Transcriptomic Analysis of Porcine Granulosa Cells Overexpressing Retinol Binding Protein 4
Abstract
1. Introduction
2. Materials and Methods
2.1. Obligatory Ethical Approval
2.2. Cell Culture
2.3. Plasmid Constructs, Transfection, and Infection
2.4. Total RNA Extraction and Real-Time Quantitative Polymerase Chain Reaction (RT-qPCR)
2.5. Western Blotting
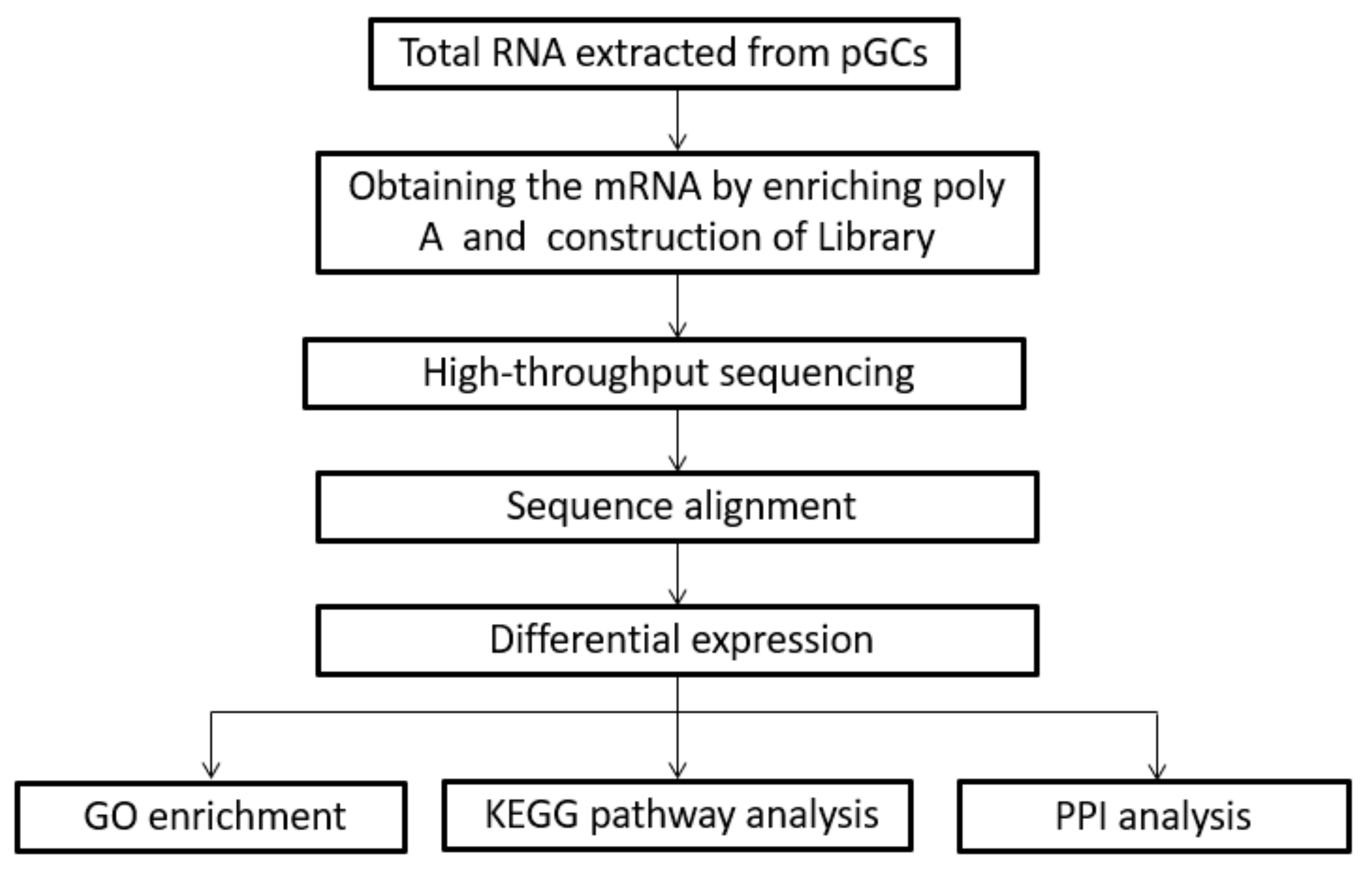
2.6. RNA-Sequencing Library Construction
2.7. Identification of Differentially Expressed Genes (DEGs)
2.8. Functional Enrichment Analysis
2.9. Statistical Analysis
3. Results
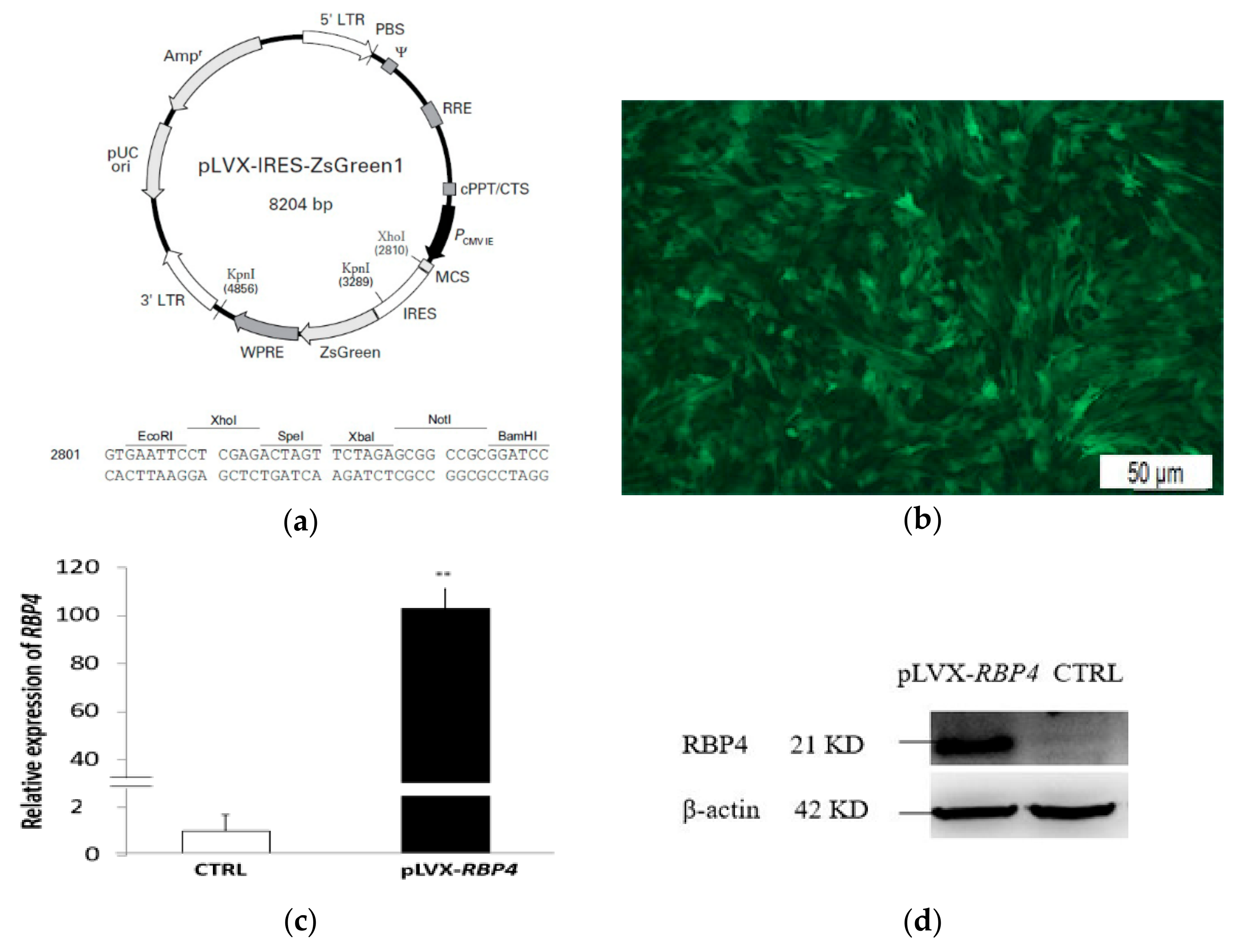
3.1. Infection Efficiency
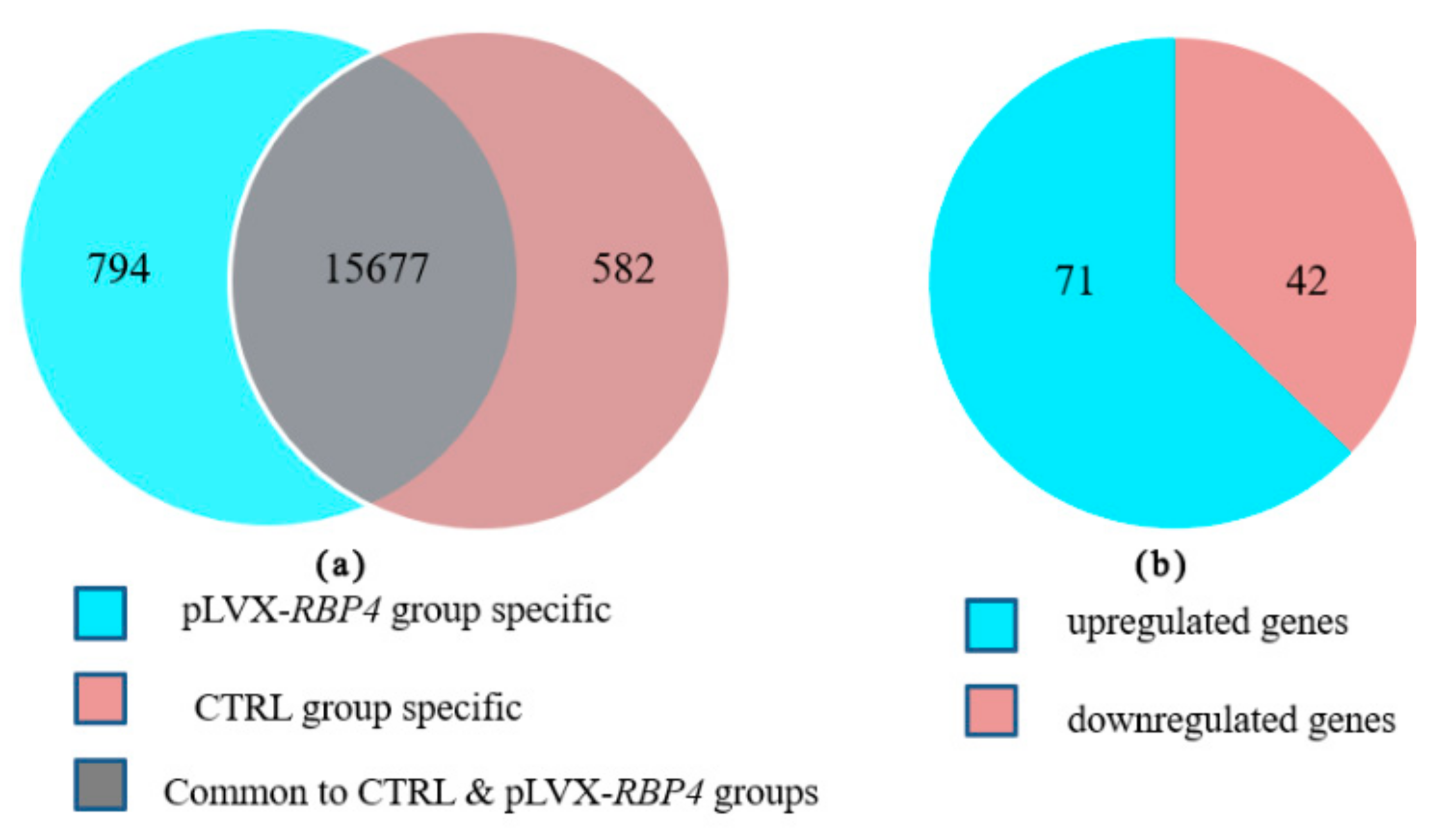
3.2. Transcriptome Profiling Analysis
3.3. Differential Expression of Messenger RNAs in Granulosa Cells (GCs)
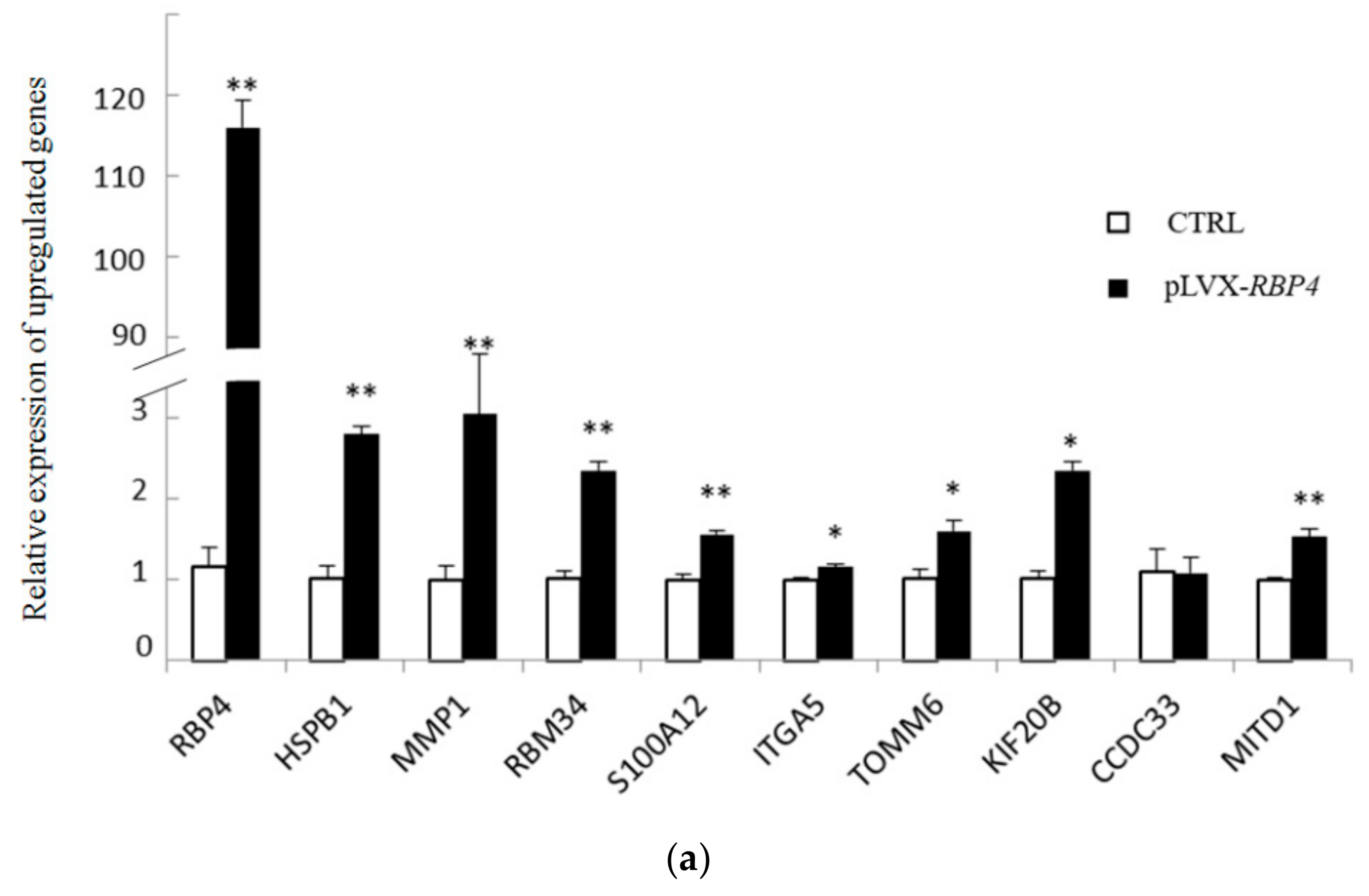
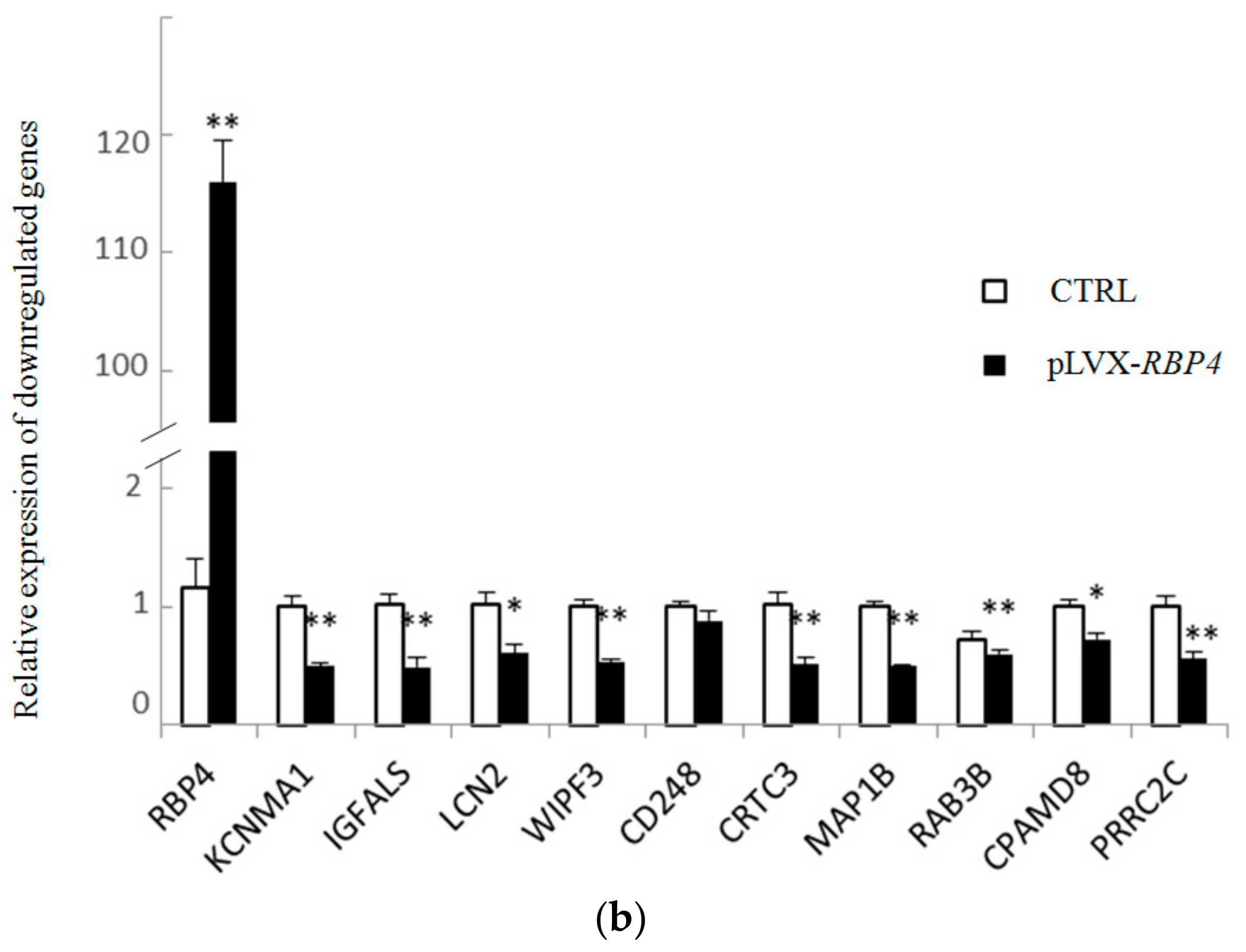
3.4. Validation of Differentially Expressed Messenger RNAs
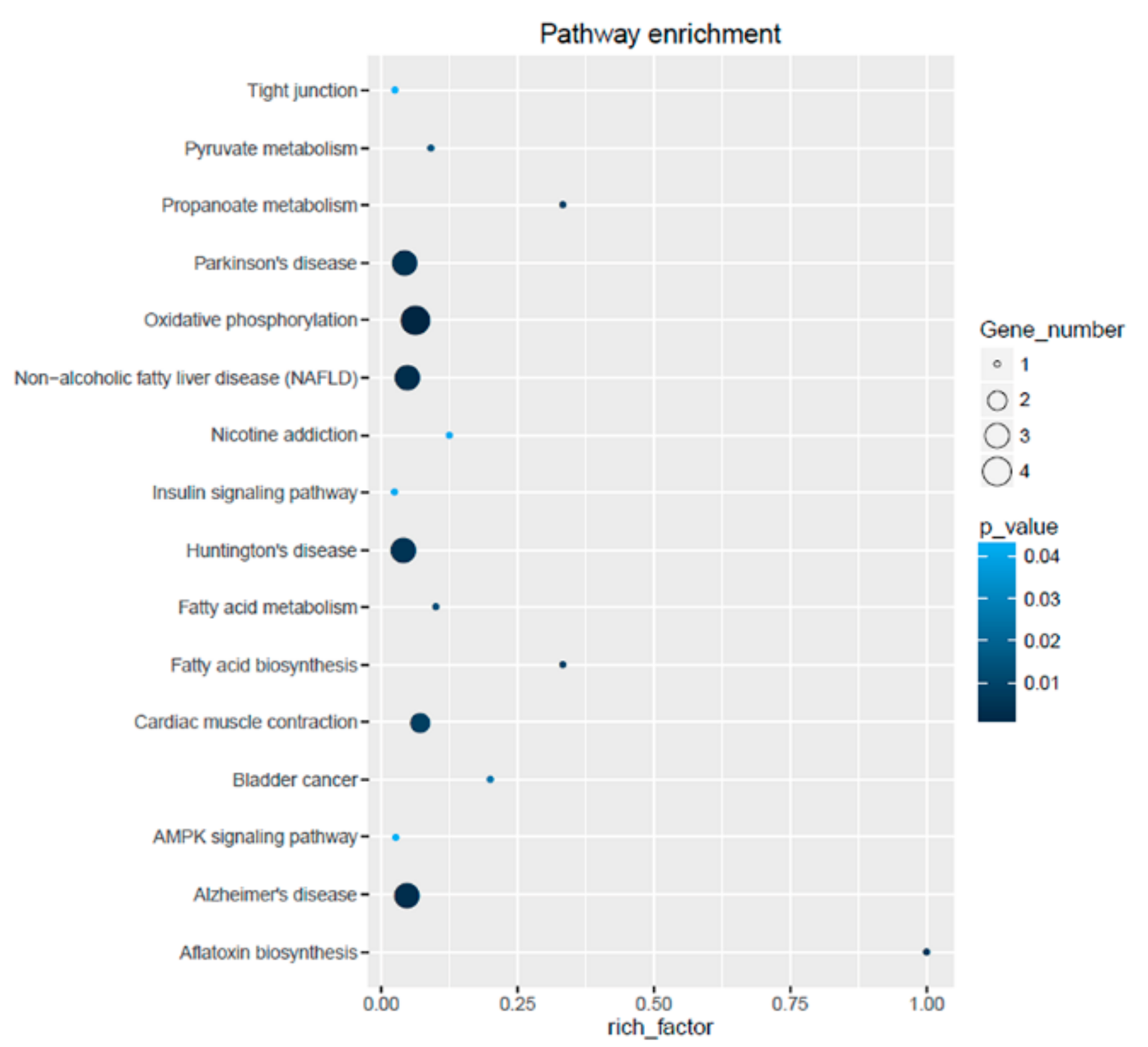
3.5. Gene Ontology Annotation and Kyoto Encyclopedia of Genes and Genomes Enrichment Analysis of Differentially Expressed Genes
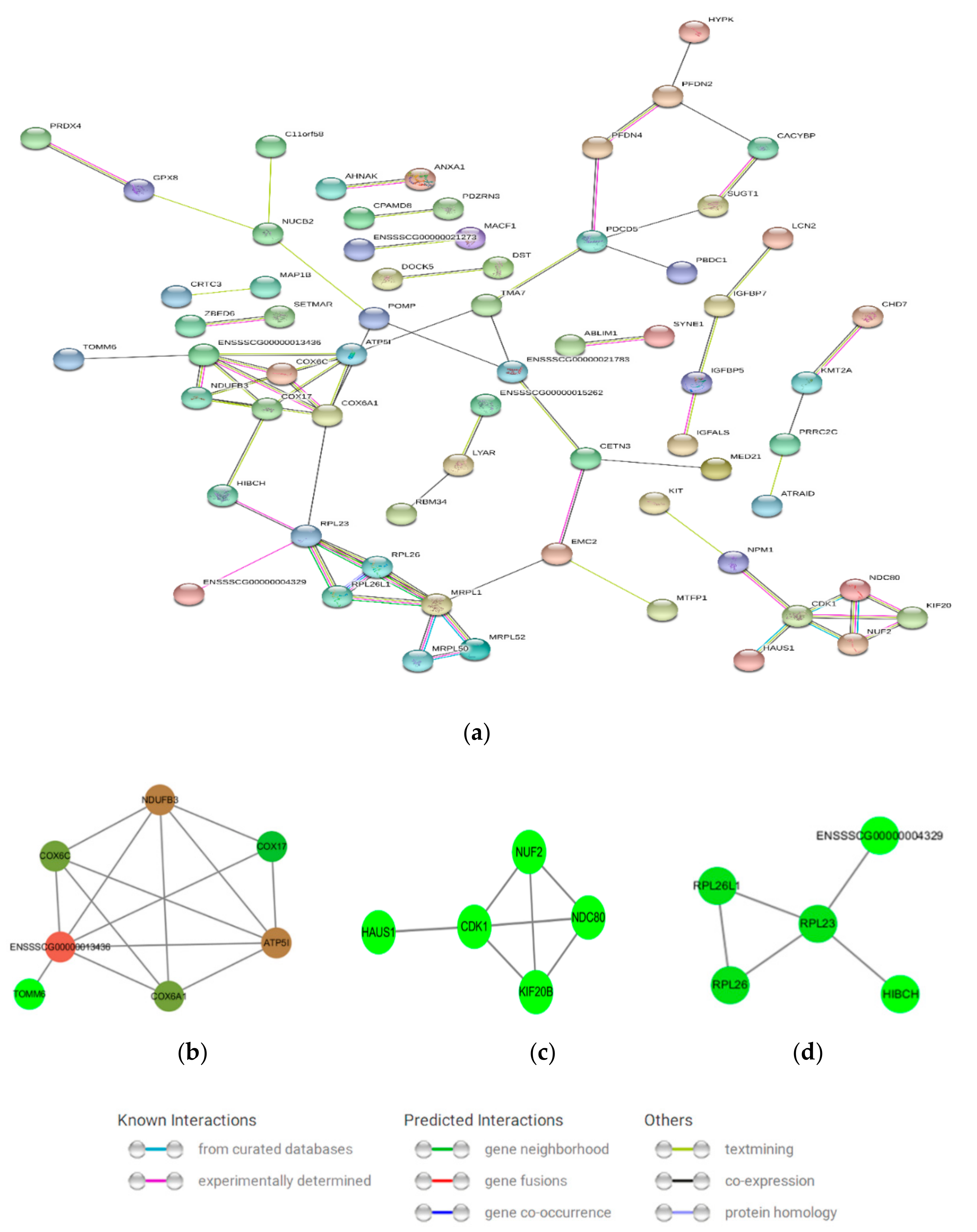
3.6. Protein–Protein Interaction (PPI) Network Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Lawrence, J.; Payton, R.; Godkin, J.; Saxton, A.; Schrick, F.; Edwards, J. Retinol improves development of bovine oocytes compromised by heat stress during maturation. J. Dairy Sci. 2004, 87, 2449–2454. [Google Scholar] [CrossRef]
- Suwa, H.; Kishi, H.; Imai, F.; Nakao, K.; Hirakawa, T.; Minegishi, T. Retinoic acid enhances progesterone production via the cAMP/PKA signaling pathway in immature rat granulosa cells. Biochem. Biophys. Rep. 2016, 8, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Elomda, A.M.; Saad, M.F.; Saeed, A.M.; Elsayed, A.; Abass, A.O.; Safaa, H.M.; Mehaisen, G.M.K. Antioxidant and developmental capacity of retinol on the in vitro culture of rabbit embryos. Zygote 2018, 26, 326–332. [Google Scholar] [CrossRef] [PubMed]
- Fujihara, M.; Yamamizu, K.; Comizzoli, P.; Wildt, D.E.; Songsasen, N. Retinoic acid promotes in vitro follicle activation in the cat ovary by regulating expression of matrix metalloproteinase 9. PLoS ONE 2018, 13, e0202759. [Google Scholar] [CrossRef] [PubMed]
- Kanai, M.; Raz, A.; Goodman, D.S. Retinol-binding protein: The transport protein for vitamin A in human plasma. J. Clin. Investig. 1968, 47, 2025–2044. [Google Scholar] [CrossRef] [PubMed]
