A Probabilistic Matrix Factorization Method for Identifying lncRNA-Disease Associations
Abstract
:1. Introduction
2. Materials
2.1. Human LncRNA-MiRNA Associations and MiRNA-Disease Associations
2.2. Human LncRNA-Disease Associations
- Step 1:
- Obtaining the set of different lncRNAs shared in both LD and LM. In addition, for convenience, we denoted the set of these shared lncRNAs as con_L.
- Step 2:
- Obtaining the set of different diseases existed in MD and LD. In addition, for convenience, we denoted the set of these shared diseases as con_D.
- Step 3:
- Obtaining the set of lncRNA-disease associations with both lncRNAs in con_L and diseases in con_D based on the set of LD.
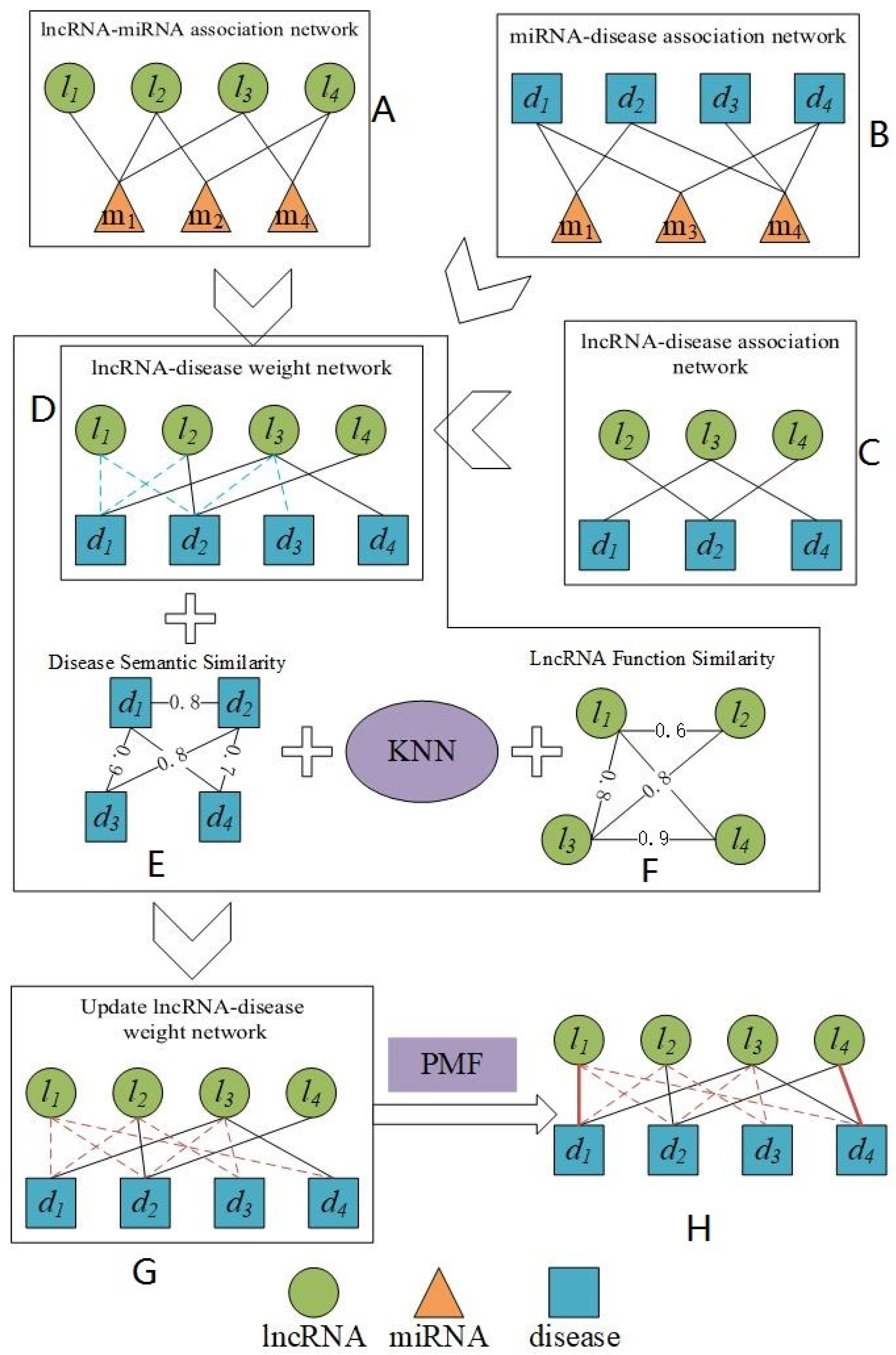
3. Methods
3.1. Construction of the lncRNA-miRNA Association Network and miRNA-Disease Association Network
- Step 1:
- Supposing that there are different lncRNAs in LM, and for convenience, we denote the set of these lncRNAs as .
- Step 2:
- Supposing that there are different diseases in MD, and for convenience, we denote the set of these diseases as .
- Step 3:
- Supposing that there are different common miRNAs existed in both MD and LM, and for convenience, we denote the set of these miRNAs as .
- Step 4:
- Hence, we can firstly obtain an lncRNA-miRNA association network , where denotes the set of experimentally verified known associations in LM. For , we define that there is an edge between and in if and only if there is an experimentally verified known associations between and in LM.
- Step 5:
- Simultaneously, we can also obtain an miRNA-disease association network , where denotes the set of experimentally verified known associations in MD. For , we define that there is an edge between and in if and only if there is an experimentally verified known associations between and in MD.
3.2. Construction of the Weighted lncRNA-Disease Association Network
- Step 1:
- Supposing that there are T different miRNAs in with experimentally verified known associations with both in LM and in MD simultaneously, and for convenience, we denote the set of these T different miRNAs as .
- Step 2:
- Supposing that is a set consisting of all miRNAs that have experimentally verified known associations with in LM, and is a set consisting of all miRNAs that have experimentally verified known associations with in MD.
