The Use of Genetic and Gene Technologies in Shaping Modern Rapeseed Cultivars (Brassica napus L.)
Abstract
1. Introduction
2. Improvement and Innovations of Canola Varieties
2.1. Higher Resolution for Rapeseed Genome Characterization
2.2. Pan-Genome
3. Breeding for Economically Important Agronomic Traits of B. napus
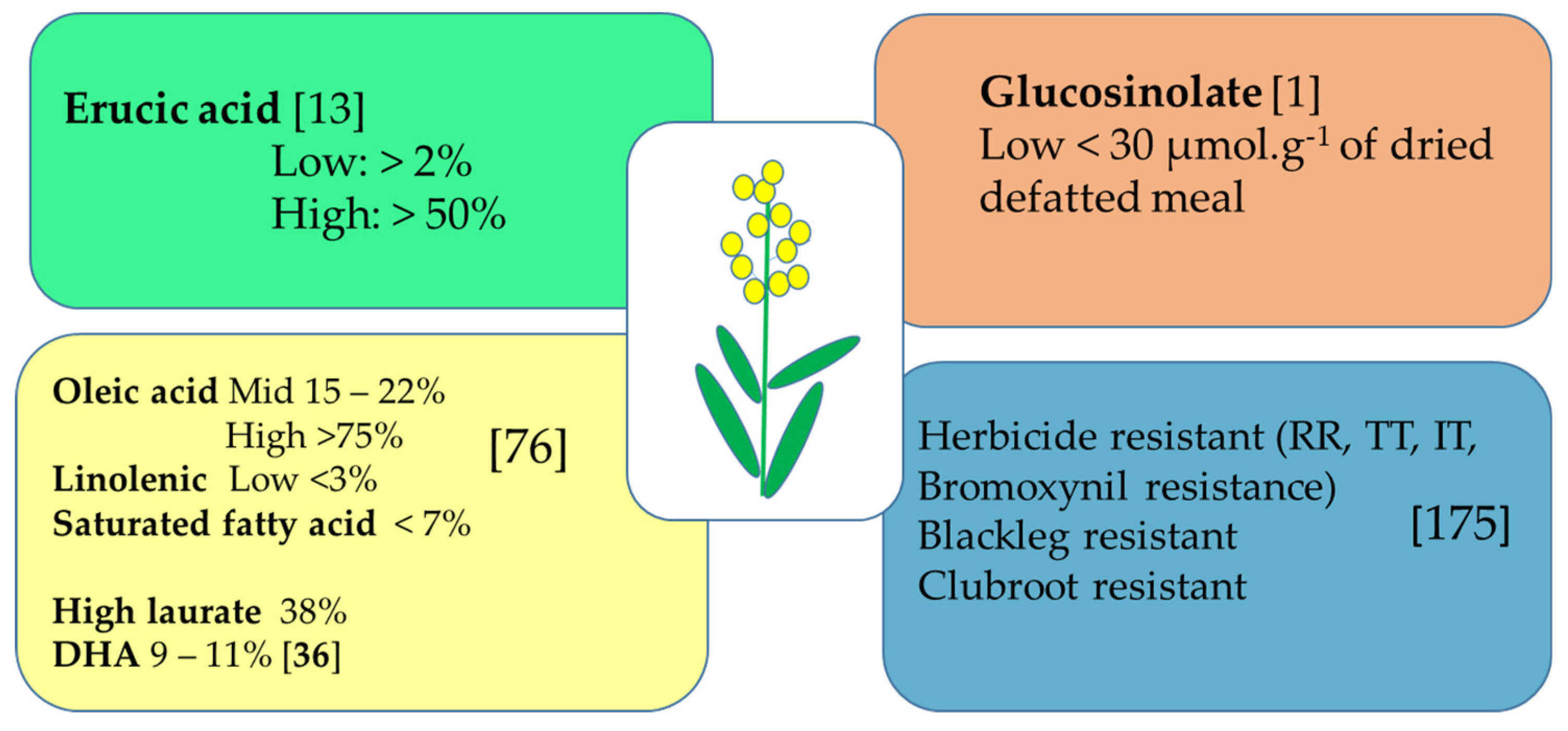
3.1. Oil Content and Specialty as Priority
3.2. Exploiting Canola Meal Potential
3.3. Disease Resistance
3.3.1. Blackleg Resistance
3.3.2. Clubroot Resistance
3.3.3. Sclerotinia Resistance
3.4. Abiotic Stress Tolerance
3.5. Herbicide Tolerance/Resistance
4. Prospects and Future Directions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- McVetty, P.B.E.; Duncan, R.W. Canola/Rapeseed: Genetics and Breeding. In Reference Module in Food Science; Elsevier: Amsterdam, The Netherlands, 2016. [Google Scholar] [CrossRef]
- OGTR. The Biology of Brassica napus L. (Canola); Aging, Department of Health and Aging, Ed.; Office of the Gene Technology Regulator: Canberra, ACT, Australia, 2008.
- Nagaharu, U. Genome-analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jpn J. Bot. 1935, 7, 389–452. [Google Scholar]
- Lu, K.; Wei, L.; Li, X.; Wang, Y.; Wu, J.; Liu, M.; Zhang, C.; Chen, Z.; Xiao, Z.; Jian, H.; et al. Whole-genome resequencing reveals Brassica napus origin and genetic loci involved in its improvement. Nat. Commun. 2019, 10, 1154. [Google Scholar] [CrossRef]
- Chalhoub, B.; Denoeud, F.; Liu, S.; Parkin, I.A.P.; Tang, H.; Wang, X.; Chiquet, J.; Belcram, H.; Tong, C.; Samans, B.; et al. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 2014, 345, 950. [Google Scholar] [CrossRef]
- Gómez-Campo, C.; Prakash, S. 2 Origin and domestication. In Developments in Plant Genetics and Breeding; Gómez-Campo, C., Ed.; Elsevier: Amsterdam, The Netherlands, 1999; Volume 4, pp. 33–58. [Google Scholar]
- Eskin, N.A.M.; Przybylski, R. RAPE SEED OIL/CANOLA. In Encyclopedia of Food Sciences and Nutrition, 2nd ed.; Caballero, B., Ed.; Academic Press: Oxford, UK, 2003; pp. 4911–4916. [Google Scholar] [CrossRef]
- Scarth, R.; McVetty, P.B.E.; Rimmer, S.R.; Stefansson, B.R. STELLAR LOW LINOLENIC-HIGH LINOLEIC ACID SUMMER RAPE. Can. J. Plant Sci. 1988, 68, 509–511. [Google Scholar] [CrossRef]
- Long, W.; Hu, M.; Gao, J.; Chen, S.; Zhang, J.; Cheng, L.; Pu, H. Identification and Functional Analysis of Two New Mutant BnFAD2 Alleles That Confer Elevated Oleic Acid Content in Rapeseed. Front. Genet. 2018, 9. [Google Scholar] [CrossRef]
- Maher, L.B.W.; Salisbury, P.; Debonte, L.; Deng, X. (Organising Committee of: Wuhan). High Oleic, low linolenic (HOLL) specialty canola development in Australia. In Proceedings of the 12th International Rapeseed Congress, Wuhan, China, 23–30 March 2007; Volume 5, pp. 22–24. [Google Scholar]
- Nadeem, M.A.; Nawaz, M.A.; Shahid, M.Q.; Doğan, Y.; Comertpay, G.; Yıldız, M.; Hatipoğlu, R.; Ahmad, F.; Alsaleh, A.; Labhane, N.; et al. DNA molecular markers in plant breeding: Current status and recent advancements in genomic selection and genome editing. Biotechnol. Biotechnol. Equip. 2018, 32, 261–285. [Google Scholar] [CrossRef]
- Spasibionek, S.; Mikołajczyk, K.; Ćwiek-Kupczyńska, H.; Piętka, T.; Krótka, K.; Matuszczak, M.; Nowakowska, J.; Michalski, K.; Bartkowiak-Broda, I. Marker assisted selection of new high oleic and low linolenic winter oilseed rape (Brassica napus L.) inbred lines revealing good agricultural value. PLoS ONE 2020, 15, e0233959. [Google Scholar] [CrossRef]
- Nath, U.K.; Wilmer, J.A.; Wallington, E.J.; Becker, H.C.; Möllers, C. Increasing erucic acid content through combination of endogenous low polyunsaturated fatty acids alleles with Ld-LPAAT + Bn-fae1 transgenes in rapeseed (Brassica napus L.). Theor. Appl. Genet. 2009, 118. [Google Scholar] [CrossRef]
- Hayward, A. Introduction-Oilseed Brassicas. In Genetics, Genomics and Breeding of Oilseed Brassicas; Edwards, D., Batley, J., Parkin, I., Kole, C., Eds.; Taylor & Francis Group: Baton Rouge, LA, USA, 2011. [Google Scholar]
- Wei, D.; Cui, Y.; He, Y.; Xiong, Q.; Qian, L.; Tong, C.; Lu, G.; Ding, Y.; Li, J.; Jung, C.; et al. A genome-wide survey with different rapeseed ecotypes uncovers footprints of domestication and breeding. J. Exp. Bot. 2017, 68, 4791–4801. [Google Scholar] [CrossRef]
- Bonnardeaux, J. Uses for Canola Meal; Department of Agriculture and Food: Perth, Western Australia, Australia, 2007.
