Dynamic Expression of Interferon Lambda Regulated Genes in Primary Fibroblasts and Immune Organs of the Chicken
Abstract
:1. Introduction
2. Material and Methods
2.1. Cells
2.2. Data mining and Bioinformatic Analysis of Chicken IFN-lambda (chIFN-λ)
2.3. Expression of Recombinant chIFN-λ3 (rchIFN-λ3)
2.4. Preparation of Primary Chicken Embryo Fibroblast
2.5. Birds and Management
2.6. RNA Extraction and Sequencing
2.7. RNA-Seq Quality
2.8. Gene Ontology (GO) and KEGG Enrichment Analysis
3. Results
3.1. Bioinformatic Analysis of chIFN-λ
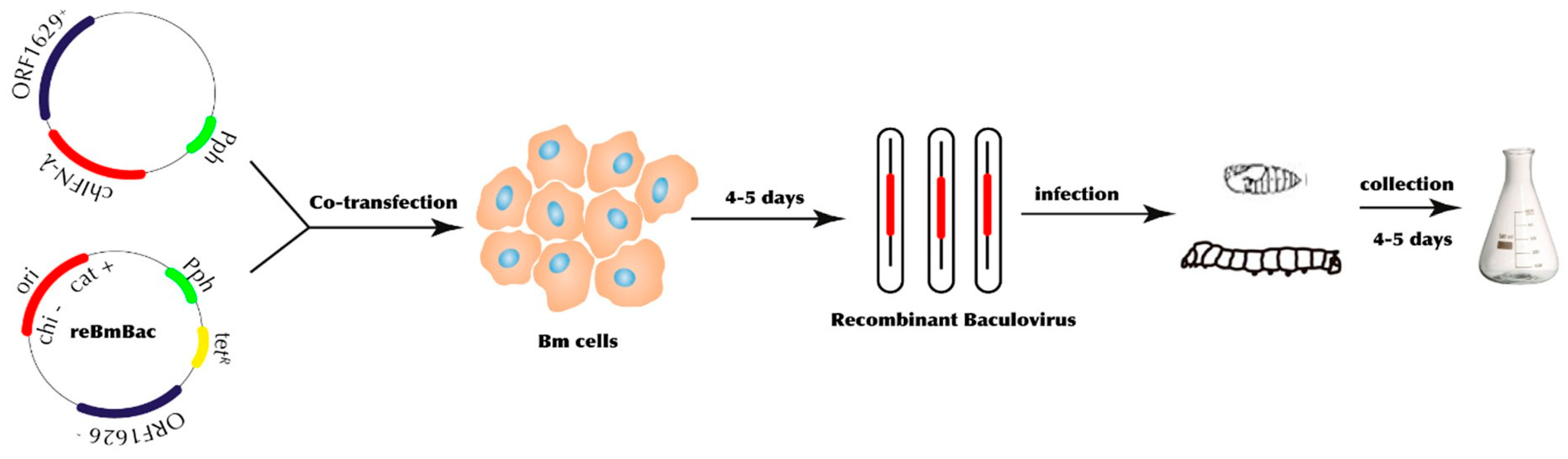
3.2. Expression of chIFN-λ in Baculovirus Expression Vector System (BEVS)
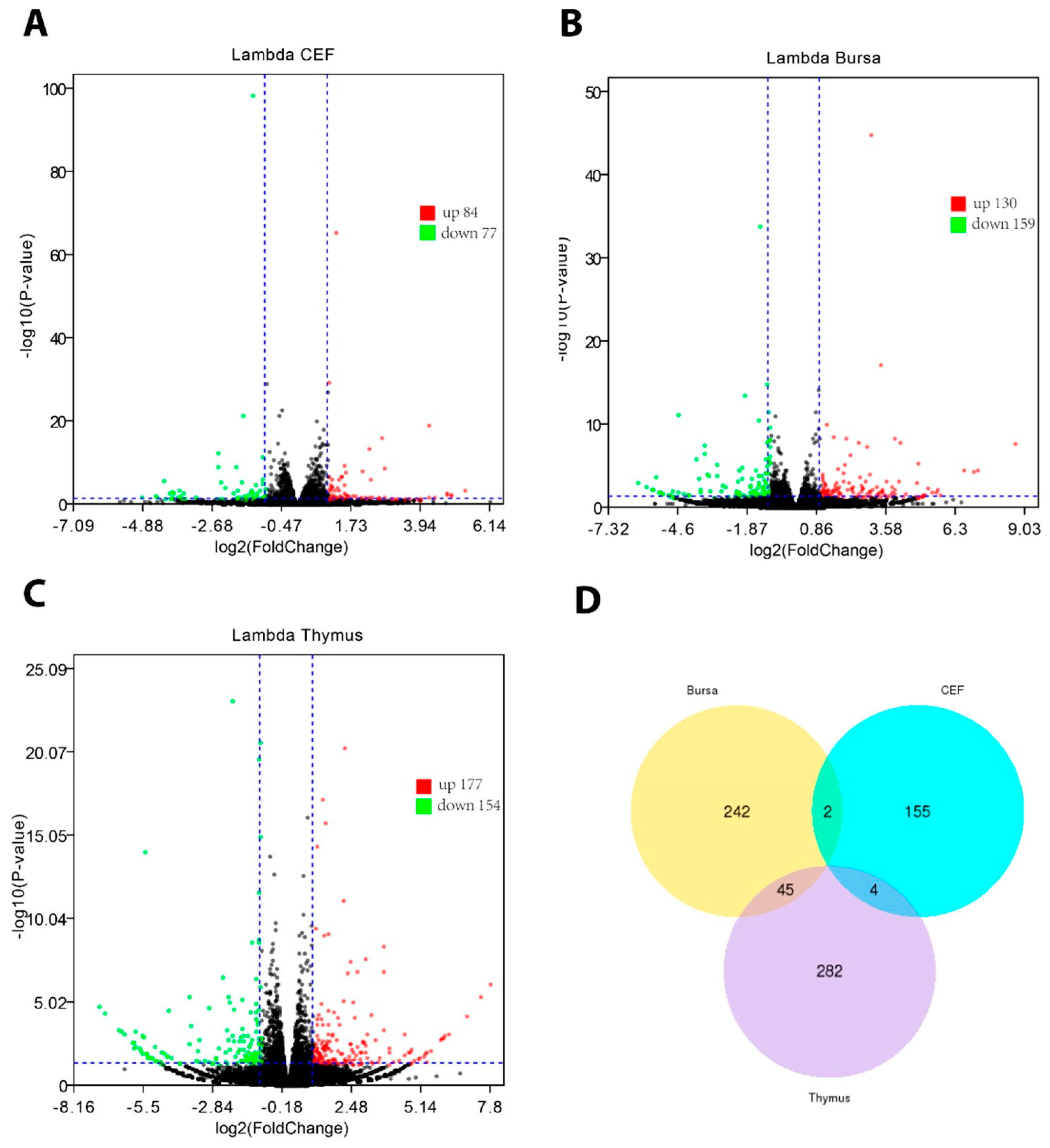
3.3. Characterization of chIFN-λ-induced Gene Expression in Chicken
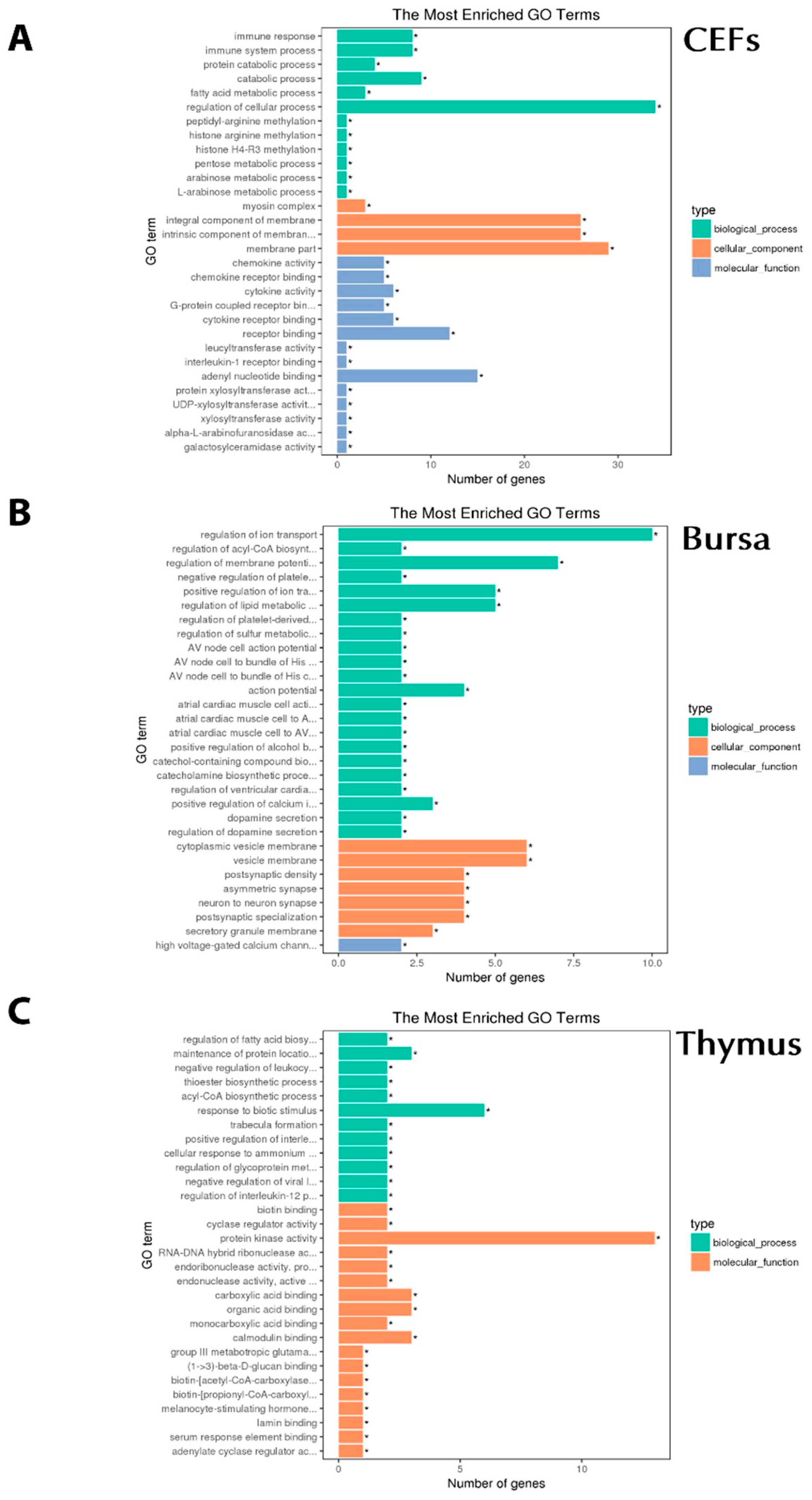
3.4. Functional Analysis of DEGs
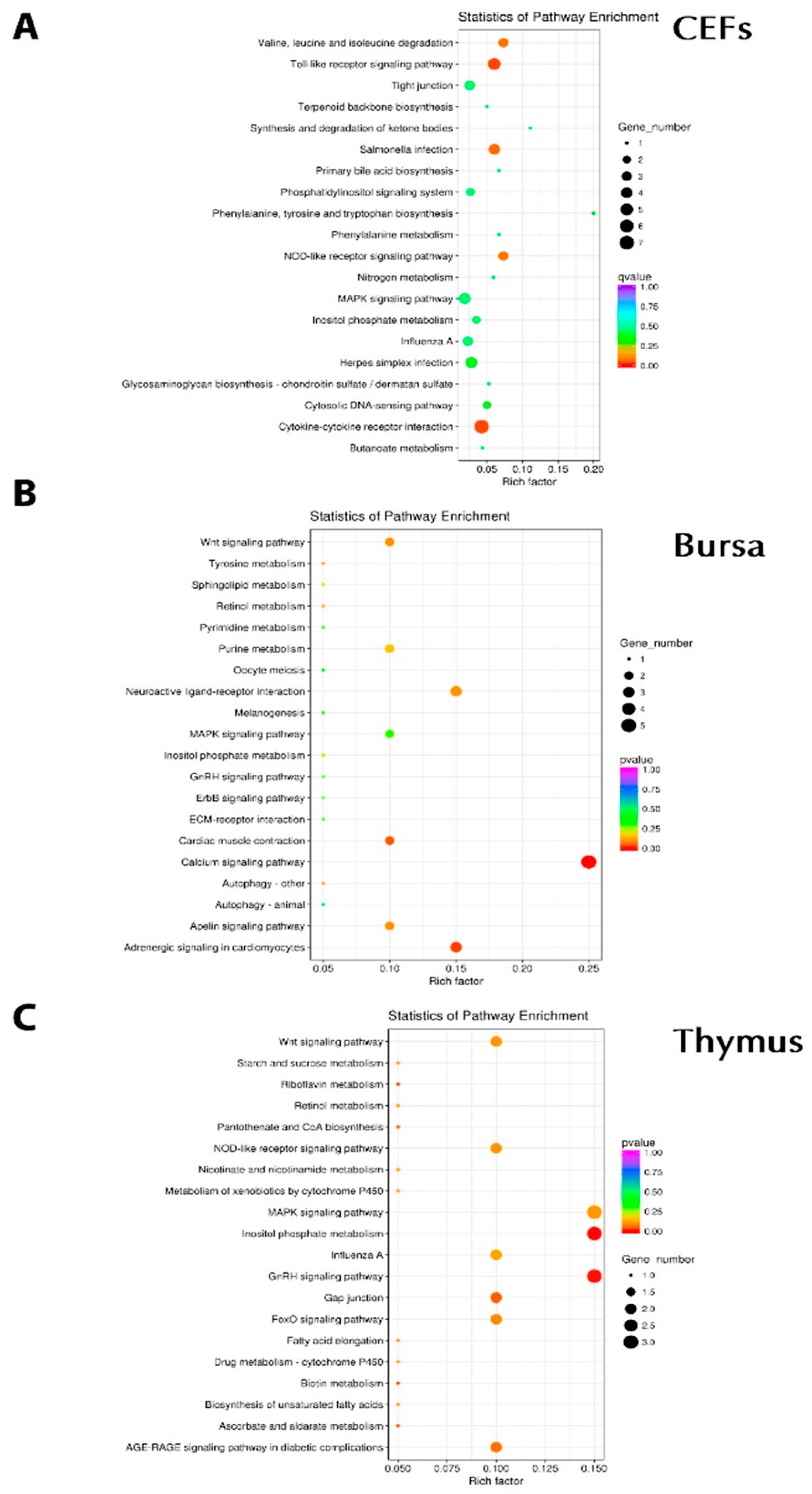
3.5. KEGG Pathway Enrichment
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Fusaro, A.; Monne, I.; Salviato, A.; Valastro, V.; Schivo, A.; Amarin, N.M.; Gonzalez, C.; Ismail, M.M.; Al-Ankari, A.-R.; Al-Blowi, M.H. Phylogeography and evolutionary history of reassortant H9N2 viruses with potential human health implications. J. Virol. 2011, 85, 8413–8421. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.-H.; Song, C.-S. H9N2 avian influenza virus in Korea: Evolution and vaccination. Clin. Exp. Vaccine Res. 2013, 2, 26–33. [Google Scholar] [CrossRef] [PubMed]
- Santhakumar, D.; Rubbenstroth, D.; Martinez-Sobrido, L.; Munir, M. Avian interferons and their antiviral effectors. Front. Immunol. 2017, 8, 49. [Google Scholar] [CrossRef] [PubMed]
- Hardy, M.P.; Sanij, E.P.; Hertzog, P.J.; Owczarek, C.M. Characterization and transcriptional analysis of the mouse Chromosome 16 cytokine receptor gene cluster. Mamm. Genome 2003, 14, 105–118. [Google Scholar] [CrossRef] [PubMed]