- Step 3:
- Let , then we can calculate the weight of in according to the following Formula (1):
3.3. Similarity Calculation
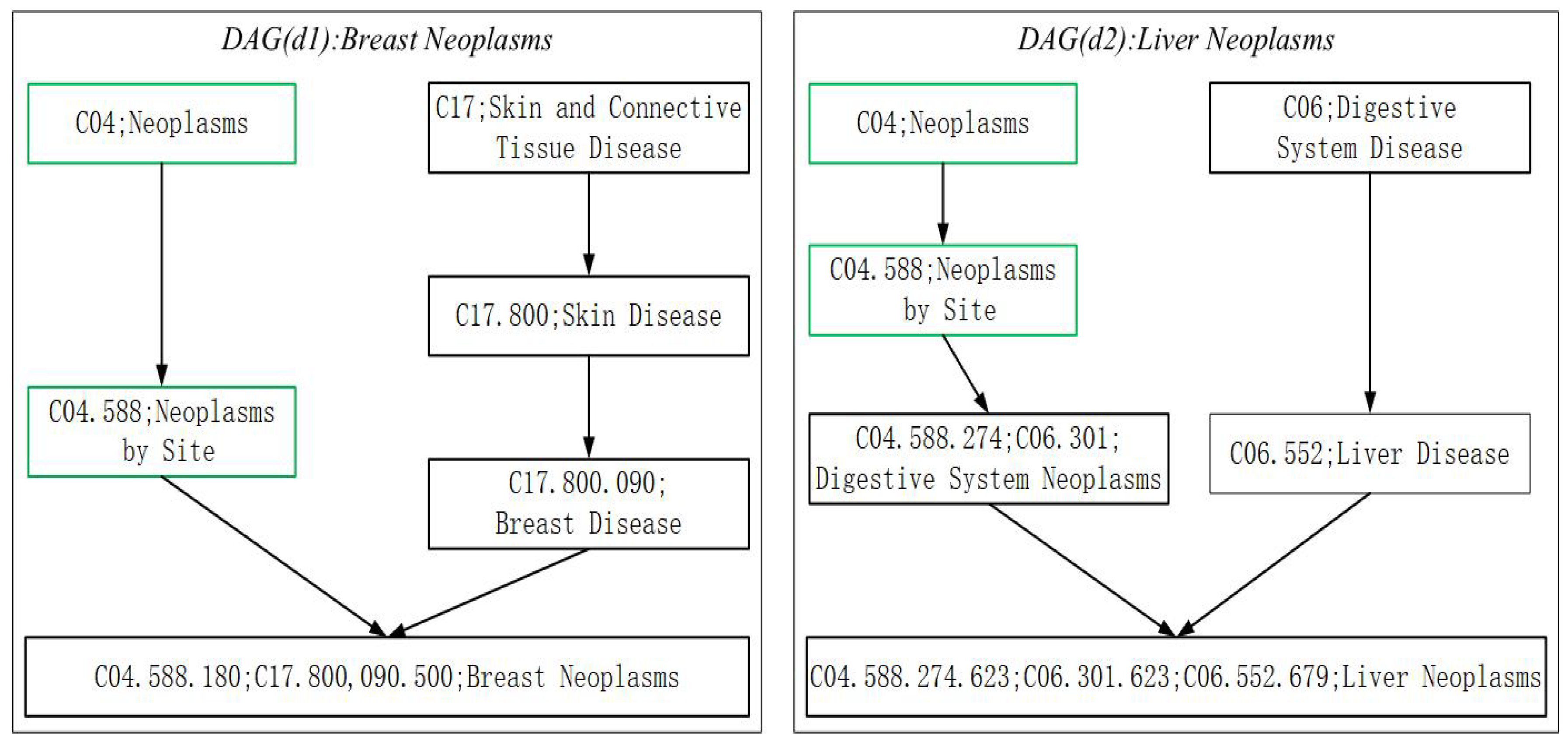
3.3.1. Disease Semantic Similarity Measure
3.3.2. LncRNA Similarity Measure
3.3.3. Weight Redistribution in Based on the KNN Algorithm
- Step 1:
- Firstly, representing the ith row of the weight matrix as , and the jth column of the weight matrix as .
- Step 2:
- Then, for any given lncRNA and any in L other than , based on above Formula (4), it is obvious that we can obtain the functional similarity easily, and moreover, after sorting these values of functional similarities between and all remaining lncRNAs other than in descending order, then we can obtain the corresponding lncRNAs from the first K elements in the sorted results. For convenience, let denote these K lncRNAs, then the qth row of can be updated according to the following Formula (6):where is a decay factor, which means that a higher decay will be assigned to if it is more dissimilar to , and is the normalization factor, which is used for normalization of the value of . Additionally, in similar way, it is obvious that the pth column of can also be updated according to the following Formula (7):where to denote the top K diseases most similar to is the decay factor, and is the normalization factor.
3.4. Construction of Our Prediction Model PMFILDA Based on
3.4.1. Standard Matrix Factorization
3.4.2. Probabilistic Matrix Factorization
3.4.3. Optimization
4. Results and Discussion
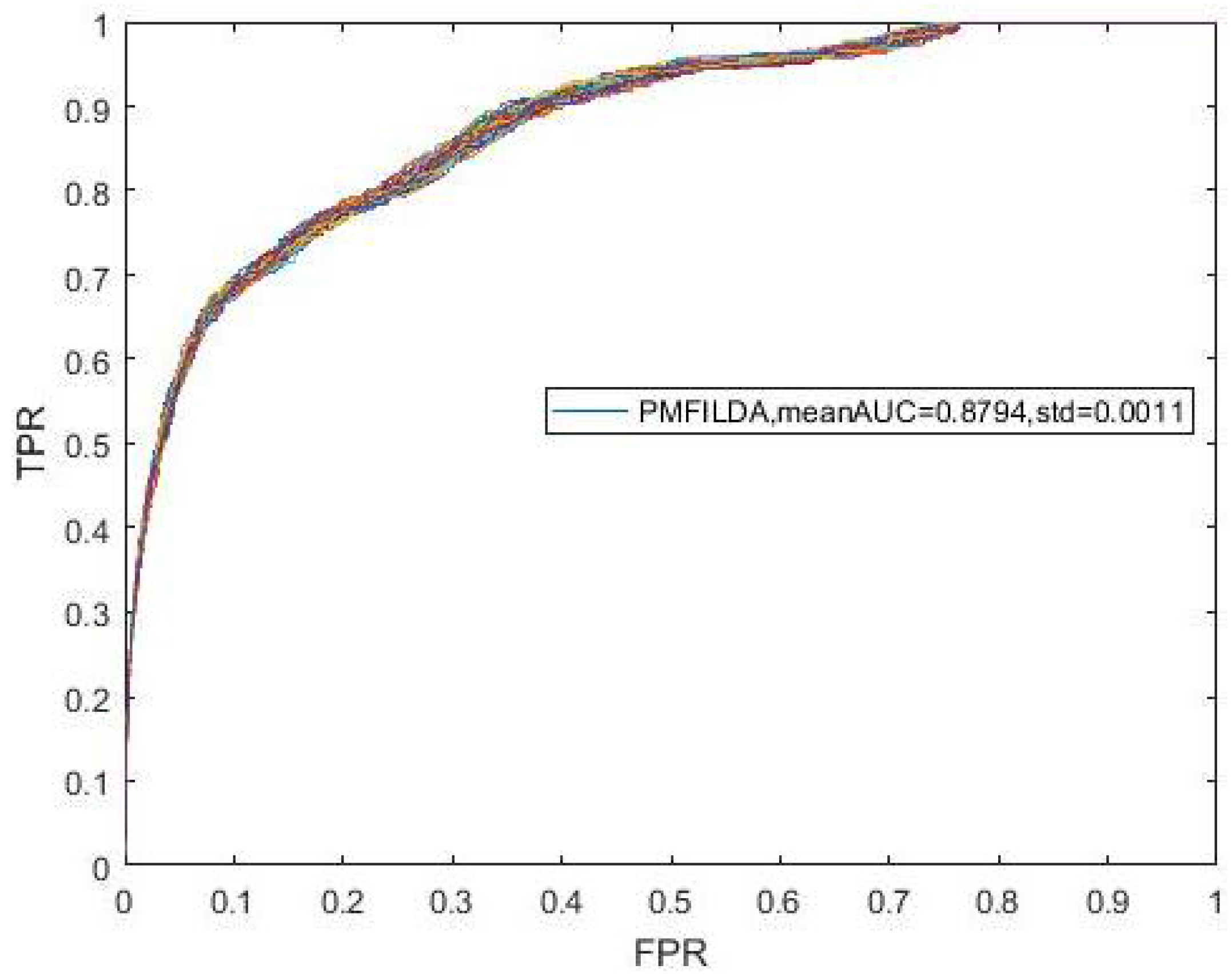
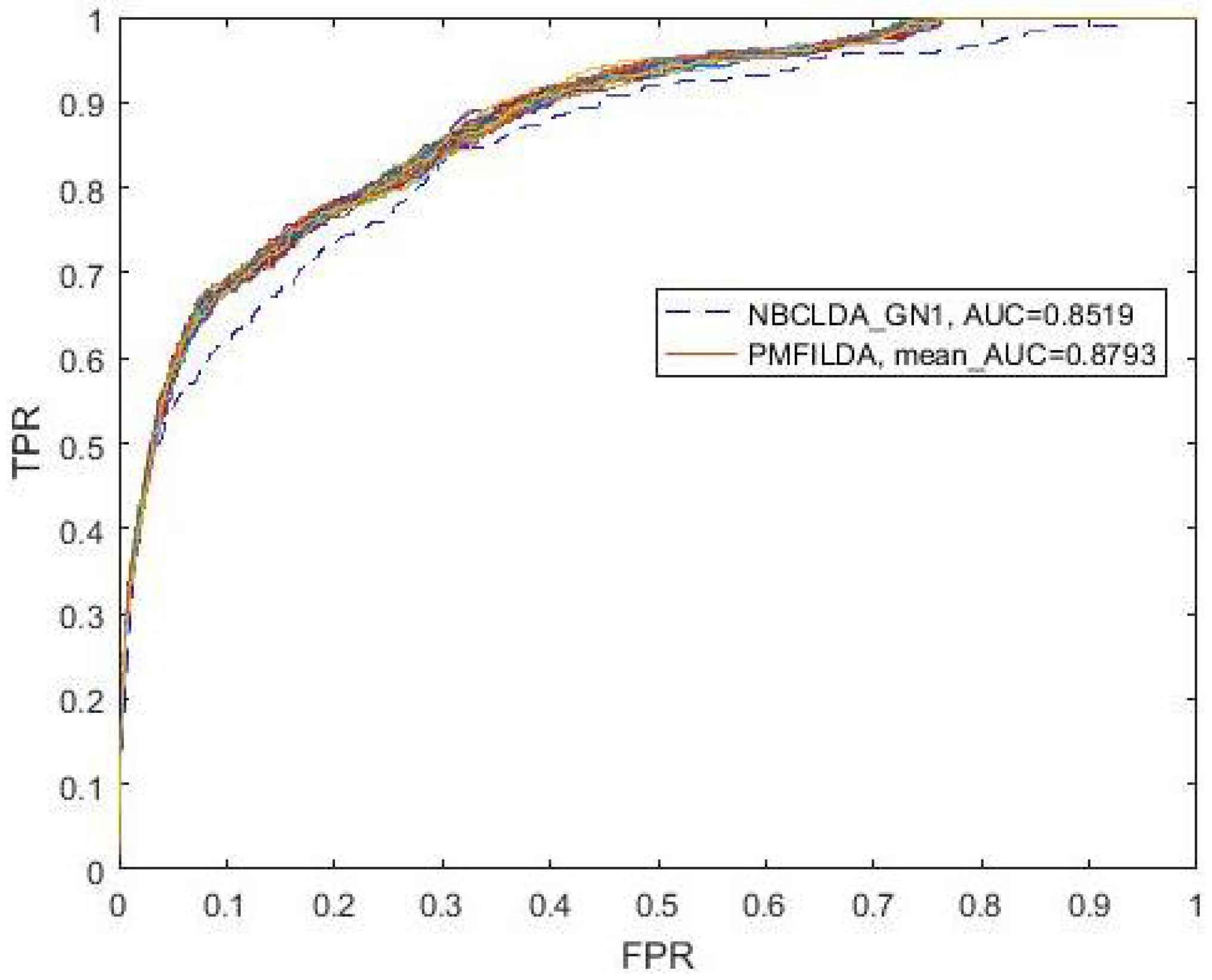
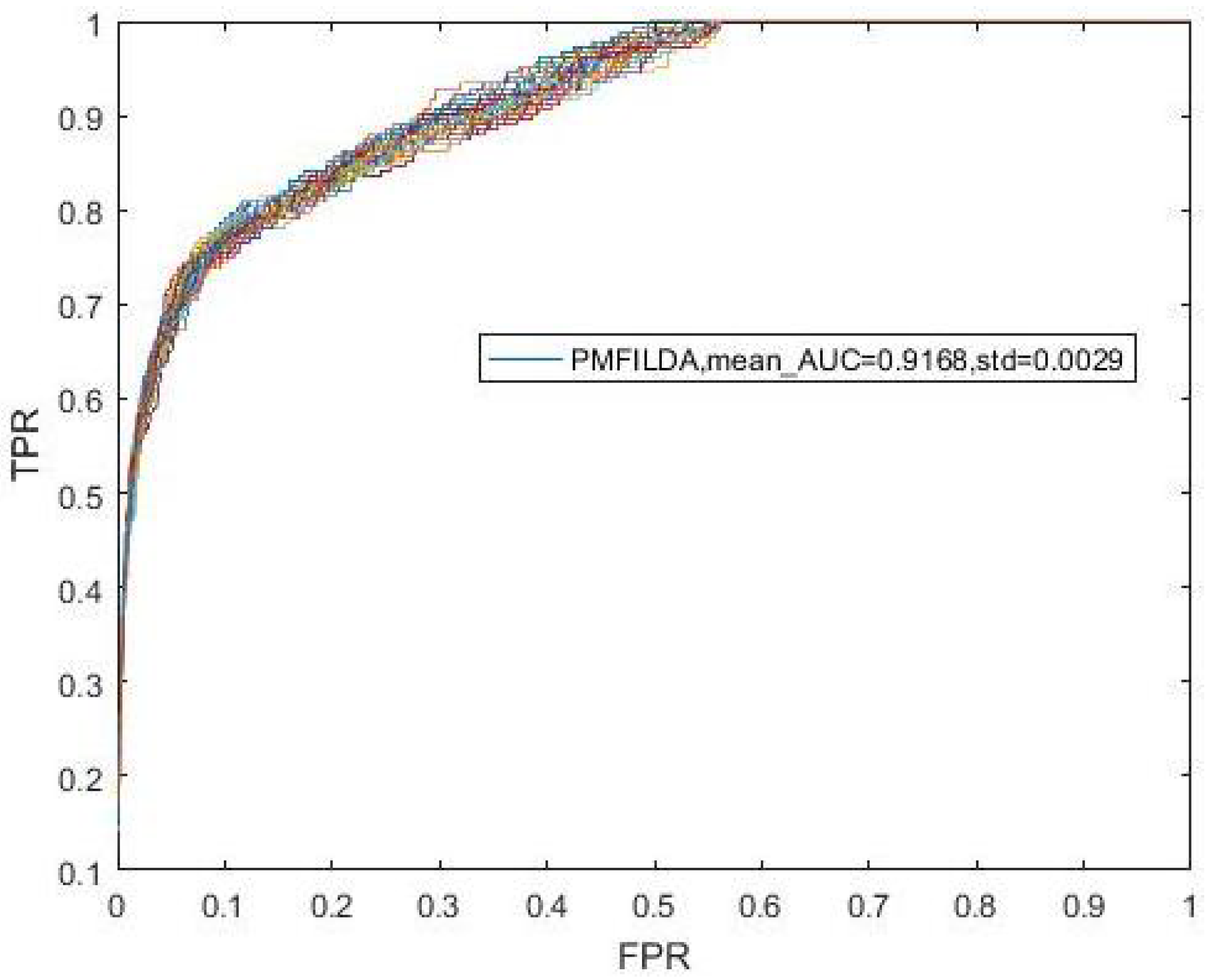
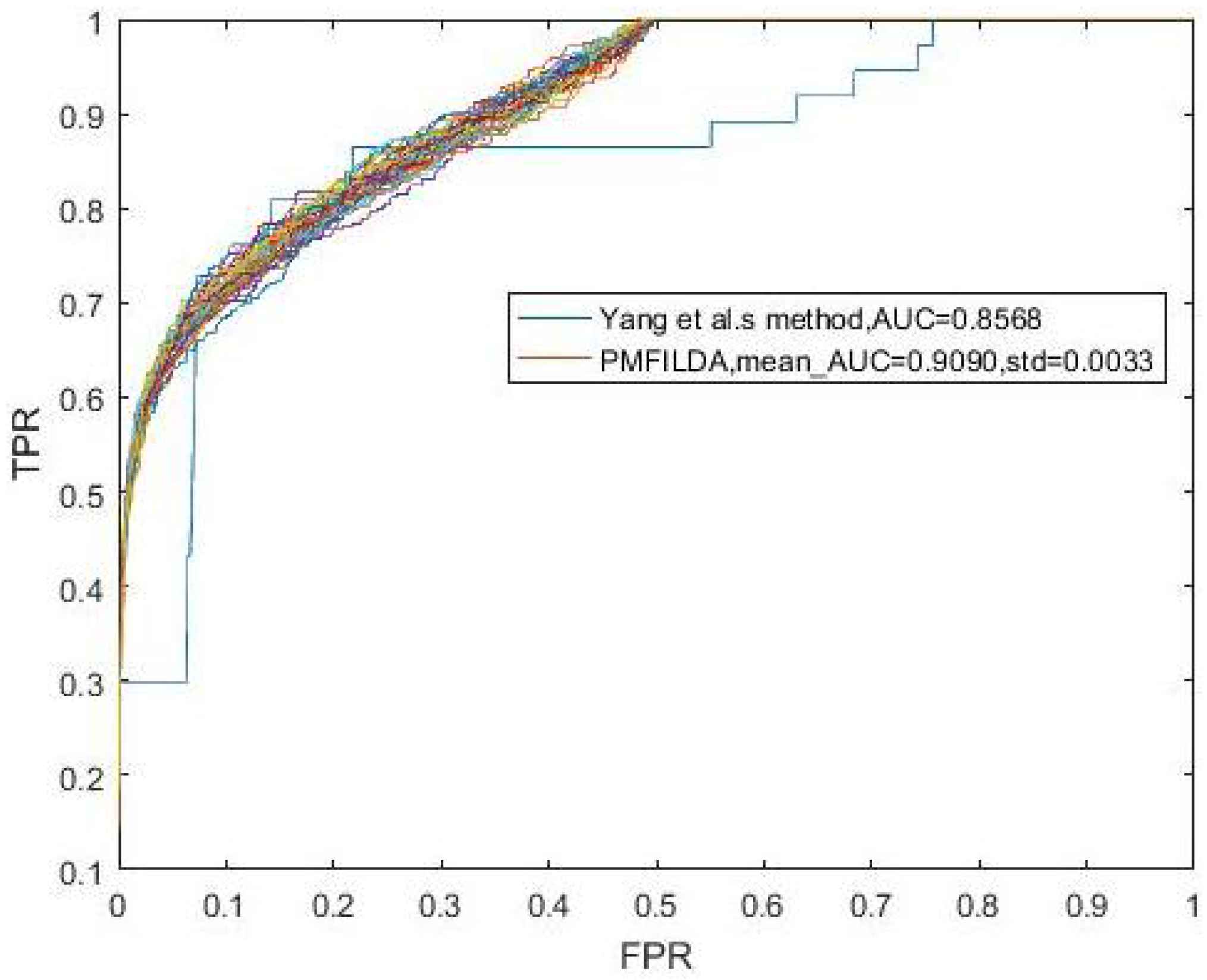
4.1. Performance Evaluation Metrics
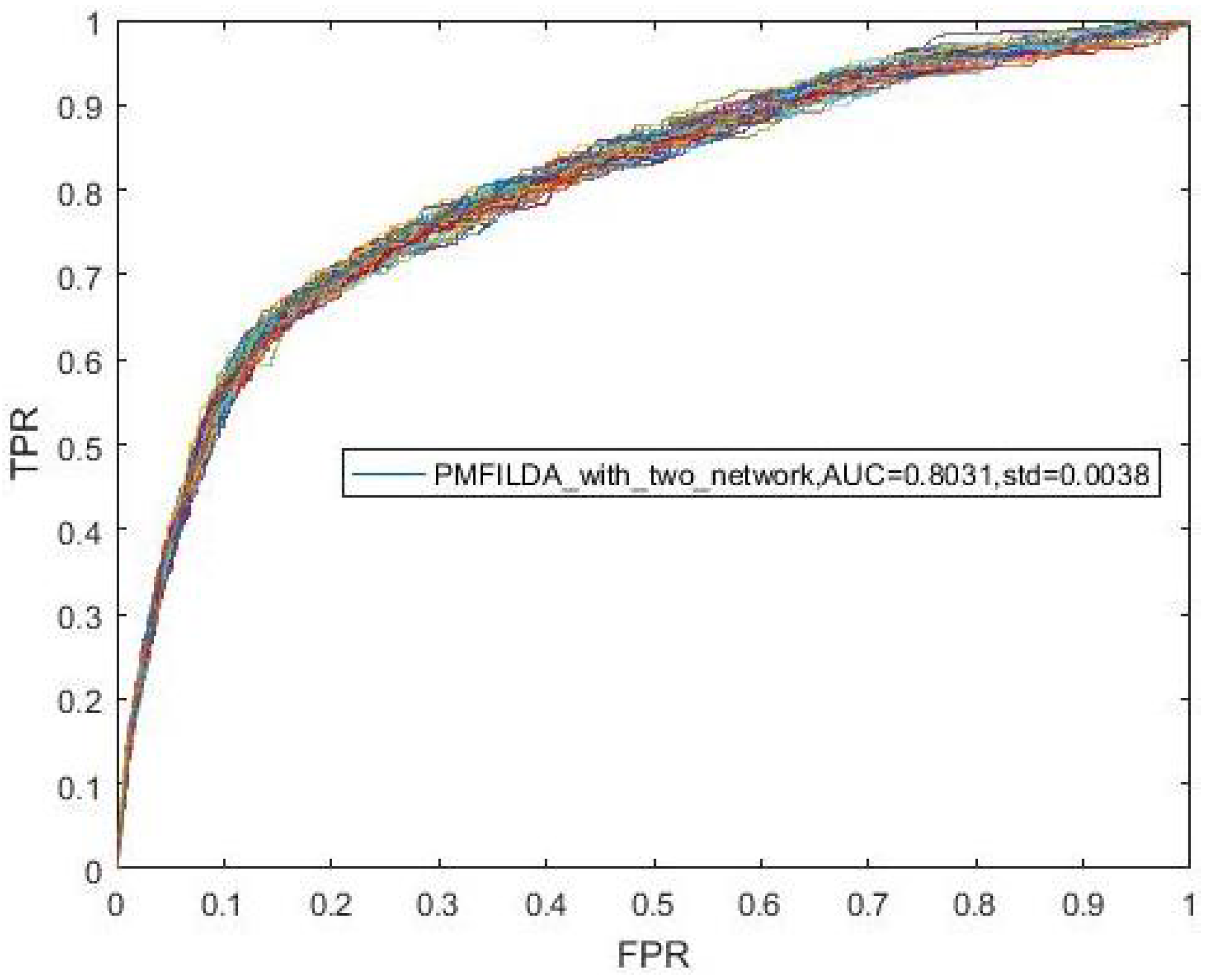
4.2. Contribution Analysis of lncRNA-Disease Associated Network
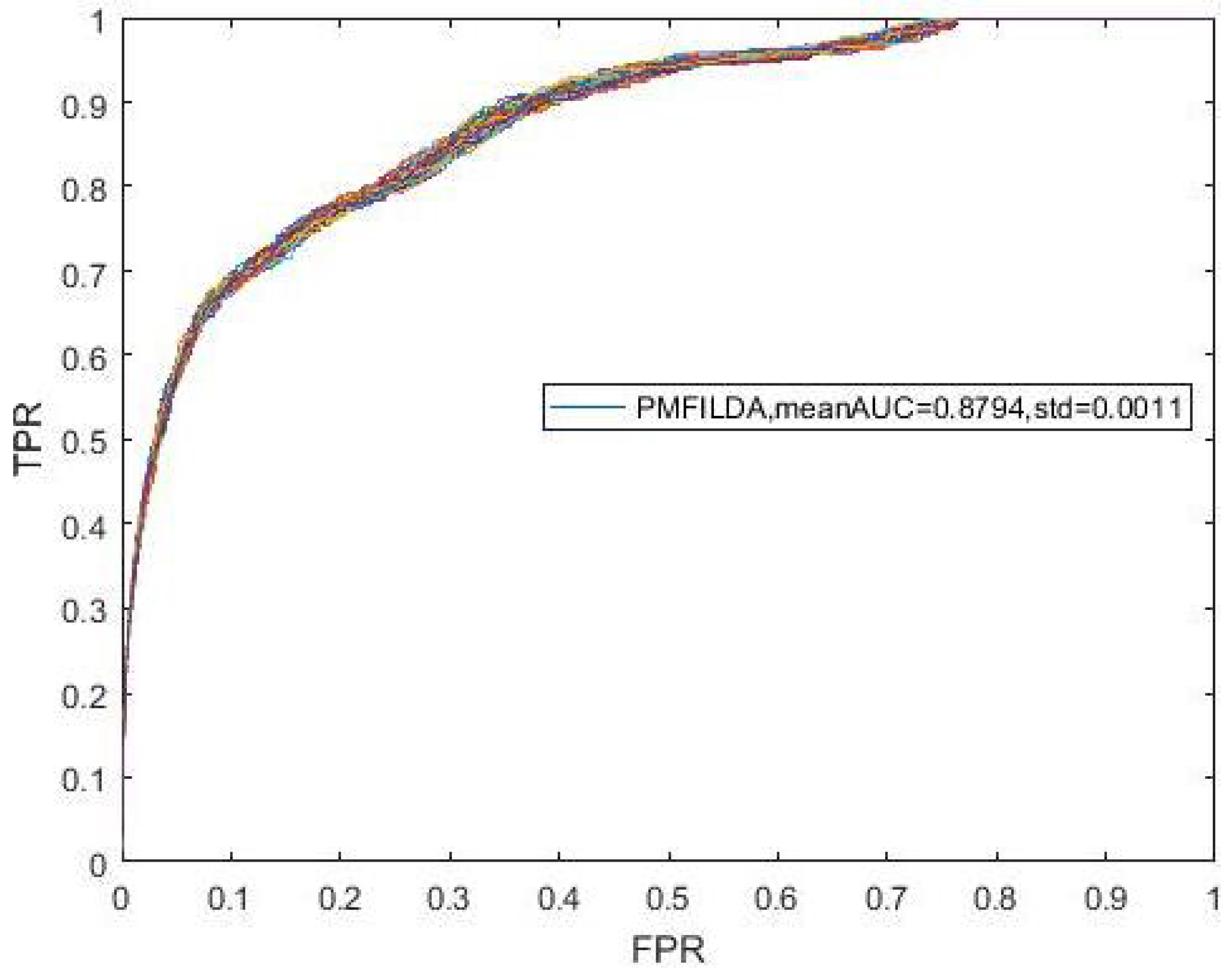
4.3. The Effects of KNN on Performance
4.4. Parameter Sensitivity Analysis
4.5. Case Studies
5. Discussion
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Guttman, M.; Russell, P.; Ingolia, N.; Weissman, J.; Lander, E. Ribosome Profiling Provides Evidence that Large Noncoding RNAs Do Not Encode Proteins. Cell 2013, 154, 240–251. [Google Scholar] [CrossRef] [PubMed]
- Vakul, M.; Yesim, G.P.; Sunil, B.; Sarath Chandra, J. Role of lncRNAs in health and disease-size and shape matter. Brief. Funct. Genom. 2015, 14, 115–129. [Google Scholar]
- Zhao, W.; Luo, J.; Jiao, S. Comprehensive characterization of cancer subtype associated long non-coding RNAs and their clinical implications. Sci. Rep. 2014, 4, 6591. [Google Scholar] [CrossRef] [PubMed]