- Nath, U.K.; Kim, H.-T.; Khatun, K.; Park, J.-I.; Kang, K.-K.; Ill-Sup, N. Modification of Fatty Acid Profiles of Rapeseed (Brassica napus L.) Oil for Using as Food, Industrial Feed-Stock and Biodiesel. Plant Breed. Biotechnol. 2016, 4, 123–134. [Google Scholar] [CrossRef]
- Palmer, C.E.D.; Keller, W.A. Future Prospects for Brassica Oilseed Improvement through Genomics. In Genetics, Genomics and Breeding of Oilseed Brassicas; Edwards, D., Batley, J., Parkin, I., Kole, C., Eds.; Taylor & Francis Group: Baton Rouge, LA, USA, 2011; pp. 1–13. [Google Scholar]
- Li, F.-W.; Harkess, A. A guide to sequence your favorite plant genomes. Appl. Plant Sci. 2018, 6, e1030. [Google Scholar] [CrossRef]
- Schiessl, S.V.; Quezada-Martinez, D.; Tebartz, E.; Snowdon, R.J.; Qian, L. The vernalisation regulator FLOWERING LOCUS C is differentially expressed in biennial and annual Brassica napus. Sci. Rep. 2019, 9, 14911. [Google Scholar] [CrossRef]
- Kyriakidou, M.; Tai, H.H.; Anglin, N.L.; Ellis, D.; Strömvik, M.V. Current Strategies of Polyploid Plant Genome Sequence Assembly. Front. Plant Sci. 2018, 9, 1660. [Google Scholar] [CrossRef]
- Yuan, Y.; Bayer, P.E.; Lee, H.T.; Edwards, D. runBNG: A software package for BioNano genomic analysis on the command line. Bioinformatics 2017, 33, 3107–3109. [Google Scholar] [CrossRef]
- Oluwadare, O.; Highsmith, M.; Cheng, J. An Overview of Methods for Reconstructing 3-D Chromosome and Genome Structures from Hi-C Data. Biol. Proced. Online 2019, 21, 7. [Google Scholar] [CrossRef]
- Malmberg, M.M.; Barbulescu, D.M.; Drayton, M.C.; Shinozuka, M.; Thakur, P.; Ogaji, Y.O.; Spangenberg, G.C.; Daetwyler, H.D.; Cogan, N.O.I. Evaluation and Recommendations for Routine Genotyping Using Skim Whole Genome Re-sequencing in Canola. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Qu, C.; Jia, L.; Fu, F.; Zhao, H.; Lu, K.; Wei, L.; Xu, X.; Liang, Y.; Li, S.; Wang, R.; et al. Genome-wide association mapping and Identification of candidate genes for fatty acid composition in Brassica napus L. using SNP markers. BMC Genom. 2017, 18, 232. [Google Scholar] [CrossRef]
- Scheben, A.; Batley, J.; Edwards, D. Genotyping-by-sequencing approaches to characterize crop genomes: Choosing the right tool for the right application. Plant Biotechnol. J. 2017, 15, 149–161. [Google Scholar] [CrossRef]
- Fikere, M.; Barbulescu, D.M.; Malmberg, M.M.; Spangenberg, G.C.; Cogan, N.O.I.; Daetwyler, H.D. Meta-analysis of GWAS in canola blackleg (Leptosphaeria maculans) disease traits demonstrates increased power from imputed whole-genome sequence. Sci. Rep. 2020, 10, 14300. [Google Scholar] [CrossRef]
- Farid, M.; Yang, R.C.; Kebede, B.; Rahman, H. Evaluation of Brassica oleracea accessions for resistance to Plasmodiophora brassicae and identification of genomic regions associated with resistance. Genome 2020, 63, 91–101. [Google Scholar] [CrossRef]
- Mason, A.S.; Higgins, E.E.; Snowdon, R.J.; Batley, J.; Stein, A.; Werner, C.; Parkin, I.A. A user guide to the Brassica 60K Illumina Infinium™ SNP genotyping array. Theor. Appl. Genet. 2017, 130, 621–633. [Google Scholar] [CrossRef] [PubMed]
- Katche, E.; Quezada-Martinez, D.; Katche, E.I.; Vasquez-Teuber, P.; Mason, A.S. Interspecific Hybridization for Brassica Crop Improvement. Crop Breed. Genet. Genom. 2019, 1, e190007. [Google Scholar] [CrossRef]
- Wang, Z.; Ma, L.-Y.; Cao, J.; Li, Y.-L.; Ding, L.-N.; Zhu, K.-M.; Yang, Y.-H.; Tan, X.-L. Recent Advances in Mechanisms of Plant Defense to Sclerotinia sclerotiorum. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Cardoza, V.S.C.N. Agrobacterium-Mediated Transformation of Canola. In Transgenic Crops of the World; Curtis, I.S., Ed.; Springer: Dordrecht, The Netherlands, 2004. [Google Scholar]
- Zhang, K.; Nie, L.; Cheng, Q.; Yin, Y.; Chen, K.; Qi, F.; Zou, D.; Liu, H.; Zhao, W.; Wang, B.; et al. Effective editing for lysophosphatidic acid acyltransferase 2/5 in allotetraploid rapeseed (Brassica napus L.) using CRISPR-Cas9 system. Biotechnol. Biofuels 2019, 12, 225. [Google Scholar] [CrossRef]
- Wu, G.; Truksa, M.; Datla, N.; Vrinten, P.; Bauer, J.; Zank, T.; Cirpus, P.; Heinz, E.; Qiu, X. Stepwise engineering to produce high yields of very long-chain polyunsaturated fatty acids in plants. Nat. Biotechnol. 2005, 23, 1013–1017. [Google Scholar] [CrossRef]
- Okuzaki, A.; Ogawa, T.; Koizuka, C.; Kaneko, K.; Inaba, M.; Imamura, J.; Koizuka, N. CRISPR/Cas9-mediated genome editing of the fatty acid desaturase 2 gene in Brassica napus. Plant Physiol. Biochem. 2018, 131, 63–69. [Google Scholar] [CrossRef]
- Petrie, J.R.; Zhou, X.-R.; Leonforte, A.; McAllister, J.; Shrestha, P.; Kennedy, Y.; Belide, S.; Buzza, G.; Gororo, N.; Gao, W.; et al. Development of a Brassica napus (Canola) Crop Containing Fish Oil-Like Levels of DHA in the Seed Oil. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Puchta, H. Applying CRISPR/Cas for genome engineering in plants: The best is yet to come. Curr. Opin. Plant Biol. 2017, 36, 1–8. [Google Scholar] [CrossRef]
- Burmistrz, M.; Krakowski, K.; Krawczyk-Balska, A. RNA-Targeting CRISPR-Cas Systems and Their Applications. Int. J. Mol. Sci. 2020, 21, 1122. [Google Scholar] [CrossRef]
- European Academies Science Advisory Council. The Regulation of Genome-Edited Plants in the European Union; EU: Geneva, Switzerland, 2020. [Google Scholar]
- Makarova, K.S.; Koonin, E.V. Annotation and Classification of CRISPR-Cas Systems. Methods Mol. Biol. 2015, 1311, 47–75. [Google Scholar] [CrossRef]
