Full-Length Multi-Barcoding: DNA Barcoding from Single Ingredient to Complex Mixtures
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Powders Preparation
2.2. DNA Preparation and Sanger Sequencing
2.3. Amplicon Libraries Preparation for SMRT Sequencing
2.4. SMRT Sequencing and Data Analysis
2.5. Clustering and Biological Identification of Resulting Reads
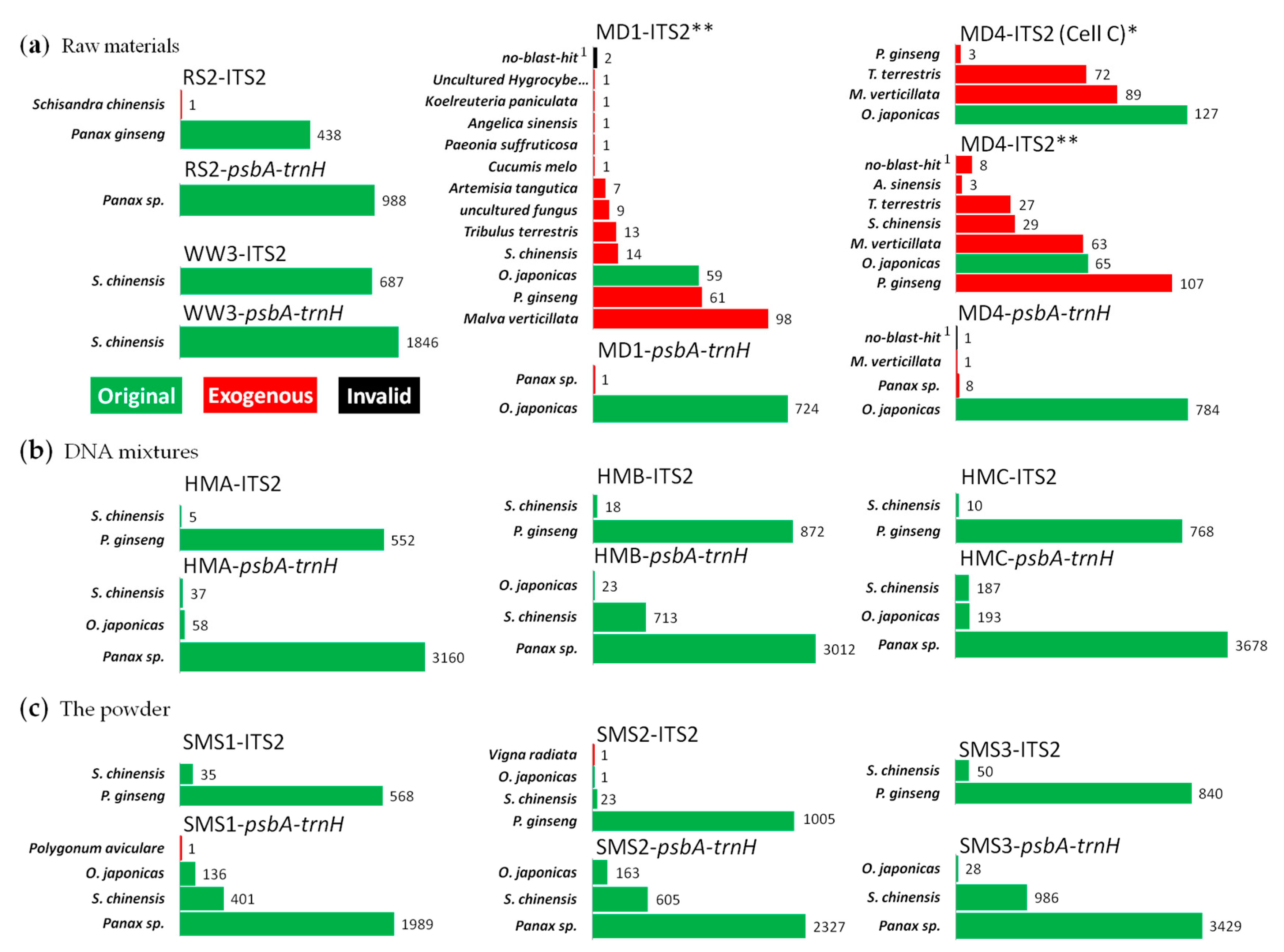
- Original: A resulting read is original if it was assigned to the biological species of a sample.
- Endogenous: A resulting read is endogenous if the species which it was assigned to could be found in the raw materials, though it was not original.
- Exogenous: A resulting read is exogenous if it was assigned to other species which was considered as contamination from the outside, but not the biological species of the raw material or the powder.
- Invalid: A resulting read is invalid if it could not be assigned to biological species due to a lower similarity with the sequences in both databases, which was also defined as noneffective, rather than the effective ones that were defined as original, endogenous or exogenous.
