IurV, Encoded by ORF VCA0231, Is Involved in the Regulation of Iron Uptake Genes in Vibrio cholerae
Abstract
1. Introduction
2. Materials and Methods
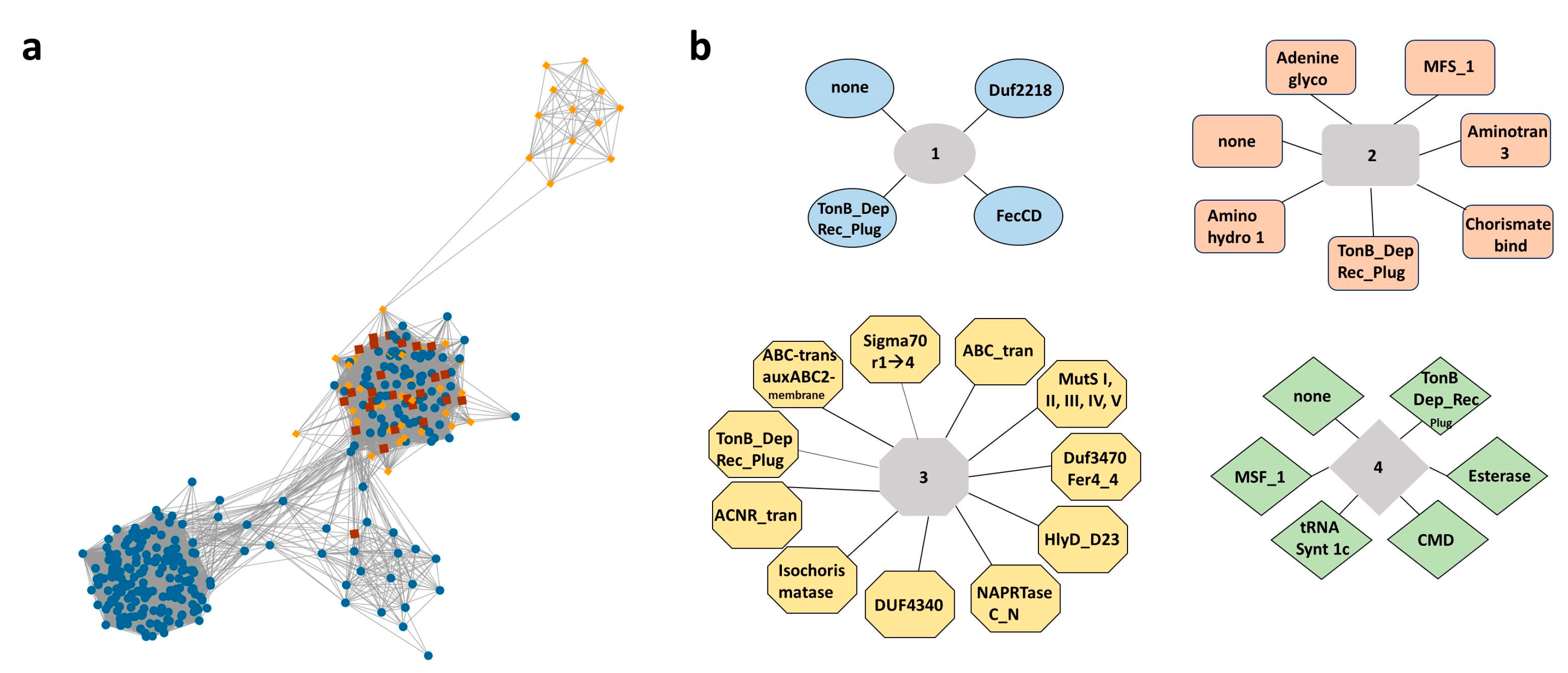
2.1. STRING and EFI-GNT Analysis
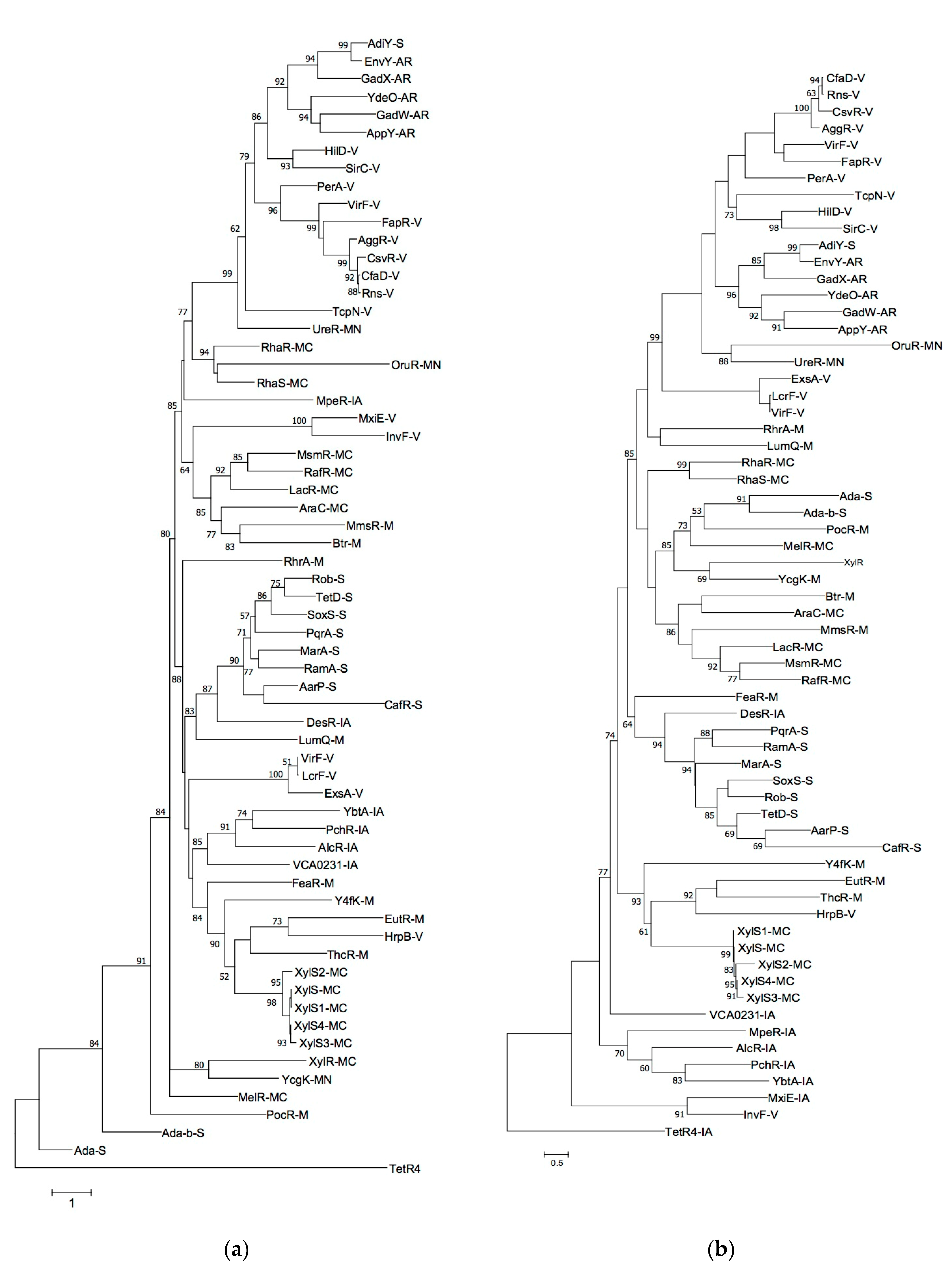
2.2. Phylogenetic Trees Construction
2.3. Construction of a V. cholerae ΔVCA0231::cat Strain
2.4. Strains Culture and RNA Extractions
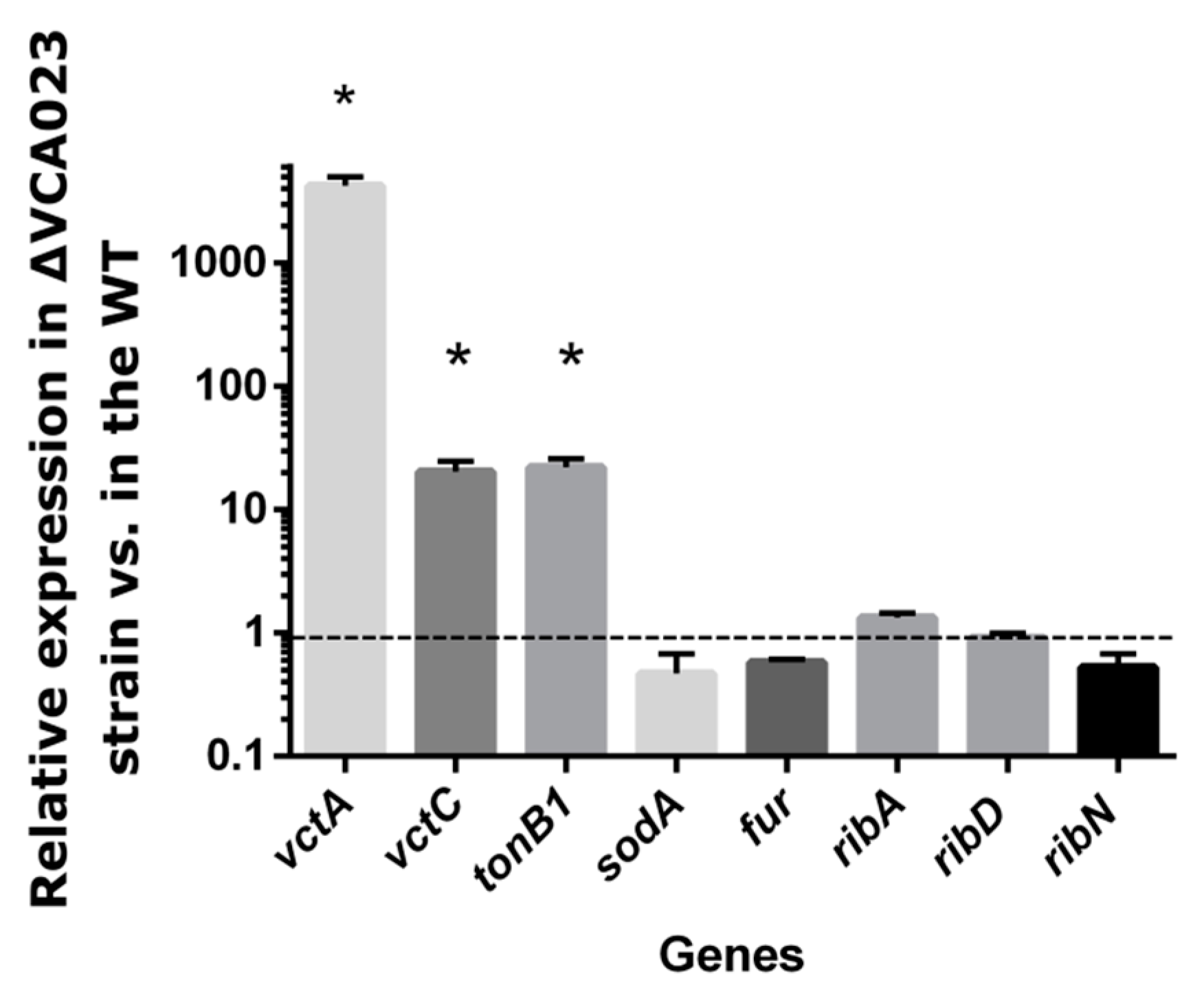
2.5. qRT-PCR
2.6. RNAseq Analysis
3. Results
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ramamurthy, T.; Mutreja, A.; Weill, F.-X.; Das, B.; Ghosh, A.; Nair, G.B. Revisiting the global epidemiology of cholera in conjuction with the genomics of Vibrio cholerae. Front. Public Health 2019, 7, 203. [Google Scholar] [CrossRef] [PubMed]
- Harris, J.B.; LaRocque, R.C.; Qadri, F.; Ryan, E.T.; Calderwood, S.B. Cholera. Lancet 2012, 379, 2466–2476. [Google Scholar] [CrossRef]
- Conner, J.G.; Teschler, J.K.; Jones, C.J.; Yildiz, F.H. Staying alive: Vibrio cholerae’s cycle of environmental survival, transmission, and dissemination. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef] [PubMed]
- Zarocostas, J. Cholera outbreak in Haiti-from 2010 to today. Lancet 2017, 389, 2274–2275. [Google Scholar] [CrossRef]
- Dureab, F.; Ismail, O.; Müller, O.; Jahn, A. Cholera outbreak in Yemen: Timeliness of reporting and response in the national electronic disease early warning system. Acta Inform. Medica 2019, 27, 85–88. [Google Scholar] [CrossRef]
- Chowdhury, F.R.; Nur, Z.; Hassan, N.; von Seidlein, L.; Dunachie, S. Pandemics, pathogenicity and changing molecular epidemiology of cholera in the era of global warming. Ann. Clin. Microbiol. Antimicrob. 2017, 16, 10. [Google Scholar] [CrossRef]
- Sánchez, M.; Sabio, L.; Gálvez, N.; Capdevila, M.; Dominguez-Vera, J.M. Iron chemistry at the service of life. IUBMB Life 2017, 69, 382–388. [Google Scholar] [CrossRef]
- Sheldon, J.R.; Laakso, H.A.; Heinrichs, D.E. Iron acquisition strategies of bacterial pathogens. Microbiol. Spectr. 2016, 4. [Google Scholar] [CrossRef]
- Cassat, J.E.; Skaar, E.P. Iron in infection and immunity. Cell Host Microbe 2013, 13, 509–519. [Google Scholar] [CrossRef]
- Peng, E.D.; Wyckoff, E.E.; Mey, A.R.; Fisher, C.R.; Payne, S.M. Nonredundant roles of iron acquisition systems in Vibrio cholerae. Infect. Immun. 2016, 84, 511–523. [Google Scholar] [CrossRef]
- Butterton, J.R.; Choi, M.H.; Watnick, P.I.; Carroll, P.A.; Calderwood, S.B. Vibrio cholerae VibF is required for vibriobactin synthesis and is a member of the family of nonribosomal peptide synthetases. J. Bacteriol. 2000, 182, 1731–1738. [Google Scholar] [CrossRef] [PubMed]
- Keating, T.A.; Marshall, C.G.; Walsh, C.T.; Keating, A.E. The structure of VibH represents nonribosomal peptide synthetase condensation, cyclization and epimerization domains. Nat. Struct. Biol. 2002, 9, 522–526. [Google Scholar] [CrossRef] [PubMed]
- Wyckoff, E.E.; Allred, B.E.; Raymond, K.N.; Payne, S.M. Catechol siderophore transport by Vibrio cholerae. J. Bacteriol. 2015, 197, 2840–2849. [Google Scholar] [CrossRef] [PubMed]
