A Guide on Deep Learning for Complex Trait Genomic Prediction
Abstract
1. Introduction
2. Genomic Prediction Principles and Current Limitations
3. Deep Learning Principles
3.1. Main Deep Learning Architectures
3.2. Algorithms and Optimization Issues
| Algorithm 1: Stochastic Gradient Descent (SGD) |
| Given the loss function l, |
| Input: Training sample (x(train), y(train)), regularization parameters , learning rate (α) and a penalization parameter. Initialize |
| Output: Model parameters |
| Initialize |
| repeat |
| Given a training set (x(train), y(train)), |
| Compute: |
| + |
| end |
| until stopping criteria/convergence; |
| where is the function gradient |
3.3. Avoiding Overfitting
4. Deep Learning Practicalities
4.1. Implementing Deep Learning
- A model is instantiated. The most usual model is “sequential”, which allows adding layers with different properties step by step.
- The architecture is defined. Here, each layer and its properties are defined. For each layer, the number of neurons, activation functions, regularization, and initialization methods are specified.
- The model is compiled. The optimizer algorithm with associated parameters (e.g., learning rate) and loss function are specified. This step allows us to symbolically define the operations (i.e., graphs) to be performed later with actual numbers.
- Training occurs. The model is fitted to the data and parameters are estimated. The number of iterations (i.e., epochs) and batch size are specified. The input and target variables need to be provided. The input data size must match that defined in step 2.
- Model predictions are validated via cross-validation.
4.2. Training and Validating Deep Learning Architectures
4.3. Feature Selection
4.4. Interpreting DL
5. Applications of Deep Learning to Genomic Prediction
6. Perspectives and Conclusion
7. Practical Recommendations
- Before starting, inspect the data, both SNPs and phenotypic distributions. Look for unexpected, weird patterns that may cause biases. Standardize the variables and targets.
- Use Keras with TensorFlow, together with Sci-Kit Learn, a collection of well documented, easy-to-use machine learning modules. Reuse, but test, available public software whenever possible.
- Exercise prudence if extremely good or poor results are obtained. Ample literature does support that differences between methods should not be dramatic.
- Do not be too ambitious. Is your data set big enough to fit such complex models?