2.6. Method Testifying by SJS
3. Results
3.1. Species Authentication by Sanger Sequencing
3.2. Analysis Results Via FLMB
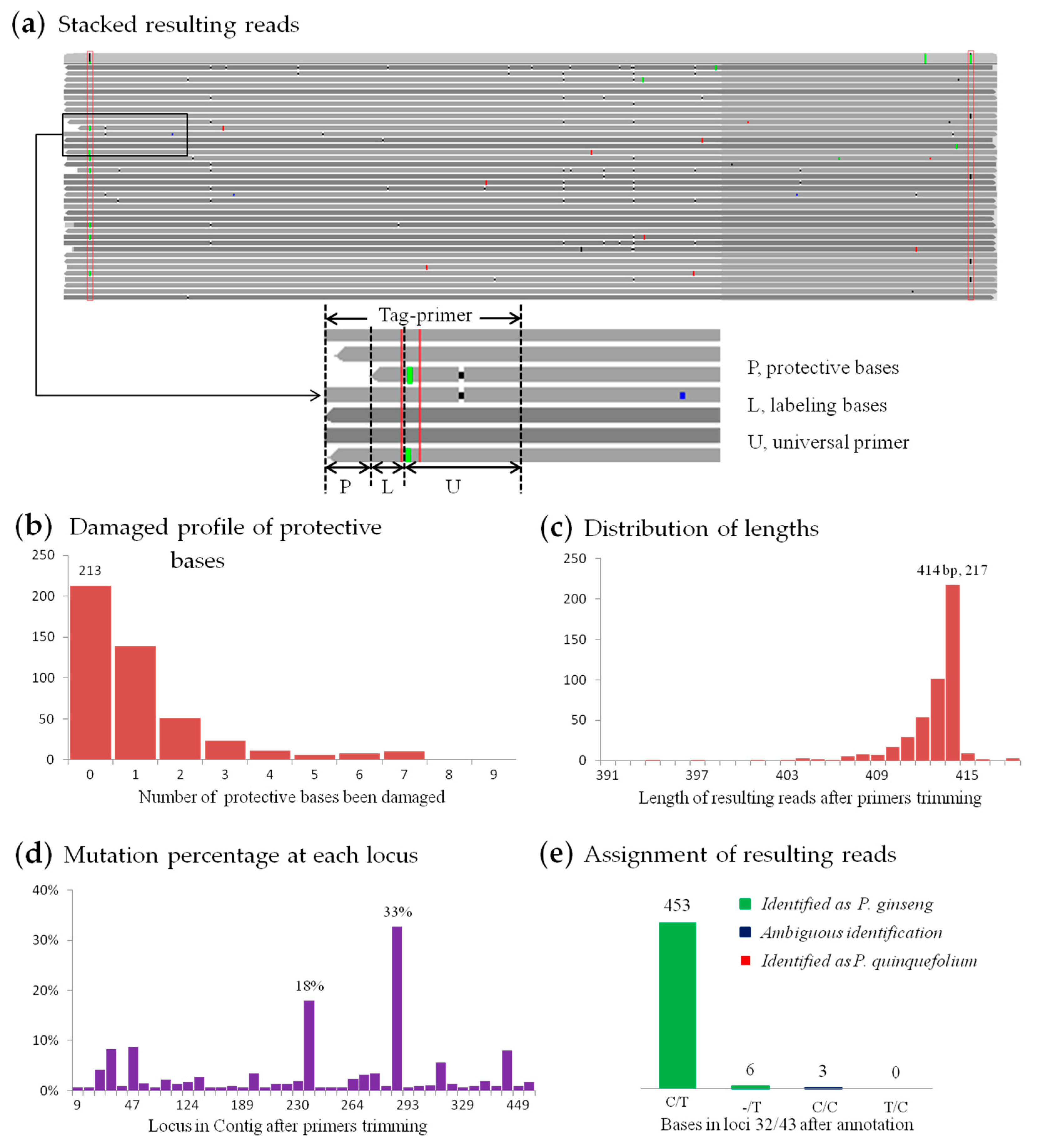
3.2.1. Data Processing of SMRT Sequencing
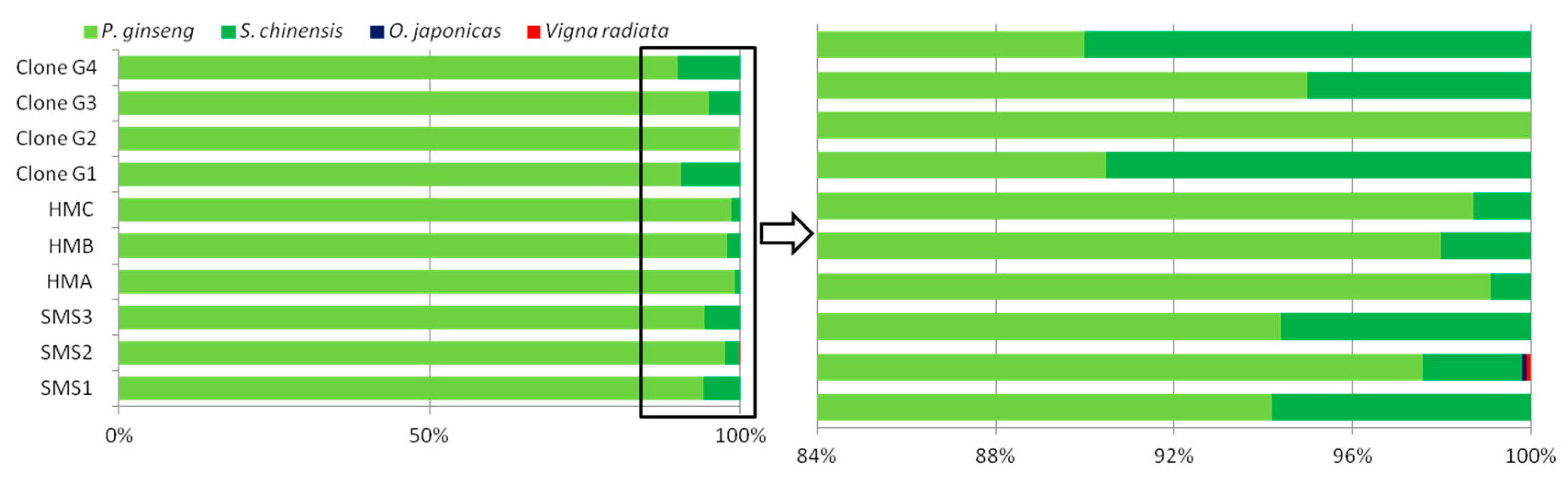
3.2.2. Species Identification of SMS by SMRT Sequencing
3.2.3. Result of Clone-Dependent Sanger Sequencing
3.2.4. Results for SJS
4. Discussion
4.1. Methodological Analysis
4.1.1. Precise Definitions from High Accuracy Resulting Reads
4.1.2. Every Resulting Reads Counts
4.1.3. Recommended Sequencing Depth
4.1.4. Combination Makers Makes Identification More Accuracy and Entirely
4.1.5. Quality Control of the Procedures
4.2. Different Sequencing Technologies Applied to DNA Barcoding
4.3. Applications and Challenges
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Hao, D.-C.; Chen, S.L.; Xiao, P.-G. Authentication of medicinal plants based on molecular biology and genomics. Pharm. Biotechnol. 2009, 16, 490–494. [Google Scholar]
- Miller, S.E. DNA barcoding and the renaissance of taxonomy. Proc. Natl. Acad. Sci. USA 2007, 104, 4775–4776. [Google Scholar] [CrossRef]
- Wallace, L.J.; Boilard, S.M.; Eagle, S.H.; Spall, J.L.; Shokralla, S.; Hajibabaei, M. DNA barcodes for everyday life: Routine authentication of natural health products. Food Res. Int. 2012, 49, 446–452. [Google Scholar] [CrossRef]
- Wu, L.; Sun, W.; Wang, B.; Zhao, H.; Li, Y.; Cai, S.; Xiang, L.; Zhu, Y.; Yao, H.; Song, J.; et al. An integrated system for identifying the hidden assassins in traditional medicines containing aristolochic acids. Sci. Rep. 2015, 5, 11318. [Google Scholar] [CrossRef]