- Wyckoff, E.E.; Mey, A.R.; Payne, S.M. Iron acquisition in Vibrio cholerae. Biometals Int. J. Role Met. Ions Biol. Biochem. Med. 2007, 20, 405–416. [Google Scholar] [CrossRef] [PubMed]
- Wyckoff, E.E.; Payne, S.M. The Vibrio cholerae VctPDGC system transports catechol siderophores and a siderophore-free iron ligand. Mol. Microbiol. 2011, 81, 1446–1458. [Google Scholar] [CrossRef]
- Mey, A.R.; Wyckoff, E.E.; Oglesby, A.G.; Rab, E.; Taylor, R.K.; Payne, S.M. Identification of the Vibrio cholerae enterobactin receptors VctA and IrgA: IrgA is not required for virulence. Infect. Immun. 2002, 70, 3419–3426. [Google Scholar] [CrossRef]
- Occhino, D.A.; Wyckoff, E.E.; Henderson, D.P.; Wrona, T.J.; Payne, S.M. Vibrio cholerae iron transport: Haem transport genes are linked to one of two sets of tonB, exbB, exbD genes. Mol. Microbiol. 1998, 29, 1493–1507. [Google Scholar] [CrossRef] [PubMed]
- Mey, A.R.; Payne, S.M. Haem utilization in Vibrio cholerae involves multiple TonB-dependent haem receptors. Mol. Microbiol. 2001, 42, 835–849. [Google Scholar] [CrossRef]
- Seliger, S.S.; Mey, A.R.; Valle, A.M.; Payne, S.M. The two TonB systems of Vibrio cholerae: Redundant and specific functions. Mol. Microbiol. 2001, 39, 801–812. [Google Scholar] [CrossRef]
- Wyckoff, E.E.; Schmitt, M.; Wilks, A.; Payne, S.M. HutZ is required for efficient heme utilization in Vibrio cholerae. J. Bacteriol. 2004, 186, 4142–4151. [Google Scholar] [CrossRef]
- Wyckoff, E.E.; Mey, A.R.; Leimbach, A.; Fisher, C.F.; Payne, S.M. Characterization of ferric and ferrous iron transport systems in Vibrio cholerae. J. Bacteriol. 2006, 188, 6515–6523. [Google Scholar] [CrossRef] [PubMed]
- Mey, A.R.; Wyckoff, E.E.; Kanukurthy, V.; Fisher, C.R.; Payne, S.M. Iron and fur regulation in Vibrio cholerae and the role of fur in virulence. Infect. Immun. 2005, 73, 8167–8178. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, B.M.; Whitmire, J.M.; Merrell, D.S. This is not your mother’s repressor: The complex role of fur in pathogenesis. Infect. Immun. 2009, 77, 2590–2601. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.M.; Quinones, M.; Pratt, J.; Ding, Y.; Waldor, M.K. Characterization of the small untranslated RNA RyhB and its regulon in Vibrio cholerae. J. Bacteriol. 2005, 187, 4005–4014. [Google Scholar] [CrossRef]
- Mey, A.R.; Craig, S.A.; Payne, S.M. Characterization of Vibrio cholerae RyhB: The RyhB regulon and role of ryhB in biofilm formation. Infect. Immun. 2005, 73, 5706–5719. [Google Scholar] [CrossRef] [PubMed]
- Fetherston, J.D.; Bearden, S.W.; Perry, R.D. YbtA, an AraC-type regulator of the Yersinia pestis pesticin/yersiniabactin receptor. Mol. Microbiol. 1996, 22, 315–325. [Google Scholar] [CrossRef]
- Anisimov, R.; Brem, D.; Heesemann, J.; Rakin, A. Transcriptional regulation of high pathogenicity island iron uptake genes by YbtA. Int. J. Med. Microbiol. 2005, 295, 19–28. [Google Scholar] [CrossRef]