- Dedicate enough time and thinking to optimize hyperparameters. Finely tune early, stopping to improve prediction performance. If the number of SNPs is too large, preselect different subsets according to the p-value or try other criteria.
- Once an optimum hyperparameter set has been decided, restart the algorithm several times to assess the influence of initial values.
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fisher, R.A. The Correlation between Relatives on the Supposition of Mendelian Inheritance. Trans. R. Soc. Edinb. 1918, 52, 399–433. [Google Scholar] [CrossRef]
- Meuwissen, T.H.E.; Hayes, B.J.; Goddard, M.E. Prediction of total genetic value using genome-wide dense marker maps. Genetics 2001, 157, 1819–1829. [Google Scholar] [PubMed]
- Gianola, D. Priors in whole-genome regression: The Bayesian alphabet returns. Genetics 2013, 194, 573–596. [Google Scholar] [CrossRef] [PubMed]
- Grattapaglia, D.; Silva-Junior, O.B.; Resende, R.T.; Cappa, E.P.; Müller, B.S.F.; Tan, B.; Isik, F.; Ratcliffe, B.; El-Kassaby, Y.A. Quantitative Genetics and Genomics Converge to Accelerate Forest Tree Breeding. Front. Plant Sci. 2018, 9, 1693. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Benyamin, B.; McEvoy, B.P.; Gordon, S.; Henders, A.K.; Nyholt, D.R.; Madden, P.A.; Heath, A.C.; Martin, N.G.; Montgomery, G.W.; et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 2010, 42, 565–569. [Google Scholar] [CrossRef] [PubMed]
- Campos, G.D.L.; Gianola, D.; Allison, D.B. Predicting genetic predisposition in humans: The promise of whole-genome markers. Nat. Rev. Genet. 2010, 11, 880–886. [Google Scholar] [CrossRef]
- Meuwissen, T.; Goddard, M. Accurate Prediction of Genetic Values for Complex Traits by Whole-Genome Resequencing. Genetics 2010, 185, 623–631. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Enciso, M.; Rincón, J.C.; Legarra, A. Sequence- vs. chip-assisted genomic selection: Accurate biological information is advised. Genet. Sel. Evol. 2015, 47, 1–14. [Google Scholar] [CrossRef]
- Heidaritabar, M.; Calus, M.P.L.; Megens, H.-J.; Vereijken, A.; Groenen, M.A.M.; Bastiaansen, J.W.M. Accuracy of genomic prediction using imputed whole-genome sequence data in white layers. J. Anim. Breed. Genet. 2016, 133, 167–179. [Google Scholar] [CrossRef]
- Ainscough, B.J.; Barnell, E.K.; Ronning, P.; Campbell, K.M.; Wagner, A.H.; Fehniger, T.A.; Dunn, G.P.; Uppaluri, R.; Govindan, R.; Rohan, T.E.; et al. A deep learning approach to automate refinement of somatic variant calling from cancer sequencing data. Nat. Genet. 2018, 50, 1735–1743. [Google Scholar] [CrossRef]