- Vanherweghem, J.L.; Tielemans, C.; Abramowicz, D.; Depierreux, M.; Vanhaelen-Fastre, R.; Vanhaelen, M.; Dratwa, M.; Richard, C.; Vandervelde, D.; Verbeelen, D.; et al. Rapidly progressive interstitial renal fibrosis in young women: Association with slimming regimen including Chinese herbs. Lancet 1993, 341, 387–391. [Google Scholar] [CrossRef]
- Chen, C.-H.; Dickman, K.G.; Moriya, M.; Zavadil, J.; Sidorenko, V.S.; Edwards, K.L.; Gnatenko, D.V.; Wu, L.; Turesky, R.J.; Wu, X.-R.; et al. Aristolochic acid-associated urothelial cancer in Taiwan. Proc. Natl. Acad. Sci. USA 2012, 109, 8241–8246. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Pang, X.; Liao, B.; Yao, H.; Song, J.; Chen, S. An authenticity survey of herbal medicines from markets in China using DNA barcoding. Sci. Rep. 2016, 6, 18723. [Google Scholar] [CrossRef]
- Chen, S.; Yao, H.; Han, J.; Liu, C.; Song, J.; Shi, L.; Zhu, Y.; Ma, X.; Gao, T.; Pang, X.; et al. Validation of the ITS2 region as a novel DNA barcode for identifying medicinal plant species. PLoS ONE 2010, 5, e8613. [Google Scholar]
- Newmaster, S.G.; Grguric, M.; Shanmughanandhan, D.; Ramalingam, S.; Ragupathy, S. DNA barcoding detects contamination and substitution in North American herbal products. BMC Med. 2013, 11, 222. [Google Scholar] [CrossRef] [PubMed]
- Song, J.; Yao, H.; Li, Y.; Li, X.; Lin, Y.; Liu, C.; Jianping, H.; Xie, C.; Chen, S. Authentication of the family polygonaceae in Chinese pharmacopoeiaby DNA barcoding technique. J. Ethnopharmacol. 2009, 124, 434–439. [Google Scholar] [CrossRef]
- DNA Barcoding System for Identifying Herbal Medicine. Available online: http://www.tcmbarcode.cn/en/ (accessed on 22 November 2017).