- Kotenko, S.V.; Gallagher, G.; Baurin, V.V.; Lewis-Antes, A.; Shen, M.; Shah, N.K.; Langer, J.A.; Sheikh, F.; Dickensheets, H.; Donnelly, R.P. IFN-λs mediate antiviral protection through a distinct class II cytokine receptor complex. Nat. Immunol. 2003, 4, 69. [Google Scholar] [CrossRef] [PubMed]
- Ank, N.; West, H.; Bartholdy, C.; Eriksson, K.; Thomsen, A.R.; Paludan, S.R. Lambda interferon (IFN-λ), a type III IFN, is induced by viruses and IFNs and displays potent antiviral activity against select virus infections in vivo. J. Virol. 2006, 80, 4501–4509. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, P. Advances in avian immunology—prospects for disease control: A review. Avian Pathol. 2010, 39, 309–324. [Google Scholar] [CrossRef] [PubMed]
- Hemann, E.A.; Gale, M., Jr.; Savan, R. Interferon lambda genetics and biology in regulation of viral control. Front. Immunol. 2017, 8, 1707. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Fu, D.; Chen, H.; Zhang, Z.; Shi, Q.; Elsayed, A.K.; Li, B. Partial antiviral activities detection of chicken Mx jointing with neuraminidase gene (NA) against Newcastle disease virus. PLoS ONE 2013, 8, e71688. [Google Scholar] [CrossRef] [PubMed]
- Reuter, A.; Soubies, S.; Härtle, S.; Schusser, B.; Kaspers, B.; Staeheli, P.; Rubbenstroth, D. Antiviral activity of lambda interferon in chickens. J. Virol. 2014, 88, 2835–2843. [Google Scholar] [CrossRef] [PubMed]
- Plachý, J.; Weining, K.C.; Kremmer, E.; Puehler, F.; Hala, K.; Kaspers, B.; Staeheli, P. Protective effects of type I and type II interferons toward Rous sarcoma virus-induced tumors in chickens. Virology 1999, 256, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Penski, N.; Härtle, S.; Rubbenstroth, D.; Krohmann, C.; Ruggli, N.; Schusser, B.; Pfann, M.; Reuter, A.; Gohrbandt, S.; Hundt, J. Highly pathogenic avian influenza viruses do not inhibit interferon synthesis in infected chickens but can override the interferon-induced antiviral state. J. Virol. 2011, 85, 7730–7741. [Google Scholar] [CrossRef] [PubMed]