- Lu, Q.; Ren, S.; Lu, M.; Zhang, Y.; Zhu, D.; Zhang, X.; Li, T. Computational prediction of associations between long non-coding RNAs and proteins. BMC Genom. 2013, 14, 651. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Xuan, Z.; Liu, C. Long non-coding RNAs and complex human diseases. Int. J. Mol. Sci. 2013, 14, 18790–18808. [Google Scholar] [CrossRef] [PubMed]
- Bussemakers, M.J.; Bokhoven, A.V.; Verhaegh, G.W.; Smit, F.P.; Karthaus, H.F.; Schalken, J.A.; Debruyne, F.M.; Ru, N.; Isaacs, W.B. DD3: A new prostate-specific gene, highly overexpressed in prostate cancer. Cancer Res. 1999, 59, 5975–5979. [Google Scholar] [PubMed]
- Managadze, D.; Rogozin, I.B.; Chernikova, D.; Shabalina, S.A.; Koonin, E.V. Negative Correlation between Expression Level and Evolutionary Rate of Long Intergenic Noncoding RNAs. Genome Biol. Evol. 2011, 3, 1390–1404. [Google Scholar] [CrossRef] [PubMed]
- Nicole, S.; Harvey, R.P.; Mattick, J.S. Long noncoding RNAs in cardiac development and pathophysiology. Circ. Res. 2012, 111, 1349–1362. [Google Scholar]
- Deeksha, B.; Shruti, K.; Saakshi, J.; Satish, S.; Kriti, K.; Chetana, S.; Sridhar, S.; Vinod, S. Conceptual approaches for lncRNA drug discovery and future strategies. Expert Opin. Drug Discov. 2012, 7, 503–513. [Google Scholar]
- Liao, Q.; Liu, C.; Yuan, X.; Kang, S.; Miao, R.; Xiao, H.; Zhao, G.; Luo, H.; Bu, D.; Zhao, H.; et al. Large-scale prediction of long non-coding RNA functions in a coding–non-coding gene co-expression network. Nucleic Acids Res. 2011, 39, 3864–3878. [Google Scholar] [CrossRef]
- Chen, X.; Yan, C.C.; Luo, C.; Ji, W.; Zhang, Y.; Dai, Q. Constructing lncRNA functional similarity network based on lncRNA-disease associations and disease semantic similarity. Sci. Rep. 2015, 5, 11338. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Sun, Y.Z.; Guan, N.; Qu, J.; Huang, Z.A.; Zhu, Z.X.; Li, J.Q. Computational models for lncRNA function prediction and functional similarity calculation. Brief. Funct. Genom. 2018, ely031. [Google Scholar] [CrossRef] [PubMed]
- Ping, P.; Wang, L.; Kuang, L.; Ye, S.; Mfb, I.; Pei, T. A Novel Method for LncRNA-Disease Association Prediction Based on an lncRNA-disease Association Network. IEEE/ACM Trans. Comput. Biol. Bioinform. 2018, 1. [Google Scholar] [CrossRef] [PubMed]
- Yao, Y.; Ma, J.; Xue, Y.; Wang, P.; Li, Z.; Liu, J.; Chen, L.; Xi, Z.; Teng, H.; Wang, Z. Knockdown of long non-coding RNA XIST exerts tumor-suppressive functions in human glioblastoma stem cells by up-regulating miR-152. Cancer Lett. 2015, 359, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Ri-Kao, Y.U.; A-Lin, J.I.; Yao, X.L.; Fang, J.J.; Jin, X.D.; Urology, D.O. Effects of long non-coding RNA-HOTAIR on the cell cycle and invasiveness of prostate cancer. Zhonghua Nan Ke Xue 2015, 21, 792–796. [Google Scholar] [PubMed]
