Complex Gene Regulation Underlying Mineral Nutrient Homeostasis in Soybean Root Response to Acidity Stress
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material and Growth Conditions
2.2. Determination of Mineral Nutrient Concentration
2.3. Transcriptome Analysis of Soybean Roots
2.4. Quantitative Real-Time PCR Analysis
2.5. Statistical Analysis
3. Results
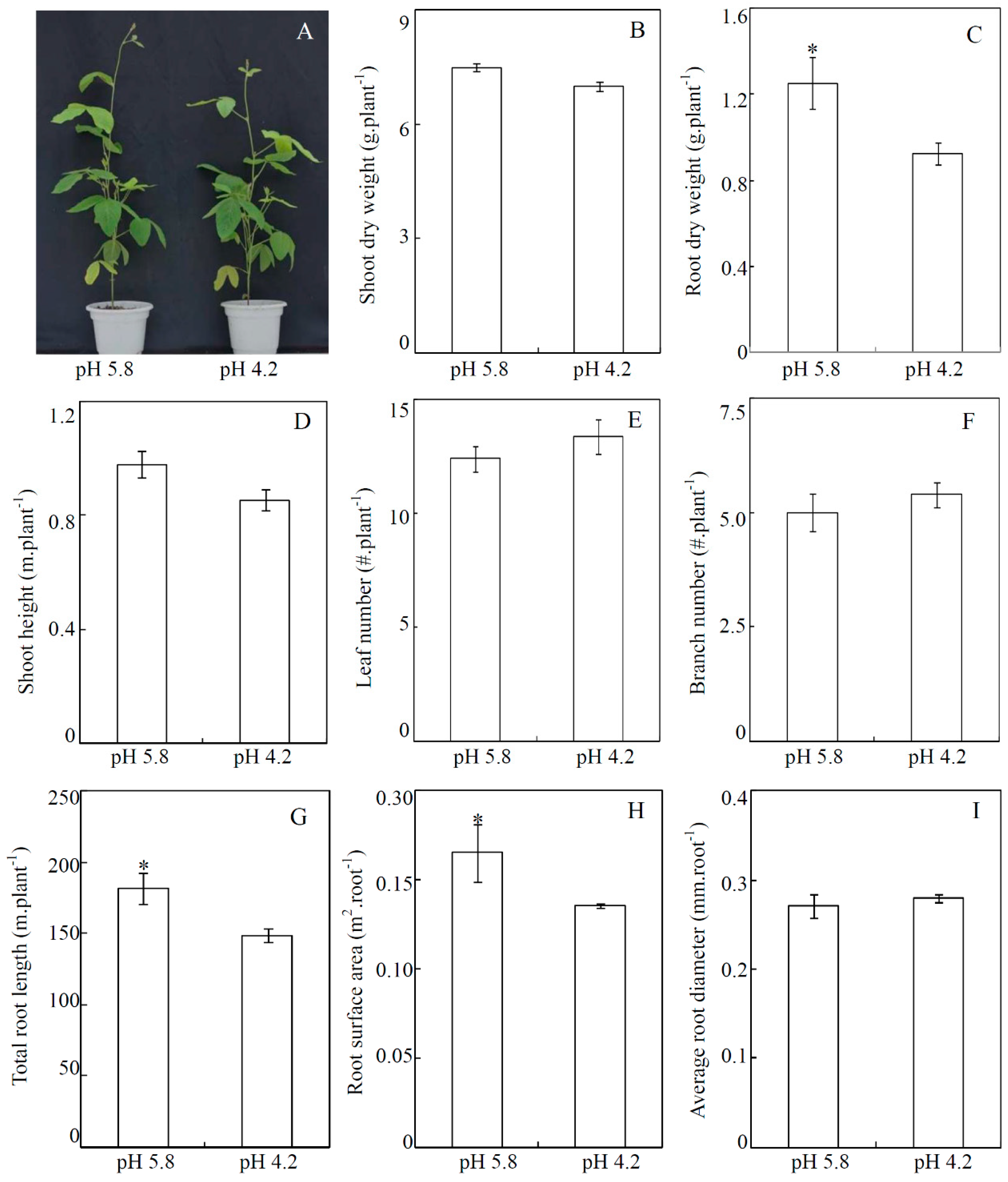
3.1. Effects of Low-pH Stress on Soybean Growth in Soils
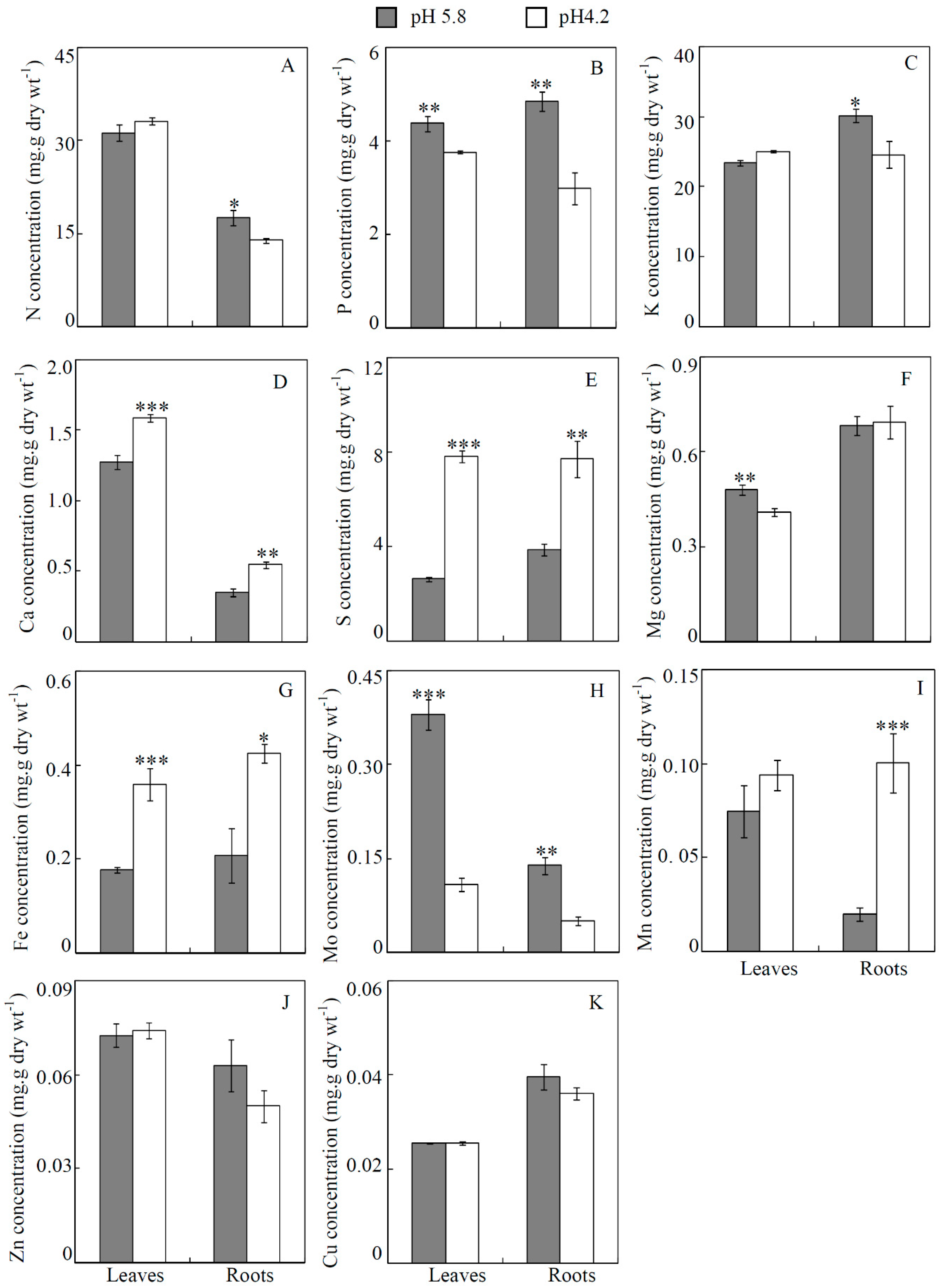
3.2. Concentration of Mineral Nutrients Affected by Acidity Stress
3.3. Growth and Mineral Nutrient Concentrations in Roots of Soybean Grown in Nutrient Solution
3.4. Transcriptomic Analysis of Soybean Roots Responding to Acidity Stress
3.5. Transcription Factors Involved in Soybean Responses to Acidity Stress
3.6. Identification of DEGs Involved in the pH Stat Pathway
3.7. DEGs Participated in Mineral Nutrients Acquisition and Translocation
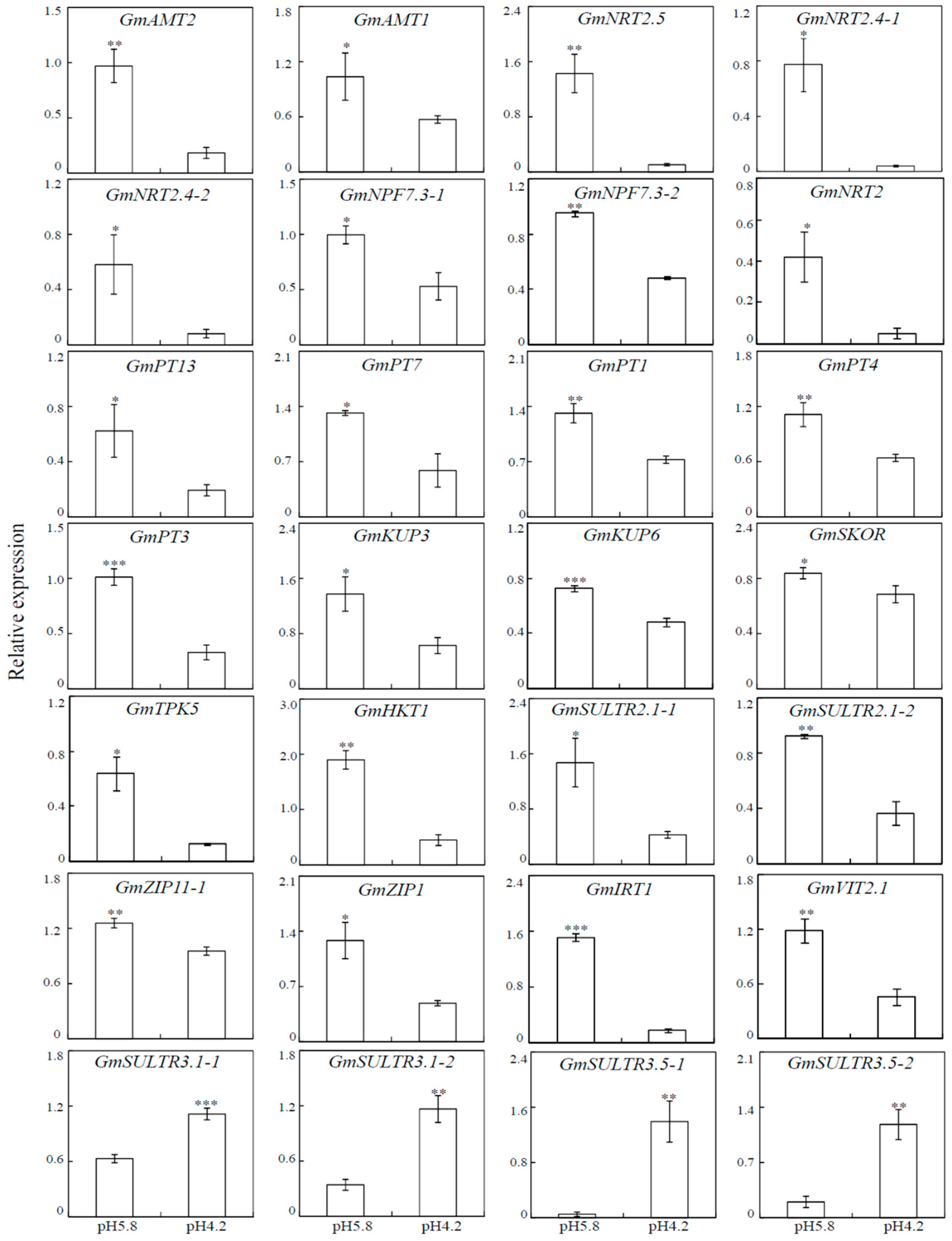
3.8. Transcripts Analysis of DEGs Involved in Nutrients Transportation Using qRT-PCR
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Von Uexküll, H.R.; Mutert, E. Global extent, development and economic impact of acid soils. Plant Soil 1995, 171, 1–15. [Google Scholar] [CrossRef]
- Kochian, L.V.; Hoekenga, O.A.; Pineros, M.A. How do crop plants tolerate acid soils? Mechanisms of aluminium tolerance and phosphorous efficiency. Annu. Rev. Plant Biol. 2004, 55, 459–493. [Google Scholar] [CrossRef] [PubMed]
- Sade, H.; Meriga, B.; Surapu, V.; Gadi, J.; Sunita, M.S.; Suravajhala, P.; Kavi Kishor, P.B. Toxicity and tolerance of aluminium in plants: Tailoring plants to suit to acid soils. Biometals 2016, 29, 187–210. [Google Scholar] [CrossRef]
- Emanuel, B.Q.; Camilo, E.M.; Ileana, E.M.; Manuel, M.E. Aluminum, a friend or foe of higher plants in acid soils. Front. Plant Sci. 2017, 1767, 1–18. [Google Scholar]
- Krug, E.C.; Frink, C.R. Acid-rain on acid soil—A new perspective. Science 1983, 221, 520–525. [Google Scholar] [CrossRef] [PubMed]
- Blake, L.; Goulding, K.W.T.; Mott, C.J.B.; Johnston, A.E. Changes in soil chemistry accompanying acidification over more than 100 years under woodland and grass at Rothamsted Experimental Station, UK. Eur. J. Soil Sci. 1999, 50, 401–412. [Google Scholar] [CrossRef]
- Xu, R.K.; Coventry, D.R.; Farhoodi, A.; Schultz, J.E. Soil acidification as influenced by crop rotations, stubble management, and application of nitrogenous fertilizer. Tarlee. South Australia. Aust. J. Soil Res. 2002, 40, 483–496. [Google Scholar] [CrossRef]
- Guo, J.H.; Liu, X.J.; Zhang, Y.; Shen, J.L.; Han, W.X.; Zhang, W.F.; Christie, P.; Goulding, K.W.T.; Vitousek, P.M.; Zhang, F.S. Significant acidification in major Chinese croplands. Science 2010, 327, 1008–1010. [Google Scholar] [CrossRef]
- Hao, T.X.; Zhu, Q.C.; Zeng, M.F.; Shen, J.B.; Shi, X.J.; Liu, X.J.; Zhang, F.S.; Wim, D.V. Quantification of the contribution of nitrogen fertilization and crop harvesting to soil acidification in a wheat-maize double cropping system. Plant Soil 2019, 434, 167–184. [Google Scholar] [CrossRef]
- Mahler, R.L.; McDole, R. Effect of soil pH on crop yield in northern Idaho. Agron. J. 1987, 79, 751–755. [Google Scholar] [CrossRef]
- Yokota, S.; Ojima, K. Physiological response of root tip of alfalfa to low pH and aluminium stress in water culture. Plant Soi1 1995, 171, 163–165. [Google Scholar]
- Koyama, H.; Toda, T.; Hara, T. Brief exposure to low-pH stress causes irreversible damage to the growing root in Arabidopsis thaliana: Pectin-Ca interaction may play an important role in proton rhizotoxicity. J. Exp. Bot. 2001, 52, 361–368. [Google Scholar] [PubMed]