- Chen, S.; Pang, X.; Song, J.; Shi, L.; Yao, H.; Han, J.; Leon, C. A renaissance in herbal medicine identification: From morphology to DNA. Biotechnol. Adv. 2014, 32, 1237–1244. [Google Scholar] [CrossRef]
- Cheng, K.-T.; Tsay, H.-S.; Chen, C.-F.; Chou, T.-W. Determination of the components in a Chinese prescription, Yu-Ping-Feng San, by RAPD analysis. Planta Med. 1998, 64, 563–565. [Google Scholar] [CrossRef] [PubMed]
- Jia, J.; Shi, L.-C.; Xu, Z.-C.; Xin, T.-Y.; Song, J.-Y.; Chen Shi, L. Identification of antler powder components based on DNA barcoding technology. Yao Xue Xue Bao Acta Pharm. Sin. 2015, 50, 1356–1361. [Google Scholar]
- Cui, Z.-H.; Jiang, C.; Li, M.-H.; Chen, M.; Zhou, L.-S.; Yuan, Y. Molecular identification of raw materials from lian qiao bai du wan. Yao Xue Xue Bao Acta Pharm. Sin. 2013, 48, 590–596. [Google Scholar]
- Shendure, J.; Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 2008, 26, 1135–1145. [Google Scholar] [CrossRef]
- Coghlan, M.L.; Haile, J.; Houston, J.; Murray, D.C.; White, N.E.; Moolhuijzen, P.; Bellgard, M.I.; Bunce, M. Deep sequencing of plant and animal DNA contained within traditional Chinese medicines reveals legality issues and health safety concerns. PLoS Genet. 2012, 8, e1002657. [Google Scholar] [CrossRef]
- Cummings, L.A.; Kurosawa, K.; Hoogestraat, D.R.; SenGupta, D.J.; Candra, F.; Doyle, M.; Thielges, S.; Land, T.A.; Rosenthal, C.A.; Hoffman, N.G.; et al. Clinical next generation sequencing outperforms standard microbiological culture for characterizing polymicrobial samples. Clin. Chem. 2016, 62, 1465–1473. [Google Scholar] [CrossRef]
- Korshunov, S.O.; Gorovtsov, A.V.; Faleeva, T.G.; Gorbov, S.N.; Kornienko, I.V. Comparison of biochemical and molecular genetic approaches for identification of environmental strains. Microbiology 2014, 83, 376–380. [Google Scholar] [CrossRef]
- Sarikhani, E.; Sagova-Mareckova, M.; Omelka, M.; Kopecky, J. The effect of peat and iron supplements on the severity of potato common scab and bacterial community in tuberosphere soil. FEMS Microbiol. Ecol. 2017, 93. [Google Scholar] [CrossRef] [PubMed]
- Evans, S.J.; Bassis, C.M.; Hein, R.; Assari, S.; Flowers, S.A.; Kelly, M.B.; Young, V.B.; Ellingrod, V.E.; McInnis, M.G. The gut microbiome composition associates with bipolar disorder and illness severity. J. Psychiatr. Res. 2017, 87, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Gopalakrishnan, V.; Spencer, C.N.; Nezi, L.; Reuben, A.; Andrews, M.C.; Karpinets, T.V.; Prieto, P.A.; Vicente, D.; Hoffman, K.; Wei, S.C.; et al. Gut microbiome modulates response to anti-PD-1 immunotherapy in melanoma patients. Science 2017, 359, 97–103. [Google Scholar] [CrossRef]
- Routy, B.; Le Chatelier, E.; Derosa, L.; Duong, C.P.M.; Alou, M.T.; Daillere, R.; Fluckiger, A.; Messaoudene, M.; Rauber, C.; Roberti, M.P.; et al. Gut microbiome influences efficacy of PD-1-based immunotherapy against epithelial tumors. Science 2017, 359, 91–97. [Google Scholar] [CrossRef]
- Sivan, A.; Corrales, L.; Hubert, N.; Williams, J.B.; Aquino-Michaels, K.; Earley, Z.M.; Benyamin, F.W.; Lei, Y.M.; Jabri, B.; Alegre, M.-L.; et al. Commensal bifidobacterium promotes antitumor immunity and facilitates anti–PD-L1 efficacy. Science 2015, 350, 1084–1089. [Google Scholar] [CrossRef]
- Pompanon, F.; Deagle, B.E.; Symondson, W.O.C.; Brown, D.S.; Jarman, S.N.; Taberlet, P. Who is eating what: Diet assessment using next generation sequencing. Mol. Ecol. 2012, 21, 1931–1950. [Google Scholar] [CrossRef]
- Zhou, X.; Li, Y.; Liu, S.; Yang, Q.; Su, X.; Zhou, L.; Tang, M.; Fu, R.; Li, J.; Huang, Q. Ultra-deep sequencing enables high-fidelity recovery of biodiversity for bulk arthropod samples without PCR amplification. Gigascience 2013, 2, 4. [Google Scholar] [CrossRef]
- Yu, D.W.; Ji, Y.; Emerson, B.C.; Wang, X.; Ye, C.; Yang, C.; Ding, Z. Biodiversity soup: Metabarcoding of arthropods for rapid biodiversity assessment and biomonitoring. Methods Ecol. Evol. 2012, 3, 613–623. [Google Scholar] [CrossRef]
- Marcelino, V.R.; Verbruggen, H. Multi-marker metabarcoding of coral skeletons reveals a rich microbiome and diverse evolutionary origins of endolithic algae. Sci. Rep. 2016, 6, 31508. [Google Scholar] [CrossRef]
- Hugenholtz, P.; Tyson, G.W. Microbiology: Metagenomics. Nature 2008, 455, 481–483. [Google Scholar] [CrossRef]
- Alanio, A.; Gits-Muselli, M.; Mercier-Delarue, S.; Dromer, F.; Bretagne, S. Diversity of pneumocystis jirovecii during infection revealed by ultra-deep pyrosequencing. Front. Microbiol. 2016, 7, 733. [Google Scholar] [CrossRef]