- Tsutsumi, C.; Okuno, M.; Tannous, L.; Piantedosi, R.; Allan, M.; Goodman, D.S.; Blaner, W.S. Retinoids and retinoid-binding protein expression in rat adipocytes. J. Biol. Chem. 1992, 267, 1805–1810. [Google Scholar] [PubMed]
- Isken, A.; Golczak, M.; Oberhauser, V.; Hunzelmann, S.; Driever, W.; Imanishi, Y.; Palczewski, K.; Von Lintig, J. RBP4 disrupts vitamin A uptake homeostasis in a STRA6-deficient animal model for Matthew-Wood Syndrome. Cell Metab. 2008, 7, 258–268. [Google Scholar] [CrossRef]
- Kawaguchi, R.; Yu, J.; Honda, J.; Hu, J.; Whitelegge, J.; Ping, P.; Wiita, P.; Bok, D.; Sun, H. A membrane receptor for retinol binding protein mediates cellular uptake of vitamin, A. Science 2007, 315, 820–825. [Google Scholar] [CrossRef]
- Yang, Q.; Graham, T.E.; Mody, N.; Preitner, F.; Peroni, O.D.; Zabolotny, J.M.; Kotani, K.; Quadro, L.; Kahn, B.B. Serum retinol binding protein 4 contributes to insulin resistance in obesity and type 2 diabetes. Nature 2005, 436, 356–362. [Google Scholar] [CrossRef]
- Li, F.; Xia, K.; Li, C.; Yang, T. Retinol-binding protein 4 as a novel risk factor for cardiovascular disease in patients with coronary artery disease and hyperinsulinemia. Am. J. Med. Sci. 2014, 348, 474–479. [Google Scholar] [CrossRef]
- Tan, B.K.; Chen, J.; Lehnert, H.; Kennedy, R.; Randeva, H.S. Raised serum, adipocyte, and adipose tissue retinol-binding protein 4 in overweight women with polycystic ovary syndrome: Effects of gonadal and adrenal steroids. J. Clin. Endocrinol. Metab. 2007, 92, 2764–2772. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Wang, Y.; Wang, Y.; Zhang, Z. Adipokine RBP4 drives ovarian cancer cell migration. J. Ovarian Res. 2018, 11, 29. [Google Scholar] [CrossRef] [PubMed]
- Marantidis, A.; Laliotis, G.P.; Avdi, M. Association of RBP4 genotype with phenotypic reproductive traits of sows. Genet. Res. Int. 2016, 2016, 4940532. [Google Scholar] [CrossRef] [PubMed]
- Brown, J.A.; Eberhardt, D.M.; Schrick, F.N.; Roberts, M.P.; Godkin, J.D. Expression of retinol-binding protein and cellular retinol-binding protein in the bovine ovary. Mol. Reprod. Dev. 2003, 64, 261–269. [Google Scholar] [CrossRef] [PubMed]
- Levi, L.; Levavi-Sivan, B.; Lubzens, E. Expression of genes associated with retinoid metabolism in the trout ovarian follicle1. Biol. Reprod. 2008, 79, 570–577. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.Z.; Denovan-Wright, E.M.; Degrave, A.; Thisse, C.; Thisse, B.; Wright, J.M. Spatio-temporal distribution of cellular retinol-binding protein gene transcripts (CRBPI and CRBPII) in the developing and adult zebrafish (Danio rerio). JBIC J. Biol. Inorg. Chem. 2004, 271, 339–348. [Google Scholar] [CrossRef]
