Overcoming Fish Defences: The Virulence Factors of Yersinia ruckeri
Abstract
1. Introduction
2. Description of Y. ruckeri, a Member of the Yersiniaceae Family
3. Y. ruckeri as the Causative Agent of Enteric Redmouth Disease
4. Y. ruckeri Has a Broad Host Range and a Broad Geographic Distribution
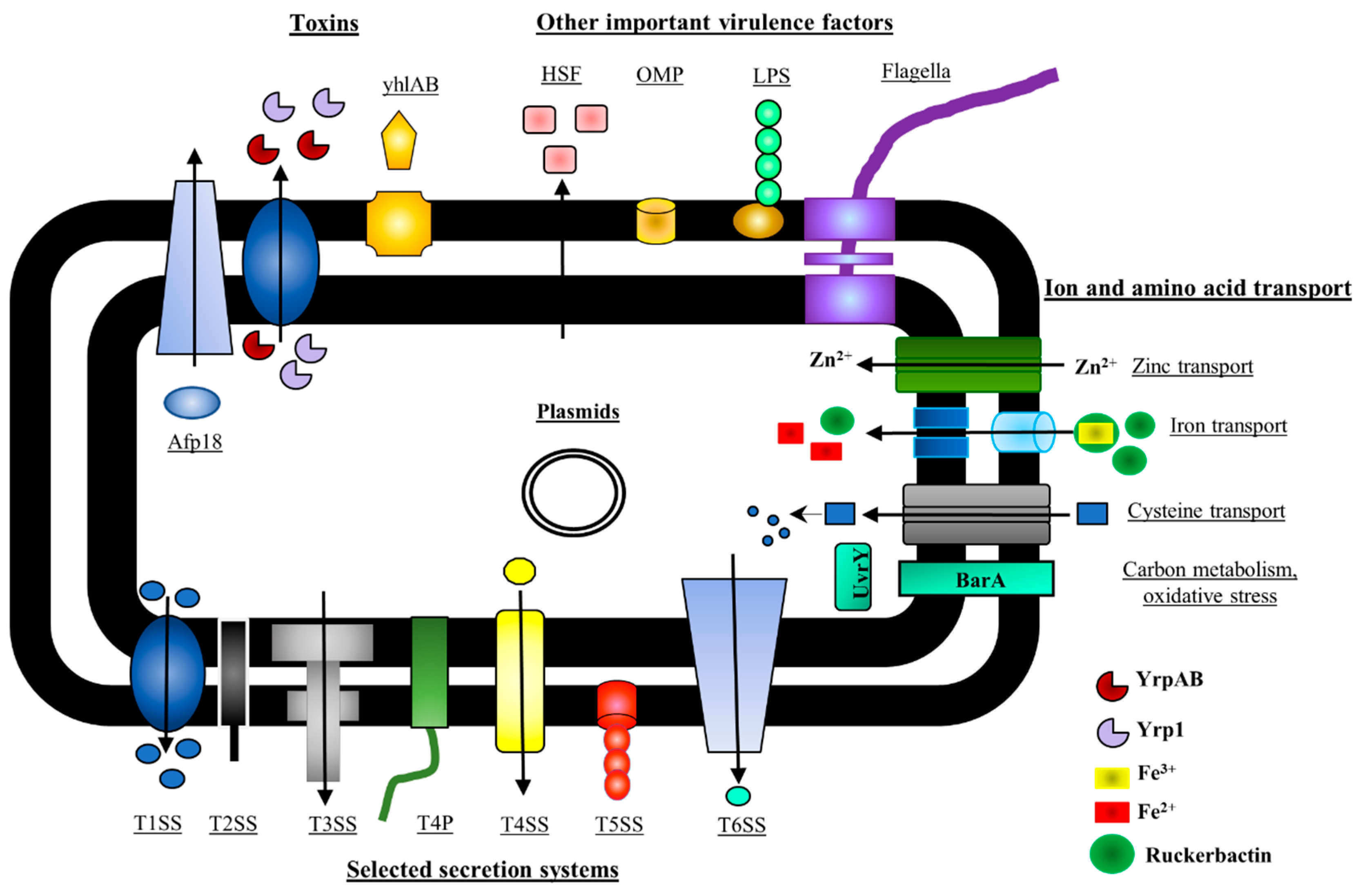
5. Y. ruckeri Pathogenicity Factors
5.1. Y. ruckeri Toxins
5.1.1. Y. ruckeri Protease 1 (Yrp1)
5.1.2. Y. ruckeri Peptidases (YrpAB)
5.1.3. Y. ruckeri Pore-Forming Toxin, yhlBA
5.1.4. Y. ruckeri Phospholipase
5.1.5. Antifeeding Prophage 18 (Afp18)
5.2. Y. ruckeri Secretion Systems
5.2.1. Y. ruckeri Type III Secretion System (T3SS)
5.2.2. Y. ruckeri Type IV Secretion Systems (T4SSs)
5.2.3. Y. ruckeri Type V Secretion Systems (T5SSs)
5.2.4. Type IV Pili
5.3. Operons Involved in Ion and Amino Acids Transport
5.3.1. Iron Acquisition System
5.3.2. The cdsAB Operon
5.3.3. znuABC and BarA-UvrY
5.4. Other Virulence Factors
5.4.1. Genes Upregulated at 18 °C
5.4.2. Heat-Sensitive Factor (HSF)
5.4.3. LPS
5.4.4. Flagella
5.4.5. Plasmids
5.4.6. TcpA
5.4.7. Outer Membrane Proteins (OMPs)
5.4.8. Biofilm as a Virulence Factor
6. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Stentiford, G.D.; Sritunyalucksana, K.; Flegel, T.W.; Williams, B.A.; Withyachumnarnkul, B.; Itsathitphaisarn, O.; Bass, D. New Paradigms to Help Solve the Global Aquaculture Disease Crisis. PLoS Pathog. 2017, 13, e1006160. [Google Scholar] [CrossRef] [PubMed]
- Walker, P.; Winton, J. Emerging viral diseases of fish and shrimp. Vet. Res. 2010, 41, 51. [Google Scholar] [CrossRef] [PubMed]
- Hjeltnes, B.; Bornø, G.; Jansen, M.; Haukaas, A.; Walde, C. The Health Situation in Norwegian Aquaculture 2016; Norwegian Veterinary Institute: Oslo, Norway, 2017. [Google Scholar]
- Austin, D.; Robertson, P.; Austin, B. Recovery of a New Biogroup of Yersinia ruckeri from Diseased Rainbow Trout (Oncorhynchus mykiss, Walbaum). Syst. Appl. Microbiol. 2003, 26, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Fouz, B.; Zarza, C.; Amaro, C. First description of non-motile Yersinia ruckeri serovar I strains causing disease in rainbow trout, Oncorhynchus mykiss (Walbaum), cultured in Spain. J. Fish Dis. 2006, 29, 339–346. [Google Scholar] [CrossRef] [PubMed]
- Wheeler, R.; Davies, R.L.; Dalsgaard, I.; Garcia, J.; Welch, T.J.; Wagley, S.; Bateman, K.S.; Verner-Jeffreys, D.W. Yersinia ruckeri biotype 2 isolates from mainland Europe and the UK likely represent different clonal groups. Dis. Aquat. Organ. 2009, 84, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Arias, C.; Olivares-Fuster, O.; Hayden, K.; Shoemaker, C.; Grizzle, J.; Klesius, P. First Report of Yersinia ruckeri Biotype 2 in the USA. J. Aquat. Anim. Health 2007, 19, 35–40. [Google Scholar] [CrossRef]
- Calvez, S.; Gantelet, H.; Blanc, G.; Douet, D.; Daniel, P. Yersinia ruckeri Biotypes 1 and 2 in France: Presence and antibiotic susceptibility. Dis. Aquat. Organ. 2014, 109, 117–126. [Google Scholar] [CrossRef]
- Bullock, G.; Stuckey, H.; Shotts, E., Jr. Enteric redmouth bacterium: Comparison of isolates from different geographic areas. J. Fish Dis. 1978, 1, 351–356. [Google Scholar] [CrossRef]
- Romalde, J.; MagariÑos, B.; Barja, J.; Toranzo, A. Antigenic and Molecular Characterization of Yersinia ruckeri Proposal for a New Intraspecies Classification. Syst. Appl. Microbiol. 1993, 16, 411–419. [Google Scholar] [CrossRef]
- O’Leary, P. Enteric Redmouth Bacterium of Salmonids: A Biochemical and Serological Comparison of Selected Isolates. Master’s Thesis, Oregon State University, Corvallis, OR, USA, 1977; p. 93. [Google Scholar]
- Stevenson, R.; Airdrie, R. Serological variation among Yersinia ruckeri strains. J. Fish Dis. 1984, 7, 247–254. [Google Scholar] [CrossRef]
- DeGrandis, S.; Krell, P.; Flett, D.; Stevenson, R. Deoxyribonucleic Acid Relatedness of Serovars of Yersinia ruckeri, the Enteric Redmouth Bacterium. J. Syst. Bacteriol. 1988, 38, 49–55. [Google Scholar] [CrossRef]
- Davies, R. O-Serotyping of Yersinia ruckeri with Special Emphasis on European Isolates. Vet. Microbiol. 1990, 22, 299–307. [Google Scholar] [CrossRef]
- Ormsby, M.; Caws, T.; Burchmore, R.; Wallis, T.; Verner-Jeffreys, D.; Davies, R. Yersinia ruckeri Isolates Recovered from Diseased Atlantic Salmon (Salmo salar) in Scotland Are More Diverse than Those from Rainbow Trout (Oncorhynchus mykiss) and Represent Distinct Subpopulations. Appl. Environ. Microbiol. 2016, 82, 5785–5794. [Google Scholar] [CrossRef] [PubMed]
- Adeolu, M.; Alnajar, S.; Naushad, S.; Gupta, R. Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: Proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Morganellaceae fam. nov., and Budviciaceae fam. nov. Int. J. Syst. Evol. Microbiol. 2016, 66, 5575–5599. [Google Scholar] [PubMed]
- Ross, A.; Rucker, R.; Ewing, W. Description of a bacterium associated with redmouth disease of rainbow trout (Salmo gairdneri). Can. J. Microbiol. 1966, 12, 763–770. [Google Scholar] [CrossRef] [PubMed]