- Chen, X.; Yan, C.C.; Zhang, X.; You, Z.H. Long non-coding RNAs and complex diseases: From experimental results to computational models. Brief. Bioinform. 2017, 18, 558–576. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Yan, G.Y. Novel human lncRNA-disease association inference based on lncRNA expression profiles. Bioinformatics 2013, 29, 2617–2624. [Google Scholar] [CrossRef]
- Huang, Y.A.; Chen, X.; You, Z.H.; Huang, D.S.; Chan, K.C.C. ILNCSIM: Improved lncRNA functional similarity calculation model. Oncotarget 2016, 7, 25902–25914. [Google Scholar] [CrossRef]
- Zhao, T.; Xu, J.; Liu, L.; Bai, J.; Xu, C.; Xiao, Y.; Li, X.; Zhang, L. Identification of cancer-related lncRNAs through integrating genome, regulome and transcriptome features. Mol. Biosyst. 2015, 11, 126–136. [Google Scholar] [CrossRef]
- Sun, J.; Shi, H.; Wang, Z.; Zhang, C.; Liu, L.; Wang, L.; He, W.; Hao, D.; Liu, S.; Zhou, M. Inferring novel lncRNA-disease associations based on a random walk model of a lncRNA functional similarity network. Mol. Biosyst. 2014, 10, 2074–2081. [Google Scholar] [CrossRef]
- Chen, X. KATZLDA: KATZ measure for the lncRNA-disease association prediction. Sci. Rep. 2014, 5, 16840. [Google Scholar] [CrossRef]
- Liu, M.X.; Chen, X.; Chen, G.; Cui, Q.H.; Yan, G.Y. A computational framework to infer human disease-associated long noncoding RNAs. PLoS ONE 2014, 9, e84408. [Google Scholar] [CrossRef] [PubMed]
- Chen, X. Predicting lncRNA-disease associations and constructing lncRNA functional similarity network based on the information of miRNA. Sci. Rep. 2015, 5, 13186. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wang, L.; Qu, J.; Guan, N.; Li, J.Q. Predicting miRNA-disease association based on inductive matrix completion. Bioinformatics 2018, 34, 4256–4265. [Google Scholar] [CrossRef]
- Chen, X.; Yin, J.; Qu, J.; Huang, L.; Wang, E. MDHGI: Matrix Decomposition and Heterogeneous Graph Inference for miRNA-disease association prediction. PLoS Comput. Biol. 2018, 14, e1006418. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Chen, X.; Yin, J. A novel computational method for the identification of potential miRNA-disease association based on symmetric non-negative matrix factorization and Kronecker regularized least square. Front. Genet. 2018, 9, 324. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Wu, Q.F.; Yan, G.Y. RKNNMDA: Ranking-based KNN for MiRNA-Disease Association prediction. RNA Biol. 2017, 14, 952–962. [Google Scholar] [CrossRef] [PubMed]
