A Novel Arsenate-Resistant Determinant Associated with ICEpMERPH, a Member of the SXT/R391 Group of Mobile Genetic Elements
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Strains
2.2. Genome Sequencing and Annotation
2.3. Phylogenetic Analysis of Core ICE Genes
2.4. Accession Number
2.5. Phenotypic Testing
3. Results and Discussion
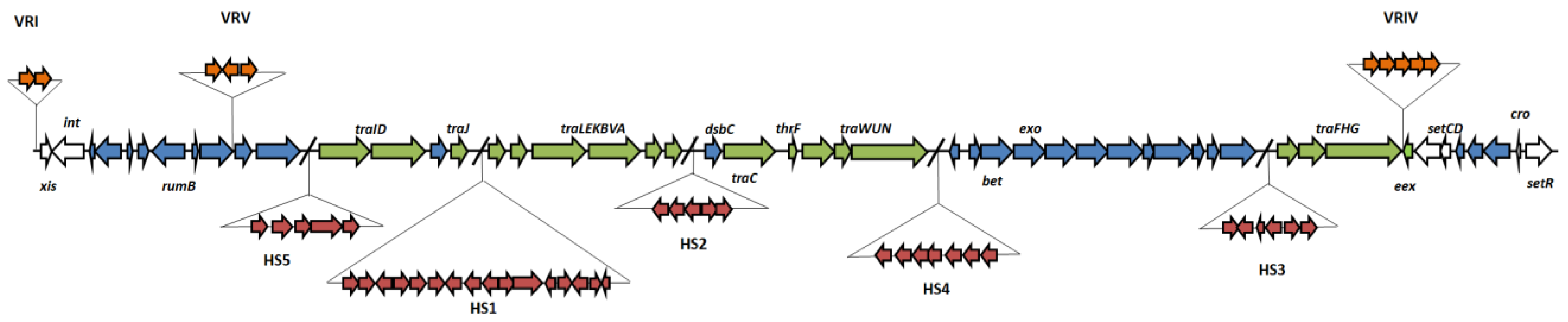
3.1. Full Sequence Analysis
3.2. Genetic Basis for Arsenic Resistance Encoded by pMERPH
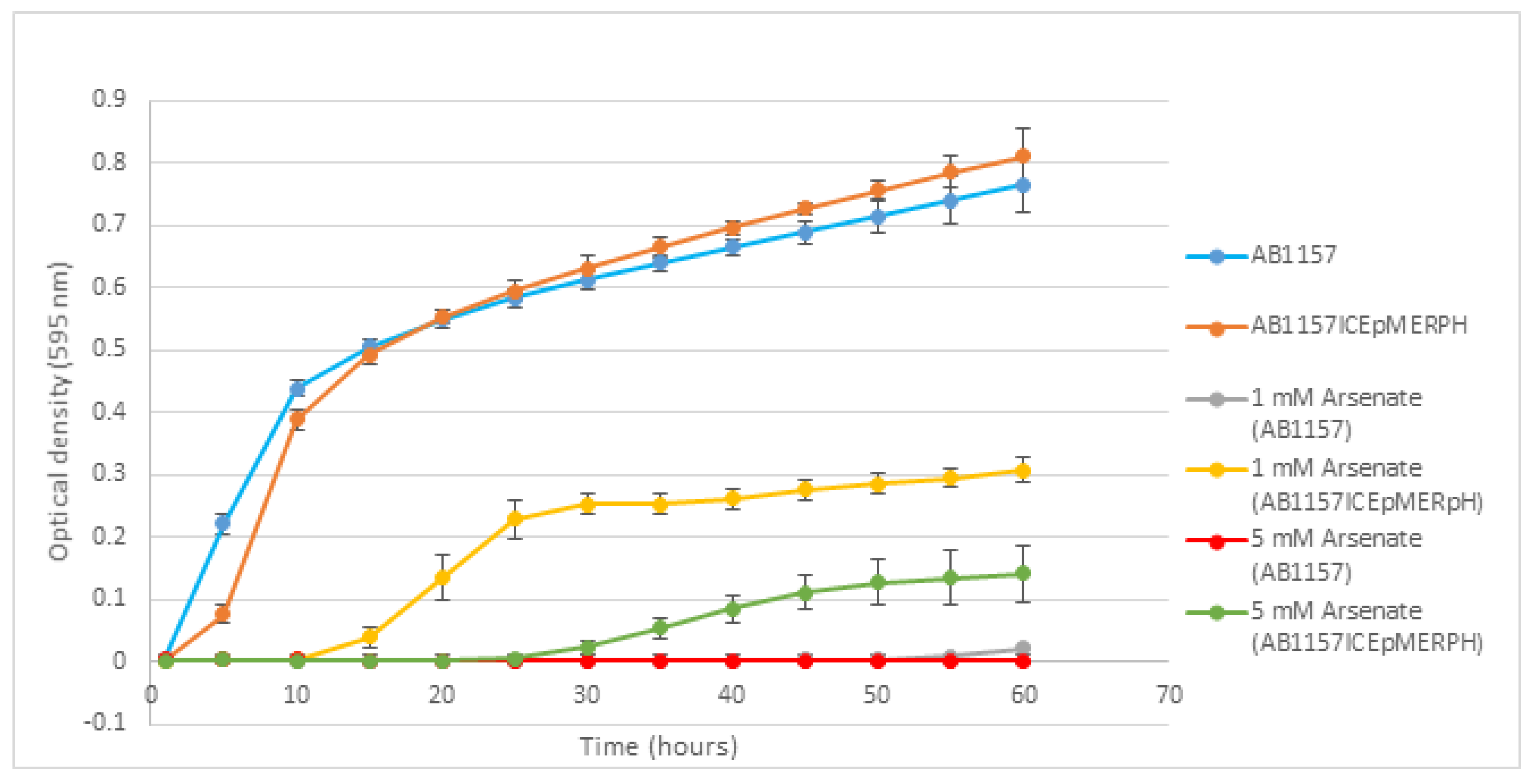
3.3. Phenotypic Testing
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wozniak, R.A.F.; Waldor, M.K. Integrative and conjugative elements: Mosaic mobile genetic elements enabling dynamic lateral gene flow. Nat. Rev. Microbiol. 2010, 8, 552–563. [Google Scholar] [CrossRef] [PubMed]
- Bi, D.; Xu, Z.; Harrison, E.M.; Tai, C.; Wei, Y.; He, X.; Jia, S.; Deng, Z.; Rajakumar, K.; Ou, H.Y. ICEberg: A web-based resource for integrative and conjugative elements found in Bacteria. Nucleic Acids Res. 2012, 40, D621–D626. [Google Scholar] [CrossRef] [PubMed]
- Bioteau, A.; Durand, R.; Burrus, V. Redefinition and Unification of the SXT/R391 Family of Integrative and Conjugative Elements. Appl. Environ. Microbiol. 2018, 84, e00485-18. [Google Scholar] [CrossRef] [PubMed]
- Boltner, D.; MacMahon, C.; Pembroke, J.T.; Strike, P.; Osborn, A.M. R391: A conjugative integrating mosaic comprised of phage, plasmid, and transposon elements. J. Bacteriol. 2002, 18, 5158–5169. [Google Scholar] [CrossRef]
- Beaber, J.W.; Hochhut, B.; Waldor, M.K. Genomic and functional analyses of SXT, an integrating antibiotic resistance gene transfer element derived from Vibrio cholerae. J. Bacteriol. 2002, 15, 4259–4269. [Google Scholar] [CrossRef]
- Ryan, M.P.; Armshaw, P.; O’Halloran, J.A.; Pembroke, J.T. Analysis and comparative genomics of R997, the first SXT/R391 integrative and conjugative element (ICE) of the Indian Sub-Continent. Sci. Rep. 2007, 7. [Google Scholar] [CrossRef]
- López-Pérez, M.; Gonzaga, A.; Rodriguez-Valera, F. Genomic diversity of “deep ecotype” Alteromonas macleodii isolates: Evidence for Pan-Mediterranean clonal frames. Genome Biol. Evol. 2013, 6, 1220–1232. [Google Scholar] [CrossRef]
- Badhai, J.; Das, S.K. Characterization of three novel SXT/R391 integrating conjugative elements ICEMfuInd1a and ICEMfuInd1b, and ICEMprChn1 identified in the genomes of Marinomonas fungiae JCM 18476T and Marinomonas profundimaris strain D104. Front. Microbiol. 2016, 7, 1896. [Google Scholar] [CrossRef]
- Dong, C.; Bai, X.; Lai, Q.; Xie, Y.; Chen, X.; Shao, Z. Draft genome sequence of Marinomonas sp. strain D104, a polycyclic aromatic hydrocarbon-degrading bacterium from the deep-sea sediment of the Arctic Ocean. Genome Announc. 2014, 1, e01211-13. [Google Scholar] [CrossRef]
- Wozniak, R.A.F.; Fouts, D.E.; Spagnoletti, M.; Colombo, M.M.; Ceccarelli, D.; Garriss, G.; Dery, C.; Burrus, V.; Waldor, M.K. Comparative ICE Genomics: Insights into the Evolution of the SXT/R391 Family of ICEs. PLoS Genet. 2009, 12. [Google Scholar] [CrossRef]