- Austin, B.; Green, M.; Rodgers, C. Morphological diversity among strains of Yersinia ruckeri. Aquaculture 1982, 27, 73–78. [Google Scholar] [CrossRef]
- Tobback, E.; Decostere, A.; Hermans, K.; Haesebrouck, F.; Chiers, K. Yersinia ruckeri infections in salmonid fish. J. Fish 2007, 30, 257–268. [Google Scholar] [CrossRef] [PubMed]
- Moreau, E.; Thomas, T.; Brevet, M.; Thorin, C.; Fournel, C.; Calvez, S. Mutations involved in the emergance of Yersinia ruckeri biotype 2 in France. Transbound. Emerg. Dis. 2019, 66, 1387–1394. [Google Scholar] [CrossRef]
- Stevenson, R.M.W.; Daly, J.G. Biochemical and Serological Characteristics of Ontario Isolates of Yersinia ruckeri. Can. J. Fish. Aquat. Sci. 1982, 39, 870–876. [Google Scholar] [CrossRef]
- Navais, R.; Méndez, J.; Cascales, D.; Reimundo, P.; Guijarro, J. The heat sensitive factor (HSF) of Yersinia ruckeri is produced by an alkyl sulphatase involved in sodium dodecyl sulphate (SDS) degradation but not in virulence. BMC Microbiol. 2014, 14, 221. [Google Scholar] [CrossRef]
- Gibello, A.; Porrero, M.C.; Blanco, M.M.; Vela, A.I.; Liébana, P.; Moreno, M.A.; Fernandez-Garayzabal, J.F.; Domínguez, L. Analysis of the gyrA Gene of Clinical Yersinia ruckeri Isolates with Reduced Susceptibility to Quinolones. Appl. Environ. Microbiol. 2004, 70, 599–602. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gibello, A.; Blanco, M.M.; Moreno, M.A.; Cutuli, M.T.; Domenech, A.; Domínguez, L.; Fernández-Garayzábal, J.F. Development of a PCR Assay for Detection of Yersinia ruckeri in Tissues of Inoculated and Naturally Infected Trout. Appl. Environ. Microbiol. 1999, 65, 346–350. [Google Scholar] [PubMed]
- Akhlaghi, M.; Yazdi, H.S. Detection and identification of virulent Yersinia ruckeri: The causative agent of enteric redmouth disease in rainbow trout (Oncorhynchus mykiss) cultured in Fars province, Iran. Iran. J. Vet. Res. 2008, 9, 347–352. [Google Scholar]
- Busch, R. Enteric redmouth disease. Mar. Fish. Rev. 1978, 40, 42–51. [Google Scholar]
- Cossarini-Dunier, M. Indirect enzyme-linked immunosorbent assay (ELISA) to titrate rainbow trout serum antibodies against two pathogens: Yersinia ruckeri and Egtved virus. Aquaculture 1985, 49, 197–208. [Google Scholar] [CrossRef]
- Ghosh, B.; Crosbie, P.; Nowak, B.; Bridle, A. A highly sensitive, non-invasive qPCR-based strategy for direct quantification of Yersinia ruckeri in fish faeces. J. Fish Dis. 2018, 41, 1421–1428. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.; Goldring, O.; Dear, G. The production and methods of use of polyclonal antisera to the pathogenic organisms Aeromonas salmonicida, Yersinia ruckeri and Renibacterium salmoninarum. J. Fish Biol. 1987, 31, 225–226. [Google Scholar] [CrossRef]
- Glenn, R.; Taylor, P.; Hanson, K. The use of a real-time PCR primer/probe set to observe infectivity of Yersinia ruckeri in Chinook salmon, Oncorhynchus tshawytscha (Walbaum, and steelhead trout, Oncorhynchus mykiss(Walbaum). J. Fish Dis. 2011, 34, 783–791. [Google Scholar] [CrossRef]
- Keeling, S.; Johnston, C.; Wallis, R.; Brosnahan, C.; Gudkovs, N.; Mcdonald, W. Development and validation of real-time PCR for the detection of Yersinia ruckeri. J. Fish Dis. 2012, 35, 119–125. [Google Scholar] [CrossRef]
- Saleh, M.; Soliman, H.; El-Matbouli, M. Loop-mediated isothermal amplification as an emerging technology for detection of Yersinia ruckeri the causative agent of enteric red mouth disease in fish. BMC Vet. Res. 2008, 4, 1–10. [Google Scholar] [CrossRef]
- Temprano, A.; Yugueros, J.; Hernanz, C.; Sánchez, M.; Berzal, B.; Luengo, J.M.; Naharro, G. Rapid identification of Yersinia ruckeri by PCR amplification of yruI-yruR quorum sensing. J. Fish Dis. 2001, 24, 253–261. [Google Scholar] [CrossRef]
- Altinok, I.; Grizzle, J.; Liu, Z. Detection of Yersinia ruckeri in rainbow trout blood by use of the polymerase chain reaction. Dis. Aquat. Organ. 2001, 44, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Noga, E.; Levine, J.; Townsend, K.; Bullis, R.; Carlson, C.; Corbett, W. Kidney biopsy: A nonlethal method for diagnosing Yersinia ruckeri infection (enteric redmouth disease) in rainbouw trout (Salmo gairdneri). Am. J. Vet. Res. 1988, 49, 363–365. [Google Scholar] [PubMed]
- Bastardo, A.; Ravelo, C.; Romalde, J. Multilocus sequence typing reveals high genetic diversity and epidemic population structure for the fish pathogen Yersinia ruckeri. Environ. Microbiol. 2012, 14, 1888–1897. [Google Scholar] [CrossRef] [PubMed]
- Gulla, S.; Mohammad, S.; Colquhoun, D. Multi-locus variable-number tandem-repeat analysis of the fish-pathogenic bacterium Yersinia ruckeri by multiplex PCR and capillary electrophoresis. J. Vis. Exp. 2019, 148, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Ewing, W.; Ross, A.; Brenner, D.; Fanning, G. Yersinia ruckeri sp. nov., the Redmouth (RM) Bacterium. Int. J. Syst. Bcteriol. 1978, 28, 37–44. [Google Scholar] [CrossRef]