- Pavlovkin, J.; Pal’ove-Balang, P.; Kolarovic, L.; Zelinová, V. Growth and functional responses of different cultivars of Lotus corniculatus to aluminum and low pH stress. J. Plant Physiol. 2009, 166, 1479–1487. [Google Scholar] [CrossRef]
- Schubert, S.; Schubert, E.; Mengel, K. Effect of low pH of the root medium on proton release, growth, and nutrient uptake of field beans (Vicia faba). Plant Soil 1990, 124, 239–244. [Google Scholar] [CrossRef]
- Yan, F.; Schubert, S.; Mengel, K. Effect of low root medium pH on net proton release, root respiration, and root growth of corn (Zea mays L.) and broad bean (Vicia faba L.). Plant Physiol. 1992, 99, 415–421. [Google Scholar] [CrossRef]
- Yan, F.; Feuerle, R.; Schäffer, S.; Fortmeier, H.; Schubert, S. Adaptation of active proton pumping and plasmalemma ATPase activity of corn roots to low root medium pH. Plant Physiol. 1998, 117, 311–319. [Google Scholar] [CrossRef]
- Kinraide, T.B. Three mechanisms for the calcium alleviation of mineral toxicities. Plant Physiol. 1998, 118, 513–520. [Google Scholar] [CrossRef]
- Kamaluddin, M.; Zwiazek, J.J. Effects of root medium pH on water transport in paper birch (Betula papyrifera) seedlings in relation to root temperature and abscisic acid treatments. Tree Physiol. 2004, 24, 1173–1180. [Google Scholar] [CrossRef] [PubMed]
- Iyer-Pascuzzi, A.S.; Jackson, T.; Cui, H.; Petricka, J.J.; Busch, W.; Tsukagoshi, H.; Benfey, P.N. Cell identity regulators link development and stress responses in the Arabidopsis root. Dev. Cell 2011, 21, 770–782. [Google Scholar] [CrossRef]
- Palóve-Balang, P.; Čiamporová, M.; Zelinová, V.; PavlovkinErika, J.; Gurinová, E.; Mistrík, I. Cellular responses of two Latin-American cultivars of Lotus corniculatus to low pH and Al stress. Cent. Eur. J. Biol. 2012, 7, 1046–1054. [Google Scholar]
- Marschner, H. Mechanisms of adaptation of plants to acid soils. Plant Soil 1991, 134, 1–20. [Google Scholar] [CrossRef]
- Zhu, Y.Y.; Di, T.J.; Xu, G.H.; Chen, X.; Zeng, H.Q.; Yan, F.; Shen, Q.R. Adaptation of plasma membrane H+-ATPase of rice roots to low pH as related to ammonium nutrition. Plant Cell Environ. 2009, 32, 1428–1440. [Google Scholar] [CrossRef]
- Bouguyon, E.; Brun, F.; Meynard, D.; Kubeš, M.; Pervent, M.; Leran, S.; Lacombe, B.; Krouk, G.; Guiderdoni, E.; Zažímalová, E.; et al. Multiple mechanisms of nitrate sensing by Arabidopsis nitrate transporter NRT1.1. Nat. Plants 2015, 1, 150–159. [Google Scholar] [CrossRef]
- Fang, X.Z.; Tian, W.H.; Liu, X.X.; Lin, X.Y.; Jin, C.W.; Zheng, S.J. Alleviation of proton toxicity by nitrate uptake specifically depends on nitrate transporter1.1 in Arabidopsis. New Phytol. 2016, 211, 149–158. [Google Scholar] [CrossRef]
- Rossini Oliva, S.; Mingorance, M.D.; Sanhueza, D.; Fry, S.C.; Leidi, E.O. Active proton efflux, nutrient retention and boron-bridging of pectin are related to greater tolerance of proton toxicity in the roots of two Erica species. Plant Physiol. Biochem. 2018, 126, 142–151. [Google Scholar] [CrossRef]
- Espen, L.; Nocito, F.F.; Cocucci, M. Effect of NO3− transport and reduction on intracellular pH: An in vivo NMR study in maize roots. J. Exp. Bot. 2004, 55, 2053–2061. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.H.; Fan, X.R.; Miller, A.J. Plant nitrogen assimilation and use efficiency. Annu. Rev. Plant Biol. 2012, 63, 153–182. [Google Scholar] [CrossRef] [PubMed]
- Crawford, N.M.; Forde, B.G. Molecular and developmental biology of inorganic nitrogen nutrition. Arabidopsis Book 2002. [Google Scholar] [CrossRef]