- Kermarrec, L.; Franc, A.; Rimet, F.; Chaumeil, P.; Humbert, J.F.; Bouchez, A. Next-generation sequencing to inventory taxonomic diversity in eukaryotic communities: A test for freshwater diatoms. Mol. Ecol. Resour. 2013, 13, 607–619. [Google Scholar] [CrossRef]
- Nguyen, T.-D.; Schmidt, B.; Zheng, Z.; Kwoh, C.-K. Efficient and accurate OTU clustering with GPU-based sequence alignment and dynamic dendrogram cutting. IEEE/ACM Trans. Comput. Biol. Bioinform. 2015, 12, 1060–1073. [Google Scholar] [CrossRef]
- Blanckenhorn, W.U.; Rohner, P.T.; Bernasconi, M.V.; Haugstetter, J.; Buser, A. Is qualitative and quantitative metabarcoding of dung fauna biodiversity feasible? Environ. Toxicol. Chem. 2016, 35, 1970–1977. [Google Scholar] [CrossRef]
- Cheng, X.; Su, X.; Chen, X.; Zhao, H.; Bo, C.; Xu, J.; Bai, H.; Ning, K. Biological ingredient analysis of traditional Chinese medicine preparation based on high-throughput sequencing: The story for Liuwei Dihuang Wan. Sci. Rep. 2014, 4, 5147. [Google Scholar] [CrossRef]
- Li, X.; Yang, Y.; Henry, R.J.; Rossetto, M.; Wang, Y.; Chen, S. Plant DNA barcoding: From gene to genome. Biol. Rev. 2015, 90, 157–166. [Google Scholar] [CrossRef]
- Singer, E.; Bushnell, B.; Coleman-Derr, D.; Bowman, B.; Bowers, R.M.; Levy, A.; Gies, E.A.; Cheng, J.-F.; Copeland, A.; Klenk, H.-P.; et al. High-resolution phylogenetic microbial community profiling. ISME J. 2016, 10, 2020–2032. [Google Scholar] [CrossRef]
- Goodwin, S.; McPherson, J.D.; McCombie, W.R. Coming of age: Ten years of next-generation sequencing technologies. Nat. Rev. Genet. 2016, 17, 333–351. [Google Scholar] [CrossRef]
- Eid, J.; Fehr, A.; Gray, J.; Luong, K.; Lyle, J.; Otto, G.; Peluso, P.; Rank, D.; Baybayan, P.; Bettman, B.; et al. Real-time DNA sequencing from single polymerase molecules. Science 2009, 323, 133–138. [Google Scholar] [CrossRef]
- Mosher, J.J.; Bowman, B.; Bernberg, E.L.; Shevchenko, O.; Kan, J.; Korlach, J.; Kaplan, L.A. Improved performance of the PacBio SMRT technology for 16S rDNA sequencing. J. Microbiol. Methods 2014, 104, 59–60. [Google Scholar] [CrossRef]
- George, N.; Flamiatos, E.; Kawasaki, K.; Kim, N.; Carriere, C.; Phan, B.; Joseph, R.; Strauss, S.; Kohli, R.; Choi, D.; et al. Oral microbiota species in acute apical endodontic abscesses. J. Oral Microbiol. 2016, 8, 30989. [Google Scholar] [CrossRef]
- Franzén, O.; Hu, J.; Bao, X.; Itzkowitz, S.H.; Peter, I.; Bashir, A. Improved OTU-picking using long-read 16S rRNA gene amplicon sequencing and generic hierarchical clustering. Microbiome 2015, 3, 43. [Google Scholar] [CrossRef]
- Jia, J.; Xu, Z.; Xin, T.; Shi, L.; Song, J. Quality control of the traditional patent medicine Yimu Wan based on SMRT sequencing and DNA barcoding. Front. Plant Sci. 2017, 8, 926. [Google Scholar] [CrossRef]
- Xin, T.; Xu, Z.; Jia, J.; Leon, C.; Hu, S.; Lin, Y.; Ragupathy, S.; Song, J.; Newmaster, S.G. Biomonitoring for traditional herbal medicinal products using DNA metabarcoding and single molecule, real-time sequencing. Acta Pharm. Sin. B 2017, 8, 488–497. [Google Scholar] [CrossRef]
- Xin, T.; Li, X.; Yao, H.; Lin, Y.; Ma, X.; Cheng, R.; Song, J.; Ni, L.; Fan, C.; Chen, S. Survey of commercial Rhodiola products revealed species diversity and potential safety issues. Sci. Rep. 2015, 5, 8337. [Google Scholar] [CrossRef]
- Chen, X.; Xiang, L.; Shi, L.; Li, G.; Yao, H.; Han, J.; Lin, Y.; Song, J.; Chen, S. Identification of crude drugs in the Japanese pharmacopoeia using a DNA barcoding system. Sci. Rep. 2017, 7, 42325. [Google Scholar] [CrossRef]
- Chinese Pharmacopoeia Commission. Pharmacopoeia of the People’s Republic of China; China Medical Science and Technology Press: Beijing, China, 2015. [Google Scholar]
- Folmer, O.; Black, M.; Hoeh, W.; Lutz, R.; Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 1994, 3, 294–299. [Google Scholar]
- Tate, J.; Simpson, B. Paraphyly of tarasa (malvaceae) and diverse origins of the polyploid species. Syst. Bot. 2003, 28, 723–738. [Google Scholar]
- Sang, T.; Crawford, D.; Stuessy, T. Chloroplast DNA phylogeny, reticulate evolution, and biogeography of Paeonia (Paeoniaceae). Am. J. Bot. 1997, 84, 1120. [Google Scholar] [CrossRef]
- Cheng, T.; Xu, C.; Lei, L.; Li, C.; Zhang, Y.; Zhou, S. Barcoding the kingdom Plantae: New PCR primers forITSregions of plants with improved universality and specificity. Mol. Ecol. Resour. 2016, 16, 138–149. [Google Scholar] [CrossRef]
- Standard Nucleotide BLAST. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi?PROGRAM=blastn&PAGE_TYPE=BlastSearch&LINK_LOC=blasthome (accessed on 1 April 2019).