- Pomp, D.; Geisert, R.D.; Yelich, J.V. Detection of transcripts for retinoic acid receptors, retinol-binding protein, and transforming growth factors during rapid trophoblastic elongation in the porcine conceptus. Biol. Reprod. 1997, 57, 286–294. [Google Scholar]
- Harney, J.P.; Ott, T.L.; Geisert, R.D.; Bazer, F.W. Retinol-binding protein gene expression in cyclic and pregnant endometrium of pigs, sheep, and cattle. Biol. Reprod. 1993, 49, 1066–1073. [Google Scholar] [CrossRef][Green Version]
- Trout, W.E.; McDonnell, J.J.; Kramer, K.K.; Baumbach, G.A.; Roberts, R.M. The retinol-binding protein of the expanding pig blastocyst: Molecular cloning and expression in trophectoderm and embryonic disc. Mol. Endocrinol. 1991, 5, 1533–1540. [Google Scholar] [CrossRef]
- Roberts, R.M.; Xie, S.; Trout, W.E. Embryo-uterine interactions in pigs during week 2 of pregnancy. J. Reprod. Fertil. Suppl. 1993, 48, 171–186. [Google Scholar]
- Albertini, D.; Combelles, C.; Benecchi, E.; Carabatsos, M. Cellular basis for paracrine regulation of ovarian follicle development. Reproduction 2001, 121, 647–653. [Google Scholar] [CrossRef]
- Sun, Y.L.; Ping, Z.G.; Li, C.J.; Sun, Y.F.; Yi, K.L.; Li, X.Y.; Wang, X.L.; Zhou, X. Comparative proteomic analysis of follicular fluids from normal and cystic follicles in sows. Reprod. Domest. Anim. 2011, 46, 889–895. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.; Zhao, Y.; Chen, S.; Chen, L.; Li, C.; Zhou, X. Regulation by FSH of the dynamic expression of retinol-binding protein 4 in the mouse ovary. Reprod. Biol. Endocrinol. 2018, 16, 25. [Google Scholar] [CrossRef] [PubMed]
- Rao, J.; Chen, J.; Bi, M.; Zhang, Y.; Chen, S.; Zhao, Y.; Wang, F.; Qiu, T.; Chen, L.; Li, C.; et al. Interaction between the expression of retinol binding protein 4 and gonadotropin receptors in follicular granulosa cells of pigs. Livest. Sci. 2019, 220, 205–210. [Google Scholar] [CrossRef]
- Li, Y.; Ganta, S.; Von Stein, F.B.; Mason, D.E.; Mitchell, B.M.; Freeman, L.C. 4-aminopyridine decreases progesterone production by porcine granulosa cells. Reprod. Biol. Endocrinol. 2003, 1, 31. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Eggers, R.; Hendriks, W.T.; Tannemaat, M.R.; Van Heerikhuize, J.J.; Pool, C.W.; Carlstedt, T.P.; Zaldumbide, A.; Hoeben, R.C.; Boer, G.J.; Verhaagen, J. Neuroregenerative effects of lentiviral vector-mediated GDNF expression in reimplanted ventral roots. Mol. Cell. Neurosci. 