- Sundaram, L.; Gao, H.; Padigepati, S.R.; McRae, J.F.; Li, Y.; Kosmicki, J.A.; Fritzilas, N.; Hakenberg, J.; Dutta, A.; Shon, J.; et al. Predicting the clinical impact of human mutation with deep neural networks. Nat. Genet. 2018, 50, 1161–1170. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Theesfeld, C.L.; Yao, K.; Chen, K.M.; Wong, A.K.; Troyanskaya, O.G. Deep learning sequence-based ab initio prediction of variant effects on expression and disease risk. Nat. Genet. 2018, 50, 1171–1179. [Google Scholar] [CrossRef] [PubMed]
- Eraslan, G.; Avsec, Ž.; Gagneur, J.; Theis, F.J. Deep learning: New computational modelling techniques for genomics. Nat. Rev. Genet. 2019, 20, 389–403. [Google Scholar] [CrossRef] [PubMed]
- Zou, J.; Huss, M.; Abid, A.; Mohammadi, P.; Torkamani, A.; Telenti, A. A primer on deep learning in genomics. Nat. Genet. 2018, 51, 12–18. [Google Scholar] [CrossRef] [PubMed]
- Gianola, D.; Okut, H.; Weigel, K.A.; Rosa, G.J. Predicting complex quantitative traits with Bayesian neural networks: A case study with Jersey cows and wheat. BMC Genet. 2011, 12, 87. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Rodríguez, P.; Gianola, D.; González-Camacho, J.M.; Crossa, J.; Manès, Y.; Dreisigacker, S. Comparison between linear and non-parametric regression models for genome-enabled prediction in wheat. G3 Genes Genomes Genet. 2012, 2, 1595–1605. [Google Scholar] [CrossRef]
- González-Recio, O.; Rosa, G.J.M.; Gianola, D. Machine learning methods and predictive ability metrics for genome-wide prediction of complex traits. Livest. Sci. 2014, 166, 217–231. [Google Scholar] [CrossRef]
- Pérez, P.; Campos, G.D.L. Genome-Wide Regression & Prediction with the BGLR Statistical Package. Genetics 2014, 198, 483–495. [Google Scholar]
- Tibshirani, R. Regression Shrinkage and Selection Via the Lasso. J. R. Stat. Soc. Ser. B 1996, 58, 267–288. [Google Scholar] [CrossRef]
- VanRaden, P.M. Efficient methods to compute genomic predictions. J. Dairy Sci. 2008, 91, 4414–4423. [Google Scholar] [CrossRef]
- White, B.W.; Rosenblatt, F. Principles of Neurodynamics: Perceptrons and the Theory of Brain Mechanisms. Am. J. Psychol. 1963, 76, 705–707. [Google Scholar] [CrossRef]
- Efron, B.; Hastie, T. Computer Age Statistical Inference: Algorithms, Evidence, and Data Science; Cambridge University Press: New York, NY, USA, 2016; Volume 5, p. 475. [Google Scholar]