- Li, J.H.; Liu, S.; Zhou, H.; Qu, L.H.; Yang, J.H. starBase v2.0: Decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014, 42, D92–D97. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Qiu, C.; Tu, J.; Geng, B.; Yang, J.; Jiang, T.; Cui, Q. HMDD v2.0: A database for experimentally supported human microRNA and disease associations. Nucleic Acids Res. 2014, 42, D1070–D1074. [Google Scholar] [CrossRef]
- Cui, T.; Zhang, L.; Huang, Y.; Yi, Y.; Tan, P.; Zhao, Y.; Hu, Y.; Xu, L.; Li, E.; Wang, D. MNDR v2.0: An updated resource of ncRNA-disease associations in mammals. Nucleic Acids Res. 2018, 46, D371–D374. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Wang, J.; Lu, M.; Song, F.; Cui, Q. Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 2010, 26, 1644–1650. [Google Scholar] [CrossRef]
- Lee, D.; Seung, H. Learning the parts of objects by non-negative matrix factorization. Nature 1999, 401, 788–791. [Google Scholar] [CrossRef]
- Mnih, A.; Salakhutdinov, R.R. Probabilistic Matrix Factorization. Adv. Neural Inf. Process. Syst. 2007, 20, 1257–1264. [Google Scholar]
- Yu, J.; Ping, P.; Wang, L.; Kuang, L.; Li, X.; Wu, Z. A Novel Probability Model for LncRNA–Disease Association Prediction Based on the Naïve Bayesian Classifier. Genes 2018, 9. [Google Scholar] [CrossRef]
- Yang, X.; Gao, L.; Guo, X.; Shi, X.; Wu, H.; Song, F.; Wang, B. A network based method for analysis of lncRNA-disease associations and prediction of lncRNAs implicated in diseases. PLoS ONE 2014, 9, e87797. [Google Scholar] [CrossRef]
- Donahue, H.J.; Genetos, D.C. Genomic approaches in breast cancer research. Brief. Funct. Genom. 2013, 12, 391–396. [Google Scholar] [CrossRef]
- Karagoz, K.; Sinha, R.; Arga, K.Y. Triple Negative Breast Cancer: A Multi-Omics Network Discovery Strategy for Candidate Targets and Driving Pathways. Omics J. Integr. Biol. 2015, 19, 115–130. [Google Scholar] [CrossRef]
- Jin, M.; Li, P.; Zhang, Q.; Yang, Z.; Shen, F. A four-long non-coding RNA signature in predicting breast cancer survival. J. Exp. Clin. Cancer Res. 2014, 33, 84. [Google Scholar]
- Xu, N.; Wang, F.; Lv, M.; Cheng, L. Microarray expression profile analysis of long non-coding RNAs in human breast cancer: A study of Chinese women. Biomed. Pharmacother. 2015, 69, 221–227. [Google Scholar] [CrossRef]
- Jadaliha, M.; Zong, X.; Malakar, P.; Ray, T.; Singh, D.K.; Freier, S.M.; Jensen, T.; Prasanth, S.G.; Karni, R.; Ray, P.S. Functional and prognostic significance of long non-coding RNA MALAT1 as a metastasis driver in ER negative lymph node negative breast cancer. Oncotarget 2016, 7, 40418–40436. [Google Scholar] [CrossRef]
- Gupta, R.A.; Nilay, S.; Wang, K.C.; Jeewon, K.; Horlings, H.M.; Wong, D.J.; Miao-Chih, T.; Tiffany, H.; Pedram, A.; Rinn, J.L. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, 1071–1076. [Google Scholar] [CrossRef]