- Karpala, A.J.; Morris, K.R.; Broadway, M.M.; McWaters, P.G.; O’Neil, T.E.; Goossens, K.E.; Lowenthal, J.W.; Bean, A.G. Molecular cloning, expression, and characterization of chicken IFN-λ. J. Interferon Cytokine Res. 2008, 28, 341–350. [Google Scholar] [CrossRef] [PubMed]
- Masuda, Y.; Matsuda, A.; Usui, T.; Sugai, T.; Asano, A.; Yamano, Y. Biological effects of chicken type III interferon on expression of interferon-stimulated genes in chickens: Comparison with type I and type II interferons. J. Vet. Med Sci. 2012, 74, 1381–1386. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wei, Y.; Li, Y.; Li, H.; Yang, X.; Yi, Y.; Zhang, Z. A Highly Efficient and Simple Construction Strategy for Producing Recombinant Baculovirus Bombyx mori Nucleopolyhedrovirus. PLoS ONE 2016, 11, e0152140. [Google Scholar] [CrossRef] [PubMed]
- Dojima, T.; Nishina, T.; Kato, T.; Uno, T.; Yagi, H.; Kato, K.; Park, E.Y. Comparison of the N-linked glycosylation of human beta1,3-N-acetylglucosaminyltransferase 2 expressed in insect cells and silkworm larvae. J. Biotechnol. 2009, 143, 27–33. [Google Scholar] [CrossRef] [PubMed]
- Usami, A.; Ishiyama, S.; Enomoto, C.; Okazaki, H.; Higuchi, K.; Ikeda, M.; Yamamoto, T.; Sugai, M.; Ishikawa, Y.; Hosaka, Y.; et al. Comparison of recombinant protein expression in a baculovirus system in insect cells (Sf9) and silkworm. J. Biochem. 2011, 149, 219–227. [Google Scholar] [CrossRef] [PubMed]
- Summers, M.D.; Smith, G.E. A Manual of Methods for Baculovirus Vectors and Insect Cell Culture Procedures; Texas Agricultural Experiment Station: College Station, TX, USA, 1987. [Google Scholar]
- Kruger, N.J. The Bradford method for protein quantitation. In The Protein Protocols Handbook; Springer: Oxford, UK, 2002; pp. 15–21. [Google Scholar]