- Osorio, C.R.; Marrero, J.; Wozniak, R.A.F.; Lemos, M.L.; Burrus, V.; Waldor, M.K. Genomic and functional analysis of ICEPdaSpa1, a fish-pathogen-derived SXT-related integrating conjugative element that can mobilize a virulence plasmid. J. Bacteriol. 2008, 9, 3353–3361. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lei, C.-W.; Zhang, A.-Y.; Wang, H.-N.; Liu, B.-H.; Yang, L.-Q.; Yang, Y.-Q. Characterization of SXT/R391 integrative and conjugative elements in Proteus mirabilis isolates from food-producing animals in China. Antimicrob. Agents Chemoth. 2016, 3, 1935–1938. [Google Scholar] [CrossRef] [PubMed]
- Pearson, M.M.; Sebaihia, M.; Churcher, C.; Quail, M.A.; Seshasayee, A.S.; Luscombe, N.M.; Abdellah, Z.; Arrosmith, C.; Atkin, B.; Chillingworth, T.; et al. Complete genome sequence of uropathogenic Proteus mirabilis, a master of both adherence and motility. J. Bacteriol. 2008, 11, 4027–4037. [Google Scholar] [CrossRef] [PubMed]
- Pembroke, J.T.; Piterina, A.V. A novel ICE in the genome of Shewanella putrefaciens W3-18-1: Comparison with the SXT/R391 ICE-like elements. FEMS Microbiol. Lett. 2006, 1, 80–88. [Google Scholar] [CrossRef][Green Version]
- Luo, P.; He, X.; Wang, Y.; Liu, Q.; Hu, C. Comparative genomic analysis of six new-found integrative conjugative elements (ICEs) in Vibrio alginolyticus. BMC Microbiol. 2016, 1, 79. [Google Scholar] [CrossRef]
- Bhotra, T.; Singh, D.V. Whole-genome sequence of Vibrio alginolyticus isolated from the mucus of the coral Fungia danai in the Andaman Sea, India’. Genome Announc. 2016, 3, e00339-16. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.B.; Yu, D.; Yue, J.J.; Kan, B. Variations in SXT elements in epidemic Vibrio cholerae O1 El Tor strains in China. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Yi, Y.; Lu, N.; Liu, F.; Li, J.; Zhang, R.; Jia, L.; Jing, H.; Xia, H.; Yang, Y.; Zhu, B.; et al. Genome sequence and comparative analysis of a Vibrio cholerae O139 strain E306 isolated from a cholera case in China. Gut Pathog. 2014, 1, 3. [Google Scholar] [CrossRef]
- Sjölund-Karlsson, M.; Reimer, A.; Folster, J.P.; Walker, M.; Dahourou, G.A.; Batra, D.G.; Martin, I.; Joyce, K.; Parsons, M.B.; Boncy, J.; et al. Drug-resistance mechanisms in Vibrio cholerae O1 outbreak strain, Haiti, 2010. Emerg. Infect. Dis. 2011, 11, 2151. [Google Scholar] [CrossRef]
- Ceccarelli, D.; Spagnoletti, M.; Hasan, N.A.; Lansing, S.; Huq, A.; Colwell, R.R. A new integrative conjugative element detected in Haitian isolates of Vibrio cholerae non-O1/non-O139. Res. Microbiol. 2013, 9, 891–893. [Google Scholar] [CrossRef]
- Chowdhury, F.; Mather, A.E.; Begum, Y.A.; Asaduzzaman, M.; Baby, N.; Sharmin, S.; Biswas, R.; Uddin, M.I.; LaRocque, R.C.; Harris, J.B.; et al. Vibrio cholerae serogroup O139: Isolation from cholera patients and asymptomatic household family members in Bangladesh between 2013 and 2014. PLoS Negl. Trop. Dis. 2015, 11, e0004183. [Google Scholar] [CrossRef] [PubMed]
- Taviani, E.; Grim, C.J.; Chun, J.; Huq, A.; Colwell, R. Genomic analysis of a novel integrative conjugative element in Vibrio cholerae. FEBS Lett. 2009, 22, 3630–3636. [Google Scholar] [CrossRef] [PubMed]
- de Oliveira Veras, A.A.; da Silva, M.L.; Gomes, J.C.M.; Dias, L.M.; de Sá, P.C.G.; Alves, J.T.C.; Castro, W.; Miranda, F.; Kazuo, E.; Marinho, D.; et al. Draft genome sequences of Vibrio fluvialis strains 560 and 539, isolated from environmental samples. Genome Announc. 2015, 1, e01344-14. [Google Scholar] [CrossRef]