- Khademi, S.; O’Connell, J.; Remis, J.; Robles-Colmenares, Y.; Miericke, L.J.; Stroud, R.M. Mechanism of ammonia transport by Amt/MEP/Rh: Structure of AmtB at 1.35 angstrom. Science 2004, 305, 1587–1594. [Google Scholar] [CrossRef]
- Ortiz-Ramirez, C.; Mora, S.I.; Trejo, J.; Pantoja, O. PvAMT1;1, a highly selective ammonium transporter that functions as H+/NH4+ symporter. J. Biol. Chem. 2011, 286, 31113–31122. [Google Scholar] [CrossRef]
- Fan, X.; Tang, Z.; Tan, Y.; Zhang, Y.; Luo, B.; Yang, M.; Lian, X.; Shen, Q.; Miller, A.J.; Xu, G. Overexpression of a pH-sensitive nitrate transporter in rice increases crop yields. Proc. Natl. Acad. Sci. USA 2016, 113, 7118–7123. [Google Scholar] [CrossRef]
- Xuan, W.; Beeckman, T.; Xu, G.H. Plant nitrogen nutrition: Sensing and signaling. Curr. Opin. Plant Biol. 2017, 39, 57. [Google Scholar] [CrossRef]
- Lager, I.; Andréasson, O.; Dunbar, T.L.; Andreasson, E.; Escobar, M.A.; Rasmusson, A.G. Changes in external pH rapidly alter plant gene expression and modulate auxin and elicitor responses. Plant Cell Environ. 2010, 33, 1513–1528. [Google Scholar] [CrossRef]
- Hu, H.Y.; He, J.; Zhao, J.J.; Ou, X.Q.; Li, H.M.; Ru, Z.G. 2018. Low pH stress responsive transcriptome of seedling roots in wheat (Triticum aestivum L.). Genes. Genom. 2010, 40, 1199–1211. [Google Scholar] [CrossRef]
- Jahan, M.A.; Harris, B.; Lowery, M.; Coburn, K.; Infante, A.M.; Percifield, R.J.; Ammer, A.G.; Kovinich, N. The NAC family transcription factor GmNAC42-1 regulates biosynthesis of the anticancer and neuroprotective glyceollins in soybean. BMC Genomics 2019, 20, 149. [Google Scholar] [CrossRef]
- Iuchi, S.; Koyama, H.; Iuchi, A.; Kobayashi, A.; Kitabayashi, S.; Kobayashi, Y.; Ikka, T.; Hirayama, T.; Shinozaki, K.; Kobayashi, M. Zinc finger protein STOP1 is critical for proton tolerance in Arabidopsis and coregulates a key gene in aluminum tolerance. Proc. Natl. Acad. Sci. USA 2007, 104, 9900–9905. [Google Scholar] [CrossRef]
- Yamaji, N.; Huang, C.F.; Nagao, S.; Yano, M.; Sato, Y.; Nagamura, Y.; Ma, J.F. A zinc finger transcription factor ART1 regulates multiple genes implicated in aluminum tolerance in rice. Plant Cell 2009, 21, 3339–3349. [Google Scholar] [CrossRef]
- Ohyama, Y.; Ito, H.; Kobayashi, Y.; Ikka, T.; Morita, A.; Kobayashi, M.; Imaizumi, R.; Aoki, T.; Komatsu, K.; Sakata, Y.; et al. Characterization of AtSTOP1 orthologous genes in tobacco and other plant species. Plant Physiol. 2013, 162, 1937–1946. [Google Scholar] [CrossRef]
- Sawaki, Y.; Kobayashi, Y.; Kihara-Doi, T.; Nishikubo, N.; Kawazu, T.; Kobayashi, M.; Iuchi, S.; Koyama, H.; Sato, S. Identification of a STOP1-like protein in Eucalyptus that regulates transcription of Al tolerance genes. Plant Sci. 2014, 223, 8–15. [Google Scholar] [CrossRef]
- Jiang, F.; Wang, T.; Wang, Y.; Kochian, L.V.; Chen, F.; Liu, J. Identification and characterization of suppressor mutants of stop1. BMC. Plant Biol. 2017, 17, 128. [Google Scholar]
- Wu, W.; Lin, Y.; Chen, Q.; Peng, W.; Peng, J.; Tian, J.; Liang, C.; Liao, H. Functional conservation and divergence of soybean GmSTOP1 members in proton and aluminum tolerance. Front. Plant Sci. 2018, 9, 570. [Google Scholar] [CrossRef]
- Zhou, Y.; Yang, Z.; Gong, L.; Liu, R.; Sun, H.; You, J. Molecular characterization of GmSTOP1 homologs in soybean under Al and proton stress. Plant Soil 2018, 427, 213–230. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, J.; Guo, J.; Zhou, F.; Singh, S.; Xu, X.; Xie, Q.; Yang, Z.; Huang, C.F. F-box protein RAE1 regulates the stability of the aluminum-resistance transcription factor STOP1 in Arabidopsis. Proc. Natl. Acad. Sci. USA 2019, 116, 319–327. [Google Scholar] [CrossRef]