- Kumar, G.; Menanteau-Ledouble, S.; Saleh, M.; El-Matbouli, M. Yersinia ruckeri, the causative agent of enteric redmouth disease in fish. Vet. Res. 2015, 46, 1–10. [Google Scholar] [CrossRef]
- Rucker, R. Redmouth disease of rainbow trout (Salmo gairdneri). Bull. Off. Int. Epizoot. 1966, 65, 825–830. [Google Scholar]
- Fuhrmann, H.; Böhm, K.; Schlotfeldt, H. An outbreak of enteric redmouth disease in West Germany. J. Fish Dis. 1983, 6, 309–311. [Google Scholar] [CrossRef]
- Bullock, G. Enteric Redmouth Disease. Fish Heal. Bull. 1989. Available online: https://apps.dtic.mil/dtic/tr/fulltext/u2/a322943.pdf (accessed on 9 September 2019).
- Bruno, D. Enteric Redmouth Disease. Aquac. Inf. Ser. 1990, 10. Available online: https://www2.gov.scot/Topics/marine/science/Publications/FRS-Reports/ais (accessed on 9 September 2019).
- Zorriehzahra, M.; Adel, M.; Delshad, S.T. Enteric redmouth disease: Past, present and future: A review. Iran. J. Fish. Sci. 2017, 16, 1135–1156. [Google Scholar]
- Gulla, S.; Barnes, A.C.; Welch, T.J.; Romalde, J.L.; Ryder, D.; Ormsby, M.J.; Carson, J.; Lagesen, K.; Verner-Jeffreys, D.W.; Davies, R.L. Multi-Locus Variable number of tandem repeat Analysis (MLVA) of Yersinia ruckeri confirms the existence of host-specificity, geographic endemism and anthropogenic dissemination of virulent clones. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [PubMed]
- McDaniel, D. Hagerman redmouth. Am. Fish. US Trout News 1971, 15, 14–28. [Google Scholar]
- Wobeser, G. An outbreak of redmouth disease in rainbow trout (Salmo gairdneri) in Saskatchewan. J. Fish. Res. Board Can. 1973, 30, 571–575. [Google Scholar] [CrossRef]
- Llewellyn, L. A bacterium with similarities to the redmouth bacterium and Serratia liquefaciens (Grimes and Hennerty) causing mortalities in hatchery reared salmonids in Australia. J. Fish Dis. 1980, 3, 29–39. [Google Scholar] [CrossRef]
- Dalsgaard, I.; From, J.; Hørlyck, V. First observation of Yersinia ruckeri in Denmark. Bull. Eur. Assoc. Fish Pathol. 1984, 4, 10. [Google Scholar]
- Lesel, R.; Lesel, M.; Gavini, F.; VuiIlaume, A. Outbreak of enteric redmouth disease, in rainbow trout, Salmo gairdneri Richardson, in France. J. Fish Dis. 1983, 6, 385–387. [Google Scholar] [CrossRef]
- Giorgetti, G.; Ceschia, G.; Bovo, G. First Isolation of Yersinia ruckeri in Farmed Rainbow Trout in Italy; Ellis, A.E., Ed.; Academic Press: London, UK, 1985; pp. 161–166. [Google Scholar]
- Sparboe, O.; Koren, C.; Hastein, T.; Poppe, T.; Stenwig, H. The first isolation of Yersinia ruckeri from farmed Norwegian salmon. Bull. Eur. Assoc. Fish Pathol. 1986, 6, 41–42. [Google Scholar]
- Grawiński, E. The occurrence of redmouth disease in the rainbow trout (Salmo gairdneri Richardson). Vet. Med. 1990, 46, 183–185. [Google Scholar]
- Oraić, D.; Zrnčić, S.; Šoštarić, B.; Bažulić, D.; Lipiej, Z. Occurence of enteric redmouth disease in rainbow trout (Oncorhynchus mykiss) on farms in Croatia. Acta Vet. Hung. 2002, 50, 283–291. [Google Scholar] [CrossRef]
- Popescu, A.; Costea, M.; Dăscălescu, P. Characterization of Yersinia ruckeri strains from salmonid populations in Romania. Rev. Rom. Med. 2008, 18. [Google Scholar]
- Anderson, C.; Knowles, G.; de Lisle, G. A survey for Yersinia ruckeri and Aeromonas salmonicida in farmed and wild fish. Surveillance 1994, 21, 39–40. [Google Scholar]
- Bragg, R.; Henton, M. Isolation of Yersinia ruckeri from rainbow trout in South Africa. Bull. Eur. Assoc. Fish Pathol. 1986, 6, 5–6. [Google Scholar]
- Enríquez, R.; Zamora, J. Isolation of Yersinia ruckeri from carp (Cyprinus carpio) in Valdivia. Arch. Med. Vet. 1987, 19, 33–36. [Google Scholar]
- Barnes, A. Enteric Redmouth Disease (ERM) (Yersinia ruckeri). In Fish Diseases and Disorders: Viral, Bacterial and Fungal Infections; Woo, P.T.K., Bruno, D.W., Eds.; CABI: Walingford, UK, 2011; Volume 3, pp. 484–511. [Google Scholar]
- Collins, R.; Foster, G.; Ross, H. Isolation of Yersinia ruckeri from farmed Atlantic Salmon (Salmo salar), and Arctic Char (Salvelinus alpinus) and a European Otter (Lutra lutra) in several water catchment areas in Scotland. Vet. Rec. 1998, 139, 169. [Google Scholar] [CrossRef]
- Gudmundsdottir, B.; Gudmundsdottir, S.; Gudmundsdottir, S.; Magnadottir, B. Yersiniosis in Atlantic cod, Gadus morhua (L.), characterization of the infective strain and host reactions. J. Fish Dis. 2014, 37, 511–519. [Google Scholar] [CrossRef]
- Navas, E.; Bohle, H.; Henríquez, P.; Grothusen, H.; Bustamante, F.; Bustos, P.; Mancilla, M. Draft Genome Sequence of the Fish Pathogen Yersinia ruckeri Strain 37551, Serotype O1b, Isolated from Diseased, Vaccinated Atlantic Salmon (Salmo salar) in Chile. Genome Announc. 2014, 2, 10–11. [Google Scholar] [CrossRef]