- Borrelli, V.M.G.; Brambilla, V.; Rogowsky, P.; Marocco, A.; Lanubile, A. The Enhancement of Plant Disease Resistance Using CRISPR/Cas9 Technology. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Hao, M.; Wang, W.; Wang, H.; Chen, F.; Chu, W.; Zhang, B.; Mei, D.; Cheng, H.; Hu, Q. An Efficient CRISPR/Cas9 Platform for Rapidly Generating Simultaneous Mutagenesis of Multiple Gene Homoeologs in Allotetraploid Oilseed Rape. Front. Plant Sci. 2018, 9, 442. [Google Scholar] [CrossRef] [PubMed]
- Zheng, M.; Zhang, L.; Tang, M.; Liu, J.; Liu, H.; Yang, H.; Fan, S.; Terzaghi, W.; Wang, H.; Hua, W. Knockout of two BnaMAX1 homologs by CRISPR/Cas9-targeted mutagenesis improves plant architecture and increases yield in rapeseed (Brassica napus L.). Plant Biotechnol. J. 2020, 18, 644–654. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Zhu, K.; Li, H.; Han, S.; Meng, Q.; Khan, S.U.; Fan, C.; Xie, K.; Zhou, Y. Precise editing of CLAVATA genes in Brassica napus L. regulates multilocular silique development. Plant Biotechnol. J. 2018, 16, 1322–1335. [Google Scholar] [CrossRef]
- Zhai, Y.; Cai, S.; Hu, L.; Yang, Y.; Amoo, O.; Fan, C.; Zhou, Y. CRISPR/Cas9-mediated genome editing reveals differences in the contribution of INDEHISCENT homologues to pod shatter resistance in Brassica napus L. Theor. Appl. Genet. 2019, 132, 2111–2123. [Google Scholar] [CrossRef]
- Sriboon, S.; Li, H.; Guo, C.; Senkhamwong, T.; Dai, C.; Liu, K. Knock-out of TERMINAL FLOWER 1 genes altered flowering time and plant architecture in Brassica napus. BMC Genet. 2020, 21, 52. [Google Scholar] [CrossRef]
- Jat, R.S.; Singh, V.V.; Sharma, P.; Rai, P.K. Oilseed brassica in India: Demand, supply, policy perspective and future potential. OCL 2019, 26, 8. [Google Scholar] [CrossRef]
- Snowdon, R.J.; Iniguez Luy, F.L. Potential to improve oilseed rape and canola breeding in the genomics era. Plant Breed. 2012, 131, 351–360. [Google Scholar] [CrossRef]
- Afzal, M.; Alghamdi, S.S.; Habib Ur Rahman, M.; Ahmad, A.; Farooq, T.; Alam, M.; Khan, I.A.; Ullah, H.; Nasim, W.; Fahad, S. Current status and future possibilities of molecular genetics techniques in Brassica napus. Biotechnol. Lett. 2018, 40, 479–492. [Google Scholar] [CrossRef]
- Witzel, K.; Neugart, S.; Ruppel, S.; Schreiner, M.; Wiesner, M.; Baldermann, S. Recent progress in the use of ‘omics technologies in brassicaceous vegetables. Front. Plant Sci. 2015, 6, 244. [Google Scholar] [CrossRef]
- Tan, C.; Liu, H.; Ren, J.; Ye, X.; Feng, H.; Liu, Z. Single-molecule real-time sequencing facilitates the analysis of transcripts and splice isoforms of anthers in Chinese cabbage (Brassica rapa L. ssp. pekinensis). BMC Plant Biol. 2019, 19, 517. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Cai, X.; Wu, J.; Liu, M.; Grob, S.; Cheng, F.; Liang, J.; Cai, C.; Liu, Z.; Liu, B.; et al. Improved Brassica rapa reference genome by single-molecule sequencing and chromosome conformation capture technologies. Hortic. Res. 2018, 5, 50. [Google Scholar] [CrossRef] [PubMed]
- Cai, G.; Yang, Q.; Chen, H.; Yang, Q.; Zhang, C.; Fan, C.; Zhou, Y. Genetic dissection of plant architecture and yield-related traits in Brassica napus. Sci. Rep. 2016, 6, 21625. [Google Scholar] [CrossRef]
- Perumal, S.; Koh, C.S.; Jin, L.; Buchwaldt, M.; Higgins, E.E.; Zheng, C.; Sankoff, D.; Robinson, S.J.; Kagale, S.; Navabi, Z.-K.; et al. A high-contiguity Brassica nigra genome localizes active centromeres and defines the ancestral Brassica genome. Nat. Plants 2020, 6, 929–941. [Google Scholar] [CrossRef] [PubMed]
- Rousseau-Gueutin, M.; Belser, C.; Silva, C.D.; Richard, G.; Istace, B.; Cruaud, C.; Falentin, C.; Boideau, F.; Boutte, J.; Delourme, R.; et al. Long-reads assembly of the Brassica napus reference genome, Darmor-bzh. bioRxiv 2020. [Google Scholar] [CrossRef]
- Lee, H.; Chawla, H.S.; Obermeier, C.; Dreyer, F.; Abbadi, A.; Snowdon, R. Chromosome-Scale Assembly of Winter Oilseed Rape Brassica napus. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Song, J.-M.; Guan, Z.; Hu, J.; Guo, C.; Yang, Z.; Wang, S.; Liu, D.; Wang, B.; Lu, S.; Zhou, R.; et al. Eight high-quality genomes reveal pan-genome architecture and ecotype differentiation of Brassica napus. Nat. Plants 2020, 6, 34–45. [Google Scholar] [CrossRef]
- Wang, W.; Guan, R.; Liu, X.; Zhang, H.; Song, B.; Xu, Q.; Fan, G.; Chen, W.; Wu, X.; Liu, X.; et al. Chromosome level comparative analysis of Brassica genomes. Plant Mol. Biol. 2019, 99, 237–249. [Google Scholar] [CrossRef]
- Qiao, J.; Zhang, X.; Chen, B.; Huang, F.; Xu, K.; Huang, Q.; Huang, Y.; Hu, Q.; Wu, X. Comparison of the cytoplastic genomes by resequencing: Insights into the genetic diversity and the phylogeny of the agriculturally important genus Brassica. BMC Genom. 2020, 21, 480. [Google Scholar] [CrossRef]
- Tettelin, H.; Masignani, V.; Cieslewicz, M.J.; Donati, C.; Medini, D.; Ward, N.L.; Angiuoli, S.V.; Crabtree, J.; Jones, A.L.; Durkin, A.S.; et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: Implications for the microbial “pan-genome”. Proc. Natl. Acad. Sci. USA 2005, 102, 13950. [Google Scholar] [CrossRef]
- Tirnaz, S.; Edwards, D.; Batley, J. The importance of plant pan-genomes in breeding. Quant. Genet. Genom. Plant Breed. 2020, 2, 27–32. [Google Scholar]
- Tao, Y.; Zhao, X.; Mace, E.; Henry, R.; Jordan, D. Exploring and Exploiting Pan-genomics for Crop Improvement. Mol. Plant 2019, 12, 156–169. [Google Scholar] [CrossRef] [PubMed]