- Lecun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Goodfellow, I.; Bengio, Y.; Courville, A. Deep Learning; MIT Press Cambridge: Cambridge, MA, USA, 2016. [Google Scholar]
- Hill, S.T.; Kuintzle, R.; Teegarden, A.; Merrill, E.; Danaee, P.; Hendrix, D.A.; Hendrix, D.A. A deep recurrent neural network discovers complex biological rules to decipher RNA protein-coding potential. Nucleic Acids Res. 2018, 46, 8105–8113. [Google Scholar] [CrossRef] [PubMed]
- Hochreiter, S.; Urgen Schmidhuber, J. Long Short-Term Memory. Neural Comput. 1997, 9, 1735–1780. [Google Scholar] [CrossRef] [PubMed]
- Patterson, J.; Gibson, A. Deep Learning: A Practitioner’s Approach; O’Reilly Media: Sebastopol, CA, USA, 2017; ISBN 978-1-4919-1425-0. [Google Scholar]
- Pouladi, F.; Salehinejad, H.; Gilani, A.M. Deep Recurrent Neural Networks for Sequential Phenotype Prediction in Genomics. arXiv 2016, arXiv:1511.02554. [Google Scholar]
- Bishop, C.M.; Lasserre, J. Generative or discriminative? Getting the best of both worlds. Bayesian Stat. 2007, 8, 3–24. [Google Scholar]
- Hinton, G.E.; Sejnowski, T.J. Optimal perceptual inference. In Proceedings of the IEEE conference on Computer Vision and Pattern Recognition, Washington, DC, USA, 19–23 June 1983; pp. 448–453. [Google Scholar]
- Hinton, G.E.; Osindero, S.; Teh, Y.-W. A Fast Learning Algorithm for Deep Belief Nets. Neural Comput. 2006, 18, 1527–1554. [Google Scholar] [CrossRef]
- Salakhutdinov, R.; Hinton, G. Deep boltzmann machines. Artif. Intell. Stat. 2009, 5, 448–455. [Google Scholar]
- Goodfellow, I.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. arXiv 2014, arXiv:1406.2661. [Google Scholar]
- Rumelhart, D.E.; Hinton, G.E.; Williams, R.J. Learning representations by back-propagating errors. Nature 1986, 323, 533–536. [Google Scholar] [CrossRef]
- Cauchy, A.-L. Methode generale pour la resolution des systemes d’equations simultanees. Compte Rendu des Seances L’Acad’emie des Sci. 1847, 25, 536–538. [Google Scholar]
- Pai, C.; Potdar, K. A Comparative Study of Categorical Variable Encoding Techniques for Neural Network Classifiers. Artic. Int. J. Comput. Appl. 2017, 175, 7–9. [Google Scholar]
- Waldmann, P. Approximate Bayesian neural networks in genomic prediction. Genet. Sel. Evol. 2018, 50, 70. [Google Scholar] [CrossRef] [PubMed]
- Bellot, P.; Campos, G.D.L.; Pérez-Enciso, M. Can Deep Learning Improve Genomic Prediction of Complex Human Traits? Genetics 2018, 210, 809–819. [Google Scholar] [CrossRef] [PubMed]