- Dalia, B.L.; Lau, S.K.; Boutros, P.C.; Fereshteh, K.; Igor, J.; Andrulis, I.L.; Ming, S.T.; Penn, L.Z. The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res. 2006, 66, 5330–5337. [Google Scholar]
- White, N.M.; Cabanski, C.R.; Silva-Fisher, J.M.; Dang, H.X.; Govindan, R.; Maher, C.A. Transcriptome sequencing reveals altered long intergenic non-coding RNAs in lung cancer. Genome Biol. 2014, 15, 429. [Google Scholar] [CrossRef]
- Prensner, J.R.; Chinnaiyan, A.M. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011, 1, 391–407. [Google Scholar] [CrossRef]
- Tony, G.; Sven, D. The hallmarks of cancer: A long non-coding RNA point of view. RNA Biol. 2012, 9, 703–719. [Google Scholar]
- Tony, G.; Monika, H.; Moritz, E.; Jeff, H.; Youngsoo, K.; Alexey, R.; Gayatri, A.; Marion, S.; Matthias, G. The noncoding RNA MALAT1 is a critical regulator of the metastasis phenotype of lung cancer cells. Cancer Res. 2013, 73, 1180–1189. [Google Scholar]
- Ji, P.; Diederichs, S.; Wang, W.; Boing, S.; Metzger, R.; Schneider, P.M.; Tidow, N.; Brandt, B.; Buerger, H.; Bulk, E.; et al. MALAT-1, a novel noncoding RNA, and thymosin beta4 predict metastasis and survival in early-stage non-small cell lung cancer. Oncogene 2003, 22, 8031. [Google Scholar] [CrossRef]
- Tano, K.; Mizuno, R.; Okada, T.; Rakwal, R.; Shibato, J.; Masuo, Y.; Ijiri, K.; Akimitsu, N. MALAT-1 enhances cell motility of lung adenocarcinoma cells by influencing the expression of motility-related genes. FEBS Lett. 2010, 584, 4575–4580. [Google Scholar] [CrossRef]
- Hrdlickova, B.; Almeida, R.C.D.; Borek, Z.; Withoff, S. Genetic variation in the non-coding genome: Involvement of micro-RNAs and long non-coding RNAs in disease. BBA Mol. Basis Dis. 2014, 1842, 1910–1922. [Google Scholar] [CrossRef]
- Sang, Y.; Zhou, F.; Wang, D.; Bi, X.; Liu, X.; Hao, Z.; Li, Q.; Zhang, W. Up-regulation of long non-coding HOTTIP functions as an oncogene by regulating HOXA13 in non-small cell lung cancer. Am. J. Transl. Res. 2015, 8, 2022. [Google Scholar]
- Xia, Y.; He, Z.; Liu, B.; Wang, P.; Chen, Y. Downregulation of Meg3 enhances cisplatin resistance of lung cancer cells through activation of the WNT/β-catenin signaling pathway. Mol. Med. Rep. 2015, 12, 4530–4537. [Google Scholar] [CrossRef]
- Berger, F.G. Interview: Screening and treatment for colorectal cancer. Colorectal Cancer 2013, 2, 117–120. [Google Scholar] [CrossRef]
- Ji, Q.; Zhang, L.; Liu, X.; Zhou, L.; Wang, W.; Han, Z.; Sui, H.; Tang, Y.; Wang, Y.; Liu, N. Long non-coding RNA MALAT1 promotes tumour growth and metastasis in colorectal cancer through binding to SFPQ and releasing oncogene PTBP2 from SFPQ/PTBP2 complex. Br. J. Cancer 2014, 111, 736. [Google Scholar] [CrossRef]