2008, 39, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Ongeri, E.M.; Verderame, M.F.; Hammond, J.M. Follicle-stimulating hormone induction of ovarian insulin-like growth factor-binding protein-3 transcription requires a TATA box-binding protein and the protein kinase A and phosphatidylinositol-3 kinase pathways. Mol. Endocrinol. 2005, 19, 1837–1848. [Google Scholar] [CrossRef][Green Version]
- Chen, S.F.; Zhou, Y.Q.; Chen, Y.R.; Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, 884–890. [Google Scholar] [CrossRef]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. clusterprofiler: An R package for comparing biological themes among gene clusters. OMICS: A J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Anders, S.; Reyes, A.; Huber, W. Detecting differential usage of exons from RNA-seq data. Nat. Précéd. 2012, 22, 2008–2017. [Google Scholar]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Lorkova, L.; Pospisilova, J.; Lacheta, J.; Leahomschi, S.; Zivny, J.; Cibula, D.; Zivny, J.; Petrak, J. Decreased concentrations of retinol-binding protein 4 in sera of epithelial ovarian cancer patients: A potential biomarker identified by proteomics. Oncol. Rep. 2012, 27, 318–324. [Google Scholar] [PubMed]
- Jeon, Y.E.; Lee, K.E.; Jung, J.A.; Yim, S.Y.; Kim, H.; Seo, S.K.; Cho, S.; Choi, Y.S.; Lee, B.S. Kisspeptin, leptin, and retinol-binding protein 4 in women with polycystic ovary syndrome. Gynecol. Obstet. Investig. 2013, 75, 268–274. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.S.; Liu, C.D.; Zhang, N.W.; Wang, S.D.; Zhang, Z.Y. Proteomics analysis for finding serum markers of ovarian cancer. Biomed. Res. Int. 2014, 2014, 179040. [Google Scholar] [CrossRef] [PubMed]
- Salilew-Wondim, D.; Wang, Q.; Tesfaye, D.; Schellander, K.; Hoelker, M.; Hossain, M.M.; Tsang, B.K. Polycystic ovarian syndrome is accompanied by repression of gene signatures associated with biosynthesis and metabolism of steroids, cholesterol and lipids. J. Ovarian Res. 2015, 8, 24. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Cao, G.; Zhang, N.; Lou, T.; Wang, Q.; Zhang, Z.; Liu, C. RBP4 regulates trophoblastic cell proliferation and invasion via the PI3K/AKT signaling pathway. Mol. Med. Rep. 2018, 18, 2873–2879. [Google Scholar] [CrossRef] [PubMed]
- Ghanem, A.; Lu, Y.H.; Cai, T.Y.; Mu, X.Z. Overexpression of RBP4 promotes proliferation, differentiation and mineralization of MC3T3-E1. Int. J. Clin. Exp. Pathol. 2017, 10, 298–305. [Google Scholar]