- Chollet, F. Keras: Deep Learning Library for Theano and Tensorflow; Manning: Shelter Island, NY, USA, 2015; p. 334. [Google Scholar]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems. arXiv 2016, arXiv:1603.04467. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Moncecchi, G.; Garreta, R. Learning Scikit-Learn: Machine Learning in Python; Packt Publishing: Birmingham, UK, 2013; ISBN 9781783281930. [Google Scholar]
- Boopathi, V.; Subramaniyam, S.; Malik, A.; Lee, G.; Manavalan, B.; Yang, D.-C. mACPpred: A Support Vector Machine-Based Meta-Predictor for Identification of Anticancer Peptides. Int. J. Mol. Sci. 2019, 20, 1964. [Google Scholar] [CrossRef]
- Tohka, J.; Moradi, E.; Huttunen, H.; Alzheimer’s Disease Neuroimaging Initiative. Comparison of Feature Selection Techniques in Machine Learning for Anatomical Brain MRI in Dementia. Neuroinformatics 2016, 14, 279–296. [Google Scholar] [CrossRef]
- Kuhn, M.; Johnson, K. Applied Predictive Modeling; Springer: New York, NY, USA, 2013; ISBN 978-1-4614-6848-6. [Google Scholar]
- Shmueli, G. To Explain or to Predict? Stat. Sci. 2010, 25, 289–310. [Google Scholar] [CrossRef]
- Sheehan, S.; Song, Y.S. Deep Learning for Population Genetic Inference. PLoS Comput. Biol. 2016, 12, 1–28. [Google Scholar] [CrossRef]
- Schwab, P.; Miladinovic, D.; Karlen, W. Granger-causal Attentive Mixtures of Experts: Learning Important Features with Neural Networks. arXiv 2018, arXiv:1802.02195. [Google Scholar]
- Dhurandhar, A.; Shanmugam, K.; Luss, R.; Olsen, P. Improving Simple Models with Confidence Profiles. arxiv 2018, arXiv:1807.07506. [Google Scholar]
- Reichstein, M.; Camps-Valls, G.; Stevens, B.; Jung, M.; Denzler, J.; Carvalhais, N.; Prabhat. Deep learning and process understanding for data-driven Earth system science. Nature 2019, 566, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Mcdowell, R.M. Genomic Selection with Deep Neural Networks. Master’s Thesis, Iowa State University, Digital Repository, Ames, IA, USA, 2016. [Google Scholar]
- Montesinos-López, A.; Montesinos-López, O.A.; Gianola, D.; Crossa, J.; Hernández-Suárez, C.M. Multi-environment Genomic Prediction of Plant Traits Using Deep Learners with Dense Architecture. G3 Genes Genomes Genet. 2018, 8, 3813–3828. [Google Scholar] [CrossRef] [PubMed]
- Montesinos-López, O.A.; Martín-Vallejo, J.; Crossa, J.; Gianola, D.; Hernández-Suárez, C.M.; Montesinos-López, A.; Juliana, P.; Singh, R. A Benchmarking Between Deep Learning, Support Vector Machine and Bayesian Threshold Best Linear Unbiased Prediction for Predicting Ordinal Traits in Plant Breeding. G3 Genes Genomes Genet. 2019, 9, 601–618. [Google Scholar] [CrossRef] [PubMed]