- Xu, B.; Yin, Z.; Chen, Y.; Wu, Y. An outbreak of a new epi- zootic in Silver Carp and Bighead Carp: Yersinia ruckeri, a new pathogen of Silver Carp and Bighead Carp. Chin. Sci. Bull. 1991, 36, 1825. [Google Scholar]
- Dwilow, A.; Souter, B.; Knight, K. Isolation of Yersinia ruckeri from burbot, Lota lota (L.) from the Mackenzie River, Canada. J. Fish Dis. 1987, 10, 315–317. [Google Scholar] [CrossRef]
- Cipriano, R.; Schill, W.; Pyle, S.; Horner, R. An epizootic in cinook salmon (Oncorhynchus tshawytscha) caused by a sorbitol-positive serovar 2 strain of Yersinia ruckeri. J. Wildl. Dis. 1986, 22, 488–492. [Google Scholar] [CrossRef]
- Huang, Y.; Jung, A.; Schäfer, W.; Mock, D.; Michael, G.B.; Runge, M.; Schwarz, S.; Steinhagen, D. Analysis of Yersinia ruckeri strains isolated from trout farms in northwest Germany. Dis. Aquat. Organ. 2015, 116, 243–249. [Google Scholar] [CrossRef]
- Valtonen, E.; Rintamaki, P.; Koskivaara, M. Occurence and pathogenicity of Yersinia ruckeri at fish farms in northern and central Finland. J. Fish Dis. 1992, 15, 163–171. [Google Scholar] [CrossRef]
- Michel, C.; Fairve, B.; de Kinkelin, P. A clinical case of enteric redmouth in minnows (Pimephales promelas) imported in Europe as bait-fish. Bull. Eur. Assoc. Fish Pathol. 1986, 6, 97–99. [Google Scholar]
- Avendaño-Herrer, R.; Tapia-Cammas, D.; Aedo, A.; Saldivia, P.; Ortega, C.; Irgang, R. Disease caused by Yersinia ruckeri serotype O2b found in Chilean-farmed coho salmon, Oncorhynchus kisutch (Walbaum, 1792). J. Fish Dis. 2016, 40, 279–285. [Google Scholar] [CrossRef]
- Berc, A.; Petrinec, Z.; Matašin, Ž.; Kozarić, Z. Yersinia ruckeri Septicaemia in Experimentally Infected Carp (Cyprinus Carpio L.) Fingerlings. Acta Vet. Hung. 2015, 47, 161–172. [Google Scholar] [CrossRef]
- Daly, J.; Lindvik, B.; Stevenson, R. Serological heterogeneity of recent isolates of Yersinia ruckeri from Ontario and British Columbia. Dis. Aquat. Organ. 1986, 1, 151–153. [Google Scholar] [CrossRef]
- McArdle, J.; Dooley-Martyn, C. Isolation of Yersinia ruckeri type I (Hagerman strain) from goldfish Carassius auratus (L.). Bull. Eur. Assoc. Fish Pathol. 1985, 5, 1–2. [Google Scholar]
- Rintamäki, P.; Valtonen, E.; Frerichs, G. Occurrence of Yersinia ruckeri infection in farmed whitefish, Coregonus peled Gmelin and Coregonus muksun Pallas, and Atlantic salmon, Salmo salar L., in northern Finland. J. Fish Dis. 1986, 9, 137–140. [Google Scholar] [CrossRef]
- Eissa, A.; Moustafa, M.; Abdelaziz, M.; Ezzeldeen, N. Yersinia ruckeri infection in cultured Nile tilapia, Oreochromis niloticus, at a semi-intensive fish farm in lower Egypt. Afr. J. Aquat. Sci. 2008, 33, 283–286. [Google Scholar] [CrossRef]
- Duman, M.; Altun, S.; Cengiz, M.; Saticioglu, I.; Buyukekiz, A.; Sahinturk, P. Genotyping and antimicrobial resistance genes of Yersinia ruckeri isolates from rainbow trout farms. Dis. Aquat. Organ. 2017, 125, 31–44. [Google Scholar] [CrossRef]
- Popović, N.; Hacmanjek, M.; Teskeredžić, E. Health status of rudd (Scardinius erythrophthalmus hesperidicus H.) in Lake Vrana on the Island of Cres, Croatia. J. Appl. Ichthyol. 2001, 17, 43–45. [Google Scholar] [CrossRef]
- Vuillaume, A.; Brun, R.; Chene, P.; Sochon, E.; Lesel, R. First isolation of Yersinia ruckeri from sturgeon, Acipenser baeri Brandt, in South West of France. Bull. Eur. Assoc. Fish Pathol. 1987, 7, 18–19. [Google Scholar]
- Korbut, R.; Mehrdana, F.; Kania, P.W.; Larsen, M.H.; Frees, D.; Dalsgaard, I.; Jørgensen, L.V.G. Antigen Uptake during Different Life Stages of Zebrafish (Danio rerio) Using a GFP-Tagged Yersinia ruckeri. PLoS ONE 2016, 11, e0158968. [Google Scholar] [CrossRef]
- Willumsen, B. Birds and wild fish as potential vectors of Yersinia ruckeri. J. Fish Dis. Dis. 1987, 40, 275–277. [Google Scholar] [CrossRef]
- De Keukeleire, S.; de Bel, A.; Jansen, Y.; Janssens, M.; Wauters, G.; Piérard, D. Yersinia ruckeri, an unusual microorganism isolated from a human wound infection. New Microbes New Infect. 2014, 2, 134–135. [Google Scholar] [CrossRef]
- Standish, I.; Erickson, S.; Kleman, E.; McCann, R.; Katona, R.; Leis, E. Yersinia ruckeri Isolated from Common Mudpuppy Necturus maculosus. J. Aquat. Anim. Health 2018, 31, 71–74. [Google Scholar] [CrossRef]
- Dudley, D.; Guentzel, M.; Ibarra, M.; Moore, B.; Sagik, B. Enumeration of potentially pathogenic bacteria from sewage sludges. Appl. Environ. Microbiol. 1980, 39, 118–126. [Google Scholar]
- Guijarro, J.; García-Torrico, A.; Cascales, D.; Méndez, J. The Infection Process of Yersinia ruckeri: Reviewing the Pieces of the Jigsaw Puzzle. Front. Cell. Infect. Microbiol. 2018, 8, 218. [Google Scholar] [CrossRef]
- Fernández, L.; Méndez, J.; Guijarro, J. Molecular virulence mechanisms of the fish pathogen Yersinia ruckeri. Vet. Microbiol. 2007, 125, 1–10. [Google Scholar] [CrossRef]
- Dahiya, I.; Stevenson, R. Yersinia ruckeri genes that attenuate survival in rainbow trout (Oncorhynchus mykiss) are identified using signature-tagged mutants. Vet. Microbiol. 2010, 144, 399–404. [Google Scholar] [CrossRef] [PubMed]