- Zheng, X.; Zhang, P.; Liao, B.; Li, J.; Liu, X.; Shi, Y.; Cheng, J.; Lai, Z.; Xu, J.; Chen, S. A Comprehensive quality evaluation system for complex herbal medicine using PacBio sequencing, PCR-Denaturing gradient gel electrophoresis, and several chemical approaches. Front. Plant. Sci. 2017, 8, 1578. [Google Scholar] [CrossRef]
- Lebedeva, L.I.; Gerasimova, T.N. Peculiarities of Philodina roseola (Ehrbg.) (Rotatoria, Bdelloida)—Growth and reproduction under various temperature conditions. Int. Rev. Gesamten Hydrobiol. Hydrogr. 1985, 70, 509–525. [Google Scholar] [CrossRef]
- Scheff, D.S.; Sehgal, B.; Subramanyam, B. Evaluating penetration ability of Plodia interpunctella (Hübner) (Lepidoptera: Pyralidae) larvae into multilayer polypropylene packages. Insects 2018, 9, 42. [Google Scholar] [CrossRef]
- McMahan, E.E.; Guédot, C. Development of Sparganothis sulfureana (Lepidoptera: Tortricidae) on cranberry cultivars. Insects 2018, 9, 4. [Google Scholar] [CrossRef]
- Amarasekare, K.G.; Shearer, P.W. Stability of Cacopsylla pyricola (Hemiptera: Psyllidae) populations in Pacific Northwest pear Orchards managed with long-term mating disruption for Cydia pomonella (Lepidoptera: Tortricidae). Insects 2017, 8, 105. [Google Scholar] [CrossRef]
- Orkunoglu-Suer, F.; Harralson, A.F.; Frankfurter, D.; Gindoff, P.; O’Brien, T.J. Targeted single molecule sequencing methodology for ovarian hyperstimulation syndrome. BMC Genom. 2015, 16, 264. [Google Scholar] [CrossRef]
- Carneiro, M.O.; Russ, C.; Ross, M.G.; Gabriel, S.B.; Nusbaum, C.; DePristo, M.A. Pacific biosciences sequencing technology for genotyping and variation discovery in human data. BMC Genom. 2012, 13, 375. [Google Scholar] [CrossRef]
- Li, Q.; Li, Y.; Song, J.; Xu, H.; Xu, J.; Zhu, Y.; Li, X.; Gao, H.; Dong, L.; Qian, J.; et al. High-accuracyde novoassembly and SNP detection of chloroplast genomes using a SMRT circular consensus sequencing strategy. New Phytol. 2014, 204, 1041–1049. [Google Scholar] [CrossRef]
- Chen, X.; Liao, B.; Song, J.; Pang, X.; Han, J.; Chen, S. A fast SNP identification and analysis of intraspecific variation in the medicinal Panax species based on DNA barcoding. Gene 2013, 530, 39–43. [Google Scholar] [CrossRef]
- Li, D.Z.; Gao, L.M.; Li, H.T.; Wang, H.; Ge, X.J.; Liu, J.Q.; Chen, Z.D.; Zhou, S.L.; Chen, S.L.; Yang, J.B.; et al. Comparative analysis of a large dataset indicates that internal transcribed spacer (ITS) should be incorporated into the core barcode for seed plants. Proc. Natl. Acad. Sci. USA 2011, 108, 19641–19646. [Google Scholar]
- Pang, X.; Liu, C.; Shi, L.; Liu, R.; Liang, D.; Li, H.; Cherny, S.S.; Chen, S. Utility of the trnH-psbA intergenic spacer region and its combinations as plant DNA barcodes: A meta-analysis. PLoS ONE 2012, 7, e48833. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L. Biological identifications through DNA barcodes. Proc. R. Soc. Lond. Ser. B Biol. Sci. 2003, 270, 313–321. [Google Scholar] [CrossRef]
- Biswas, I.; Maguin, E.; Ehrlich, S.D.; Gruss, A. A 7-base-pair sequence protects DNA from exonucleolytic degradation in Lactococcus lactis. Proc. Natl. Acad. Sci. USA 1995, 92, 2244–2248. [Google Scholar] [CrossRef]
- Liu, W.; Kim, H.J.; Lucchetta, E.M.; Du, W.; Ismagilov, R.F. Isolation, incubation, and parallel functional testing and identification by FISH of rare microbial single-copy cells from multi-species mixtures using the combination of chemistrode and stochastic confinement. Lab Chip 2009, 9, 2153–2162. [Google Scholar] [CrossRef]
- Stoeckle, M.Y.; Gamble, C.C.; Kirpekar, R.; Young, G.; Ahmed, S.; Little, D.P. Commercial teas highlight plant DNA barcode identification successes and obstacles. Sci. Rep. 2011, 1, 42. [Google Scholar] [CrossRef]
- Chang, C.-H.; Jang-Liaw, N.-H.; Lin, Y.-S.; Fang, Y.-C.; Shao, K.-T. Authenticating the use of dried seahorses in the traditional Chinese medicine market in Taiwan using molecular forensics. J. Food Drug Anal. 2013, 21, 310–316. [Google Scholar] [CrossRef][Green Version]
- Yan, D.; Luo, J.Y.; Han, Y.M.; Peng, C.; Dong, X.P.; Chen, S.L.; Sun, L.G.; Xiao, X.H. Forensic DNA barcoding and bio-response studies of animal horn products used in traditional medicine. PLoS ONE 2013, 8, e55854. [Google Scholar] [CrossRef]