- Khaki, S.; Wang, L. Crop Yield Prediction Using Deep Neural Networks. arXiv 2019, arXiv:1902.02860. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wang, D. Application of deep learning in genomic selection. In Proceedings of the 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Kansas City, MO, USA, 13–16 November 2017; p. 2280. [Google Scholar]
- Rachmatia, H.; Kusuma, W.A.; Hasibuan, L.S. Prediction of maize phenotype based on whole-genome single nucleotide polymorphisms using deep belief networks Related content Prediction of maize phenotype based on whole-genome single nucleotide polymorphisms using deep belief networks. J. Phys. Conf. 2017, 835, 12003. [Google Scholar] [CrossRef]
- Ma, W.; Qiu, Z.; Song, J.; Cheng, Q.; Ma, C. DeepGS: Predicting phenotypes from genotypes using Deep Learning. Planta 2017. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Adv. Neural Inf. Process. Syst. 2012, 60, 1097–1105. [Google Scholar] [CrossRef]
- Pattanayak, S. Pro Deep Learning with TensorFlow; Apress: New York, NY, USA, 2017; ISBN 978-1-4842-3095-4. [Google Scholar]
- Veerkamp, R.F.; Bouwman, A.C.; Schrooten, C.; Calus, M.P.L. Genomic prediction using preselected DNA variants from a GWAS with whole-genome sequence data in Holstein-Friesian cattle. Genet. Sel. Evol. 2016, 48, 1–14. [Google Scholar] [CrossRef]
| Term | Definition |
|---|---|
| Activation function | The mathematical function f that produces neuron’s output f(w’x + b). |
| Backpropagation | Backpropagation is an efficient algorithm to compute the loss, it propagates the error at the output layer level backward. |
| Batch | In stochastic gradient Ddescent (SGD) algorithm, each of the sample partitions within a given epoch. |
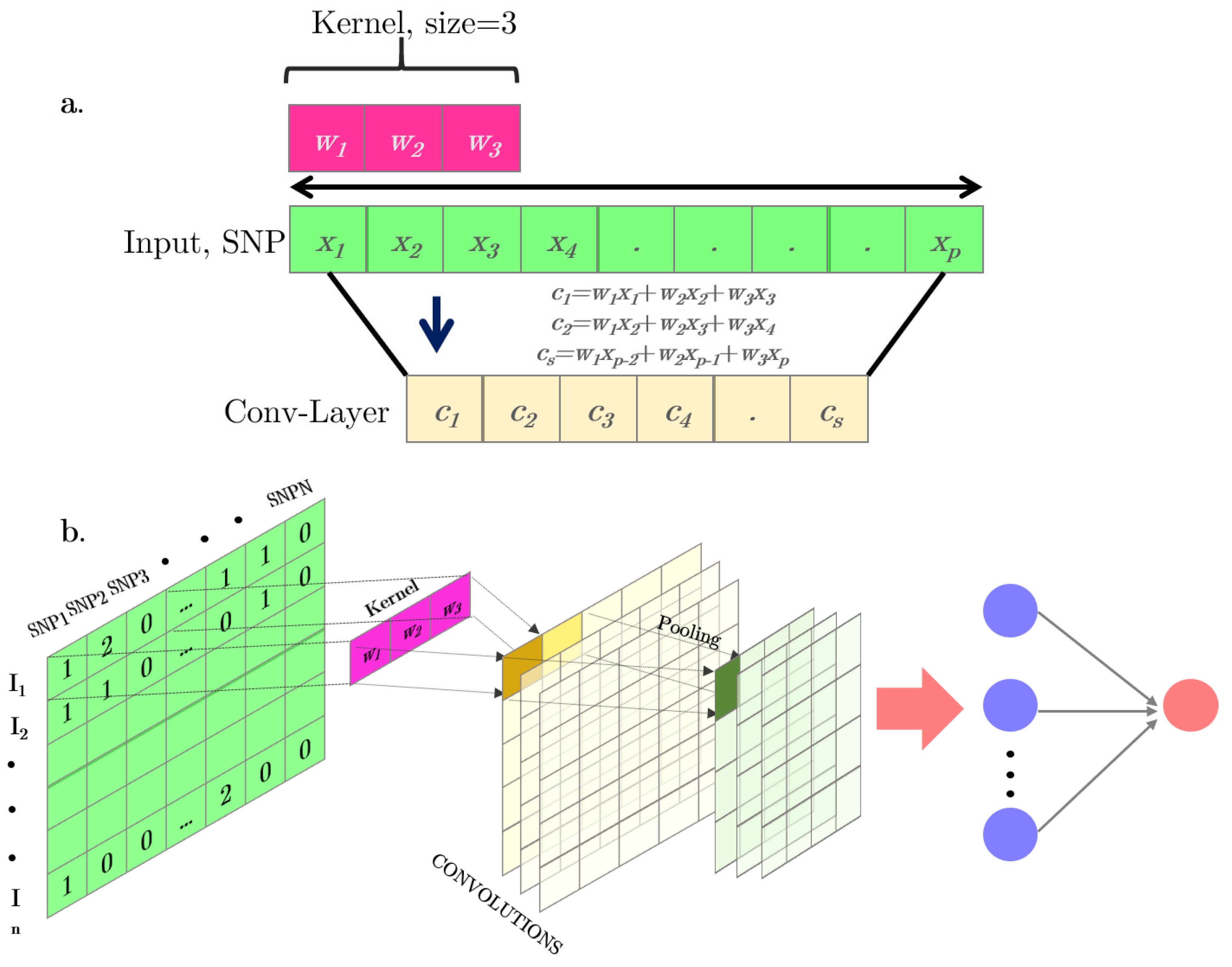
| Convolution | Mathematically, a convolution is defined as an “integral transform” between two functions, where one of the functions must be a kernel. The discrete version of the operation is simply the weighting sum of several copies of the original function (f) shifting over the kernel. |
| Convolutional neural network | A CNN is a special case of neural networks which uses convolution instead a full matrix multiplication in the hidden layers. A typical CNN is made up of dense fully connected layers and “convolutional layers”. |