- Fernández, L.; Marquez, I.; Guijarro, J. Identification of specific In Vivo-induced (ivi) genes in Yersinia ruckeri and analysis of ruckerbactin, a catecholate siderophore iron acquisition system. Appl. Environ. Microbiol. 2004, 70, 5199–5207. [Google Scholar] [CrossRef] [PubMed]
- Kumar, G.; Hummel, K.; Ahrens, M.; Menanteau-Ledouble, S.; Welch, T.J.; Eisenacher, M.; Razzazi-Fazeli, E.; El-Matbouli, M. Shotgun proteomic analysis of Yersinia ruckeri strains under normal and iron-limited conditions. Vet. Res. 2016, 47, 100. [Google Scholar] [CrossRef] [PubMed]
- Kumar, G.; Hummel, K.; Welch, T.; Razzazi-Fazeli, E.; El-Matbouli, M. Global proteomic profiling of Yersinia ruckeri strains. Vet. Res. 2017, 48, 55. [Google Scholar] [CrossRef] [PubMed]
- Secades, P.; Guijarro, J. Purification and Characterization of an Extracellular Protease from the Fish pathogen Yersinia ruckeri and effect of culture condition on production. Appl. Environ. Microbiol. 1999, 65, 3969–3975. [Google Scholar] [PubMed]
- Balakrishnan, L.; Hughes, C.; Koronakis, V. Substrate-triggered recruitment of the TolC channel-tunnel during type I export of hemolysin by Escherichia coli. J. Mol. Biol. 2001, 313, 501–510. [Google Scholar] [CrossRef]
- Fernandez, L.; Secades, P.; Lopez, J.; Marquez, I.; Guijarro, J. Isolation and analysis of a protease gene with an ABC transport system in the fish pathogen Yersinia ruckeri: Insertional mutagenesis and involvement in virulence. Microbiology 2002, 148, 2233–2243. [Google Scholar] [CrossRef][Green Version]
- Fernandez, L.; Lopez, J.; Secades, P.; Menendez, A.; Marquez, I.; Guijarro, J. In Vitro and In Vivo Studies of the Yrp1 Protease from Yersinia ruckeri and Its Role in Protective Immunity against Enteric Red Mouth Disease of Salmonids. Appl. Environ. Microbiol. 2003, 69, 7328–7335. [Google Scholar] [CrossRef]
- Navais, R.; Méndez, J.; Pérez-Pascual, D.; Cascales, D.; Guijarro, J. The yrpAB operon of Yersinia ruckeri encoding two putative U32 peptidases is involved in virulence and induced under microaerobic conditions. Virulence 2014, 5, 619–624. [Google Scholar] [CrossRef]
- Fernández, L.; Prieto, M.; Guijaro, J. The iron- and temperature-regulated haemolysin YhlA is a virulence factor of Yersinia ruckeri. Microbiology 2007, 153, 483–489. [Google Scholar] [CrossRef]
- Leo, J.; Grin, I.; Linke, D. Type V secretion: Mechanism(s) of autotransport through the bacterial outer membrane. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2012, 367, 1088–1101. [Google Scholar] [CrossRef] [PubMed]
- Chain, P.S.; Carniel, E.; Larimer, F.W.; Lamerdin, J.; Stoutland, P.O.; Regala, W.M.; Georgescu, A.M.; Vergez, L.M.; Land, M.L.; Motin, V.L.; et al. Insights into the evolution of Yersinia pestis through whole-genome comparison with Yersinia pseudotuberculosis. Proc. Natl. Acad. Sci. USA 2004, 101, 13826–13831. [Google Scholar] [CrossRef] [PubMed]
- Flores-Diaz, M.; Monturiol-Gross, L.; Naylor, C.; Alape-Giron, A.; Flieger, A. Bacterial Sphingomyelinases and Phospholipases as Virulence Factors. Microbiol. Mol. Biol. Rev. MMBR 2016, 80, 597–628. [Google Scholar] [CrossRef] [PubMed]
- Romalde, J.L.; Toranzo, A.E. Pathological activities of Yersinia ruckeri, the enteric redmouth (ERM) bacterium. FEMS Microbiol. Lett. 1993, 112, 291–299. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Romalde, J.; Santos, Y.; Toranzo, A. Presence of skin permeability factors in the extracellular products of Yersinia ruckeri. Curr. Microbiol. 1992, 24, 263–267. [Google Scholar] [CrossRef]
- Welch, T.J.; Verner-Jeffreys, D.W.; Dalsgaard, I.; Wiklund, T.; Evenhuis, J.P.; Cabrera, J.A.G.; Hinshaw, J.M.; Drennan, J.D.; LaPatra, S.E. Independent Emergence of Yersinia Ruckeri Biotype 2 in the United States and Europe. Appl. Environ. Microbiol. 2011, 77, 3493–3499. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jozwick, A.; Graf, J.; Welch, T. The flagellar master operon flhDC is a pleiotropic regulator involved in motility and virulence of the fish pathogen Yersinia ruckeri. J. Appl. Microbiol. 2016, 122, 578–588. [Google Scholar] [CrossRef] [PubMed]
- Jozwick, A.; LaPatra, S.; Graf, J.; Welch, T. Flagellar regulation mediated by the Rcs pathway is required for virulence in the fish pathogen Yersinia ruckeri. Fish Shellfish Immunol. 2019, 91, 306–314. [Google Scholar] [CrossRef]
- Heymann, J.B.; Bartho, J.D.; Rybakova, D.; Venugopal, H.P.; Winkler, D.C.; Sen, A.; Hurst, M.R.H.; Mitra, A.K. Three-dimensional Structure of the Toxin-delivery Particle Antifeeding Prophage of Serratia entomophila. J. Biol. Chem. 2013, 288, 25276–25284. [Google Scholar] [CrossRef]
- Jank, T.; Eckerle, S.; Steinemann, M.; Trillhaase, C.; Schimpl, M.; Wiese, S.; Van Aalten, D.M.F.; Driever, W.; Aktories, K. Tyrosine glycosylation of Rho by Yersinia toxin impairs blastomere cell behaviour in zebrafish embryos. Nat. Commun. 2015, 6, 1–16. [Google Scholar] [CrossRef]
