1. Introduction
Chromosome rearrangements are known to be one of the main etiological factors contributing to hypoprolificacy in domestic species, especially the domestic pig [
1,
2,
3]. To date over 200 distinct chromosome rearrangements have been identified in the domestic pig. The vast majority are balanced reciprocal translocations, making up over 90% of described rearrangements [
4]. It is estimated that chromosome rearrangements in domestic pigs occur spontaneously in one of 200 live births [
5,
6]. The prevalence of chromosome rearrangements among swine herds is thought to be between 0.5% and 1.5%, dependent on the intensity of cytogenetic screening within these populations [
3,
5,
6]. Although reciprocal translocations are quite prevalent throughout swine herds and the genetic and genomic reasons for this are poorly understood.
Reciprocal translocations are rearrangements involving two non-homologous chromosomes which break simultaneously, and subsequently misrepair, resulting in an exchange of chromosome segments. Research that involves the human genome has suggested that translocation breakpoints occurred nonrandomly for each individual chromosome pair, with specific chromosomes and cytogenetic landmarks being particularly susceptible to breakage. Various chromosomal features recognizable in a karyotype such as the total length of chromosomes and chromosome arms, chromosome morphology, as well as, chromatin density (heterochromatic and euchromatic), and the presence of common fragile sites have all been suggested to influence the frequency at which chromosome regions rearrange [
7,
8,
9,
10].
Despite the considerable number of chromosome rearrangements identified in the domestic pig, the characteristics of these rearrangements is largely unknown. Some work suggests that there is a non-random distribution of translocation breakpoints across chromosomes in the pig genome [
1,
11]. Amongst the first five reciprocal translocations identified in pigs, 2 breakpoints appeared on chromosome 6, and three breakpoints were found on chromosome 14 [
11,
12,
13,
14]. The presence of multiple breakpoints in close proximity on chromosome 14 led to the suggestion that regions of fragility promoting chromosome breakage may be present in the pig karyotype, and that there may be a non-random distribution of translocation breakpoints [
14,
15]. In addition, many breakpoints are known to overlap with common fragile sites, regions of the chromosomes which are susceptible to breakage under exposure of specific chemical stressors [
16].
Although a handful of chromosomes have been examined in some detail, no comprehensive analysis of translocation breakpoints across the pig karyotype has ever been conducted. The identification of chromosome rearrangements in swine herds has increased in the past two decades with the continuation of a large screening programs at the National Veterinary School of Toulouse in France, and at the University of Guelph in Canada. With over 190 unique reciprocal chromosome translocations identified in the domestic pig, it is now easier to observe patterns in the number of breakpoints on chromosomes, chromosome arms, and cytogenetic landmarks.
Using 195 reciprocal chromosome rearrangements identified in our lab and those reported in the literature we performed a comprehensive analysis of the translocation breakpoints at the whole chromosome and cytogenetic band levels. Our observation of rearrangements breakpoints lead us to add empirical evidence that breakpoints are nonrandomly distributed across chromosomes, and cytogenetic bands.
4. Discussion and Conclusions
Observing the distribution of translocation breakpoints across chromosomes, chromosome arms, and cytogenetic bands revealed the chromosome regions most susceptible to rearrangement, and physical features that are associated with higher breakpoint number and translocation frequency. Translocation breakpoints are nonrandomly distributed across chromosomes and chromosome arms, with particular chromosomes appearing to be far more susceptible to rearrangement than others. In addition, the length of cytogenetic bands, G-banding (heterochromatin and euchromatin), and the presence of fragile sites were all found to be associated with a higher number of breakpoints on cytogenetic bands. In particular we observed that G-negative bands had high translocation frequencies on average, with those G-negative bands with common fragile sites having the highest translocation frequencies of all bands on average.
Taking the chromosome as an individual unit, we found that chromosomes do not rearrange in direct proportion to their length. Although longer chromosomes tended to have more breakpoints in general, length appeared to have no relationship with whether chromosomes had a deficiency or a surplus of breakpoints. The physical length of chromosomes has long been suggested to influence the number of translocation breakpoints on each chromosome in the pig [
4,
21], and human [
9,
22] genomes. The rationale behind this is that longer chromosomes should have more opportunities for breakage, and therefore should break and translocate more often. Taken simply as a numerical value, this is generally true in the pig, however once length is accounted for and we see no evidence that longer physical length is associated with higher translocation frequency. Both long and short chromosomes have high (chromosomes 12 and 14) and low (chromosomes 2 and 18) translocation frequencies. Those chromosomes with the highest translocation frequencies appear to have some feature that promotes more frequent rearrangement. This suggests that chromosome features beyond the simple physical length and breakpoint number should be considered for potential roles in promoting translocation events.
Observing the number of breakpoints relative to the physical length of chromosome arms yielded slightly different results. Longer chromosome arms tended to have more translocation breakpoints, and chromosome arms typically rearranged in proportion to their length. As with whole chromosomes, longer chromosome arms have been previously suggested to be predisposed to having more translocation breakpoints [
10]. Examining the translocation frequencies of the p and q arms of each chromosome, however, revealed a different trend. We observed that these chromosome arm pairs may have considerably different translocation frequencies from one another even though they are part of the same chromosome, for example, chromosome arm 12q has a translocation frequency 3.4x higher than chromosome arm 12p. Differences in breakpoint number between chromosome arm pairs has been previously established [
8], however this difference in translocation frequency is perhaps more unexpected, as length is taken into account, and suggests that the factors that promote rearrangement on each chromosome are not necessarily present across the entire chromosome, and may be more localized to specific regions.