- Asselin, É.; Xiao, C.W.; Wang, Y.F.; Tsang, B.K. Mammalian follicular development and atresia: Role of apoptosis. Neurosignals 2000, 9, 87–95. [Google Scholar] [CrossRef]
- Cree-Green, M.; Newcomer, B.R.; Coe, G.; Newnes, L.; Baumgartner, A.; Brown, M.S.; Pyle, L.; Reusch, J.E.; Nadeau, K.J. Peripheral insulin resistance in obese girls with hyperandrogenism is related to oxidative phosphorylation and elevated serum free fatty acids. Am. J. Physiol. Metab. 2015, 308, E726–E733. [Google Scholar] [CrossRef]
- Das, D.; Arur, S. Conserved insulin signaling in the regulation of oocyte growth, development, and maturation. Mol. Reprod. Dev. 2017, 84, 444–459. [Google Scholar] [CrossRef] [PubMed]
- Itami, N.; Munakata, Y.; Shirasuna, K.; Kuwayama, T.; Iwata, H. Promotion of glucose utilization by insulin enhances granulosa cell proliferation and developmental competence of porcine oocyte grown in vitro. Zygote 2017, 25, 65–74. [Google Scholar] [CrossRef] [PubMed]
- Shafiee, M.N.; Seedhouse, C.; Mongan, N.; Chapman, C.; Deen, S.; Abu, J.; Atiomo, W. Up-regulation of genes involved in the insulin signalling pathway (IGF1, PTEN and IGFBP1) in the endometrium may link polycystic ovarian syndrome and endometrial cancer. Mol. Cell. Endocrinol. 2016, 424, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Wang, S.; Zhang, Z.; Lin, Q.; Liu, Y.; Xiao, Y.; Xiao, K.; Wang, Z. Defective insulin signaling and the protective effects of dimethyldiguanide during follicular development in the ovaries of polycystic ovary syndrome. Mol. Med. Rep. 2017, 16, 8164–8170. [Google Scholar] [CrossRef] [PubMed]
- Elis, S.; Desmarchais, A.; Maillard, V.; Uzbekova, S.; Monget, P.; Dupont, J. Cell proliferation and progesterone synthesis depend on lipid metabolism in bovine granulosa cells. Theriogenology 2015, 83, 840–853. [Google Scholar] [CrossRef]
- Xing, Y.; Liu, Y.; Liu, X.; Wang, S.; Li, P.; Lin, X.; Sui, C.; Xu, C.; Qi, B.; Tong, Q. Effects of Gui Zhu Yi Kun formula on the P53/AMPK pathway of autophagy in granulosa cells of rats with polycystic ovary syndrome. Exp. Ther. Med. 2017, 13, 3567–3573. [Google Scholar] [CrossRef] [PubMed]
- Yong, E.L.; Baird, D.T.; Yates, R.; Hillier, S.G.; Reichert, L.E. Hormonal regulation of the growth and steroidogenic function of human granulosa cells. J. Clin. Endocrinol. Metab. 1992, 74, 842–849. [Google Scholar] [CrossRef] [PubMed]