- Yang, X.; Pan, J.; Wang, Y.; Shen, X. Type VI Secretion Systems Present New Insights on Pathogenic Yersinia. Front. Cell. Infect. Microbiol. 2018, 8, 260. [Google Scholar] [CrossRef] [PubMed]
- Hurst, M.; Beard, S.; Jackson, T.; Jones, S. Isolation and characterization of the Serratia entomophila antifeeding prophage. FEMS Microbiol. Lett. 2007, 270, 42–48. [Google Scholar] [CrossRef] [PubMed]
- Puhar, A. Type III secretion system. Curr. Biol. 2014, 24, R784–R791. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.; Wang, K.-Y.; Wang, J.; Chen, D.-F.; Huang, X.-L.; Ouyang, P.; Geng, Y.; He, Y.; Zhou, Y.; Min, J. Genome sequence of the fish pathogen Yersinia ruckeri SC09 provides insights into niche adaptation and pathogenic mechanism. Int. J. Mol. Sci. 2016, 17, 557. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, D.; Du, D.; Li, S.; Yan, W. Advances in the Assembly Model of Bacterial Type IVB Secretion Systems. Appl. Sci. 2018, 8, 2368. [Google Scholar] [CrossRef]
- Alvarez-Martinez, C.; Christie, P. Biological diversity of prokaryotic type IV secretion systems. Microbiol Mol Biol Rev. 2009, 73, 775–808. [Google Scholar] [CrossRef] [PubMed]
- Méndez, J.; Fernández, L.; Menéndez, A.; Reimundo, P.; Pérez-Pascual, D.; Navais, R.; Guijarro, J.A. A chromosomally located traHIJKCLMN operon encoding a putative type IV secretion system is involved in the virulence of Yersinia ruckeri. Appl. Environ. Microbiol. 2009, 75, 937–945. [Google Scholar] [CrossRef] [PubMed]
- Wrobel, A.; Ottoni, C.; Leo, J.; Linke, D. pYR4 From a Norwegian Isolate of Yersinia ruckeri Is a Putative Virulence Plasmid Encoding Both a Type IV Pilus and a Type IV Secretion System. Front. Cell. Infect. Microbiol. 2018, 8, 1–14. [Google Scholar] [CrossRef]
- Cascales, D.; Guijarro, J.; Garcia-Torrico, A.; Méndez, J. Comparative genome analysis reveals important genetic differences among serotype O1 and serotype O2 strains of Y. ruckeri and provides insights into host adaptation and virulence. MicrobiologyOpen 2017, 6. [Google Scholar] [CrossRef]
- Fan, E.; Chauhan, N.; Udatha, D.; Leo, J.; Linke, D. Type V Secretion Systems in Bacteria. Microbiol. Spectr. 2016, 4, 1–24. [Google Scholar]
- Chauhan, N.; Wrobel, A.; Skurnik, M.; Leo, J. Yersinia adhesins: An arsenal for infection. Proteom. Clin. Appl. 2016, 10, 949–963. [Google Scholar] [CrossRef] [PubMed]
- Leo, J.; Oberhettinger, P.; Schütz, M.; Linke, D. The inverse autotransporter family: Intimin, invasin and related proteins. Int. J. Med Microbiol. 2015, 305, 276–282. [Google Scholar] [CrossRef] [PubMed]
- Heinz, E.; Stubenrauch, C.J.; Grinter, R.; Croft, N.P.; Purcell, A.W.; Strugnell, R.A.; Dougan, G.; Lithgow, T. Conserved features in the structure, mechanism and biogenesis of the inverse autotransporter protein family. Genome Biol. Evol. 2016, 8, 1690–1705. [Google Scholar] [CrossRef] [PubMed]
- Wrobel, A.; Ottoni, C.; Leo, J.C.; Gulla, S.; Linke, D. The repeat structure of two paralogous genes, Yersinia ruckeri Invasin (yrInv) and a ‘Y. ruckeri Invasin-like molecule’, (yrIlm) sheds light on the evolution of adhesive capacities of a fish pathogen. J. Struct. Biol. 2017, 201, 171–183. [Google Scholar] [CrossRef] [PubMed]
- Leo, J.; Oberhettinger, P.; Chaubey, M.; Schütz, M.; Kühner, D.; Bertsche, U.; Schwarz, H.; Götz, F.; Autenrieth, I.B.; Coles, M.; et al. The Intimin periplasmic domain mediates dimerisation and binding to peptidoglycan. Mol. Microbiol. 2015, 95, 80–100. [Google Scholar] [CrossRef]
- Craig, L.; Forest, K.; Maier, B. Type IV pili: Dynamics, biophysics and functional consequences. Nat. Rev. Microbiol. 2019, 17, 429–440. [Google Scholar] [CrossRef]
- Ahmed, E.; Holmström, S. Siderophores in environmental research: Roles and applications. Microb. Biotechnol. 2014, 7, 196–208. [Google Scholar] [CrossRef]
- Méndez, J.; Reimundo, P.; Pérez-Pascual, D.; Navais, R.; Gómez, E.; Guijarro, A. A Novel cdsAB Operon Is Involved in the Uptake of L-Cysteine and Participates in the Pathogenesis of Yersinia ruckeri. J. Bacteriol. 2011, 193, 944–951. [Google Scholar] [CrossRef]
- Patzer, S.; Hantke, K. The ZnuABC high-affinity zinc uptake system and its regulator Zur in Escherichia coli. Mol. Microbiol. 1998, 28, 1199–1210. [Google Scholar] [CrossRef]
- Campoy, S.; Jara, M.; Busquets, N.; de Rozas, A.; Badiola, I.; Barbe, J. Role of the high-affinity zinc uptake znuACB system in Salmonella enterica serovar Typhimurium virulence. Infect. Immun. 2002, 70, 4721–4725. [Google Scholar] [CrossRef]