- Smati, M.; Magistro, G.; Adiba, S.; Wieser, A.; Picard, B.; Schubert, S.; Denamur, E. Strain-specific impact of the high-pathogenicity island on virulence in extra-intestinal pathogenic Escherichia coli. Int. J. Med. Microbiol. 2017, 307, 44–56. [Google Scholar] [CrossRef]
- Heinrichs, D.E.; Poole, K. Cloning and sequence analysis of a gene (pchR) encoding an AraC family activator of pyochelin and ferripyochelin receptor synthesis in Pseudomonas aeruginosa. J. Bacteriol. 1993, 175, 5882–5889. [Google Scholar] [CrossRef]
- Lin, P.-C.; Youard, Z.A.; Reimmann, C. In vitro-binding of the natural siderophore enantiomers pyochelin and enantiopyochelin to their AraC-type regulators PchR in Pseudomonas. Biometals 2013, 26, 1067–1073. [Google Scholar] [CrossRef]
- Beaumont, F.C.; Kang, H.Y.; Brickman, T.J.; Armstrong, S.K. Identification and characterization of alcR, a gene encoding an AraC-like regulator of alcaligin siderophore biosynthesis and transport in Bordetella pertussis and Bordetella bronchiseptica. J. Bacteriol. 1998, 180, 862–870. [Google Scholar] [CrossRef] [PubMed]
- Pradel, E.; Guiso, N.; Locht, C. Identification of AlcR, an AraC-type regulator of alcaligin siderophore synthesis in Bordetella bronchiseptica and Bordetella pertussis. J. Bacteriol. 1998, 180, 871–880. [Google Scholar] [CrossRef] [PubMed]
- Hollander, A.; Mercante, A.D.; Shafer, W.M.; Cornelissen, C.N. The iron-repressed, AraC-like regulator MpeR activates expression of fetA in Neisseria gonorrhoeae. Infect. Immun. 2011, 79, 4764–4776. [Google Scholar] [CrossRef]
- Tanabe, T.; Takata, N.; Naka, A.; Moon, Y.-H.; Nakao, H.; Inoue, Y.; Narimatsu, S.; Yamamoto, S. Identification of an AraC-like regulator gene required for induction of the 78-kDa ferrioxamine B receptor in Vibrio vulnificus. FEMS Microbiol. Lett. 2005, 249, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Tanabe, T.; Funahashi, T.; Miyamoto, K.; Tsujibo, H.; Yamamoto, S. Identification of genes, desR and desA, required for utilization of desferrioxamine B as a xenosiderophore in Vibrio furnissii. Biol. Pharm. Bull. 2011, 34, 570–574. [Google Scholar] [CrossRef][Green Version]
- Funahashi, T.; Tanabe, T.; Maki, J.; Miyamoto, K.; Tsujibo, H.; Yamamoto, S. Identification and characterization of Aeromonas hydrophila genes encoding the outer membrane receptor of ferrioxamine B and an AraC-type transcriptional regulator. Biosci. Biotechnol. Biochem. 2014, 78, 1777–1787. [Google Scholar] [CrossRef][Green Version]
- Sepúlveda-Cisternas, I.; Salazar, J.C.; García-Angulo, V.A. Overview on the bacterial iron-riboflavin metabolic axis. Front. Microbiol. 2018, 9, 1478. [Google Scholar] [CrossRef]
- Sepúlveda-Cisternas, I.; Lozano Aguirre, L.; Fuentes Flores, A.; Vásquez Solis de Ovando, I.; García-Angulo, V.A. Transcriptomics reveals a cross-modulatory effect between riboflavin and iron and outlines responses to riboflavin biosynthesis and uptake in Vibrio cholerae. Sci. Rep. 2018, 8, 3149. [Google Scholar] [CrossRef]