- Bayer, P.E.; Golicz, A.A.; Scheben, A.; Batley, J.; Edwards, D. Plant pan-genomes are the new reference. Nat. Plants 2020, 6, 914–920. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.; Zhang, N.; Severing, E.I.; Nijveen, H.; Cheng, F.; Visser, R.G.F.; Wang, X.; de Ridder, D.; Bonnema, G. Beyond genomic variation—Comparison and functional annotation of three Brassica rapa genomes: A turnip, a rapid cycling and a Chinese cabbage. BMC Genom. 2014, 15, 250. [Google Scholar] [CrossRef] [PubMed]
- Golicz, A.A.; Batley, J.; Edwards, D. Towards plant pangenomics. Plant Biotechnol. J. 2016, 14, 1099–1105. [Google Scholar] [CrossRef] [PubMed]
- Hurgobin, B.; Golicz, A.A.; Bayer, P.E.; Chan, C.-K.K.; Tirnaz, S.; Dolatabadian, A.; Schiessl, S.V.; Samans, B.; Montenegro, J.D.; Parkin, I.A.P.; et al. Homoeologous exchange is a major cause of gene presence/absence variation in the amphidiploid Brassica napus. Plant Biotechnol. J. 2018, 16, 1265–1274. [Google Scholar] [CrossRef]
- Bayer, P.E.; Golicz, A.A.; Tirnaz, S.; Chan, C.-K.K.; Edwards, D.; Batley, J. Variation in abundance of predicted resistance genes in the Brassica oleracea pangenome. Plant Biotechnol. J. 2019, 17, 789–800. [Google Scholar] [CrossRef]
- Dolatabadian, A.; Bayer, P.E.; Tirnaz, S.; Hurgobin, B.; Edwards, D.; Batley, J. Characterization of disease resistance genes in the Brassica napus pangenome reveals significant structural variation. Plant Biotechnol. J. 2020, 18, 969–982. [Google Scholar] [CrossRef]
- Chittem, K.; Yajima, W.R.; Goswami, R.S.; del Río Mendoza, L.E. Transcriptome analysis of the plant pathogen Sclerotinia sclerotiorum interaction with resistant and susceptible canola (Brassica napus) lines. PLoS ONE 2020, 15, e0229844. [Google Scholar] [CrossRef]
- Galindo-González, L.; Manolii, V.; Hwang, S.-F.; Strelkov, S.E. Response of Brassica napus to Plasmodiophora brassicae Involves Salicylic Acid-Mediated Immunity: An RNA-Seq-Based Study. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Sun, Q.; Lin, L.; Liu, D.; Wu, D.; Fang, Y.; Wu, J.; Wang, Y. CRISPR/Cas9-Mediated Multiplex Genome Editing of the BnWRKY11 and BnWRKY70 Genes in Brassica napus L. Int. J. Mol. Sci. 2018, 19, 2716. [Google Scholar] [CrossRef] [PubMed]
- Shahbandeh, M. USDA Foreign Agricultural Service, US Department of Agriculture, Consumption of Vegetable Oils Worldwide from 2013/14 to 2019/2020, by Oil Type (in Million Metric Tons); US Department of Agriculture: Washington, DC, USA, 2020.
- Przybylski, R. Canola/Rapeseed Oil. In Vegetable Oils in Food Technology; Gunstone, F.D., Ed.; Wiley-Blackwell: Oxford, UK, 2011. [Google Scholar] [CrossRef]
- Aukema, H.; Campbell, L. 9—Oil Nutrition and Utilization. In Canola; Daun, J.K., Eskin, N.A.M., Hickling, D., Eds.; AOCS Press: Urbana, IL, USA, 2011. [Google Scholar] [CrossRef]
- Lin, L.; Allemekinders, H.; Dansby, A.; Campbell, L.; Durance-Tod, S.; Berger, A.; Jones, P.J. Evidence of health benefits of canola oil. Nutr. Rev. 2013, 71, 370–385. [Google Scholar] [CrossRef] [PubMed]
- Scarth, R.; McVetty, P. Designer oil canola –A review of new food-grade brassica oils with focus on high oleic, low linolenic types. In Proceedings of the 10th International GCIRC Congress, Canberra, Australia, 26–29 September 1999; p. 10. [Google Scholar]
- Salisbury, P.A.; Cowling, W.A.; Potter, T.D. Continuing innovation in Australian canola breeding. Crop Pasture Sci. 2016, 67, 266–272. [Google Scholar] [CrossRef]
- Canola Council of Canada. Resilient. 2015. Available online: https://www.canolacouncil.org/media/575447/ccc_2015_resilient.pdf (accessed on 12 July 2020).
- He, M.; Qin, C.-X.; Wang, X.; Ding, N.-Z. Plant Unsaturated Fatty Acids: Biosynthesis and Regulation. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Xu, L.; Zeng, W.; Li, J.; Liu, H.; Yan, G.; Si, P.; Yang, C.; Shi, Y.; He, Q.; Zhou, W. Characteristics of membrane-bound fatty acid desaturase (FAD) genes in Brassica napus L. and their expressions under different cadmium and salinity stresses. Environ. Exp. Bot. 2019, 162, 144–156. [Google Scholar] [CrossRef]
- von Borcke, L. High Oleic Acid Oilseed: Oil Seed Rape FAD2 Mutants with Increased Mono-Unsaturated Fatty Acid Content. Granted Patent AU 2012223022, 2 March 2012. International Publication No. WO 2012/117256, US Publication No. US 2014/0150132. [Google Scholar]
- Xue, Y.; Chen, B.; Wang, R.; Win, A.N.; Li, J.; Chai, Y. Genome-Wide Survey and Characterization of Fatty Acid Desaturase Gene Family in Brassica napus and Its Parental Species. Appl. Biochem. Biotechnol. 2018, 184, 582–598. [Google Scholar] [CrossRef]
- Zhu, Q.; King, G.J.; Liu, X.; Shan, N.; Borpatragohain, P.; Baten, A.; Wang, P.; Luo, S.; Zhou, Q. Identification of SNP loci and candidate genes related to four important fatty acid composition in Brassica napus using genome wide association study. PLoS ONE 2019, 14, e0221578. [Google Scholar] [CrossRef]
- Zhai, Y.; Yu, K.; Cai, S.; Hu, L.; Amoo, O.; Xu, L.; Yang, Y.; Ma, B.; Jiao, Y.; Zhang, C.; et al. Targeted mutagenesis of BnTT8 homologs controls yellow seed coat development for effective oil production in Brassica napus L. Plant Biotechnol. J. 2020, 18, 1153–1168. [Google Scholar] [CrossRef]
- Walsh, T.A.; Bevan, S.A.; Gachotte, D.J.; Larsen, C.M.; Moskal, W.A.; Merlo, P.A.O.; Sidorenko, L.V.; Hampton, R.E.; Stoltz, V.; Pareddy, D.; et al. Canola engineered with a microalgal polyketide synthase-like system produces oil enriched in docosahexaenoic acid. Nat. Biotechnol. 2016, 34, 881–887. [Google Scholar] [CrossRef]
- USDA (US Department of Agriculture). Determination of Nonregulated Status for Nuseed DHA Canola; USDA, Ed.; USDA: Washington, DC, USA, 2018.