| Dropout | Dropout means that a given percentage of neurons output is set to zero. The percentage is kept constant, but the specific neurons are randomly sampled in every iteration. The goal of dropout is to avoid overfitting. |
| Early stopping | An anti-overfitting strategy that consists of stopping the algorithm before it converges. |
| Epoch | In SGD and related algorithms, an iteration comprising all batches in a given partition. In the next epoch, a different partition is employed. |
| Kernel = Filter = Tensor | In DL terminology, the kernel is a multidimensional array of weights. |
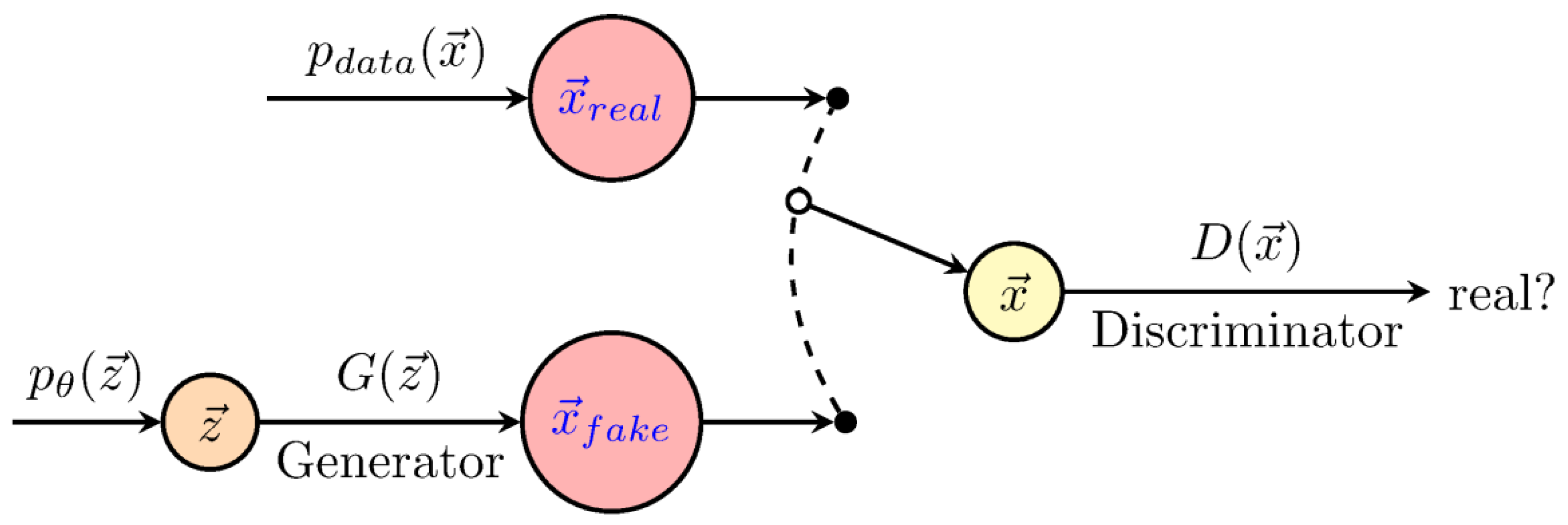
| Generative adversarial network (GAN) | GANs are based on a simple idea: train two networks simultaneously, the generator (G), which defines a probability distribution based on the information from the samples, and the discriminator (D), which distinguishes data produced by G from the real data. |
| Learning rate | Specify the speed of gradient update (α in Algorithm 1). |
| Loss | Loss function measures how differences between observed and predicted target variables are quantified. |
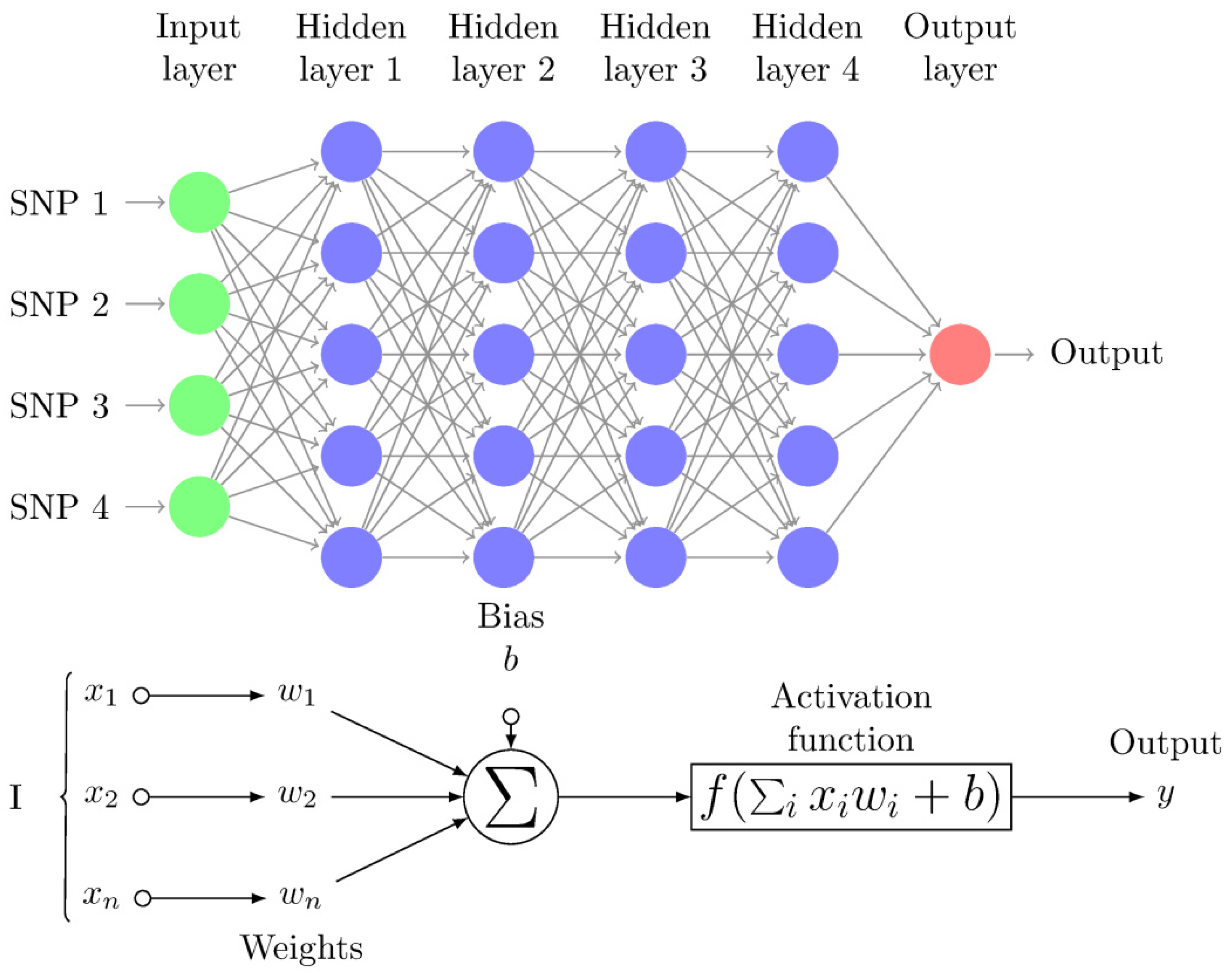
| Neuron | The basic unit of a DL algorithm. A “neuron” takes as input a list of variable values (x) multiplied by “weights” (w) and, as output, produces a non-linear transformation f(w’x + b) where f is the activation function and b is the bias. Both w and b need to be estimated for each neuron such that the loss is minimized across the whole set of neurons. |
| Neuron layer | “Neurons” are arranged in layers, i.e., groups of neurons that take the output of previous group of neurons as input (Figure 1). |
| Multilayer perceptron (MLP) | Multilayer perceptron network is one of the most popular NN architectures, which consists of a series of fully connected layers, called input, hidden, and output layers. The layers are connected by a directed graph. |
| Optimizer | An algorithm to find weights (w and b) that minimize the loss function. Most DL optimizers are based on stochastic gradient descent (SGD). |
| Pooling | A pooling function substitutes the output of a network at a certain location with a summary statistic of the neighboring outputs. This is one of the crucial steps on the CNN architecture. The most common pooling operations are maximum, mean, and median. |
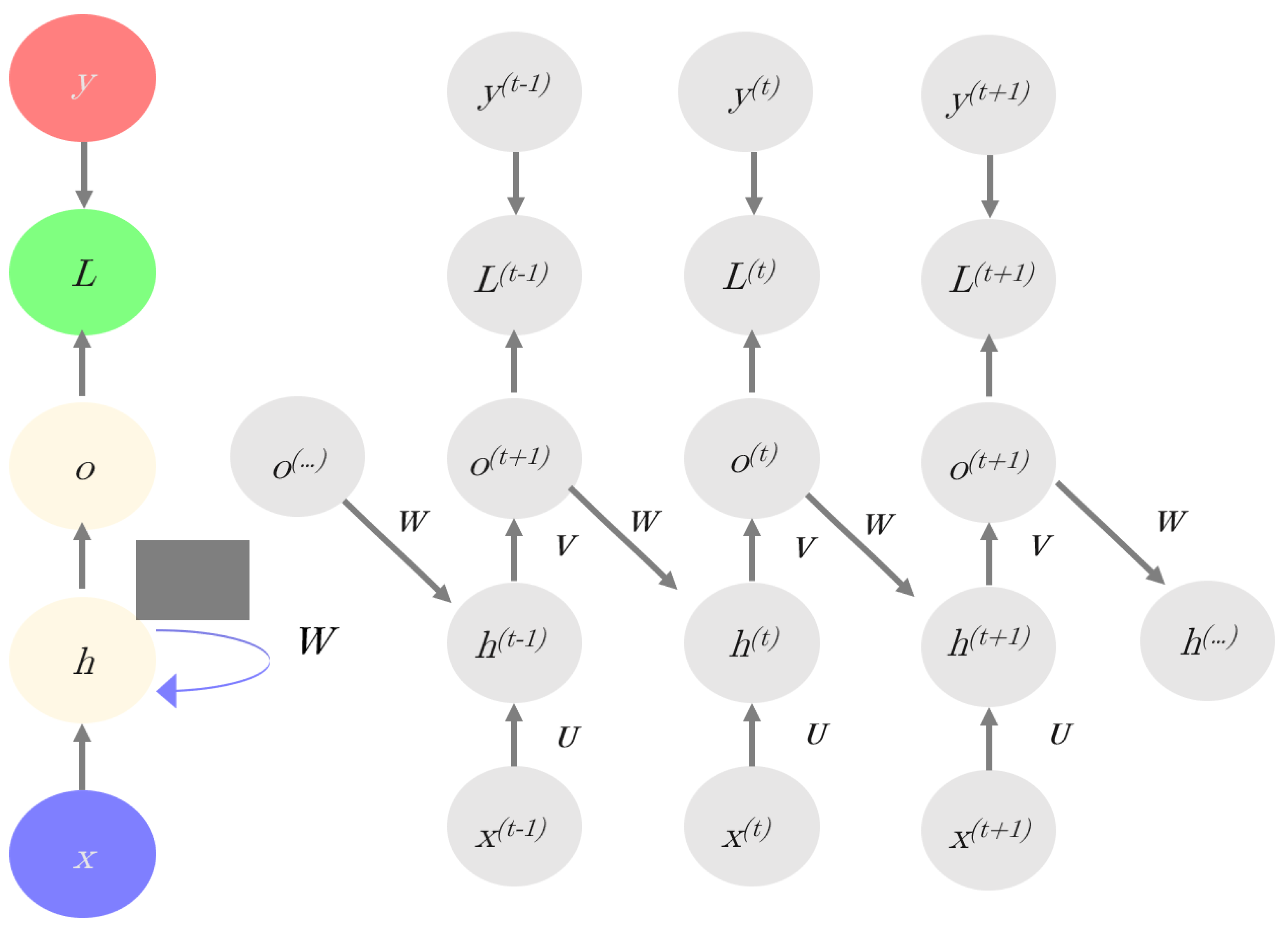