- Rivera-Chávez, F.; Mekalanos, J.J. Cholera toxin promotes pathogen acquisition of host-derived nutrients. Nature 2019, 572, 244–248. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Zallot, R.; Oberg, N.; Gerlt, J.A. The EFI web resource for genomic enzymology tools: Leveraging protein, genome, and metagenome databases to discover novel enzymes and metabolic pathways. Biochemistry 2019, 58, 4169–4182. [Google Scholar] [CrossRef] [PubMed]
- Ibarra, J.A.; Pérez-Rueda, E.; Segovia, L.; Puente, J.L. The DNA-binding domain as a functional indicator: The case of the AraC/XylS family of transcription factors. Genetica 2008, 133, 65–76. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Guindon, S.; Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 2003, 52, 696–704. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Sawitzke, J.A.; Thomason, L.C.; Bubunenko, M.; Li, X.; Costantino, N.; Court, D.L. Recombineering: Using drug cassettes to knock out genes in vivo. Methods Enzymol. 2013, 533, 79–102. [Google Scholar] [CrossRef]
- Santiviago, C.A.; Reynolds, M.M.; Porwollik, S.; Choi, S.-H.; Long, F.; Andrews-Polymenis, H.L.; McClelland, M. Analysis of pools of targeted Salmonella deletion mutants identifies novel genes affecting fitness during competitive infection in mice. PLoS Pathog. 2009, 5, e1000477. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- Cisternas, I.S.; Torres, A.; Flores, A.F.; Angulo, V.A.G. Differential regulation of riboflavin supply genes in Vibrio cholerae. Gut Pathog. 2017, 9, 10. [Google Scholar] [CrossRef][Green Version]
- Wattam, A.R.; Davis, J.J.; Assaf, R.; Boisvert, S.; Brettin, T.; Bun, C.; Conrad, N.; Dietrich, E.M.; Disz, T.; Gabbard, J.L.; et al. Improvements to PATRIC, the all-bacterial bioinformatics database and analysis resource center. Nucleic Acids Res. 2017, 45, D535–D542. [Google Scholar] [CrossRef]
- Langmead, B.; Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 2012, 9, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, S.; Chan, C.-K.K. Analysis of RNA-Seq data using TopHat and Cufflinks. Methods Mol. Biol. 2016, 1374, 339–361. [Google Scholar] [CrossRef]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Gene Ontology Consortium. The Gene Ontology Resource: 20 years and still GOing strong. Nucleic Acids Res. 2019, 47, D330–D338. [Google Scholar] [CrossRef] [PubMed]
- Gerlt, J.A.; Bouvier, J.T.; Davidson, D.B.; Imker, H.J.; Sadkhin, B.; Slater, D.R.; Whalen, K.L. Enzyme Function Initiative-Enzyme Similarity Tool (EFI-EST): A web tool for generating protein sequence similarity networks. Biochim. Biophys. Acta 2015, 1854, 1019–1037. [Google Scholar] [CrossRef]
- Zallot, R.; Oberg, N.O.; Gerlt, J.A. “Democratized” genomic enzymology web tools for functional assignment. Curr. Opin. Chem. Biol. 2018, 47, 77–85. [Google Scholar] [CrossRef]
- Ferguson, A.D.; Deisenhofer, J. TonB-dependent receptors-structural perspectives. Biochim. Biophys. Acta 2002, 1565, 318–332. [Google Scholar] [CrossRef]
- Gallegos, M.T.; Schleif, R.; Bairoch, A.; Hofmann, K.; Ramos, J.L. Arac/XylS family of transcriptional regulators. Microbiol. Mol. Biol. Rev. 1997, 61, 393–410. [Google Scholar] [CrossRef]