- Ge, J.; Wen, Z.; Wang, X.; Hu, S.; Liu, Y.; Kong, X.; Chen, H.; Bu, Z. Generating Vesicular Stomatitis Virus Pseudotype Bearing the Severe Acute Respiratory Syndrome Coronavirus Spike Envelope Glycoprotein for Rapid and Safe Neutralization Test or Cell-Entry Assay. Ann. N. Y. Acad. Sci. 2006, 1081, 246–248. [Google Scholar] [CrossRef] [PubMed]
- Swayne, D.E. Laboratory Manual for the Isolation and Identification of Avian Pathogens; American Association of Avian Pathologists: Jacksonville, FL, USA, 1998. [Google Scholar]
- Rio, D.C.; Ares, M.; Hannon, G.J.; Nilsen, T.W. Purification of RNA using TRIzol (TRI reagent). Cold Spring Harb. Protoc. 2010, 2010, pdb. prot5439. [Google Scholar] [CrossRef]
- Marcello, T.; Grakoui, A.; Barba–Spaeth, G.; Machlin, E.S.; Kotenko, S.V.; Macdonald, M.R.; Rice, C.M. Interferons α and λ inhibit hepatitis C virus replication with distinct signal transduction and gene regulation kinetics. Gastroenterology 2006, 131, 1887–1898. [Google Scholar] [CrossRef] [PubMed]
- Schilling, M.A.; Katani, R.; Memari, S.; Cavanaugh, M.; Buza, J.; Radzio-Basu, J.; Mpenda, F.N.; Deist, M.S.; Lamont, S.J.; Kapur, V. Transcriptional Innate Immune Response of the Developing Chicken Embryo to Newcastle Disease Virus Infection. Front. Genet. 2018, 9, 61. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.L.; Lombardi, G.; Foster, G.R. Type I interferons and the innate immune response—more than just antiviral cytokines. Mol. Immunol. 2005, 42, 869–877. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, P.; Poh, T.Y.; Rothwell, L.; Avery, S.; Balu, S.; Pathania, U.S.; Hughes, S.; Goodchild, M.; Morrell, S.; Watson, M. A genomic analysis of chicken cytokines and chemokines. J. Interferon Cytokine Res. 2005, 25, 467–484. [Google Scholar] [CrossRef] [PubMed]
- Goossens, K.E.; Karpala, A.J.; Ward, A.; Bean, A.G. Characterisation of chicken ZAP. Dev. Comp. Immunol. 2014, 46, 373–381. [Google Scholar] [CrossRef] [PubMed]