- Li, Y.; Li, Y.; Fernandez Crespo, R.; Leanse, L.G.; Langford, P.R.; Bossé, J.T. Characterization of the Actinobacillus pleuropneumoniae SXT-related integrative and conjugative element ICE Apl2 and analysis of the encoded FloR protein: Hydrophobic residues in transmembrane domains contribute dynamically to florfenicol and chloramphenicol efflux. J. Antimicrob. Chemother. 2017, 1, 57–65. [Google Scholar]
- Okada, K.; Natakuathung, W.; Na-Ubol, M.; Roobthaisong, A.; Wongboot, W.; Maruyama, F.; Nakagawa, I.; Chantaroj, S.; Hamada, S. Characterization of 3 Megabase-Sized Circular Replicons from Vibrio cholerae. Emerg. Infect. Dis. 2015, 7, 1262–1263. [Google Scholar] [CrossRef] [PubMed]
- Poulin-Laprade, D.; Matteau, D.; Jacques, P.-E.; Rodrigue, S.; Burrus, V. Transfer activation of SXT/R391 integrative and conjugative elements: Unraveling the SetCD regulon. Nucleic Acids Res. 2015, 4, 2045–2056. [Google Scholar] [CrossRef] [PubMed]
- López-Pérez, M.; Ramon-Marco, N.; Rodriguez-Valera, F. Networking in microbes: Conjugative elements and plasmids in the genus Alteromonas. BMC Genomics 2017, 1, 36. [Google Scholar] [CrossRef]
- Frey, K.; Bishop-Lilly, K.; Daligault, H.; Davenport, K.; Bruce, D.; Chain, P.; Coyne, S.; Chertkov, O.; Freitas, T.; Jaissle, J.; et al. Full-genome assembly of reference strain Providencia stuartii ATCC 33672. Genome Announc. 2014, 5, e01082-14. [Google Scholar] [CrossRef]
- Kuleshov, K.V.; Kostikova, A.; Pisarenko, S.V.; Kovalev, D.A.; Tikhonov, S.N.; Savelieva, I.V.; Saveliev, V.N.; Vasilieva, O.V.; Zinich, L.S.; Pidchenko, N.N.; et al. Comparative genomic analysis of two isolates of Vibrio cholerae O1 Ogawa El Tor isolated during outbreak in Mariupol in 2011. Infect. Genet. Evol. 2016, 44, 471–478. [Google Scholar] [CrossRef]
- Marin, M.A.; Fonseca, E.L.; Andrade, B.N.; Cabral, A.C.; Vicente, A.C.P. Worldwide occurrence of integrative conjugative element encoding multidrug resistance determinants in epidemic Vibrio cholerae O1. PLoS ONE 2014, 9, e108728. [Google Scholar] [CrossRef]
- Li, X.Y.; Du, Y.; Du, P.C.; Dai, H.; Fang, Y.J.; Li, Z.P.; Lv, N.; Zhu, B.L.; Kan, B.; Wang, D.C. SXT/R391 integrative and conjugative elements in Proteus species reveal abundant genetic diversity and multidrug resistance. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Bie, L.Y.; Wu, H.; Wang, X.H.; Wang, M.Y.; Xu, H. Identification and characterization of new members of the SXT/R391 family of integrative and conjugative elements (ICEs) in Proteus mirabilis. Int. J. Antimicrob. Agents 2017, 2, 242–246. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Wang, Y.; Li, Z.; Liu, Z.; Li, X.; Diao, B.; Kan, B.; Wang, D. Distribution and genetic characteristics of SXT/R391 integrative conjugative elements in Shewanella spp. from China. Front. Microbiol. 2018, 9, 920. [Google Scholar] [CrossRef] [PubMed]
- Siebor, E.; de Curraize, C.; Neuwirth, C. Genomic context of resistance genes within a French clinical MDR Proteus mirabilis: Identification of the novel genomic resistance island GI Pmi 1. J. Antimicrob. Chemother. 2018, 7, 1808–1811. [Google Scholar] [CrossRef]