- Li, Y.; Li, Y.; Chen, W.; He, F.; Tan, Z.; Zheng, J.; Wang, W.; Zhao, Q.; Li, J. NEAT expression is associated with tumor recurrence and unfavorable prognosis in colorectal cancer. Oncotarget 2015, 6, 27641–27650. [Google Scholar] [CrossRef]
- Wu, Y.; Yang, L.; Zhao, J.; Li, C.; Nie, J.; Liu, F.; Zhuo, C.; Zheng, Y.; Li, B.; Wang, Z. Nuclear-enriched abundant transcript 1 as a diagnostic and prognostic biomarker in colorectal cancer. Mol. Cancer 2015, 14, 191. [Google Scholar] [CrossRef]
- Sun, J.; Ding, C.; Yang, Z.; Liu, T.; Zhang, X.; Zhao, C.; Wang, J. The long non-coding RNA TUG1 indicates a poor prognosis for colorectal cancer and promotes metastasis by affecting epithelial-mesenchymal transition. J. Transl. Med. 2016, 14, 42. [Google Scholar] [CrossRef]
- Chen, X.; Xie, D.; Wang, L.; Zhao, Q.; Liu, H. BNPMDA: Bipartite Network Projection for MiRNA-Disease Association prediction. Bioinformatics 2018, 34, 3178–3186. [Google Scholar] [CrossRef]
- Chen, X.; Huang, L. LRSSLMDA: Laplacian Regularized Sparse Subspace Learning for MiRNA-Disease Association prediction. PLoS Comput. Biol. 2017, 13, e1005912. [Google Scholar] [CrossRef]
- Chen, X.; Huang, L.; Xie, D.; Zhao, Q. EGBMMDA: Extreme Gradient Boosting Machine for MiRNA-Disease Association prediction. Cell Death Dis. 2018, 9, 3. [Google Scholar] [CrossRef]
- Zhao, H.; Kuang, L.; Feng, X.; Wang, L. Inferring microRNA-disease associations based on Weighted Interactive Network. Int. J. Mol. Sci. 2019, 20, 110. [Google Scholar] [CrossRef]
- Zhao, H.; Kuang, L.; Wang, L.; Ping, P.; Xuan, Z.; Pei, T.; Wu, Z. Prediction of microRNA-disease associations based on distance correlation set. BMC Bioinform. 2018, 19, 141. [Google Scholar] [CrossRef]
- Zou, Q.; Li, J.; Song, L.; Zeng, X.; Wang, G. Similarity computation strategies in the microRNA-disease network: A survey. Brief. Funct. Genom. 2015, 15, 55–64. [Google Scholar] [CrossRef]
- Zeng, X.; Zhang, X.; Zou, Q. Integrative approaches for predicting microRNA function and prioritizing disease-related microRNA using biological interaction networks. Brief. Bioinform. 2016, 17, 193–203. [Google Scholar] [CrossRef]

| Methods | AUCs | Methods | AUCs | Methods | AUCs |
|---|---|---|---|---|---|
| PMFILDA | 0.8793 | PMFILDA | 0.9169 | PMFILDA | 0.9090 |
| 0.8519 | HGLDA | 0.8519 | Method of Yang et al. | 0.8568 |
| KNN | K-Means | |
|---|---|---|
| Mean_AUC | 0.8794 | 0.8589 |
| STD | 0.0278 | 0.0011 |
| Diseases | lncRNAs | Evidence (PUBMED) |
|---|---|---|
| Breast Cancer | MALAT1 | 22492512, 22996375, 24499465, 27250026, 27777857, 27191888 |
| Breast Cancer | HOTAIR | 24499465, 20930520, 21925379, 20393566, 19182780, 21903344 |
| Breast Cancer | H19 | 22996375, 21489289, 14729626, 16707459, 21748294, 18794369 |