- Conover, C.A.; Faessen, G.F.; Ilg, K.E.; Chandrasekher, Y.A.; Christiansen, M.; Overgaard, M.T.; Oxvig, C.; Giudice, L.C. Pregnancy-associated plasma protein-a is the insulin-like growth factor binding protein-4 protease secreted by human ovarian granulosa cells and is a marker of dominant follicle selection and the corpus luteum. Endocrinology 2001, 142, 2155–2158. [Google Scholar] [CrossRef]
- Spicer, L.; Aad, P. Insulin-like growth factor (IGF) 2 stimulates steroidogenesis and mitosis of bovine granulosa cells through the IGF1 receptor: Role of follicle-stimulating hormone and IGF2 receptor1. Biol. Reprod. 2007, 77, 18–27. [Google Scholar] [CrossRef]
- Alemi Yahya, M.; Meisel Sharon, S.; Hantisteanu, S.; Hallak, M.; Werner, H.; Bruchim, I. The proliferative effect of dendritic cells in ovarian cancer and the relationship with the Igf signaling pathway. Harefuah 2019, 158, 30–34. [Google Scholar]
- Ellis, R.J. The general concept of molecular chaperones. Philos. Trans. R Soc. Lond. B Biol. Sci. 1993, 339, 257–261. [Google Scholar] [PubMed]
- Velázquez, M.M.; Alfaro, N.S.; Salvetti, N.R.; Stangaferro, M.L.; Rey, F.; Panzani, C.G.; Ortega, H.H. Levels of heat shock protein transcripts in normal follicles and ovarian follicular cysts. Reprod. Biol. 2011, 11, 276–283. [Google Scholar] [CrossRef]
- Malumbres, M.; Barbacid, M. Cell cycle, CDKs and cancer: A changing paradigm. Nat. Rev. Cancer 2009, 9, 153–166. [Google Scholar] [CrossRef] [PubMed]
- Hatzirodos, N.; Hummitzsch, K.; Irving-Rodgers, H.F.; Harland, M.L.; Morris, S.E.; Rodgers, R.J. Transcriptome profiling of granulosa cells from bovine ovarian follicles during atresia. BMC Genom. 2014, 15, 40. [Google Scholar]
| Gene Symbol | Gene Name | Description and Summary of the Function |
|---|---|---|
| Upregulated | ||
| RBP4 | Plasma retinol binding protein 4 | Plays a role in retinol transport; involved in insulin resistance and diabetes |
| HSPB1 | Heat shock protein 27 | Provides thermotolerance in vivo and support cell survival under stress conditions |
| MMP1 | Matrix metallopeptidase 1 | Involved in the breakdown of extracellular matrix |
| RBM34 | RNA binding motif protein 34 | Ubiquitous expression in testis, lymph node, and other tissues |
| S100A12 | S100 calcium-binding protein A12 | Involved in the regulation of cell cycle progression and differentiation and specific calcium-dependent signal transduction |
| ITGA5 | Integrin α-5 | Participates in cell-surface mediated signaling |
| TOMM6 | Mitochondrial import receptor subunit TOM6 | May play a role in breast carcinogenesis |