- Sugimoto, Y.; Maruyama, F.; Suzuki, S. Draft genome sequence of a Shewanella halifaxensis strain isolated from the intestine of Marine Red Seabream (Pagrus major), which includes an integrative conjugative element with macrolide resistance genes. Genome Announc. 2018, 16, e00297-18. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Jia, H.; Cui, G.; Tong, H.; Wei, J.; Shao, D.; Liu, K.; Qiu, Y.; Li, B.; Ma, Z. ICEAplChn1, a novel SXT/R391 integrative conjugative element (ICE), carrying multiple antibiotic resistance genes in Actinobacillus pleuropneumoniae. Vet. Microbiol. 2018, 220, 18–23. [Google Scholar] [CrossRef] [PubMed]
- Lei, C.-W.; Chen, Y.-P.; Kang, Z.-Z.; Kong, L.-H.; Wang, H.-N. Characterization of a novel SXT/R391 Integrative and Conjugative Element carrying cfr, blaCTX-M-65, fosA3 and aac(6′)-Ib-cr in Proteus mirabilis. Antimicrob Agents Chemoth. 2018, 9, 00849-18. [Google Scholar] [CrossRef]
- Di Noto, G.P.; Jara, E.; Iriarte, A.; Centrón, D.; Quiroga, C. Genome analysis of a clinical isolate of Shewanella sp. uncovered an active hybrid integrative and conjugative element carrying an integron platform inserted in a novel genomic locus. Microbiol. 2016, 8, 1335–1345. [Google Scholar] [CrossRef]
- Villeneuve, C.; Martineau, C.; Mauffrey, F.; Villemur, R. Complete genome sequences of Methylophaga sp. strain JAM1 and Methylophaga sp. strain JAM7. J. Bacteriol. 2012, 194, 4126–4127. [Google Scholar] [CrossRef]
- Xu, M.; Fang, Y.; Liu, J.; Chen, X.; Sun, G.; Guo, J.; Hua, Z.; Tu, Q.; Wu, L.; Zhou, J.; et al. Draft genome sequence of Shewanella decolorationis S12, a dye-degrading bacterium isolated from a wastewater treatment plant. Genome Announc. 2013, 6, e00993-13. [Google Scholar] [CrossRef]
- Deng, Y.; Chen, C.; Zhao, Z.; Zhao, J.; Jacq, A.; Huang, X.; Yang, Y. The RNA chaperone Hfq is involved in colony morphology, nutrient utilization and oxidative and envelope stress response in Vibrio alginolyticus. PLoS ONE 2016, 9, e0163689. [Google Scholar] [CrossRef] [PubMed]
- Azarian, T.; Ali, A.; Johnson, J.A.; Mohr, D.; Prosperi, M.; Veras, N.M.; Jubair, M.; Strickland, S.L.; Rashid, M.H.; Alam, M.T.; et al. Phylodynamic analysis of clinical and environmental Vibrio cholerae isolates from Haiti reveals diversification driven by positive selection. MBio 2014, 6, e01824-14. [Google Scholar] [CrossRef] [PubMed]
- Bishop-Lilly, K.A.; Johnson, S.L.; Verratti, K.; Luu, T.; Khiani, A.; Awosika, J.; Mokashi, V.P.; Chain, P.S.; Sozhamannan, S. Genome sequencing of 15 clinical Vibrio isolates, including 13 non-O1/non-O139 serogroup strains. Genome Announc. 2014, 5, e00893-14. [Google Scholar] [CrossRef] [PubMed]
- Orata, F.D.; Kirchberger, P.C.; Méheust, R.; Barlow, E.J.; Tarr, C.L.; Boucher, Y. The dynamics of genetic interactions between Vibrio metoecus and Vibrio cholerae, two close relatives co-occurring in the environment. Genome Biol. Evol. 2015, 10, 2941–2954. [Google Scholar] [CrossRef] [PubMed]
- Zhu, C.; Sun, B.; Liu, T.; Zheng, H.; Gu, W.; He, W.; Sun, F.; Wang, Y.; Yang, M.; Bei, W.; et al. Genomic and transcriptomic analyses reveal distinct biological functions for cold shock proteins (Vpa CspA and Vpa CspD) in Vibrio parahaemolyticus CHN25 during low-temperature survival. BMC genomics 2017, 1, 436. [Google Scholar]
- Cui, Y.; Yang, X.; Didelot, X.; Guo, C.; Li, D.; Yan, Y.; Zhang, Y.; Yuan, Y.; Yang, H.; Wang, J.; et al. Epidemic clones, oceanic gene pools, and eco-LD in the free living marine pathogen Vibrio parahaemolyticus. Mol. Biol. Evol. 2015, 6, 1396–1410. [Google Scholar] [CrossRef]