- OGTR. Issue of Licence DIR 155 to Nuseed Pty Ltd for the Commercial Release of GM Canola; Department of Health and Aging, Ed.; OGTR: Canberra, Australia, 2018.
- Warner, D.J.; Lewis, K.A. Evaluation of the Risks of Contaminating Low Erucic Acid Rapeseed with High Erucic Rapeseed and Identification of Mitigation Strategies. Agriculture 2019, 9, 190. [Google Scholar] [CrossRef]
- Ogunkunle, O.; Ahmed, N.A. A review of global current scenario of biodiesel adoption and combustion in vehicular diesel engines. Energy Rep. 2019, 5, 1560–1579. [Google Scholar] [CrossRef]
- Liu, J.; Hao, W.; Liu, J.; Fan, S.; Zhao, W.; Deng, L.; Wang, X.; Hu, Z.; Hua, W.; Wang, H. A Novel Chimeric Mitochondrial Gene Confers Cytoplasmic Effects on Seed Oil Content in Polyploid Rapeseed (Brassica napus). Mol. Plant 2019, 12, 582–596. [Google Scholar] [CrossRef] [PubMed]
- Wanasundara, J.P.D.; Tan, S.; Alashi, A.M.; Pudel, F.; Blanchard, C. Chapter 18—Proteins From Canola/Rapeseed: Current Status. In Sustainable Protein Sources; Nadathur, S.R., Wanasundara, J.P.D., Scanlin, L., Eds.; Academic Press: San Diego, CA, USA, 2017. [Google Scholar] [CrossRef]
- Hein, T. Canola Meal in Dairy Rations. 2020. Available online: https://www.allaboutfeed.net/New-Proteins/Articles/2020/7/Canola-meal-in-dairy-rations-618105E/ (accessed on 3 August 2020).
- Skugor, A.; Kjos, N.P.; Sundaram, A.Y.M.; Mydland, L.T.; Ånestad, R.; Tauson, A.-H.; Øverland, M. Effects of long-term feeding of rapeseed meal on skeletal muscle transcriptome, production efficiency and meat quality traits in Norwegian Landrace growing-finishing pigs. PLoS ONE 2019, 14, e0220441. [Google Scholar] [CrossRef] [PubMed]
- Slominski, B.A. Developments in the breeding of low fibre rapeseed/canola. J. Anim. Feed Sci. 1997, 6, 303–318. [Google Scholar] [CrossRef][Green Version]
- Jensen, S.K.; Liu, Y.-G.; Eggum, B.O. The influence of seed size and hull content on the composition and digestibility of rapeseeds in rats. Anim. Feed Sci. Technol. 1995, 54, 9–19. [Google Scholar] [CrossRef]
- OGTR. The Biology of Brassica Napus L. (Canola) and Brassica Juncea (L.) Czern. & Coss. (Indian mustard); Health ADO, Ed.; OGTR: Canberra, Australia, 2017.
- Sønderby, I.E.; Geu-Flores, F.; Halkier, B.A. Biosynthesis of glucosinolates—Gene discovery and beyond. Trends Plant Sci. 2010, 15, 283–290. [Google Scholar] [CrossRef]
- Zang, Y.X.; Kim, H.U.; Kim, J.A.; Lim, M.H.; Jin, M.; Lee, S.C.; Kwon, S.J.; Lee, S.I.; Hong, J.K.; Park, T.H.; et al. Genome-wide identification of glucosinolate synthesis genes in Brassica rapa. FEBS J. 2009, 276, 3559–3574. [Google Scholar] [CrossRef]
- Piślewska-Bednarek, M.; Nakano, R.T.; Hiruma, K.; Pastorczyk, M.; Sanchez-Vallet, A.; Singkaravanit-Ogawa, S.; Ciesiołka, D.; Takano, Y.; Molina, A.; Schulze-Lefert, P.; et al. Glutathione Transferase U13 Functions in Pathogen-Triggered Glucosinolate Metabolism. Plant Physiol. 2018, 176, 538–551. [Google Scholar] [CrossRef]
- Kittipol, V.; He, Z.; Wang, L.; Doheny-Adams, T.; Langer, S.; Bancroft, I. Genetic architecture of glucosinolate variation in Brassica napus. J. Plant Physiol. 2019, 240, 152988. [Google Scholar] [CrossRef]
- Geu-Flores, F.; Nielsen, M.T.; Nafisi, M.; Møldrup, M.E.; Olsen, C.E.; Motawia, M.S.; Halkier, B.A. Glucosinolate engineering identifies a γ-glutamyl peptidase. Nat. Chem. Biol. 2009, 5, 575–577. [Google Scholar] [CrossRef]
- Blažević, I.; Montaut, S.; Burčul, F.; Olsen, C.E.; Burow, M.; Rollin, P.; Agerbirk, N. Glucosinolate structural diversity, identification, chemical synthesis and metabolism in plants. Phytochemistry 2020, 169, 112100. [Google Scholar] [CrossRef]
- Burow, M.; Halkier, B.A. How does a plant orchestrate defense in time and space? Using glucosinolates in Arabidopsis as case study. Curr. Opin. Plant Biol. 2017, 38, 142–147. [Google Scholar] [CrossRef]
- Campbell, L.; Rempel, C.B.; Wanasundara, J.P.D. Canola/Rapeseed Protein: Future Opportunities and Directions-Workshop Proceedings of IRC 2015. Plants 2016, 5, 17. [Google Scholar] [CrossRef]
- Knutt, C. Protein Industries Canada Invests to Improve Canola Protein Content. Available online: https://www.proteinindustriescanada.ca/news/protein-industries-canada-invests-to-improve-canola-protein-content (accessed on 27 July 2020).