- Williams, J.H.; Phillips, T.D.; Jolly, P.E.; Stiles, J.K.; Jolly, C.M.; Aggarwal, D. Human aflatoxicosis in developing countries: A review of toxicology, exposure, potential health consequences, and interventions. Am. J. Clin. Nutr. 2004, 80, 1106–1122. [Google Scholar] [CrossRef]
- Pawlowski, J.; Audic, S.; Adl, S.; Bass, D.; Belbahri, L.; Berney, C.; Bowser, S.S.; Cepicka, I.; Decelle, J.; Dunthorn, M.; et al. CBOL protist working group: Barcoding eukaryotic richness beyond the animal, plant, and fungal kingdoms. PLoS Biol. 2012, 10, e1001419. [Google Scholar] [CrossRef]
- Newmaster, S.G.; Fazekas, A.J.; Ragupathy, S. DNA barcoding in land plants: Evaluation of rbcL in a multigene tiered approach. Can. J. Bot. 2006, 84, 335–341. [Google Scholar] [CrossRef]
- Kumeta, Y.; Maruyama, T.; Asama, H.; Yamamoto, Y.; Hakamatsuka, T.; Goda, Y. Species identification of Asini Corii Collas (donkey glue) by PCR amplification of cytochrome b gene. J. Nat. Med. 2014, 68, 181–185. [Google Scholar] [CrossRef]
- Chen, S. I19 Major achievements of traditional medicine in treating epidemic and chronic diseases. Biochem. Pharmacol. 2017, 139, 110. [Google Scholar] [CrossRef]
- Ni, Q.; Wang, J.; Li, E.-Q.; Zhao, A.-B.; Yu, B.; Wang, M.; Huang, C.-R. Study on the protective effect of the mixture of Shengmai powder and danshen decoction on the myocardium of diabetic cardiomyopathy in the rat model. Chin. J. Integr. Med. 2011, 17, 116–125. [Google Scholar] [CrossRef]
- Zhao, Z.; Wang, H.; Yuan, H.; Zhang, C.; Bao, G.; Zhao, Z.; Song, B.; Bao, L. Anti-inflammatory, diuretic and removing urinary calculus actions of Sanweijili powder. Mod. J. Integr. Tradit. Chin. West. Med. 2009, 23, 2766–2767. [Google Scholar]
- Ko, R. Safety of ethnic & imported herbal and dietary supplements. Clin. Toxicol. 2006, 44, 611–616. [Google Scholar]




| Formula | Samples (and Its Biological Origin) | DNA ID | Notes |
|---|---|---|---|
| Sheng-Mai-San | Ginseng Radix et Rhizoma (Panax ginseng) | RS1 | Company A |
| RS2 | Company A | ||
| RS3 | Company A | ||
| Ophiopogonis Radix (Ophiopogon japonicus) | MD1 | ‘Zhe’ O. japonicus; Cixi, Zhejiang province; producing area | |
| MD2 | ‘Zhe’ O. japonicus; Xiangshan, Zhejiang province; wildness | ||
| MD3 | ‘Chuan’ O. japonicus; Santai, Sichuan province; Market B | ||
| MD4 | ‘Chuan’ O. japonicus; Sichuan province; Company C | ||
| Schisandrae Chinensis Fructus (Schisandra chinensis) | WW1 | Company C | |
| WW2 | Company C | ||
| WW3 | Company C | ||
| Sheng-Mai-San (SMS) | SMS1 | Mixed powder | |
| SMS2 | Mixed powder | ||
| SMS3 | Mixed powder | ||
| DNA mixture (HSM) | HMA | DNA Volume of MD1-RS1-WW1 = 3:3:2 | |
| HMB | DNA Volume of MD2-RS2-WW2 = 3:3:2 | ||
| HMC | DNA Volume of MD3-RS3-WW3 = 3:3:2 | ||
| Sanwei-Jili-San | Malvae Fructus (Malva veriticillata) | DK | Company D |
| Tribuli Fructus (Tribulus terrestris) | JL | Company D | |
| Chinese fresh-water crab (Eriocheir sinensis) | FH | A local supermarket | |
| Sanwei-Jili-San (SJS) | SJS1 | Mixed powder | |
| SJS2 | Mixed powder | ||
| SJS3 | Mixed powder | ||
| DNA mixture (JH) | JHA | DNA Volume of DK-JL-FH = 3:5:3 | |
| JHB | DNA Volume of DK-JL-FH = 3:5:3 | ||
| JHC | DNA Volume of DK-JL-FH = 3:5:3 |
| Raw Data | Circular Consensus Sequence Filtration | Extraction | |
|---|---|---|---|
| Reads | 57,147 | 36,497 | 25,877 |
| Total bases | 1.784 × 109 | 19,308,858 | 14,117,133 |
| Data size | 24.4 GB | 21.3 MB | 13.7 MB |
| Mean length | 31,217 bp | 529 bp | 546 bp |
| Length range | 0~70 kb | 200~799 bp | 348~799 bp |
| SMRT Cell | DNA ID | Pair of Tag-Primers # | Resulting Reads | No-Blast-Hit1 Resulting Reads | Effective Rate |
|---|---|---|---|---|---|
| Cell A | HM1 | T17 | 3255 | 0 | 100.00% |
| HM2 | T18 | 3748 | 0 | 100.00% | |
| HM3 | T19 | 4058 | 0 | 100.00% | |
| SMS1 | T13 | 2527 | 0 | 100.00% | |
| SMS2 | T14 | 3095 | 0 | 100.00% | |
| SMS3 | T15 | 4443 | 0 | 100.00% | |
| SMS1 | i13 | 603 | 0 | 100.00% | |
| SMS2 | i14 | 1030 | 0 | 100.00% | |
| SMS3 | i15 | 890 | 0 | 100.00% | |
| HMA | i17 | 557 | 0 | 100.00% | |
| HMB | i18 | 890 | 0 | 100.00% | |
| HMC | i19 | 778 | 0 | 100.00% | |
| Cell B | RS1 | i5 | 279 | 0 | 100.00% |
| RS2 | i6 | 439 | 0 | 100.00% | |
| RS3 | i7 | 462 | 0 | 100.00% | |
| WW1 | i9 | 431 | 0 | 100.00% | |
| WW2 | i10 | 706 | 0 | 100.00% | |
| WW3 | i11 | 686 | 0 | 100.00% | |