- Kalburge, S.; Polson, S.; Crotty, K.B.; Katz, L.; Turnsek, M.; Tarr, C.; Martinez-Urtaza, J.; Boyd, E. Complete genome sequence of Vibrio parahaemolyticus environmental strain UCM-V493. Genome Announc. 2014, 2, e00159-14. [Google Scholar] [CrossRef]
- Danin-Poleg, Y.; Raz, N.; Roig, F.J.; Amaro, C.; Kashi, Y. Draft genome sequence of environmental bacterium Vibrio vulnificus CladeA-yb158. Genome Announc. 2015, 4, e00754-15. [Google Scholar] [CrossRef]
- Roig, F.J.; González-Candelas, F.; Sanjuán, E.; Fouz, B.; Feil, E.J.; Llorens, C.; Baker-Austin, C.; Oliver, J.D.; Danin-Poleg, Y.; Gibas, C.J.; et al. Phylogeny of Vibrio vulnificus from the analysis of the core-genome: Implications for intra-species taxonomy. Front. Microbiol. 2018, 8, 2613. [Google Scholar] [CrossRef]
- Tiruvayipati, S.; Bhassu, S.; Kumar, N.; Baddam, R.; Shaik, S.; Gurindapalli, A.K.; Thong, K.L.; Ahmed, N. Genome anatomy of the gastrointestinal pathogen, Vibrio parahaemolyticus of crustacean origin. Gut Pathog. 2013, 1, 37. [Google Scholar] [CrossRef]
- Yang, Y.-T.; Chen, I.-T.; Lee, C.-T.; Chen, C.-Y.; Lin, S.-S.; Hor, L.-I.; Tseng, T.-C.; Huang, Y.-T.; Sritunyalucksana, K.; Thitamadee, S.; et al. Draft genome sequences of four strains of Vibrio parahaemolyticus, three of which cause early mortality syndrome/acute hepatopancreatic necrosis disease in shrimp in China and Thailand. Genome Announc. 2014, 5, e00816-14. [Google Scholar] [CrossRef] [PubMed]
- Kumar, R.; Chang, C.-C.; Ng, T.H.; Ding, J.-Y.; Tseng, T.-C.; Lo, C.-F.; Wang, H.-C. Draft genome sequence of Vibrio parahaemolyticus strain M1-1, which causes acute hepatopancreatic necrosis disease in shrimp in Vietnam. Genome Announc. 2018, 3, e01468-17. [Google Scholar] [CrossRef] [PubMed]
- Ryan, M.P.; Armshaw, P.; Pembroke, J.T. New and emerging SXT/R391 integrative conjugative elements as vehicles for stable mobile element transfer and spread of antibiotic resistance in both human and animals. In Antimicrobial Research: Novel Bioknowledge and Educational Programs; Méndez-Vilas, A., Ed.; Formatex Research Center: Badajoz, Spain, 2017; Volume 6, pp. 593–598. [Google Scholar]
- McGrath, B.M.; Pembroke, J.T. Detailed analysis of the insertion site of the mobile elements R997, pMERPH, R392, R705 and R391 in E. coli K12. FEMS Microbiol. Lett. 2004, 237, 19–26. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Burrus, V.; Marrero, J.; Waldor, M.K. The current ICE age: Biology and evolution of SXT-related integrating conjugative elements. Plasmid 2006, 55, 173–183. [Google Scholar] [CrossRef]
- Hedges, R.W.; Datta, N.; Coetzee, J.N.; Dennison, S. R factors from Proteus morganii. J. Gen. Microbiol. 1973, 77, 249–259. [Google Scholar] [CrossRef]
- Armshaw, P.; Pembroke, J.T. Examination of the cell sensitizing gene orf43 of ICE R391 suggests a role in ICE transfer enhancement to recipient cells. FEMS Microbiol. Lett. 2015, 362, 1–7. [Google Scholar] [CrossRef]
- Armshaw, P.; Pembroke, J.T. Control of expression of the ICE R391 encoded UV-inducible cell-sensitising function. BMC Microbiol. 2013, 13, 195. [Google Scholar] [CrossRef] [PubMed]