- Jayawardhane, K.N.; Singer, S.D.; Ozga, J.A.; Rizvi, S.M.; Weselake, R.J.; Chen, G. Seed-specific down-regulation of Arabidopsis CELLULOSE SYNTHASE 1 or 9 reduces seed cellulose content and differentially affects carbon partitioning. Plant Cell Rep. 2020, 39, 953–969. [Google Scholar] [CrossRef]
- Jiang, J.; Zhu, S.; Yuan, Y.; Wang, Y.; Zeng, L.; Batley, J.; Wang, Y.-P. Transcriptomic comparison between developing seeds of yellow- and black-seeded Brassica napus reveals that genes influence seed quality. BMC Plant Biol. 2019, 19, 203. [Google Scholar] [CrossRef]
- Facciotti, D. Production of Improved Rapeseed Exhibiting Yellow-Seed Coat. US Patent 006547711B2, 15 April 2003. [Google Scholar]
- Chungu, C.; Rakow, G.; Raney, J.P.; Kubik Thomas, J. Low Fiber Yellow Canola Seeds Comprising High Oleic, Low Linolenic Oil. World Intellectual Property Organization WO2007/016521A2, 8 February 2007. [Google Scholar]
- Zhang, X.; Fernando, W.G.D. Insights into fighting against blackleg disease of Brassica napus in Canada. Crop Pasture Sci. 2017, 69, 40–47. [Google Scholar] [CrossRef]
- Diaz, C.; Cevallos, F.; Damicone, J. Characterization of the Race Structure of Leptosphaeria maculans Causing Blackleg of Winter Canola in Oklahoma and Kansas. Plant Dis. 2019, 103, 2353–2358. [Google Scholar] [CrossRef]
- Hwang, S.F.; Strelkov, S.E.; Ahmed, H.U.; Manolii, V.P.; Zhou, Q.; Fu, H.; Turnbull, G.; Fredua-Agyeman, R.; Feindel, D. Virulence and inoculum density-dependent interactions between clubroot resistant canola (Brassica napus) and Plasmodiophora brassicae. Plant Pathol. 2017, 66, 1318–1328. [Google Scholar] [CrossRef]
- Rempel, C.B.; Hutton, S.N.; Jurke, C.J. Clubroot and the importance of canola in Canada. Can. J. Plant Pathol. 2014, 36, 19–26. [Google Scholar] [CrossRef]
- Denton-Giles, M.; Derbyshire, M.C.; Khentry, Y.; Buchwaldt, L.; Kamphuis, L.G. Partial stem resistance in Brassica napus to highly aggressive and genetically diverse Sclerotinia sclerotiorum isolates from Australia. Can. J. Plant Pathol. 2018, 40, 551–561. [Google Scholar] [CrossRef]
- Aldrich-Wolfe, L.; Travers, S.; Nelson, B.D., Jr. Genetic Variation of Sclerotinia sclerotiorum from Multiple Crops in the North Central United States. PLoS ONE 2015, 10, e0139188. [Google Scholar] [CrossRef]
- Fu, F.; Zhang, X.; Liu, F.; Peng, G.; Yu, F.; Fernando, D. Identification of resistance loci in Chinese and Canadian canola/rapeseed varieties against Leptosphaeria maculans based on genome-wide association studies. BMC Genom. 2020, 21, 501. [Google Scholar] [CrossRef]
- Raman, R.; Diffey, S.; Barbulescu, D.M.; Coombes, N.; Luckett, D.; Salisbury, P.; Cowley, R.; Marcroft, S.; Raman, H. Genetic and physical mapping of loci for resistance to blackleg disease in canola (Brassica napus L.). Sci. Rep. 2020, 10, 4416. [Google Scholar] [CrossRef]
- Rong, S.; Feng, J.; Li, Q.; Fei, W.; Ahmed, H.U.; Liang, Y.; Hwang, S.-F.; Strelkov, S.E. Pathogenic variability and prevalence of Avr genes in Leptosphaeria maculans populations from Alberta, Canada. J. Plant Dis. Prot. 2015, 122, 161–168. [Google Scholar] [CrossRef]
- Zhang, X.; Peng, G.; Kutcher, H.R.; Balesdent, M.-H.; Delourme, R.; Fernando, W.G.D. Breakdown of Rlm3 resistance in the Brassica napus–Leptosphaeria maculans pathosystem in western Canada. Eur. J. Plant Pathol. 2016, 145, 659–674. [Google Scholar] [CrossRef]
- Van De Wouw, A.P.; Marcroft, S.J.; Howlett, B.J. Blackleg disease of canola in Australia. Crop Pasture Sci. 2016, 67, 273–283. [Google Scholar] [CrossRef]
- Raman, H.; Raman, R.; Diffey, S.; Qiu, Y.; McVittie, B.; Barbulescu, D.M.; Salisbury, P.A.; Marcroft, S.; Delourme, R. Stable Quantitative Resistance Loci to Blackleg Disease in Canola (Brassica napus L.) Over Continents. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Zhou, T.; Xu, W.; Hirani, A.H.; Liu, Z.; Tuan, P.A.; Ayele, B.T.; Daayf, F.; McVetty, P.B.E.; Duncan, R.W.; Li, G. Transcriptional Insight Into Brassica napus Resistance Genes LepR3 and Rlm2-Mediated Defense Response Against the Leptosphaeria maculans Infection. Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Zou, Z.; Liu, F.; Selin, C.; Fernando, W.G.D. Generation and Characterization of a Virulent Leptosphaeria maculans Isolate Carrying a Mutated AvrLm7 Gene Using the CRISPR/Cas9 System. Front. Microbiol. 2020, 11, 1969. [Google Scholar] [CrossRef]
- Wallenhammar, A.-C. Prevalence of Plasmodiophora brassicae in a spring oilseed rape growing area in central Sweden and factors influencing soil infestation levels. Plant Pathol. 1996, 45, 710–719. [Google Scholar] [CrossRef]
- Hirani, A.H.; Gao, F.; Liu, J.; Fu, G.; Wu, C.; McVetty, P.B.E.; Duncan, R.W.; Li, G. Combinations of Independent Dominant Loci Conferring Clubroot Resistance in All Four Turnip Accessions (Brassica rapa) From the European Clubroot Differential Set. Front. Plant Sci. 2018, 9. [Google Scholar] [CrossRef]
- Manzanares-Dauleux, M.J.; Delourme, R.; Baron, F.; Thomas, G. Mapping of one major gene and of QTLs involved in resistance to clubroot in Brassica napus. Theor. Appl. Genet. 2000, 101, 885–891. [Google Scholar] [CrossRef]
- Zhou, L.; Zhang, L.; He, Y.; Liu, F.; Li, M.; Wang, Z.; Ji, G. Isolation and characterization of bacterial isolates for biological control of clubroot on Chinese cabbage. Eur. J. Plant Pathol. 2014, 140, 159–168. [Google Scholar] [CrossRef][Green Version]
- Sedaghatkish, A.; Gossen, B.D.; Yu, F.; Torkamaneh, D.; McDonald, M.R. Whole-genome DNA similarity and population structure of Plasmodiophora brassicae strains from Canada. BMC Genom. 2019, 20, 744. [Google Scholar] [CrossRef]
- Dixon, G. The Occurrence and Economic Impact of Plasmodiophora brassicae and Clubroot Disease. J. Plant Growth Regul. 2009, 28, 194–202. [Google Scholar] [CrossRef]