- Dahiya, I.; Stevenson, R. The ZnuABC operon is important for Yersinia ruckeri infections of rainbow trout, Oncorhynchus mykiss (Walbaum). J. Fish Dis. 2010, 33, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Pernestig, A.; Georgellis, D.; Romeo, T.; Suzuki, K.; Tomenius, H.; Normark, S.; Melefors, O. The Escherichia coli BarA-UvrY two-component system is needed for efficient switching between glycolytic and gluconeogenic carbon sources. J. Bacteriol. 2003, 185, 843–853. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, I.; Stevenson, R. The UvrY response regulator of the BarA—UvrY two-component system contributes to Yersinia ruckeri infection of rainbow trout (Oncorhynchus mykiss). Arch. Microbiol. 2010, 192, 541–547. [Google Scholar] [CrossRef] [PubMed]
- Mendez, J.; Cascales, D.; Garcia-Torrico, A.; Guijarro, J. Temperature-Dependent Gene Expression in Yersinia ruckeri: Tracking Specific Genes by Bioluminescence during In Vivo Colonization. Front. Microbiol. 2018, 9, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Furones, M.; Gilpin, M.; Alderman, D.; Munn, C. Virulence of Yersinia ruckeri serotype I strains is associated with a heat sensitive factor (HSF) in cell extracts. FEMS Microbiol. Lett. 1990, 66, 339–343. [Google Scholar] [CrossRef][Green Version]
- Altinok, I.; Ozturk, R.; Kahraman, U.; Capkin, E. Protection of rainbow trout against yersiniosis by lpxD mutant Yersinia ruckeri. Fish Shellfish Immunol. 2016, 55, 21–27. [Google Scholar] [CrossRef]
- Cornelis, G.R.; Boland, A.; Boyd, A.P.; Geuijen, C.; Iriarte, M.; Neyt, C.; Sory, M.-P.; Stainier, I. The Virulence Plasmid of Yersinia, an Antihost Genome. Microbiol. Mol. Biol. Rev. 1998, 62, 1315–1352. [Google Scholar]
- Mühlenkamp, M.; Oberhettinger, P.; Leo, J.; Linke, D.; Schütz, M. Yersinia adhesin A (YadA)—Beauty & beast. Int. J. Med. Microbiol. 2015, 305, 252–258. [Google Scholar]
- McNally, A.; Thomson, N.; Reuter, S.; Wren, B. ‘Add, stir and reduce’: Yersinia spp. as model bacteria for pathogen evolution. Nat. Rev. Microbiol. 2016, 14, 177–190. [Google Scholar] [CrossRef]
- Rajanna, C.; Revazishvili, T.; Rashid, M.H.; Chubinidze, S.; Bakanidze, L.; Tsanava, S.; Imnadze, P.; Bishop-Lilly, K.A.; Sozhamannan, S.; Gibbons, H.S.; et al. Characterization of pPCP1 Plasmids in Yersinia pestis Strains Isolated from the Former Soviet Union. Int. J. Microbiol. 2010, 2010, 1–9. [Google Scholar] [CrossRef]
- Eppinger, M.; Radnedge, L.; Andersen, G.; Vietri, N.; Severson, G.; Mou, S.; Ravel, J.; Worsham, P.L. Novel plasmids and resistance phenotypes in Yersinia pestis: Unique plasmid inventory of strain Java 9 mediates high levels of arsenic resistance. PLoS ONE 2012, 7, e32911. [Google Scholar] [CrossRef] [PubMed]
- De Grandis, S.; Stevenson, R. Variations in plasmid profiles and growth characteristics of Yersinia ruckeri strains. FEMS Microbiol. Lett. 1982, 15, 199–202. [Google Scholar] [CrossRef][Green Version]
- Stave, J.; Cook, T.; Roberson, B. Chemiluminescent responses of striped bass, Morone saxatilis (Walbaum), phagocytes to strains of Yersinia ruckeri. J. Fish Dis. 1987, 10, 1–10. [Google Scholar] [CrossRef]
- Toranzo, A.; Barja, J.; Colwell, R.; Hetrick, F. Characterization of Plasmids in Bacterial Fish Pathogens. Infect. Immun. 1983, 39, 184–192. [Google Scholar] [PubMed]
- Garcia, J.; Dominguez, L.; Larsen, J.; Pedersen, K. Ribotyping and plasmid profiling of Yersinia ruckeri. J. Appl. Microbiol. 1998, 85, 949–955. [Google Scholar] [CrossRef] [PubMed]
- Guilvout, I.; Quilici, M.; Rabot, S.; Lesel, R.; Mazigh, D. BamHI restriction endonuclease analysis of Yersinia ruckeri plasmids and their relatedness to the genus Yersinia 42- to 47-megadalton plasmid. Appl. Environ. Microbiol. 1988, 54, 2594–2597. [Google Scholar] [PubMed]
- De Grandis, S.; Stevenson, R. Antimicrobial susceptibility patterns and R plasmid-mediated resistance of the fish pathogen Yersinia ruckeri. Antimicrob. Agents Chemother. 1985, 27, 938–942. [Google Scholar] [CrossRef]
- Welch, T.J.; Fricke, W.F.; McDermott, P.F.; White, D.G.; Rosso, M.-L.; Rasko, D.A.; Mammel, M.K.; Eppinger, M.; Rosovitz, M.; Wagner, D.; et al. Multiple Antimicrobial Resistance in Plague: An Emerging Public Health Risk. PLoS ONE 2007, 2, e309. [Google Scholar] [CrossRef]
- Liu, T.; Wang, E.; Wei, W.; Wang, K.; Yang, Q.; Ai, X. TcpA, a novel Yersinia ruckeri TIR-containing virulent protein mediates immune evasion by targeting MyD88 adaptors. Fish Shellfish Immunol. 2019, 94, 58–65. [Google Scholar] [CrossRef]
- Rollauer, S.; Sooreshjani, M.; Noinaj, N.; Buchanan, S. Outer membrane protein biogenesis in Gram-negative bacteria. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370, 20150023. [Google Scholar] [CrossRef]
- Wang, E.; Qin, Z.; Yu, Z.; Ai, X.; Wang, K.; Yang, Q.; Liu, T.; Chen, D.; Geng, Y.; Huang, X.; et al. Molecular Characterization, Phylogenetic, Expression, and Protective Immunity Analysis of OmpF, a Promising Candidate Immunogen Against Yersinia ruckeri Infection in Channel Catfish. Front. Immunol. 2018, 9, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Ormsby, M.; Grahame, E.; Burchmore, R.; Davies, R. Comparative bioinformatic and proteomic approaches to evaluate the outer membrane proteome of the fish pathogen Yersinia ruckeri. J. Proteomics 2019, 199, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Coquet, L.; Cosette, P.; Junter, G.-A.; Beucher, E.; Saiter, J.-M.; Jouenne, T. Adhesion of Yersinia ruckeri to fish farm materials: Influence of cell and material surface properties. Colloids Surfaces B Biointerfaces 2002, 26, 373–378. [Google Scholar] [CrossRef]