| Breast Cancer | MEG3 | 27166155, 14602737, 22393162, 22487937 |
| Breast Cancers | GAS5 | 27034004, 18836484, 20673990, 22487937, 22664915, 26662314 |
| Breast Cancer | PTPRG-AS1 | 26409453 |
| Breast Cancer | NEAT1 | 25417700, 27147820, 21532345, 27556296 |
| Breast Cancer | PVT1 | 24780616, 17908964, 25122612, 26889781 |
| Breast Cancer | CDKN2B-AS1 | 17440112, 20956613, 20453838, 20956613 |
| Breast Cancer | TUG1 | 27791993 |
| Breast Cancer | XIST | 17545591, 27248326, 18006640, 19440381, 24141629, 26637364 |
| Breast Cancer | ZFAS1 | 21460236 |
| Lung Cancer | H19 | 27186394, 26729200 |
| Lung Cancer | HOTAIR | 27186394, 26729200, 24757675, 23668363, 27270317 |
| Lung Cancer | MALAT1 | 25217850, 20937273, 20937273, 27777857 |
| Lung Cancer | HOTTIP | 27347311, 26265284 |
| Lung Cancer | MEG3 | 14602737, 26059239 |
| Lung Cancer | CDKN2B-AS1 | 27307748, 26729200, 26453113, 25964559, 25889788 |
| Lung Cancer | GAS5 | 27631209, 26634743, 24357161 |
| Lung Cancer | CCAT1 | 25129441 |
| Lung Cancer | XIST | 27501756, 26339353 |
| Lung Cancer | CASC2 | 26790438 |
| Lung Cancer | PVT1 | 26908628, 26729200, 25400777 |
| Lung Cancer | ZNRD1-AS1 | 27166266 |
| Lung Cancer | NEAT1 | 27351135, 27270317, 25889788 |
| Lung Cancer | TUG1 | 24853421, 27485439 |
| Colorectal Cancer | H19 | 8564957, 22427002, 11120891, 26989025, 19926638, 26068968 |
| Colorectal Cancer | HOTTIP | 26617875, 26678886, 27546609 |
| Colorectal Cancer | XIST | 17143621 |
| Colorectal Cancer | NEAT1 | 26314847, 26552600 |
| Colorectal Cancer | MEG3 | 25636452, 26934323 |
| Colorectal Cancer | TUG1 | 26856330, 27421138 |
| Colorectal Cancer | PVT1 | 26990997, 24196785 |
| Colorectal Cancer | CCAT1 | 23416875, 26064266, 26823726, 24594601, 23594791, 26752646 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xuan, Z.; Li, J.; Yu, J.; Feng, X.; Zhao, B.; Wang, L. A Probabilistic Matrix Factorization Method for Identifying lncRNA-Disease Associations. Genes 2019, 10, 126. https://doi.org/10.3390/genes10020126
Xuan Z, Li J, Yu J, Feng X, Zhao B, Wang L. A Probabilistic Matrix Factorization Method for Identifying lncRNA-Disease Associations. Genes. 2019; 10(2):126. https://doi.org/10.3390/genes10020126
Chicago/Turabian StyleXuan, Zhanwei, Jiechen Li, Jingwen Yu, Xiang Feng, Bihai Zhao, and Lei Wang. 2019. "A Probabilistic Matrix Factorization Method for Identifying lncRNA-Disease Associations" Genes 10, no. 2: 126. https://doi.org/10.3390/genes10020126
APA StyleXuan, Z., Li, J., Yu, J., Feng, X., Zhao, B., & Wang, L. (2019). A Probabilistic Matrix Factorization Method for Identifying lncRNA-Disease Associations. Genes, 10(2), 126. https://doi.org/10.3390/genes10020126






