang, W.; Wang, W. Potential functions of microRNAs in starch metabolism and development revealed by miRNA transcriptome profiling of cassava cultivars and their wild progenitor. BMC Plant Biol. 2015, 15, 33. [Google Scholar] [CrossRef]
- Preston, J.C.; Hileman, L.C. Functional Evolution in the Plant SQUAMOSA-PROMOTER BINDING PROTEIN-LIKE (SPL) Gene Family. Front. Plant Sci. 2013, 4, 80. [Google Scholar] [CrossRef]
- Smith, M.A.; Gesell, T.; Stadler, P.F.; Mattick, J.S. Widespread purifying selection on RNA structure in mammals. Nucleic Acids Res. 2013, 41, 8220–8236. [Google Scholar] [CrossRef]

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Suksamran, R.; Saithong, T.; Thammarongtham, C.; Kalapanulak, S. Genomic and Transcriptomic Analysis Identified Novel Putative Cassava lncRNAs Involved in Cold and Drought Stress. Genes 2020, 11, 366. https://doi.org/10.3390/genes11040366
Suksamran R, Saithong T, Thammarongtham C, Kalapanulak S. Genomic and Transcriptomic Analysis Identified Novel Putative Cassava lncRNAs Involved in Cold and Drought Stress. Genes. 2020; 11(4):366. https://doi.org/10.3390/genes11040366
Chicago/Turabian StyleSuksamran, Rungaroon, Treenut Saithong, Chinae Thammarongtham, and Saowalak Kalapanulak. 2020. "Genomic and Transcriptomic Analysis Identified Novel Putative Cassava lncRNAs Involved in Cold and Drought Stress" Genes 11, no. 4: 366. https://doi.org/10.3390/genes11040366
APA StyleSuksamran, R., Saithong, T., Thammarongtham, C., & Kalapanulak, S. (2020). Genomic and Transcriptomic Analysis Identified Novel Putative Cassava lncRNAs Involved in Cold and Drought Stress. Genes, 11(4), 366. https://doi.org/10.3390/genes11040366