- Hatakeyama, K.; Suwabe, K.; Tomita, R.N.; Kato, T.; Nunome, T.; Fukuoka, H.; Matsumoto, S. Identification and Characterization of Crr1a, a Gene for Resistance to Clubroot Disease (Plasmodiophora brassicae Woronin) in Brassica rapa L. PLoS ONE 2013, 8, e54745. [Google Scholar] [CrossRef]
- Ueno, H.; Matsumoto, E.; Aruga, D.; Kitagawa, S.; Matsumura, H.; Hayashida, N. Molecular characterization of the CRa gene conferring clubroot resistance in Brassica rapa. Plant Mol. Biol. 2012, 80, 621–629. [Google Scholar] [CrossRef]
- Chang, A.; Lamara, M.; Wei, Y.; Hu, H.; Parkin, I.A.P.; Gossen, B.D.; Peng, G.; Yu, F. Clubroot resistance gene Rcr6 in Brassica nigra resides in a genomic region homologous to chromosome A08 in B. rapa. BMC Plant Biol. 2019, 19, 224. [Google Scholar] [CrossRef]
- Nguyen, M.L.; Monakhos, G.F.; Komakhin, R.A.; Monakhos, S.G. The New Clubroot Resistance Locus Is Located on Chromosome A05 in Chinese Cabbage (Brassica rapa L.). Russ. J. Genet. 2018, 54, 296–304. [Google Scholar] [CrossRef]
- Hejna, O.; Havlickova, L.; He, Z.; Bancroft, I.; Curn, V. Analysing the genetic architecture of clubroot resistance variation in Brassica napus by associative transcriptomics. Mol. Breed. 2019, 39, 112. [Google Scholar] [CrossRef]
- Zhou, Q.; Galindo-González, L.; Hwang, S.-F.; Strelkov, S.E. Application of genomics and transcriptomics to accelerate development of clubroot resistant canola. Can. J. Plant Pathol. 2020. [Google Scholar] [CrossRef]
- Strelkov, S.E.; Hwang, S.-F. Clubroot in the Canadian canola crop: 10 years into the outbreak. Can. J. Plant Pathol. 2014, 36, 27–36. [Google Scholar] [CrossRef]
- Boland, G.J.; Hall, R. Index of plant hosts of Sclerotinia sclerotiorum. Can. J. Plant Pathol. 1994, 16, 93–108. [Google Scholar] [CrossRef]
- Guyon, K.; Balagué, C.; Roby, D.; Raffaele, S. Secretome analysis reveals effector candidates associated with broad host range necrotrophy in the fungal plant pathogen Sclerotinia sclerotiorum. BMC Genom. 2014, 15, 336. [Google Scholar] [CrossRef]
- Liu, S.; Wang, H.; Zhang, J.; Fitt, B.D.L.; Xu, Z.; Evans, N.; Liu, Y.; Yang, W.; Guo, X. In vitro mutation and selection of doubled-haploid Brassica napus lines with improved resistance to Sclerotinia sclerotiorum. Plant Cell Rep. 2005, 24, 133–144. [Google Scholar] [CrossRef]
- Mullins, E.; Quinlan, C.; Jones, P. Isolation of Mutants Exhibiting Altered Resistance to Sclerotinia sclerotiorum from Small M2 Populations of an Oilseed Rape (Brassica napus) Variety. Eur. J. Plant Pathol. 1999, 105, 465–475. [Google Scholar] [CrossRef]
- Auclair, J.; Boland, G.J.; Cober, E.; Graef, G.L.; Steadman, J.R.; Zilka, J.; Rajcan, I. Development of a new field inoculation technique toassess partial resistance in soybean to Sclerotinia sclerotiorum. Can. J. Plant Sci. 2004, 84, 57–64. [Google Scholar] [CrossRef]
- You, M.P.; Uloth, M.B.; Li, X.X.; Banga, S.S.; Banga, S.K.; Barbetti, M.J. Valuable New Resistances Ensure Improved Management of Sclerotinia Stem Rot (Sclerotinia sclerotiorum) in Horticultural and Oilseed Brassica Species. J. Phytopathol. 2016, 164, 291–299. [Google Scholar] [CrossRef]
- Wang, Z.; Wan, L.; Xin, Q.; Chen, Y.; Zhang, X.; Dong, F.; Hong, D.; Yang, G. Overexpression of OsPGIP2 confers Sclerotinia sclerotiorum resistance in Brassica napus through increased activation of defense mechanisms. J. Exp. Bot. 2018, 69, 3141–3155. [Google Scholar] [CrossRef]
- Falak, I.; Charne, D.R.; Patel, J.D.; Tulsieram, L. Sclerotinia-Resistant Brassica. US Patent US008558067B2, 15 October 2013. [Google Scholar]
- Li, J.; Zhao, Z.; Hayward, A.; Cheng, H.; Fu, D. Integration analysis of quantitative trait loci for resistance to Sclerotinia sclerotiorum in Brassica napus. Euphytica 2015, 205, 483–489. [Google Scholar] [CrossRef]
- Lohani, N.; Jain, D.; Singh, M.B.; Bhalla, P.L. Engineering Multiple Abiotic Stress Tolerance in Canola, Brassica napus. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Khanzada, H.; Wassan, G.M.; He, H.; Mason, A.S.; Keerio, A.A.; Khanzada, S.; Faheem, M.; Solangi, A.M.; Zhou, Q.; Fu, D.; et al. Differentially evolved drought stress indices determine the genetic variation of Brassica napus at seedling traits by genome-wide association mapping. J. Adv. Res. 2020, 24, 447–461. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.N.; Zhang, J.; Luo, T.; Liu, J.; Ni, F.; Rizwan, M.; Fahad, S.; Hu, L. Morpho-physiological and biochemical responses of tolerant and sensitive rapeseed cultivars to drought stress during early seedling growth stage. Acta Physiol. Plant. 2019, 41, 25. [Google Scholar] [CrossRef]
- Wani, S.H.; Kumar, V.; Shriram, V.; Sah, S.K. Phytohormones and their metabolic engineering for abiotic stress tolerance in crop plants. Crop J. 2016, 4, 162–176. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, Y.; Liu, Y.; Zhou, W.; Yan, B.; Yang, J.; Shen, Y. Overexpression of BcHsfA1 transcription factor from Brassica campestris improved heat tolerance of transgenic tobacco. PLoS ONE 2018, 13, e0207277. [Google Scholar] [CrossRef]
- Yang, H.; Wu, J.-J.; Tang, T.; Liu, K.-D.; Dai, C. CRISPR/Cas9-mediated genome editing efficiently creates specific mutations at multiple loci using one sgRNA in Brassica napus. Sci. Rep. 2017, 7, 7489. [Google Scholar] [CrossRef]
- Wu, J.; Yan, G.; Duan, Z.; Wang, Z.; Kang, C.; Guo, L.; Liu, K.; Tu, J.; Shen, J.; Yi, B.; et al. Roles of the Brassica napus DELLA Protein BnaA6.RGA, in modulating drought tolerance by interacting with the ABA signaling component BnaA10.ABF2. Front. Plant Sci. 2020, 11. [Google Scholar] [CrossRef]
- Walsh, M.; Newman, P.; Powles, S. Targeting weed seeds in-crop: A New Weed Control Paradigm for Global Agriculture. Weed Technol. 2013, 27, 431–436.178. [Google Scholar] [CrossRef]
- Manitoba Agriculture and Resource Development Office. Guidelines: Crop Production Costs; Manitoba Agriculture and Resource Development, Ed.; Manitoba Agriculture and Resource Development Office: Winnipeg, MB, Canada, 2020.