- Coquet, L.; Cosette, P.; Quillet, L.; Petit, F.; Junter, G.; Jouenne, T. Occurrence and Phenotypic Characterization of Yersinia ruckeri Strains with Biofilm-Forming Capacity in a Rainbow Trout Farm. Appl. Environ. Microbiol. 2002, 68, 470–475. [Google Scholar] [CrossRef] [PubMed]
- Coquet, L.; Cosette, P.; Dé, E.; Galas, L.; Vaudry, H.; Rihouey, C.; Lerouge, P.; Junter, G.A.; Jouenne, T. Immobilization induces alterations in the outer membrane protein pattern of Yersinia ruckeri. J. Proteome Res. 2005, 4, 1988–1998. [Google Scholar] [CrossRef]
- Menanteau-Ledouble, S.; Lawrence, M.; El-matbouli, M. Invasion and replication of Yersinia ruckeri in fish cell cultures. BMC Vet. Res. 2018, 14, 81. [Google Scholar] [CrossRef]
- Tobback, E.; Decostere, A.; Hermans, K.; van den Broeck, W.; Haesebrouck, F.; Chiers, K. In Vitro markers for virulence in Yersinia ruckeri. J. Fish Dis. 2010, 33, 197–209. [Google Scholar] [CrossRef]
- Delshad, S.; Soltanian, S.; Sharifiyazdi, H.; Bossier, P. Effect of quorum quenching bacteria on growth, virulence factors and biofilm formation of Yersinia ruckeri in vitro and an in vivo evaluation of their probiotic effect in rainbow trout. J. Fish Dis. 2018, 41, 1429–1438. [Google Scholar] [CrossRef]
- Ohtani, M.; Villumsen, K.; Strøm, H.; Raida, M. 3D visualization of the initial Yersinia ruckeri infection route in rainbow trout (Oncorhynchus mykiss) by optical projection tomography. PLoS ONE 2014, 9, e89672. [Google Scholar] [CrossRef]
- Ohtani, M.; Kasper, R.; Koppang, E.; Raida, M. Global 3D Imaging of Yersinia ruckeri Bacterin Uptake in Rainbow Trout Fry. PLoS ONE 2014, 10, e0117263. [Google Scholar] [CrossRef]
| Year | |||||||
|---|---|---|---|---|---|---|---|
| 1977 | 1978 | 1984 | 1988 | 1990 | 1993 | 2016 | |
| Serotype name | I | I | I | - | O1 | O1a | O1 |
| II | II | II | II | O2 | O2a,b,c | O2 | |
| - | III | III | III | O1 | O1b | O1 | |
| - | - | IV | - | - | - | - | |
| - | - | V | V | O5 | O3 | O5 | |
| - | - | VI | VI | O6 | O4 | O6 | |
| - | - | - | - | O7 | - | O7 | |
| O8 | |||||||
| Common Fish Name | Scientific Name | Reference |
|---|---|---|
| Arctic char | Salvelinus alpinus | [60] |
| Atlantic cod | Gadus morhua | [61] |
| Atlantic salmon | Salmo salar | [15,60,62] |
| Bighead carp | Aristichthys nobilis | [63] |
| Burbot | Lota lota | [64] |
| Brook trout | Salvelinus fontinalis | [21,65] |
| Brown trout | Salmo trutta | [66,67] |
| Coalfish | Pollachius virens | [68] |
| Coho salmon | Oncorhynchus kisutch | [69] |
| Common carp | Cyprinus carpio | [70] |
| Chinook salmon | Oncorhynchus tshawytscha | [65] |
| Cisco | Coregonus artedii | [21] |
| Cutthroat trout | Salmo clarkii | [71] |
| Eel | Anguilla anguilla | [41] |
| Fathead minnow | Pimephales promelas | [68] |
| Goldfish | Carassius auratus auratus | [72] |
| Muksun | Coregonus muksun | [73] |
| Nile tilapia | Oreochromis niloticus | [74] |
| Perch | Perca fluviatilis | [67] |
| Peled | Coregonus peled | [73] |
| Rainbow trout | Oncorhynchus mykiss | [17,38,41,66,67,71,75] |
| Rudd | Scardinius erythrophthalmus | [76] |
| Sockeye salmon | Oncorhynchus nerka | [38] |
| Sole | Solea solea | [68] |
| Silver carp | Hypophthalmichthys molitrix | [63] |
| Sturgeon | Acipenser baeri | [77] |
| Turbot | Scophthalmus maximus | [68] |
| Zebrafish | Danio rerio | [78] |
| Name | Scientific Name | Reference |
|---|---|---|
| Muskrat | Ondatra zibethica | [21] |
| Eurasian otter | Lutra lutra | [60] |
| Sea gulls and other birds | Larus spp. | [79] |
| Falco spp. | [39] | |
| Humans (wound infection) | Homo sapiens | [80] |
| Common mudpuppy | Necturus maculosus | [81] |
| Turtles | Cheloniidae | [39] |
| Water Sewage sludge | - | [82] |
| Aquatic invertebrates | - | [79] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wrobel, A.; Leo, J.C.; Linke, D. Overcoming Fish Defences: The Virulence Factors of Yersinia ruckeri. Genes 2019, 10, 700. https://doi.org/10.3390/genes10090700
Wrobel A, Leo JC, Linke D. Overcoming Fish Defences: The Virulence Factors of Yersinia ruckeri. Genes. 2019; 10(9):700. https://doi.org/10.3390/genes10090700
Chicago/Turabian StyleWrobel, Agnieszka, Jack C. Leo, and Dirk Linke. 2019. "Overcoming Fish Defences: The Virulence Factors of Yersinia ruckeri" Genes 10, no. 9: 700. https://doi.org/10.3390/genes10090700
APA StyleWrobel, A., Leo, J. C., & Linke, D. (2019). Overcoming Fish Defences: The Virulence Factors of Yersinia ruckeri. Genes, 10(9), 700. https://doi.org/10.3390/genes10090700