- Armshaw, P.; Pembroke, J.T. Generation and analysis of an ICE R391 deletion library identifies genes involved in the element encoded UV-inducible cell-sensitising function. FEMS Microbiol. Lett. 2013, 342, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Ryan, M.P.; Armshaw, P.; Pembroke, J.T. SXT/R391 integrative and conjugative elements [ICEs] encode a novel “trap-door” strategy for mobile element escape. Front. Microbiol. 2016, 7, 829. [Google Scholar] [CrossRef] [PubMed]
- Xiong, A.; Jayaswal, R.K. Molecular characterization of a chromosomal determinant conferring resistance to zinc and cobalt ions in Staphylococcus aureus. J. Bacteriol. 1998, 180, 4024–4029. [Google Scholar]
- Kunito, T.; Kusano, T.; Oyaizu, H.; Senoo, K.; Kanazawa, S.; Matsumoto, S. Cloning and Sequence Analysis of czc Genes in Alcaligenes sp. Strain CT14. Biosci. Biotechnol. Biochem. 2009, 60, 699–704. [Google Scholar] [CrossRef] [PubMed]
- Peters, S.E.; Hobman, J.L.; Strike, P.; Ritchie, D.A. Novel mercury resistance determinants carried by IncJ plasmids pMERPH and R391. Mol. Gen. Genet. 1991, 228, 294–299. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Blanco, A.; Lemos, M.L.; Osorio, C.R. Integrating Conjugative Elements as vectors of antibiotic, Mercury, and Quaternary Ammonium compound resistance in marine aquaculture environments. Antimicrob. Agents Chemother. 2012, 56, 2619–2626. [Google Scholar] [CrossRef] [PubMed]
- Parmeciano DI Noto, G.; Iriarte, A.; Ramírez, M.S.; Centrón, D.; Quiroga, C. ICE SXT vs. ICESh95: Co-existence of Integrative and Conjugative Elements and Competition for a New Host. Sci. Rep. 2019, 9, 8045. [Google Scholar] [CrossRef] [PubMed]
- Davies, M.R.; Shera, J.; Van Domselaar, G.H.; Sriprakash, K.S.; McMillan, D.J. A Novel Integrative Conjugative Element mediates genetic transfer from Group G Streptococcus to other ß-Hemolytic Streptococci. J. Bacteriol. 2009, 191, 2257–2265. [Google Scholar] [CrossRef] [PubMed]
- Ryan, M.P.; Pembroke, J.T.; Adley, C.C. Novel Tn4371-ICE like element in Ralstonia pickettii and Genome mining for comparative elements. BMC Microbiol. 2009, 9, 242. [Google Scholar] [CrossRef]
- Jones, P.D. Water quality and fisheries in the Mersey estuary, England: A historical perspective. Mar. Pollut. Bull. 2006, 53, 144–154. [Google Scholar] [CrossRef]
- Wilke, A.; Zagnitko, O.; Formsma, K.; Aziz, R.K.; Kubal, M.; Vonstein, V.; Stevens, R.; McNeil, L.K.; Edwards, R.A.; Pusch, G.D.; et al. The RAST Server: Rapid Annotations using Subsystems Technology. BMC Genom. 2008, 9, 75. [Google Scholar]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Murphy, D.; Pembroke, J.T. Transfer of the IncJ plasmid R391 to recombination deficient Escherichia coli K12: Evidence that R391 behaves as a conjugal transposon. FEMS Microbiol. Lett. 2002, 134, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Oden, K.L.; Gladysheva, T.B.; Rosen, B.P. Arsenate reduction mediated by the plasmid-encoded ArsC protein is coupled to glutathione. Mol. Microbiol. 1994, 12, 301–306. [Google Scholar] [CrossRef] [PubMed]
- Anes, J.; McCusker, M.P.; Fanning, S.; Martins, M. The ins and outs of RND efflux pumps in Escherichia coli. Front. Microbiol. 2015, 6, 587. [Google Scholar] [CrossRef] [PubMed]
- Panne, D.; Müller, S.A.; Wirtz, S.; Engel, A.; Bickle, T.A. The McrBCx restriction endonuclease assembles into a ring structure in the presence of G nucleotides. EMBO J. 2001, 20, 3210–3217. [Google Scholar] [CrossRef] [PubMed]