- Graham, P.H.; Vance, C.P. Legumes: Importance and constraints to greater use. Plant Physiol. 2003, 131, 872–877. [Google Scholar] [CrossRef]
- Wilson, R.F. Soybean: Market driven research needs. In Genetics and Genomics of Soybean. Plant Genetics and Genomics: Crops and Models; Stacey, G., Ed.; Springer: New York, NY, USA, 2008; Volume 2, pp. 3–15. [Google Scholar]
- Lu, S.J.; Zhao, X.H.; Hu, Y.L.; Liu, S.L.; Nan, H.Y.; Li, X.M.; Fang, C.; Cao, D.; Shi, X.Y.; Kong, L.P.; et al. Natural variation at the soybean j locus improves adaptation to the tropics and enhances yield. Nat. Genet. 2017, 49, 773. [Google Scholar] [CrossRef]
- Embrapa-cpac. Relatório técnico annual; Pmpresa brasileira de pesquisa agropecufiria, centro de pesquisa agropecuziria dos cerrados distrito federal: Distrito Federal, Brazil, 1979; p. 195. [Google Scholar]
- Spehar, C.R. Impact of strategic genes in soybean on agricultural development in the Brazilian tropical savannah. Field Crops Res. 1995, 41, 141–146. [Google Scholar] [CrossRef]
- Liang, C.; Wang, J.; Zhao, J.; Tian, J.; Liao, H. Control of phosphate homeostasis through gene regulation in crops. Curr. Opin. Plant Biol. 2014, 21, 59–66. [Google Scholar] [CrossRef]
- Chen, Z.C.; Liao, H. Organic acid anions: An effective defensive weapon for plants against aluminum toxicity and phosphorus deficiency in acidic soils. J. Genet. Genomics 2016, 43, 631–638. [Google Scholar] [CrossRef] [PubMed]
- Reis, A.R.D.; Lisboa, L.A.M.; Reis, H.P.G.; Barcelos, J.P.Q.; Santos, E.F.; Santini, J.M.K.; Meyer-Sand, B.R.V.; Putti, F.F.; Galindo, F.S.; Kaneko, F.H.; et al. Depicting the physiological and ultrastructural responses of soybean plants to Al stress conditions. Plant Physiol. Biochem. 2018, 130, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.C.; Chu, S.J.; Guo, Y.M.; Ji, Y.J.; Hu, D.Q.; Cheng, J.; Lu, G.H.; Yang, R.W.; Tang, C.Y.; Qi, J.L.; et al. Novel mechanisms for organic acid-mediated aluminium tolerance in roots and leaves of two contrasting soybean genotypes. AoB Plants 2017, 9, plx064. [Google Scholar] [CrossRef]
- Liu, N.; Song, F.; Zhu, X.; You, J.; Yang, Z.; Li, X. Salicylic acid alleviates aluminum toxicity in soybean roots through modulation of reactive oxygen species metabolism. Front. Chem. 2017, 5, 96. [Google Scholar] [CrossRef] [PubMed]
- Wu, W.; Lin, Y.; Liu, P.; Chen, Q.; Tian, J.; Liang, C. Association of extracellular dNTP utilization with a GmPAP1-like protein identified in cell wall proteomic analysis of soybean roots. J. Exp. Bot. 2018, 69, 603–617. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Pertea, M.; Pertea, G.M.; Antonescu, C.M.; Chang, T.C.; Mendel, J.T.; Salzberg, S.L. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 2015, 33, 290–295. [Google Scholar] [CrossRef]
- Sawaki, Y.; Iuchi, S.; Kobayashi, Y.; Kobayashi, Y.; Ikka, T.; Sakurai, N.; Fujita, M.; Shinozaki, K.; Shibata, D.; Kobayashi, M.; et al. STOP1 regulates multiple genes that protect arabidopsis from proton and aluminum toxicities. Plant Physiol. 2009, 150, 281–294. [Google Scholar] [CrossRef] [PubMed]
- Eswaran, H.; Reich, P.; Beinroth, F. Global distribution of soils with acidity. In Plant-Soil Interaction at Low pH: Sustainable Agriculture and Forestry Production; Moniz, A.C., Furlani, A.M.C., Schaffert, R.E., Fageria, N.K., Rosolem, C.A., Cantarella, H., Eds.; Brazilian Soil Science Society: Campinas, Brazil, 1997; pp. 159–164. [Google Scholar]
- Ham, B.K.; Chen, J.; Yan, Y.; Lucas, W.J. Insights into plant phosphate sensing and signaling. Curr. Opin. Biotechnol. 2018, 49, 1–9. [Google Scholar] [CrossRef]