| MD1 | T1 | 725 | 0 | 100.00% | |
| MD2 | T2 | 578 | 0 | 100.00% | |
| MD3 | T3 | 749 | 0 | 100.00% | |
| MD4 | T4 | 794 | 1 | 99.87% | |
| MD1 ** | i1 | 268 | 2 | 99.25% | |
| MD2 ** | i2 | 512 | 7 | 98.63% | |
| MD3 ** | i3 | 568 | 14 | 97.54% | |
| MD4 ** | i4 | 302 | 8 | 97.35% | |
| WW1 | T9 | 1276 | 0 | 100.00% | |
| WW2 | T10 | 445 | 0 | 100.00% | |
| WW3 | T11 | 1846 | 0 | 100.00% | |
| RS1 | T5 | 911 | 0 | 100.00% | |
| RS2 | T6 | 988 | 0 | 100.00% | |
| RS3 | T7 | 1617 | 0 | 100.00% | |
| Cell C | MD3 * | i3 | 291 | 0 | 100.00% |
| JL | i39 | 777 | 0 | 100.00% | |
| DK | i40 | 1396 | 0 | 100.00% | |
| FH | C9 | 2067 | 0 | 100.00% | |
| SJS1 | i23 | 1977 | 0 | 100.00% | |
| SJS2 | i22 | 3091 | 0 | 100.00% | |
| SJS3 | i21 | 1559 | 0 | 100.00% | |
| SJS1 | C1 | 898 | 2 | 99.78% | |
| SJS2 | C2 | 1041 | 0 | 100.00% | |
| SJS3 | C3 | 324 | 0 | 100.00% | |
| JHC | C6 | 1074 | 7 | 99.35% | |
| JHC | i32 | 2182 | 0 | 100.00% | |
| JHB | i31 | 1751 | 0 | 100.00% | |
| JHA | i24 | 1348 | 0 | 100.00% | |
| Total | 60,232 | 41 | 99.93% | ||
| Briefing | Reference | |
|---|---|---|
| Sanger sequencing | The trnH-psbA could distinguish 18 species of Polygonaceae and their adulterants including 10 species that recorded in Chinese pharmacopoeia. | [10] |
| The discrimination ability of ITS2, a most suitable region for DNA barcoding, were tested by more than 6600 plant samples and its successful identification rate was 92.7% at the species level. | [8] | |
| Most (26/44) of the North American herbal products tested contained DNA barcodes from plant species not listed on the labels. Sequences of different species had yield up from the same sample product. | [9] | |
| DNA barcoding was used to authenticate the components of antler powder in the market, while a few samples containing multi-species were analyzed by cloning method. | [14] | |
| Short-read NGS | Amplicons of trnL and 16S from 15 TCM samples were sequenced in platform of Roche GS Junior, and over 49,000 amplicon sequence reads were generated. Many trnL sequence reads could not been precisely identified (most were just assigned to genera level), due to the limitation of read-length and reference database. | [17] |
| HTS data for 30 Liuwei-Dihuang-Wan samples were generated based on 454 GS FLX Titanium sequencing. On averages, 3 and 2.4 prescribed species could be detected from a sample based on ITS2 and trnL, respectively. Vigna genus, a possible contaminated species, was detected in all three reference samples. | [34] | |
| Long-read NGS | A total of 3703 and 4810 CCS reads from two reference and three commercial Yimu-Wan samples were mapped to the ITS2 and psbA-trnH regions, respectively. SMRT sequencing provides an affordable way to monitor the legality and safety of traditional patent medicines. | [42] |
| A comprehensive quality evaluation system for herbal medicine was established, by combining two genetic-based approaches—third-generation sequencing and denaturing gradient gel electrophoresis, with analytical chemistry approaches. | [52] | |
| SMS and SJS have been analyzed through FLMB by an adequate and well-controlled methodological analysis, which shows perfect interpretation for DNA barcoding. | This study |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, P.; Liu, C.; Zheng, X.; Wu, L.; Liu, Z.; Liao, B.; Shi, Y.; Li, X.; Xu, J.; Chen, S. Full-Length Multi-Barcoding: DNA Barcoding from Single Ingredient to Complex Mixtures. Genes 2019, 10, 343. https://doi.org/10.3390/genes10050343
Zhang P, Liu C, Zheng X, Wu L, Liu Z, Liao B, Shi Y, Li X, Xu J, Chen S. Full-Length Multi-Barcoding: DNA Barcoding from Single Ingredient to Complex Mixtures. Genes. 2019; 10(5):343. https://doi.org/10.3390/genes10050343
Chicago/Turabian StyleZhang, Peng, Chunsheng Liu, Xiasheng Zheng, Lan Wu, Zhixiang Liu, Baosheng Liao, Yuhua Shi, Xiwen Li, Jiang Xu, and Shilin Chen. 2019. "Full-Length Multi-Barcoding: DNA Barcoding from Single Ingredient to Complex Mixtures" Genes 10, no. 5: 343. https://doi.org/10.3390/genes10050343
APA StyleZhang, P., Liu, C., Zheng, X., Wu, L., Liu, Z., Liao, B., Shi, Y., Li, X., Xu, J., & Chen, S. (2019). Full-Length Multi-Barcoding: DNA Barcoding from Single Ingredient to Complex Mixtures. Genes, 10(5), 343. https://doi.org/10.3390/genes10050343