- Mattijssen, S.; Pruijn, G.J. Viperin, a key player in the antiviral response. Microbes Infect. 2012, 14, 419–426. [Google Scholar] [CrossRef] [PubMed]
- Yount, J.S.; Moltedo, B.; Yang, Y.-Y.; Charron, G.; Moran, T.M.; López, C.B.; Hang, H.C. Palmitoylome profiling reveals S-palmitoylation–dependent antiviral activity of IFITM3. Nat. Chem. Biol. 2010, 6, 610. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Markosyan, R.M.; Zheng, Y.-M.; Golfetto, O.; Bungart, B.; Li, M.; Ding, S.; He, Y.; Liang, C.; Lee, J.C. IFITM proteins restrict viral membrane hemifusion. PLoS Pathogens 2013, 9, e1003124. [Google Scholar] [CrossRef] [PubMed]
- Amini-Bavil-Olyaee, S.; Choi, Y.J.; Lee, J.H.; Shi, M.; Huang, I.-C.; Farzan, M.; Jung, J.U. The antiviral effector IFITM3 disrupts intracellular cholesterol homeostasis to block viral entry. Cell Host Microbe 2013, 13, 452–464. [Google Scholar] [CrossRef] [PubMed]
- Heidari, M.; Sarson, A.J.; Huebner, M.; Sharif, S.; Kireev, D.; Zhou, H. Marek’s disease virus–induced immunosuppression: Array analysis of chicken immune response gene expression profiling. Viral Immunol. 2010, 23, 309–319. [Google Scholar] [CrossRef] [PubMed]
- Munoz, I.; Berges, M.; Bonsergent, C.; Cormier-Aline, F.; Quéré, P.; Sibille, P. Cloning, expression and functional characterization of chicken CCR6 and its ligand CCL20. Mol. Immunol. 2009, 47, 551–559. [Google Scholar] [CrossRef] [PubMed]
- Iellem, A.; Mariani, M.; Lang, R.; Recalde, H.; Panina-Bordignon, P.; Sinigaglia, F.; D’Ambrosio, D. Unique chemotactic response profile and specific expression of chemokine receptors CCR4 and CCR8 by CD4+ CD25+ regulatory T cells. J. Exp. Med. 2001, 194, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Hughes, S.; Haynes, A.; O’Regan, M.; Bumstead, N. Identification, mapping, and phylogenetic analysis of three novel chicken CC chemokines. Immunogenetics 2001, 53, 674–683. [Google Scholar] [CrossRef] [PubMed]