- Vitorello, V.A.; Capaldi, F.R.; Stefanuto, V.A. Recent advances in aluminum toxicity and resistance in higher plants. Braz. J. Plant. Physiol. 2005, 17, 129–143. [Google Scholar] [CrossRef]
- Shavrukov, Y.; Hirai, Y. Good and bad protons: Genetic aspects of acidity stress responses in plants. J. Exp. Bot. 2016, 67, 15–30. [Google Scholar] [CrossRef]
- Loqué, D.; Lalonde, S.; Looger, L.L.; Von-Wirén, N.; Frommer, W.B. A cytosolic trans-activation domain essential for ammonium uptake. Nature 2007, 446, 195–198. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Loqué, D.; Ye, F.H.; Frommer, W.B.; Von-Wirén, N. Nitrogen-dependent posttranscriptional regulation of the ammonium transporter atamt1;1. Plant Physiol. 2007, 143, 732–744. [Google Scholar] [CrossRef] [PubMed]
- Yuan, L.; Graff, L.; Loqué, D.; Kojima, S.; Tsuchiya, Y.N.; Takahashi, H.; Von-Wirén, N. AtAMT1;4, a pollen-specific high-affinity ammonium transporter of the plasma membrane in Arabidopsis. Plant Cell Physiol. 2009, 50, 13–25. [Google Scholar] [CrossRef]
- Gu, R.; Duan, F.; An, X.; Zhang, F.; Von-Wirén, N.; Yuan, L. Characterization of AMT-mediated high-affinity ammonium uptake in roots of maize (Zea mays L.). Plant Cell Physiol. 2013, 54, 1515–1524. [Google Scholar] [CrossRef] [PubMed]
- Giehl, R.F.H.; Laginha, A.M.; Duan, F.; Rentsch, D.; Yuan, L.; Von-Wirén, N. A critical role of AMT2;1 in root-to-shoot translocation of ammonium in Arabidopsis. Mol. Plant 2017, 10, 1449–1460. [Google Scholar] [CrossRef]
- Lin, S.H.; Kuo, H.F.; Canivenc, G.; Lin, C.S.; Lepetit, M.; Hsu, P.K.; Tillard, P.; Lin, H.L.; Wang, Y.Y.; Tsai, C.B.; et al. Mutation of the Arabidopsis NRT1.5 nitrate transporter causes defective root-to-shoot nitrate transport. Plant Cell 2008, 20, 2514–2528. [Google Scholar] [CrossRef] [PubMed]
- Yan, N. Structural biology of the major facilitator superfamily transporters. Annu. Rev. Biophys. 2015, 44, 257–283. [Google Scholar] [CrossRef] [PubMed]
- Meng, S.; Peng, J.S.; He, Y.N.; Zhang, G.B.; Yi, H.Y.; Fu, Y.L.; Gong, J.M. Arabidopsis NRT1.5 mediates the suppression of nitrate starvation-induced leaf senescence by modulating foliar potassium level. Mol. Plant 2016, 9, 461–470. [Google Scholar] [CrossRef]
- Li, H.; Yu, M.; Du, X.Q.; Wang, Z.F.; Wu, W.H.; Quintero, F.J.; Jin, X.H.; Li, H.D.; Wang, Y. NRT1.5/NPF7.3 functions as a proton-coupled H+/K+ antiporter for K+ loading into the xylem in Arabidopsis. Plant Cell 2017, 29, 2016–2026. [Google Scholar] [CrossRef]
- Qin, L.; Zhao, J.; Tian, J.; Chen, L.Y.; Sun, Z.A.; Guo, Y.X.; Lu, X.; Gu, M.; Xu, G.H.; Liao, H. The high-affinity phosphate transporter GmPT5 regulates phosphate transport to nodules and nodulation in soybean. Plant Physiol. 2012, 159, 1634–1643. [Google Scholar] [CrossRef]
- Gaymard, F.; Pilot, G.; Lacombe, B.; Bouchez, D.; Bruneau, D.; Boucherez, J.; Michaux-Ferrière, N.; Thibaud, J.B.; Sentenac, H. Identification and disruption of a plant shaker-like outward channel involved in K+ release into the xylem sap. Cell 1998, 94, 647–655. [Google Scholar] [CrossRef]
- Wang, Y.; Wu, W.H. Potassium transport and signaling in higher plants. Annu. Rev. Plant Biol. 2013, 64, 451–476. [Google Scholar] [CrossRef]
- Véry, A.A.; Nieves-Cordones, M.; Daly, M.; Khan, I.; Fizames, C.; Sentenac, H. Molecular biology of K+ transport across the plant cell membrane: What do we learn from comparison between plant species? J. Plant Physiol. 2014, 171, 748–769. [Google Scholar] [CrossRef]
- Li, W.H.; Xua, G.H.; Abdel, A.; Yua, L. Plant HAK/KUP/KT K+ transporters: Function and regulation. Semin. Cell Dev. Biol. 2018, 74, 133–141. [Google Scholar] [CrossRef]