- Egan, S.M. Growing repertoire of AraC/XylS activators. J. Bacteriol. 2002, 184, 5529–5532. [Google Scholar] [CrossRef]
- García Angulo, V.A.; Bonomi, H.R.; Posadas, D.M.; Serer, M.I.; Torres, A.G.; Zorreguieta, A.; Goldbaum, F.A. Identification and characterization of RibN, a novel family of riboflavin transporters from Rhizobium leguminosarum and other proteobacteria. J. Bacteriol. 2013, 195, 4611–4619. [Google Scholar] [CrossRef]
- Salvail, H.; Massé, E. Regulating iron storage and metabolism with RNA: An overview of posttranscriptional controls of intracellular iron homeostasis. Wiley Interdiscip. Rev. RNA 2012, 3, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Chareyre, S.; Mandin, P. Bacterial iron homeostasis regulation by sRNAs. Microbiol. Spectr. 2018, 6. [Google Scholar] [CrossRef]
- Martinez-Guerrero, C.E.; Ciria, R.; Abreu-Goodger, C.; Moreno-Hagelsieb, G.; Merino, E. GeConT 2: Gene context analysis for orthologous proteins, conserved domains and metabolic pathways. Nucleic Acids Res. 2008, 36, W176–W180. [Google Scholar] [CrossRef] [PubMed]
- Heinrichs, D.E.; Poole, K. PchR, a regulator of ferripyochelin receptor gene (fptA) expression in Pseudomonas aeruginosa, functions both as an activator and as a repressor. J. Bacteriol. 1996, 178, 2586–2592. [Google Scholar] [CrossRef] [PubMed]
- Mercante, A.D.; Jackson, L.; Johnson, P.J.T.; Stringer, V.A.; Dyer, D.W.; Shafer, W.M. MpeR regulates the mtr efflux locus in Neisseria gonorrhoeae and modulates antimicrobial resistance by an iron-responsive mechanism. Antimicrob. Agents Chemother. 2012, 56, 1491–1501. [Google Scholar] [CrossRef]
- Fantappiè, L.; Scarlato, V.; Delany, I. Identification of the in vitro target of an iron-responsive AraC-like protein from Neisseria meningitidis that is in a regulatory cascade with Fur. Microbiol. Read. Engl. 2011, 157, 2235–2247. [Google Scholar] [CrossRef] [PubMed]
- Monteverde, D.R.; Gómez-Consarnau, L.; Suffridge, C.; Sañudo-Wilhelmy, S.A. Life’s utilization of B vitamins on early Earth. Geobiology 2016. [Google Scholar] [CrossRef]
- Baichoo, N.; Wang, T.; Ye, R.; Helmann, J.D. Global analysis of the Bacillus subtilis Fur regulon and the iron starvation stimulon. Mol. Microbiol. 2002, 45, 1613–1629. [Google Scholar] [CrossRef]
- Crossley, R.A.; Gaskin, D.J.H.; Holmes, K.; Mulholland, F.; Wells, J.M.; Kelly, D.J.; van Vliet, A.H.M.; Walton, N.J. Riboflavin biosynthesis is associated with assimilatory ferric reduction and iron acquisition by Campylobacter jejuni. Appl. Environ. Microbiol. 2007, 73, 7819–7825. [Google Scholar] [CrossRef]
- Berges, M.; Michel, A.-M.; Lassek, C.; Nuss, A.M.; Beckstette, M.; Dersch, P.; Riedel, K.; Sievers, S.; Becher, D.; Otto, A.; et al. Iron regulation in Clostridioides difficile. Front. Microbiol. 2018, 9, 3183. [Google Scholar] [CrossRef]