| Recurrent neural Network (RNN) | RNN architecture considers information from multiple previous layers. In an RNN, the current hidden layer is a nonlinear function of both the previous layer(s) and the current input (x). The model has memory since the bias term is based on the “past”. These networks can be used in temporal-like data structures. |
| Stochastic gradient descent (SGD) | An optimizing algorithm that consists of randomly partitioning the whole dataset into subsets called “batches” or “minibatches” and updates the gradient using only that data subset. The next batch is used in the next iteration. |
| Weight regularization | An excess of parameters (weights, w) may produce the phenomenon called “overfitting”, which means that the model adjusts to the observed data very well, but prediction of new unobserved data is very poor. To avoid this, weights are estimated subject to constraints, a strategy called “penalization” or “regularization”. The two most frequent regularizations are the L1 and L2 norms, which set restrictions on the sum of absolute values of w (L1) or of the square values (L2). |
| Study | Species | Approx. N | Approx. No. SNPs | Performance * |
|---|---|---|---|---|
| Mcdowell [51] | Arabidopsis, Maize, wheat | 270–400 | 70–1k | MLP ≥ PL |
| Liu and Wang [55] | Soybean | 5k | 4k | CNN > RR-BLUP, Lasso-Bayes, Bayes A |
| Rachmatia et al. [56] | Maize | 300 | 1k | PL > DBN |
| Bellot et al. [38] | Human | 100k | 10k–50k | PL ≥ CNN > MLP |
| Ma et al. [57] | Wheat | 2k | 33k | CNN ~ PL ~ GBLUP > MLP |
| Montesinos-López et al. [52] | Maize, wheat | 250–2k | 12k–160k | GBLUP > MLP |
| Montesinos-López et al. [53] | Wheat | 800–4k | 2k | GBLUP > MLP |
| Khaki and Wang [54] | Maize | 2k genotypes (150k samples) | 20k | DL > PL |
| Waldmann [37] | pig | 3226 (simulated) 3534 (real) | 10k, 50k | DL > GBLUP/BayesLasso |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pérez-Enciso, M.; Zingaretti, L.M. A Guide on Deep Learning for Complex Trait Genomic Prediction. Genes 2019, 10, 553. https://doi.org/10.3390/genes10070553
Pérez-Enciso M, Zingaretti LM. A Guide on Deep Learning for Complex Trait Genomic Prediction. Genes. 2019; 10(7):553. https://doi.org/10.3390/genes10070553
Chicago/Turabian StylePérez-Enciso, Miguel, and Laura M. Zingaretti. 2019. "A Guide on Deep Learning for Complex Trait Genomic Prediction" Genes 10, no. 7: 553. https://doi.org/10.3390/genes10070553
APA StylePérez-Enciso, M., & Zingaretti, L. M. (2019). A Guide on Deep Learning for Complex Trait Genomic Prediction. Genes, 10(7), 553. https://doi.org/10.3390/genes10070553