- McLeod, R. Annual Costs of Weeds in Australia; eSYS Development Pty Limited: Canberra, Australia, 2018. [Google Scholar]
- Beckie, H.J.; Flower, K.C.; Ashworth, M.B. Farming without Glyphosate? Plants 2020, 9. [Google Scholar] [CrossRef]
- Duke, S.O.; Powles, S.B. Glyphosate: A once-in-a-century herbicide. Pest Manag. Sci. 2008, 64, 319–325. [Google Scholar] [CrossRef]
- Asaduzzaman, M.; Pratley, J.E.; Luckett, D.; Lemerle, D.; Wu, H. Weed management in canola (Brassica napus L): A review of current constraints and future strategies for Australia. Arch. Agron. Soil Sci. 2020, 66, 427–444. [Google Scholar] [CrossRef]
- Brookes, G.; Barfoot, P. Environmental impacts of genetically modified (GM) crop use 1996–2018: Impacts on pesticide use and carbon emissions. GM Crops Food 2020, 11, 215–241. [Google Scholar] [CrossRef] [PubMed]
- Green, J.M. Current state of herbicides in herbicide-resistant crops. Pest Manag. Sci. 2014, 70, 1351–1357. [Google Scholar] [CrossRef] [PubMed]
- Wallace, D.B.; Laima, S.K. Development of Triazine Resistance in Crops by Classical Plant Breeding. Weed Sci. 1987, 35, 9–11. [Google Scholar]
- Funke, T.; Han, H.; Healy-Fried, M.L.; Fischer, M.; Schönbrunn, E. Molecular basis for the herbicide resistance of Roundup Ready crops. Proc. Natl. Acad. Sci. USA 2006, 103, 13010–13015. [Google Scholar] [CrossRef]
- ISAAA’s GM Approval Database. 2020. Available online: http://www.isaaa.org/gmapprovaldatabase/ (accessed on 24 July 2020).
- ACIL Tasman Pty Ltd. GM Canola: An Information Package; Department of Agriculture, Fisheries and Forestry Australia: Canberra, Australia, 2007. Available online: https://www.agriculture.gov.au/sites/default/files/sitecollectiondocuments/ag-food/biotech/gm-canola-info-package.pdf (accessed on 15 August 2020).
- Kirkegaard, J.A.; Lilley, J.M.; Morrison, M.J. Drivers of trends in Australian canola productivity and future prospects. Crop Pasture Sci. 2016, 67, i–ix. [Google Scholar] [CrossRef]
- Benbrook, C.M. Trends in glyphosate herbicide use in the United States and globally. Environ. Sci. Eur. 2016, 28, 3. [Google Scholar] [CrossRef]
- Fernandez-Cornejo, J.; Wechsler, S.; Milkove, D. The Adoption of Genetically Engineered Alfalfa, Canola, and Sugarbeets in the United States. Agriculture, U. S. Department of Agriculture, Ed.; 2016. Available online: https://www.ers.usda.gov/webdocs/publications/81176/eib-163_summary.pdf?v=0 (accessed on 17 July 2020).
- Aghazadeh, R.; Zamani, M.; Motallebi, M.; Moradyar, M. Agrobacterium -Mediated Transformation of the Oryza sativa Thaumatin-Like Protein to Canola (R Line Hyola308) for Enhancing Resistance to Sclerotinia sclerotiorum. Iran. J. Biotechnol. 2017, 15, 201–207. [Google Scholar] [CrossRef]
- Sutherland, S. Canola Weed Management. 1999. Available online: http://www.australianoilseeds.com/__data/assets/pdf_file/0012/2712/Chapter_12__Canola_Weed_Management.pdf (accessed on 17 July 2020).
- Heap, I.; Duke, S.O. Overview of glyphosate-resistant weeds worldwide. Pest Manag. Sci. 2018, 74, 1040–1049. [Google Scholar] [CrossRef]
- Antier, C.K.P.; Reboud, X.; Ulber, L.; Baret, P.V.; Messéan, A. Glyphosate Use in the European Agricultural Sector and a Framework for Its Further Monitoring. Sustainability 2020, 12, 5682. [Google Scholar] [CrossRef]
- Department of Primary Industries and Regional Development (DPIRD). Canola Variety Sowing Guide for Western Australia; DPIRD: South Perth, WA, Australia, 2019. Available online: https://www.agric.wa.gov.au/sites/gateway/files/DPIRD%20canola%20variety%20sowing%20guide%202019%20Bulletin%204897.pdf (accessed on 17 July 2020).
- Idnurm, A.; Urquhart, A.S.; Vummadi, D.R.; Chang, S.; Van de Wouw, A.P.; López-Ruiz, F.J. Spontaneous and CRISPR/Cas9-induced mutation of the osmosensor histidine kinase of the canola pathogen Leptosphaeria maculans. Fungal Biol. Biotechnol. 2017, 4, 12. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.; Zhang, X.; Peng, G.; Falk, K.C.; Strelkov, S.E.; Gossen, B.D. Genotyping-by-sequencing reveals three QTL for clubroot resistance to six pathotypes of Plasmodiophora brassicae in Brassica rapa. Sci. Rep. 2017, 7, 4516. [Google Scholar] [CrossRef] [PubMed]
- Fu, F.; Liu, X.; Wang, R.; Zhai, C.; Peng, G.; Yu, F.; Fernando, W.G.D. Fine mapping of Brassica napus blackleg resistance gene Rlm1 through bulked segregant RNA sequencing. Sci. Rep. 2019, 9, 14600. [Google Scholar] [CrossRef] [PubMed]
- Hopkins, R.J.; Dam, N.M.V.; Loon, J.J.A.V. Role of Glucosinolates in Insect-Plant Relationships and Multitrophic Interactions. Annu. Rev. Entomol. 2009, 54, 57–83. [Google Scholar] [CrossRef]
- Bell, L. The Biosynthesis of Glucosinolates: Insights, Inconsistencies, and Unknowns. In Annual Plant Reviews Online; Roberts, J.A., Ed.; 2019. [Google Scholar] [CrossRef]
- Lin, P.; Qin, S.; Pu, Q.; Wang, Z.; Wu, Q.; Gao, P.; Schettler, J.; Guo, K.; Li, R.; Li, G.; et al. CRISPR-Cas13 Inhibitors Block RNA Editing in Bacteria and Mammalian Cells. Mol. Cell 2020, 78, 850–861.e855. [Google Scholar] [CrossRef]
- Mackay, T.F. The Genetic Architecture of Quantitative Traits. Annu. Rev. Genet. 2001, 35, 303–339. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ton, L.B.; Neik, T.X.; Batley, J. The Use of Genetic and Gene Technologies in Shaping Modern Rapeseed Cultivars (Brassica napus L.). Genes 2020, 11, 1161. https://doi.org/10.3390/genes11101161
Ton LB, Neik TX, Batley J. The Use of Genetic and Gene Technologies in Shaping Modern Rapeseed Cultivars (Brassica napus L.). Genes. 2020; 11(10):1161. https://doi.org/10.3390/genes11101161
Chicago/Turabian StyleTon, Linh Bao, Ting Xiang Neik, and Jacqueline Batley. 2020. "The Use of Genetic and Gene Technologies in Shaping Modern Rapeseed Cultivars (Brassica napus L.)" Genes 11, no. 10: 1161. https://doi.org/10.3390/genes11101161
APA StyleTon, L. B., Neik, T. X., & Batley, J. (2020). The Use of Genetic and Gene Technologies in Shaping Modern Rapeseed Cultivars (Brassica napus L.). Genes, 11(10), 1161. https://doi.org/10.3390/genes11101161