| KIF20B | kinesin-like protein 20A | Could promote cancer progression and is essential for cytokinesis in cell cycle |
| CCDC33 | Coiled-coil domain-containing protein 33 | May be regulated by hormone and nutrition |
| MITD1 | Microtubule interacting and trafficking domain containing 1 | Participates in the abscission phase of cytokinesis |
| Downregulated | ||
| KCNMA1 | Potassium calcium-activated channel subfamily M α 1 | Relates with several cancers, such as breast carcinoma, prostate cancer, and cervical cancers |
| IGFALS | Insulin like growth factor binding protein acid labile subunit | Act as IGFBPs, binding IGFs exerts its function |
| LCN2 | Lipocalin-2 | Estradiol can regulate the expression of LCN2 and is linked to IR |
| WIPF3 | Wasl interacting protein family member 3 | May be a regulator of cytoskeletal organization and play a role in spermatogenesis |
| CD248 | Endosialin | Essential for embryo development |
| CRTC3 | Creb-regulated transcription coactivator 3 | Transcriptional coactivator; plays roles in adipose development and energy metabolism |
| MAP1B | Microtubule-associated protein 1B | Plays an important role in development and function of the nervous system |
| RAB3B | Ras-related protein-3B | May play a role in regulating GLUT4 translocation in adipocytes |
| CPAMD8 | C3 and pzp-like α-2-macroglobulin domain-containing protein 8 | May be associated with cataract development |
| PRRC2C | Proline rich coiled-coil 2C | Involved in the development of basement membrane |
| GO Terms | Gene Name | Gene Ratio | p-Value |
|---|---|---|---|
| Biological process (BP) | |||
| GO:0006457-protein folding | HSPB1, PFDN4, PFDN2, PRDX4 | 6.3% | 0.000427 |
| GO:0006887-exocytosis | KIT, RAB3B | 2.7% | 0.00471 |
| GO:0032652-regulation of interleukin-1 production | HSPB1, ANXA1 | 10.5% | 0.00508 |
| GO:0016477-cell migration | KIT, PEAK1, CD248 | 1.1% | 0.00512 |
| GO:0032612-interleukin-1 production | HSPB1, ANXA1 | 10.0% | 0.00562 |
| GO:0048870-cell motility | KIT, PEAK1, CD248 | 1.0% | 0.00688 |
| GO:0051674-localization of cell | KIT, PEAK1, CD248 | 1.0% | 0.00688 |
| GO:0040011-locomotion | KIT, PEAK1, CD248 | 0.9% | 0.0107 |
| GO:0006928-movement of cell or subcellular component | KIT, PEAK1, CD248 | 0.9% | 0.0117 |
| GO:0000278-mitotic cell cycle | MITD1, CDK1, NUF2, RPL26 | 6.3% | 0.0127 |
| Molecular function (MF) | |||
| GO:0005520-insulin-like growth factor binding | IGFBP5, IGFALS | 12.5% | 0.000330 |