- Jiménez-Munguía, I.; Calderón-Santiago, M.; Rodríguez-Franco, A.; Priego-Capote, F.; Rodríguez-Ortega, M.J. Multi-omic profiling to assess the effect of iron starvation in Streptococcus pneumoniae TIGR4. PeerJ 2018, 6, e4966. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Lonergan, Z.R.; Gonzalez-Gutierrez, G.; Nairn, B.L.; Maxwell, C.N.; Zhang, Y.; Andreini, C.; Karty, J.A.; Chazin, W.J.; Trinidad, J.C.; et al. Multi-metal restriction by calprotectin impacts de novo flavin biosynthesis in Acinetobacter baumannii. Cell Chem. Biol. 2019, 26, 745–755.e7. [Google Scholar] [CrossRef] [PubMed]
| ORF | Gene Name | Log2 Fold Change | p Value | Annotation |
|---|---|---|---|---|
| VC2691 | cpxP | −5.95515 | <5.00E−05 | P pilus assembly/Cpx signaling pathway, periplasmic inhibitor/zinc−resistance associated protein |
| VC0633 | OmpU | −5.92287 | <5.00E−05 | Outer membrane porin OmpU |
| VCA0261−0 | −5.08589 | <5.00E−05 | Hypotetical protein, Hypotetical protein | |
| VCA0262 | −3.93658 | 0.0024 | Hypotetical protein | |
| VC1587 | −3.42537 | <5.00E−05 | Membrane protein | |
| VCA0615 | msrB | −3.27937 | <5.00E−05 | Peptide−methionine (S)−S−oxide reductase MsrA/Peptide−methionine (R)−S−oxide reductase MsrB |
| VCA0258 | −3.22844 | <5.00E−05 | Hemolysin | |
| VC2216 | copG | −2.95846 | <5.00E−05 | CopG protein |
| VCA0650−1 | −2.94231 | <5.00E−05 | D−alanine−D−alanine ligase, Hipotetical protein | |
| VCA0139 | −2.84816 | <5.00E−05 | Acyl−CoA synthetase | |
| VC2215 | −2.5293 | <5.00E−05 | Lead, cadmium, zinc and mercury transporting ATPase; Copper−translocating P−type ATPase | |
| VCA0029 | −2.48946 | <5.00E−05 | Transcriptional regulator, IclR family | |
| VC0232 | −2.40928 | <5.00E−05 | Hypotetical protein | |
| VCA0752 | −2.16805 | <5.00E−05 | Thioredoxin 2 | |
| VCA0648−9 | −2.08498 | <5.00E−05 | Hypotetical protein, Hypotetical protein | |
| VCA1011 | −2.07667 | <5.00E−05 | Glutaredoxin | |
| VC1044−5 | −2.00207 | <5.00E−05 | Hypotetical protein, RNA polymerase ECF−type sigma factor | |
| VCA0646−7 | −1.97651 | <5.00E−05 | Putative hemolysin | |
| VC1723 | −1.88711 | <5.00E−05 | COG0398: uncharacterized membrane protein | |
| VC0143 | −1.88157 | <5.00E−05 | Hypotetical protein | |
| VCA0151 | −1.72402 | <5.00E−05 | Predicted ferric reductase | |
| VC0285 | 1.68838 | <5.00E−05 | 4−hydroxy−2−oxoglutarate aldolase; 2−dehydro−3−deoxyphosphogluconate aldolase | |
| VCA0687 | fbpC | 1.75742 | <5.00E−05 | ABC transporter, ATP−binding protein (cluster 1, maltose/g3p/polyamine/iron); ABC transporter, ATP−binding protein (cluster 10, nitrate/sulfonate/bicarbonate) | fbpC |
| VC1589 | 1.76693 | <5.00E−05 | α−acetolactate decarboxylase | |
| VC2076−77−78 | feoABC | 1.81165 | <5.00E−05 | Ferrous iron transport protein C, Ferrous iron transporter FeoB, Ferrous iron transporter−associated protein FeoA |
| VCA0686 | 1.82212 | <5.00E−05 | ABC transporter, permease protein 1 (cluster 1, maltose/g3p/polyamine/iron)/ABC transporter, permease protein 2 (cluster 1, maltose/g3p/polyamine/iron) | |
| VC1334 | 1.82445 | <5.00E−05 | Tripartite tricarboxylate transporter TctC family | |
| VC1266−7 | 1.84393 | <5.00E−05 | Iron−regulated protein A precursor, Hypotetical protein | |