- Hawkesford, M.J. Transporter gene families in plants: The sulphate transporter gene family—Redundancy or specialization? Physiol. Plant 2003, 117, 155–163. [Google Scholar] [CrossRef]
- Fitzpatrick, K.L.; Tyerman, S.D.; Kaiser, B.N. Molybdate transport through the plant sulfate transporter SHST1. FEBS Lett. 2008, 582, 1508–1513. [Google Scholar] [CrossRef] [PubMed]
- Cao, M.J.; Wang, Z.; Wirtz, M.; Hell, R.; Oliver, D.J.; Xiang, C.B. SULTR3;1 is a chloroplast-localized sulfate transporter in Arabidopsis thaliana. Plant J. 2013, 73, 607–616. [Google Scholar] [CrossRef] [PubMed]
- Kataoka, T.; Hayashi, N.; Yamaya, T.; Takahashi, H. Root-to-shoot transport of sulfate in Arabidopsis. Evidence for the role of SULTR3;5 as a component of low-affinity sulfate transport system in the root vasculature. Plant Physiol. 2004, 136, 4198–4204. [Google Scholar] [CrossRef] [PubMed]
- Zuber, H.; Davidian, J.C.; Aubert, G.; Aimé, D.; Belghazi, M.; Lugan, R.; Heintz, D.; Wirtz, M.; Hell, R.; Thompson, R.; et al. The seed composition of Arabidopsis mutants for the group 3 sulfate transporters indicates a role in sulfate translocation within developing seeds. Plant Physiol. 2010, 154, 913–926. [Google Scholar] [CrossRef]
- Ishimaru, Y.; Takahashi, R.; Bashir, K.; Shimo, H.; Senoura, T.; Sugimoto, K.; Ono, K.; Yano, M.; Ishikawa, S.; Arao, T.; et al. Characterizing the role of rice NRAMP5 in manganese, iron and cadmium transport. Sci. Rep. 2012, 2, 286. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, A.; Yamaji, N.; Yokosho, K.; Ma, J.F. Nramp5 is a major transporter responsible for manganese and cadmium uptake in rice. Plant Cell 2012, 24, 2155–2167. [Google Scholar] [CrossRef]
- Milner, M.J.; Seamon, J.; Craft, E.; Kochian, L.V. Transport properties of members of the ZIP family in plants and their role in Zn and Mn homeostasis. J. Exp. Bot. 2013, 64, 369–381. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Yamaji, N.; Yamane, M.; Kashino-Fujii, M.; Sato, K.; Ma, J.F. The HvNRAMP5 transporter mediates uptake of cadmium and manganese, but not iron. Plant Physiol. 2016, 172, 1899–1910. [Google Scholar] [CrossRef] [PubMed]
- Peng, F.; Wang, C.; Zhu, J.S.; Zeng, J.; Kang, H.Y.; Fan, X.; Sha, L.; Zhang, H.Q.; Zhou, Y.H.; Wang, Y. Expression of TpNRAMP5, a metal transporter from Polish wheat (Triticum polonicum L.), enhances the accumulation of Cd, Co and Mn in transgenic Arabidopsis plants. Planta 2018, 247, 1395–1406. [Google Scholar] [CrossRef] [PubMed]
- Montanini, B.; Blaudez, D.; Jeandroz, S.; Sanders, D.; Chalot, M. Phylogenetic and functional analysis of the Cation Diffusion Facilitator (CDF) family: Improved signature and prediction of substrate specificity. BMC Genomics 2007, 8, 107–122. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Liu, B. Identification of a rice metal tolerance protein OsMTP11 as a manganese transporter. PLoS ONE 2017, 12, e0174987. [Google Scholar] [CrossRef] [PubMed]
- Roberts, J.K.; Hooks, M.A.; Miaullis, A.P.; Edwards, S.; Webster, C. Contribution of malate and amino acid metabolism to cytoplasmic pH regulation in hypoxic maize root tips studied using nuclear magnetic resonance spectroscopy. Plant Physiol. 1992, 98, 480–487. [Google Scholar] [CrossRef] [PubMed]
- Sakano, K. Revision of biochemical pH-stat: Involvement of alternative pathway metabolisms. Plant Cell Physiol. 1998, 39, 467–473. [Google Scholar] [CrossRef]
- Bown, A.W.; Shelp, B.J. The metabolism and functions of g-aminobutyric acid. Plant Physiol. 1997, 115, 1–5. [Google Scholar] [CrossRef]
- Bouche, N.; Fromm, H. GABA in plants: Just a metabolite? Trends Plant Sci. 2004, 9, 110–115. [Google Scholar] [CrossRef]