- Keestra, A.M.; de Zoete, M.R.; Bouwman, L.I.; Vaezirad, M.M.; van Putten, J.P. Unique features of chicken Toll-like receptors. Dev. Comp. Immunol. 2013, 41, 316–323. [Google Scholar] [CrossRef] [PubMed]
- Nerren, J.R.; He, H.; Genovese, K.; Kogut, M.H. Expression of the avian-specific toll-like receptor 15 in chicken heterophils is mediated by gram-negative and gram-positive bacteria, but not TLR agonists. Vet. Immunol. Immunopathol. 2010, 136, 151–156. [Google Scholar] [CrossRef] [PubMed]
- Oven, I.; Rus, K.R.; Dušanić, D.; Benčina, D.; Keeler, C.L.; Narat, M. Diacylated lipopeptide from Mycoplasma synoviae mediates TLR15 induced innate immune responses. Vet. Res. 2013, 44, 99. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Jiang, J.; Tien, P.; Liu, W.; Li, J. IFN-λ: A new spotlight in innate immunity against influenza virus infection. Protein Cell 2018, 9, 832–837. [Google Scholar]
- Cao, Y.; Huang, Y.; Xu, K.; Liu, Y.; Li, X.; Xu, Y.; Zhong, W.; Hao, P. Differential responses of innate immunity triggered by different subtypes of influenza a viruses in human and avian hosts. BMC Med. Genom. 2017, 10, 70. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Zou, T.; Hu, X.; Jin, H. Type III interferon gene expression in response to influenza virus infection in chicken and duck embryonic fibroblasts. Mol. Immunol. 2015, 68, 657–662. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.-D.; Feng, N.; Sen, A.; Balan, M.; Tseng, H.-C.; McElrath, C.; Smirnov, S.V.; Peng, J.; Yasukawa, L.L.; Durbin, R.K. Distinct roles of type I and type III interferons in intestinal immunity to homologous and heterologous rotavirus infections. PLoS Pathogens 2016, 12, e1005600. [Google Scholar]
- Hernández, P.P.; Mahlakõiv, T.; Yang, I.; Schwierzeck, V.; Nguyen, N.; Guendel, F.; Gronke, K.; Ryffel, B.; Hölscher, C.; Dumoutier, L. Interferon-λ and interleukin 22 act synergistically for the induction of interferon-stimulated genes and control of rotavirus infection. Nat. Immunol. 2015, 16, 698. [Google Scholar] [CrossRef] [PubMed]