| VC2761 | 1.84679 | <5.00E−05 | Multidrug resistance transporter, Bcr/CflA family | |
| VC1543−44−45−46 | exbB2 exbD2 tonB2 VC1543 | 1.88124 | <5.00E−05 | TPR domain protein, putative component of TonB system, putative TonB−dependent receptor, Ferric siderophore transport system, biopolymer transport protein ExbB |
| VC0284 | 2.02153 | <5.00E−05 | Ferrichrome−iron receptor | |
| VCA1041 | 2.0355 | <5.00E−05 | Phosphosugar mutase of unknown sugar (see annotation) | |
| VC1572 | 2.12029 | <5.00E−05 | Hypotetical protein | |
| VC1265 | 2.15881 | <5.00E−05 | Probable thiol oxidoreductase with 2 cytochrome c heme−binding sites | |
| VC0364 | bfd | 2.22671 | <5.00E−05 | Bacterioferritin−associated ferredoxin |
| VC0286 | gntU | 2.22969 | <5.00E−05 | Low−affinity gluconate/H+ symporter GntU |
| VCA0976 | 2.23539 | <5.00E−05 | Hypotetical protein | |
| VCA0984 | ildD | 2.27194 | <5.00E−05 | L−lactate dehydrogenase | lldD |
| VC1332 | 2.34818 | <5.00E−05 | Tripartite tricarboxylate transporter TctA family | |
| VCA0983 | 2.43475 | <5.00E−05 | L−lactate permease | |
| VCA0227 | vctP | 2.47329 | <5.00E−05 | Ferric vibriobactin, enterobactin transport system, substrate−binding protein VctP |
| VC0091 | 2.48987 | <5.00E−05 | Uncharacterized SAM−dependent O−methyltransferase | |
| VC1547−8 | exbB2 VC1548 | 2.54473 | <5.00E−05 | MotA/TolQ/ExbB proton channel family protein, TonB system biopolymer transport component; Chromosome segregation ATPase |
| VC0608 | fbpA | 2.57038 | <5.00E−05 | Ferric iron ABC transporter, iron−binding protein |
| VC0199 | 2.6252 | <5.00E−05 | Efflux ABC transporter, permease/ATP−binding protein Atu2242 | |
| VC1333 | 2.66616 | <5.00E−05 | Tripartite tricarboxylate transporter TctB family | |
| VC0201−2−3 | fhuAC VC0203 | 2.67288 | <5.00E−05 | Ferrichrome transport ATP−binding protein FhuC, Ferrichrome−binding periplasmic protein precursor, Ferrichrome transport system permease protein FhuB |
| VC1264 | irpA | 2.6755 | <5.00E−05 | Iron−regulated protein A precursor |
| VC1573 | fumC | 2.68696 | <5.00E−05 | Fumarate hydratase class II (EC 4.2.1.2) | fumC |
| VC2209 | vibF | 2.71161 | <5.00E−05 | Non−ribosomal peptide synthetase modules, siderophore biosynthesis | vibF |
| VCA0230 | vctC | 2.71848 | <5.00E−05 | Ferric vibriobactin, enterobactin transport system, ATP−binding protein |
| VC0474 | irgB | 2.842 | <5.00E−05 | Iron−regulated virulence regulatory protein irgB |
| VC2210 | viuB | 3.05349 | <5.00E−05 | Vibriobactin utilization protein ViuB |
| VCA0914 | hutB | 3.1391 | <5.00E−05 | Hemin ABC transporter, permease protein |
| VC2211 | viuA | 3.17699 | <5.00E−05 | Ferric vibriobactin receptor ViuA |
| VC0776−7−8 | viuPDG | 3.36327 | <5.00E−05 | Ferric vibriobactin, enterobactin transport system, substrate−binding protein ViuP, permease protein ViuD, permease protein ViuG |
| VCA0977 | 3.59744 | <5.00E−05 | ABC transporter, ATP−binding protein (cluster 5, nickel/peptides/opines)/ABC transporter, ATP−binding protein (cluster 5, nickel/peptides/opines) | |