- Bustos, R.; Castrillo, G.; Linhares, F.; Puga, M.I.; Rubio, V.; Pérez-Pérez, J.; Solano, R.; Leyva, A.; Paz-Ares, J. A central regulatory system largely controls transcriptional activation and repression responses to phosphate starvation in Arabidopsis. PLoS Genet. 2010, 6, e1001102. [Google Scholar] [CrossRef]
- Kobayashi, Y.; Ohyama, Y.; Kobayashi, Y.; Ito, H.; Iuchi, S.; Fujita, M.; Zhao, C.R.; Tanveer, T.; Ganesan, M.; Kobayashi, M.; et al. STOP2 activates transcription of several genes for Al-and low pH-tolerance that are regulated by STOP1 in Arabidopsis. Mol. Plant 2014, 7, 311–322. [Google Scholar] [CrossRef] [PubMed]


| Total Expressed Genes | Up-Regulated Genes | Down-Regulated Genes | |
|---|---|---|---|
| pH 5.8 | 42,621 | ||
| pH 4.2 | 41,857 | ||
| DEGs * | 974 | 419 | 555 |
| Number | Gene ID | Expression Level at pH 5.8 | Expression Level at pH 4.2 | Log2 Fold Change (pH 4.2/pH 5.8) | p Value | Description | Gene Name |
|---|---|---|---|---|---|---|---|
| 1 | Glyma.16G041200 | 243.18 | 823.46 | 1.8 | 8.49 × 10−12 | glutamate dehydrogenase 1 | GmGDH1 |
| 2 | Glyma.09G262900 | 122.93 | 43.47 | −1.5 | 2.20×10−5 | NAD-dependent malic enzyme | GmME1-1 |
| 3 | Glyma.04G086300 | 37.51 | 198.71 | 2.4 | 2.91×10−6 | NAD-dependent malic enzyme | GmME1-2 |
| 4 | Glyma.13G231700 | 972.9 | 3302.8 | 1.8 | 1.74×10−6 | pyruvate decarboxylase 1 | GmPCD1 |
| 5 | Glyma.18G204200 | 1282.4 | 3518.3 | 1.5 | 2.88×10−5 | pyruvate decarboxylase 2 | GmPCD2-1 |
| 6 | Glyma.07G153100 | 934.5 | 3283.6 | 1.8 | 1.57×10−5 | pyruvate decarboxylase 2 | GmPCD2-2 |
| 7 | Glyma.04G240800 | 1201.0 | 4375.6 | 1.9 | 5.47×10−8 | alcohol dehydrogenase 1 | GmADH1-1 |
| 8 | Glyma.14G121200 | 63.3 | 399.0 | 2.7 | 5.31×10−16 | alcohol dehydrogenase 1-like | GmADH1-2 |
| 9 | Glyma.06G122600 | 3010.3 | 10795.3 | 1.8 | 2.14×10−7 | alcohol dehydrogenase 1 | GmADH1-3 |
| 10 | Glyma.18G200300 | 467.7 | 187.2 | −1.3 | 3.59×10−5 | alcohol dehydrogenase-like 4 | GmADH4 |
| Number | Gene ID | Expression Level at pH5.8 | Expression Level at pH4.2 | Log2 Fold Change (pH 4.2/pH 5.8) | p Value | Description | Gene Name |
|---|---|---|---|---|---|---|---|
| 1 | Glyma.10G132300 | 306.82 | 127.02 | −1.27 | 2.8 × 10−4 | ammonium transporter 1 | GmAMT1 |
| 2 | Glyma.01G123400 | 81.10 | 6.82 | −3.57 | 6.37 × 10−11 | ammonium transporter 2-like | GmAMT2 |
| 3 | Glyma.05G070600 | 194.19 | 62.38 | −1.64 | 1.25 × 10−4 | protein NRT1/PTR FAMILY 7.3-like | GmNPF7.3-1 |
| 4 | Glyma.17G153300 | 611.91 | 209.01 | −1.55 | 9.05 × 10−4 | protein NRT1/PTR FAMILY 7.3-like | GmNPF7.3-2 |
| 5 | Glyma.13G323800 | 5449.11 | 476.64 | −3.52 | 3.00 × 10−23 | NRT2 protein | GmNRT2 |
| 6 | Glyma.12G176900 | 2041.80 | 13.51 | −7.24 | 6.20 × 10−66 | high affinity nitrate transporter 2.4-like | GmNRT2.4-1 |
| 7 | Glyma.11G195200 | 8226.27 | 426.09 | −4.27 | 4.28 × 10−31 | high affinity nitrate transporter 2.4 | GmNRT2.4-2 |
| 8 | Glyma.18G141900 | 1306.71 | 33.46 | −5.29 | 1.53 × 10−33 | high affinity nitrate transporter 2.5-like | GmNRT2.5 |
| 9 | Glyma.03G122500 | 352.71 | 882.01 | 1.32 | 8.93 × 10−4 | protein NRT1/PTR FAMILY 3.1 | GmNPF3.1 |
| 10 | Glyma.18G126500 | 23.47 | 77.52 | 1.72 | 4.03 × 10−5 | protein NRT1/PTR FAMILY 4.6 | GmNPF4.6 |
| 11 | Glyma.02G005800 | 665.66 | 6.09 | −6.77 | 1.42 × 10−5 | inorganic phosphate transporter 1-7-like | GmPT1 |
| 12 | Glyma.07G222700 | 188.38 | 63.69 | −1.56 | 2.98 × 10−5 | inorganic phosphate transporter 1-9-like | GmPT3 |
| 13 | Glyma.10G006700 | 1478.73 | 379.51 | −1.96 | 4.63 × 10−6 | inorganic phosphate transporter 1-3 | GmPT4 |