- Romaniuk, K.; Golec, P.; Dziewit, L. Insight Into the Diversity and Possible Role of Plasmids in the Adaptation of Psychrotolerant and Metalotolerant Arthrobacter spp. to Extreme Antarctic Environments. Front. Microbiol. 2018, 9, 3144. [Google Scholar] [CrossRef] [PubMed]
- Bhatt, K.; Timsit, E.; Rawlyk, N.; Potter, A.; Liljebjelke, K. Integrative Conjugative Element ICEHs1 encodes for antimicrobial resistance and metal tolerance in Histophilus somni. Front. Vet. Sci. 2018, 5, 153. [Google Scholar] [CrossRef]
- Harada, S.; Ishii, Y.; Saga, T.; Tateda, K.; Yamaguchi, K. Chromosomally encoded blaCMY-2 located on a novel SXT/R391-related Integrating Conjugative Element in a Proteus mirabilis clinical isolate. Antimicrob. Agents Chemother. 2010, 54, 3545–3550. [Google Scholar] [CrossRef]
- Balado, M.; Lemos, M.L.; Osorio, C.R. Integrating conjugative elements of the SXT/R391 family from fish-isolated Vibrios encode restriction-modification systems that confer resistance to bacteriophages. FEMS Microbiol. Ecol. 2013, 83, 457–467. [Google Scholar] [CrossRef]
- Raleigh, E.A.; Wilson, G. Escherichia coli K-12 restricts DNA containing 5-methylcytosine. Proc. Natl. Acad. Sci. USA 1986, 83, 9070–9074. [Google Scholar] [CrossRef]
- Carraro, N.; Poulin, D.; Burrus, V. Replication and active partition of Integrative and Conjugative Elements [ICEs] of the SXT/R391 Family: The line between ICEs and Conjugative plasmids is getting thinner. PLoS Genet. 2015, 11, e1005298. [Google Scholar] [CrossRef]
- Ryan, M.P.; Slattery, S.; Pembroke, J.T. Conservation of Mercury resistance determinants amongst ICE-like mobile bacterial genetic elements: Comparative analysis and dissection of function. In Understanding Microbial Pathogens: Current Knowledge and Educational Ideas on Antimicrobial Research; Méndez-Vilas, A., Ed.; Formatex Research Center: Badajoz, Spain, 2018; Volume 7, pp. 121–126. [Google Scholar]
- Chen, J.; Yoshinaga, M.; Garbinski, L.D.; Rosen, B.P. Synergistic interaction of glyceraldehydes-3-phosphate dehydrogenase and ArsJ, a novel organoarsenical efflux permease, confers arsenate resistance. Mol. Microbiol. 2016, 100, 945–953. [Google Scholar] [CrossRef] [PubMed]
- Zegers, I.; Martins, J.C.; Willem, R.; Wyns, L.; Messens, J. Arsenate reductase from S. aureus plasmid pI258 is a phosphatase drafted for redox duty. Nat. Struct. Biol. 2001, 8, 843–847. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Madegowda, M.; Bhattacharjee, H.; Rosen, B.P. ArsP: A methylarsenite efflux permease. Mol. Microbiol. 2015, 98, 625–635. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Wu, S.; Lilley, R.M.; Zhang, R. The diversity of membrane transporters encoded in bacterial arsenic-resistance operons. Peer J. 2015, 3, e943. [Google Scholar] [CrossRef]
- Fekih, I.B.; Zhang, C.; Li, Y.P.; Zhao, Y.; Alwathnani, H.A.; Saquib, Q.; Rensing, C.; Cervantes, C. Distribution of arsenic resistance genes in prokaryotes. Front. Microbiol. 2018, 9, 2473. [Google Scholar] [CrossRef]
- Diorio, C.; Cai, J.; Marmor, J.; Shinder, R.; DuBow, M.S. An Escherichia coli chromosomal ars operon homolog is functional in arsenic detoxification and is conserved in gram-negative bacteria. J. Bacteriol. 1995, 8, 2050–2056. [Google Scholar] [CrossRef] [PubMed]