| GO:0044183-protein binding involved in protein folding | HSPB1, PFDN2 | 20.0% | 0.00137 |
| GO:0042803-protein homodimerization activity | HSPB1, MITD1, CACYBP, AIMP1, PRDX4 | 3.3% | 0.00148 |
| GO:0003735-structural constituent of ribosome | MRPL52, RPL26, RPL26L1, RPL23 | 4.4% | 0.00155 |
| GO:0019838-growth factor binding | IGFBP5, IGFALS | 5.7% | 0.00161 |
| GO:0004713-protein tyrosine kinase activity | KIT, PEAK1 | 4.3% | 0.00277 |
| GO:0016684-oxidoreductase activity, acting on peroxide as acceptor | GPX8, PRDX4 | 9.1% | 0.00675 |
| GO:0016209-antioxidant activity | GPX8, PRDX4 | 5.7% | 0.0166 |
| GO: 0051082-unfolded protein binding | PFDN4, PFDN2 | 5.7% | 0.0166 |
| GO:0046983-protein dimerization activity | HSPB1, MITD1, CACYBP, AIMP1, PRDX4 | 1.9% | 0.0169 |
| Cellular component (CC) | |||
| GO:0015934-large ribosomal subunit | MRPL52, RPL26, RPL26L1, RPL23 | 7.8% | 0.000231 |
| GO:0044391-ribosomal subunit | MRPL52, RPL26, RPL26L1, RPL23 | 4.9% | 0.00135 |
| GO:0005739-mitochondrion | MRPL50, NDUFB3, COX17, MTFP1, MRPL52, COX6C, ATP5I, PFDN2, PRDX4 | 1.9% | 0.00141 |
| GO:0005840-ribosome | MRPL52, RPL26, RPL26L1, RPL23 | 3.9% | 0.00327 |
| GO:0044429-mitochondrial part | NDUFB3, COX17, MTFP1, MRPL52, COX6C, ATP5I | 2.2% | 0.00490 |
| GO:0031967-organelle envelope | NDUFB3, CACYBP, COX17, MTFP1, COX6C, ATP5I | 2.2% | 0.00567 |
| GO:0031975-envelope | NDUFB3, CACYBP, COX17, MTFP1, COX6C, ATP5I | 2.2% | 0.00567 |
| GO: 0005740-mitochondrial envelope | NDUFB3, COX17, MTFP1, COX6C, ATP5I | 2.5% | 0.00660 |
| GO: 0031970-organelle envelope lumen | CACYBP, COX17 | 9.5% | 0.00700 |
| GO:0005743-mitochondrial inner membrane | NDUFB3, MTFP1, COX6C, ATP5I | 3.1% | 0.00749 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, Y.; Li, C.; Zhou, X. Transcriptomic Analysis of Porcine Granulosa Cells Overexpressing Retinol Binding Protein 4. Genes 2019, 10, 615. https://doi.org/10.3390/genes10080615
Zhao Y, Li C, Zhou X. Transcriptomic Analysis of Porcine Granulosa Cells Overexpressing Retinol Binding Protein 4. Genes. 2019; 10(8):615. https://doi.org/10.3390/genes10080615
Chicago/Turabian StyleZhao, Yun, Chunjin Li, and Xu Zhou. 2019. "Transcriptomic Analysis of Porcine Granulosa Cells Overexpressing Retinol Binding Protein 4" Genes 10, no. 8: 615. https://doi.org/10.3390/genes10080615
APA StyleZhao, Y., Li, C., & Zhou, X. (2019). Transcriptomic Analysis of Porcine Granulosa Cells Overexpressing Retinol Binding Protein 4. Genes, 10(8), 615. https://doi.org/10.3390/genes10080615