| 14 | Glyma.10G186500 | 919.56 | 381.20 | −1.27 | 1.82 × 10−4 | inorganic phosphate transporter 1-7-like | GmPT7 |
| 15 | Glyma.20G204000 | 294.20 | 71.42 | −2.04 | 2.17 × 10−5 | inorganic phosphate transporter 1-7-like | GmPT13 |
| 16 | Glyma.12G133400 | 96.70 | 22.67 | −2.09 | 1.87 × 10−7 | sodium transporter HKT1-like | GmHKT1 |
| 17 | Glyma.16G046200 | 669.36 | 304.90 | −1.13 | 3.12 × 10−5 | potassium transporter 4-like | GmKUP3 |
| 18 | Glyma.02G033600 | 566.51 | 275.46 | −1.04 | 2.81 × 10−4 | Potassium transporter 6 | GmKUP6 |
| 19 | Glyma.03G223900 | 43.03 | 17.06 | −1.33 | 1.01 × 10−3 | two-pore potassium channel 5-like | GmTPK5 |
| 20 | Glyma.02G243400 | 692.88 | 298.10 | −1.21 | 3.79 × 10−5 | potassium channel SKOR-like | GmSKOR |
| 21 | Glyma.06G143800 | 100.89 | 249.30 | 1.31 | 4.16 × 10−6 | potassium transporter 2-like | GmKT2 |
| 22 | Glyma.11G238400 | 70.15 | 18.35 | −1.93 | 2.75 × 10−6 | sulfate transporter 2.1-like | GmSULTR2.1-1 |
| 23 | Glyma.08G138600 | 78.51 | 29.44 | −1.42 | 1.81 × 10−4 | sulfate transporter 2.1-like | GmSULTR2.1-2 |
| 24 | Glyma.18G019000 | 123.53 | 26.66 | −2.21 | 6.41 × 10−8 | sulfate transporter 2.1-like | GmSULTR2.1-3 |
| 25 | Glyma.19G159000 | 126.03 | 411.14 | 1.71 | 4.51 × 10−6 | sulfate transporter 3.1-like | GmSULTR3.1-1 |
| 26 | Glyma.02G145100 | 83.76 | 214.91 | 1.36 | 1.38 × 10−6 | sulfate transporter 3.1-like | GmSULTR3.1-2 |
| 27 | Glyma.03G156700 | 378.82 | 1484.45 | 1.97 | 2.27 × 10−6 | sulfate transporter 3.1-like | GmSULTR3.1-3 |
| 28 | Glyma.09G188700 | 4.38 | 64.18 | 3.87 | 4.31 × 10−13 | sulfate transporter 3.5 | GmSULTR3.5-1 |
| 29 | Glyma.07G088200 | 89.10 | 448.25 | 2.33 | 7.3 × 10−4 | sulfate transporter 3.5 | GmSULTR3.5-2 |
| 30 | Glyma.13G004400 | 1633.97 | 480.53 | −1.77 | 7.75 × 10−11 | zinc transporter 1-like isoform X2 | GmZIP1 |
| 31 | Glyma.13G338300 | 495.89 | 159.55 | −1.64 | 2.28 × 10−7 | zinc transporter 1-like | GmZIP2 |
| 32 | Glyma.08G328000 | 77.30 | 20.91 | −1.89 | 2.66 × 10−5 | zinc transporter 11 | GmZIP11-1 |
| 33 | Glyma.15G036200 | 208.80 | 85.91 | −1.28 | 3.16 × 10−5 | Zinc transporter 1 | GmZIP11-2 |
| 34 | Glyma.09G122600 | 469.38 | 132.66 | −1.82 | 1.22 × 10−4 | metal tolerance protein 10-like | GmMTP7-1 |
| 35 | Glyma.08G164800 | 71.85 | 220.35 | 1.62 | 7.03 × 10−7 | metal tolerance protein 10-like | GmMTP7-2 |
| 36 | Glyma.06G115800 | 1674.89 | 353.75 | −2.24 | 3.48 × 10−6 | metal transporter Nramp5-like | GmNramp5 |
| 37 | Glyma.07G223200 | 10.03 | 1.80 | −2.48 | 2.79 × 10−4 | zinc transporter 10-like precursor | GmIRT1 |
| 38 | Glyma.05G121300 | 68.95 | 21.61 | −1.67 | 1.65 × 10−4 | vacuolar iron transporter homolog 4-like | GmVIT2.1 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, Q.; Wu, W.; Zhao, T.; Tan, W.; Tian, J.; Liang, C. Complex Gene Regulation Underlying Mineral Nutrient Homeostasis in Soybean Root Response to Acidity Stress. Genes 2019, 10, 402. https://doi.org/10.3390/genes10050402
Chen Q, Wu W, Zhao T, Tan W, Tian J, Liang C. Complex Gene Regulation Underlying Mineral Nutrient Homeostasis in Soybean Root Response to Acidity Stress. Genes. 2019; 10(5):402. https://doi.org/10.3390/genes10050402
Chicago/Turabian StyleChen, Qianqian, Weiwei Wu, Tong Zhao, Wenqi Tan, Jiang Tian, and Cuiyue Liang. 2019. "Complex Gene Regulation Underlying Mineral Nutrient Homeostasis in Soybean Root Response to Acidity Stress" Genes 10, no. 5: 402. https://doi.org/10.3390/genes10050402
APA StyleChen, Q., Wu, W., Zhao, T., Tan, W., Tian, J., & Liang, C. (2019). Complex Gene Regulation Underlying Mineral Nutrient Homeostasis in Soybean Root Response to Acidity Stress. Genes, 10(5), 402. https://doi.org/10.3390/genes10050402