| VC0200 | fhuA | 3.62978 | <5.00E−05 | Ferrichrome−iron receptor |
| VCA0540 | focA | 3.73459 | <5.00E−05 | Formate efflux transporter FocA |
| VC0772 | vibE | 3.80191 | <5.00E−05 | 2,3−dihydroxybenzoate−AMP ligase of siderophore biosynthesis @ 2,3−dihydroxybenzoate−AMP ligase |
| VC0771 | vibB | 4.12473 | <5.00E−05 | Isochorismatase of siderophore biosynthesis |
| VCA0907 | hutZ | 4.18631 | <5.00E−05 | Pyridoxamine 5’−phosphate oxidase−related putative heme iron utilization protein |
| VCA0908 | hutX | 4.24657 | <5.00E−05 | Putative heme iron utilization protein |
| VCA0909 | hutW | 4.29654 | <5.00E−05 | Radical SAM family protein HutW, similar to coproporphyrinogen III oxidase, oxygen−independent, associated with heme uptake |
| VC0775 | 4.40511 | <5.00E−05 | Amide synthase component of siderophore synthetase | |
| VC1688 | mntX | 4.6546 | <5.00E−05 | Manganese transport protein MntX |
| VCA0576 | hutA | 4.70136 | <5.00E−05 | TonB−dependent heme and hemoglobin receptor HutA; TonB−dependent hemin, ferrichrome receptor |
| VCA0910−1−2 | tonB1 exbB1 exbD1 | 4.74152 | <5.00E−05 | Putative TonB−dependent receptor, Ferric siderophore transport system, biopolymer transport protein ExbB, Biopolymer transport protein ExbD1 |
| VCA0913 | 4.82235 | <5.00E−05 | Periplasmic hemin−binding protein | |
| VC0774 | vibA | 4.90504 | <5.00E−05 | 2,3−dihydro−2,3−dihydroxybenzoate dehydrogenase of siderophore biosynthesis @ 2,3−dihydro−2,3−dihydroxybenzoate dehydrogenase |
| VC0773 | vibC | 5.3735 | <5.00E−05 | Isochorismate synthase @ Isochorismate synthase of siderophore biosynthesis |
| VCA0233 | 5.66124 | <5.00E−05 | Hypothetical protein colocalized with Enterobactin receptor VctA | |
| VBIVibCho83274_2864 | vctA | 8.67608 | <5.00E−05 | Enterobactin receptor VctA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sachman-Ruiz, B.; Ibarra, J.A.; Estrada-de los Santos, P.; Torres Muñoz, A.; Giménez, B.; Salazar, J.C.; García-Angulo, V.A. IurV, Encoded by ORF VCA0231, Is Involved in the Regulation of Iron Uptake Genes in Vibrio cholerae. Genes 2020, 11, 1184. https://doi.org/10.3390/genes11101184
Sachman-Ruiz B, Ibarra JA, Estrada-de los Santos P, Torres Muñoz A, Giménez B, Salazar JC, García-Angulo VA. IurV, Encoded by ORF VCA0231, Is Involved in the Regulation of Iron Uptake Genes in Vibrio cholerae. Genes. 2020; 11(10):1184. https://doi.org/10.3390/genes11101184
Chicago/Turabian StyleSachman-Ruiz, Bernardo, José Antonio Ibarra, Paulina Estrada-de los Santos, Alexia Torres Muñoz, Begoña Giménez, Juan Carlos Salazar, and Víctor Antonio García-Angulo. 2020. "IurV, Encoded by ORF VCA0231, Is Involved in the Regulation of Iron Uptake Genes in Vibrio cholerae" Genes 11, no. 10: 1184. https://doi.org/10.3390/genes11101184
APA StyleSachman-Ruiz, B., Ibarra, J. A., Estrada-de los Santos, P., Torres Muñoz, A., Giménez, B., Salazar, J. C., & García-Angulo, V. A. (2020). IurV, Encoded by ORF VCA0231, Is Involved in the Regulation of Iron Uptake Genes in Vibrio cholerae. Genes, 11(10), 1184. https://doi.org/10.3390/genes11101184