- Fensterl, V.; Sen, G.C. Interferons and viral infections. Biofactors 2009, 35, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Rathinam, V.A.; Fitzgerald, K.A. Innate immune sensing of DNA viruses. Virology 2011, 411, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Saito, A.; Cai, L.; Matsuhisa, K.; Ohtake, Y.; Kaneko, M.; Kanemoto, S.; Asada, R.; Imaizumi, K. Neuronal activity-dependent local activation of dendritic unfolded protein response promotes expression of brain-derived neurotrophic factor in cell soma. J. Neurochem. 2018, 144, 35–49. [Google Scholar] [CrossRef] [PubMed]

© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Arslan, M.; Yang, X.; Santhakumar, D.; Liu, X.; Hu, X.; Munir, M.; Li, Y.; Zhang, Z. Dynamic Expression of Interferon Lambda Regulated Genes in Primary Fibroblasts and Immune Organs of the Chicken. Genes 2019, 10, 145. https://doi.org/10.3390/genes10020145
Arslan M, Yang X, Santhakumar D, Liu X, Hu X, Munir M, Li Y, Zhang Z. Dynamic Expression of Interferon Lambda Regulated Genes in Primary Fibroblasts and Immune Organs of the Chicken. Genes. 2019; 10(2):145. https://doi.org/10.3390/genes10020145
Chicago/Turabian StyleArslan, Mehboob, Xin Yang, Diwakar Santhakumar, Xingjian Liu, Xiaoyuan Hu, Muhammad Munir, Yinü Li, and Zhifang Zhang. 2019. "Dynamic Expression of Interferon Lambda Regulated Genes in Primary Fibroblasts and Immune Organs of the Chicken" Genes 10, no. 2: 145. https://doi.org/10.3390/genes10020145
APA StyleArslan, M., Yang, X., Santhakumar, D., Liu, X., Hu, X., Munir, M., Li, Y., & Zhang, Z. (2019). Dynamic Expression of Interferon Lambda Regulated Genes in Primary Fibroblasts and Immune Organs of the Chicken. Genes, 10(2), 145. https://doi.org/10.3390/genes10020145