- Ji, G.; Silver, S. Reduction of arsenate to arsenite by the ArsC protein of the arsenic resistance operon of Staphylococcus aureus plasmid pI258. Proc. Natl. Acad. Sci. USA 1992, 20, 9474–9478. [Google Scholar] [CrossRef] [PubMed]
- Rosenstein, R.; Peschel, A.; Wieland, B.; Götz, F. Expression and regulation of the antimonite, arsenite, and arsenate resistance operon of Staphylococcus xylosus plasmid pSX267. J. Bacteriol. 1992, 11, 3676–3683. [Google Scholar] [CrossRef] [PubMed]
- Hedges, R.; Baumberg, S. Resistance to arsenic compounds conferred by a plasmid transmissible between strains of Escherichia coli. J. Bacteriol. 1973, 1, 459. [Google Scholar]
- Bruhn, D.F.; Li, J.; Silver, S.; Roberta, F.; Rosen, B.P. The arsenical resistance operon of IncN plasmid R46. FEMS Microbiol. Lett. 1996, 2–3, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Silver, S.; Phung, L.T. Bacterial heavy metal resistance: New surprises. Annu. Rev. Microb. 1996, 1, 753–789. [Google Scholar] [CrossRef] [PubMed]
- Tuffin, I.M.; de Groot, P.; Deane, S.M.; Rawlings, D.E. An unusual Tn21-like transposon containing an ars operon is present in highly arsenic-resistant strains of the biomining bacterium Acidithiobacillus caldus. Microbiol. 2005, 9, 3027–3039. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.; Postis, V.L.; Rahman, M.; Wright, G.S.; Roach, P.C.; Deacon, S.E.; Ingram, J.C.; Henderson, P.J.; Findlay, J.B.; Phillips, S.E.; et al. Investigation of the structure and function of a Shewanella oneidensis arsenical-resistance family transporter. Mol. Membr. Biol. 2008, 8, 691–701. [Google Scholar] [CrossRef] [PubMed]
- Xu, C.; Shi, W.; Rosen, B.P. The chromosomal arsR gene of Escherichia coli encodes a trans-acting metalloregulatory protein. J. Biol. Chem. 1996, 5, 2427–2432. [Google Scholar] [CrossRef] [PubMed]

| Strain | Genotype | Source |
|---|---|---|
| AB1157 | F−, thr-1, araC14, leuB6,∆(gpt-proA)62, lacY1, tsx-33, qsr’-0, glnV44, galK2, λ-, Rac-0,hisG4, rfbC1, mgl-51, rpoS396, rpsL31 (StrR), kdgK51, xylA5, mtl-1, argE3, thi-1 | E. coli genetic stock center (CGSC), Yale University, New Haven, Connecticut, USA |
| NCIMB12504 (K802) | F-, lacY1 or Δ(cod-lacI)6, glnX44(AS), galK2(Oc), galT22, λ-, e14-, mcrA0, rfbC1, metB1, mcrB1, hsdR2 | NCIMB |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ryan, M.P.; Slattery, S.; Pembroke, J.T. A Novel Arsenate-Resistant Determinant Associated with ICEpMERPH, a Member of the SXT/R391 Group of Mobile Genetic Elements. Genes 2019, 10, 1048. https://doi.org/10.3390/genes10121048
Ryan MP, Slattery S, Pembroke JT. A Novel Arsenate-Resistant Determinant Associated with ICEpMERPH, a Member of the SXT/R391 Group of Mobile Genetic Elements. Genes. 2019; 10(12):1048. https://doi.org/10.3390/genes10121048
Chicago/Turabian StyleRyan, Michael P., Shannon Slattery, and J. Tony Pembroke. 2019. "A Novel Arsenate-Resistant Determinant Associated with ICEpMERPH, a Member of the SXT/R391 Group of Mobile Genetic Elements" Genes 10, no. 12: 1048. https://doi.org/10.3390/genes10121048
APA StyleRyan, M. P., Slattery, S., & Pembroke, J. T. (2019). A Novel Arsenate-Resistant Determinant Associated with ICEpMERPH, a Member of the SXT/R391 Group of Mobile Genetic Elements. Genes, 10(12), 1048. https://doi.org/10.3390/genes10121048