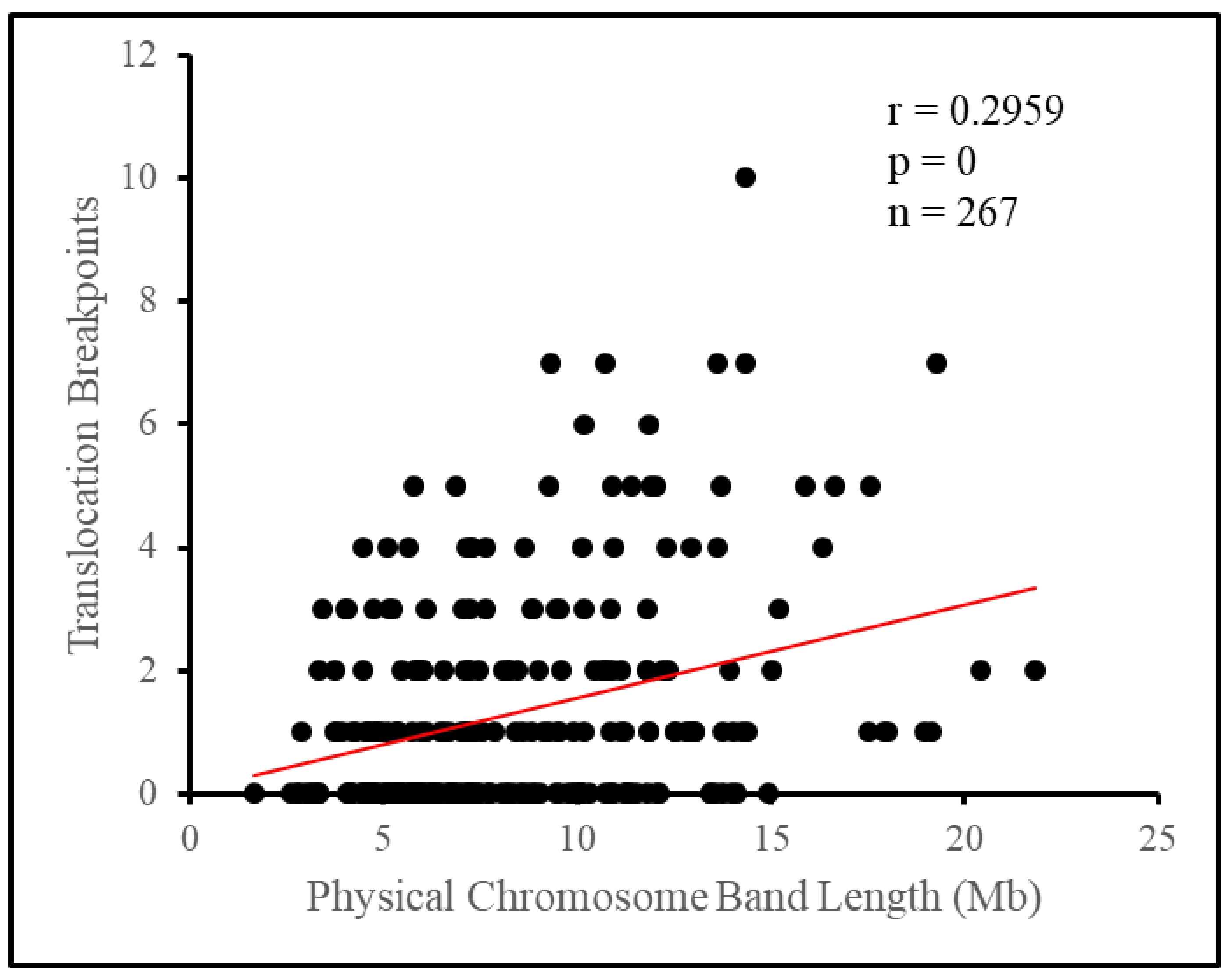
Breaking the chromosome down further into cytogenetic bands, we observed the appearance of a non-random distribution of breakpoints. Attempting to fit the number of breakpoints per band under a Poisson distribution yielded a poor fit, as a surplus of bands had no breakpoints at all, or four or more breakpoints, while there was a deficiency of bands with one to three breakpoints. As such, there tended to be a clustering of breakpoints on relatively few bands, with 12.4% of bands, those with four or more breakpoints, having 47.7% of all breakpoints between them. Length appeared to influence some of this difference as longer bands tended to have more breakpoints, however, this influence appeared small overall. Translocation breakpoints in the pig genome have previously been suggested to be distributed nonrandomly based on empirical evidence [
15]. In the human genome, breakpoints have previously been shown to be nonrandomly distributed across cytogenetic bands, with clusters of breakpoints on individual bands being referred to as hotspots for rearrangement [
7,
9,
10,
23]. Clustering of breakpoints in the pig and human genomes occur in a variety of positions, with no apparent influence due to the relative position to the centromere on the chromosome arm [
24]. The clustering of breakpoints in a few cytogenetic bands in distinct chromosomal positions, such as 1q17 and 15q13, suggests that individual bands may have some feature that promotes rearrangement events.
We found that G-negative bands were strongly associated with higher breakpoint numbers and translocation frequency. G-negative bands as a whole had a translocation frequency five times greater than that of G-positive bands. These results are in agreement with several studies of translocation breakpoints in humans which consistently indicate that cytogenetic G-negative bands have a higher density of breakpoints than G-positive bands [
7,
8,
9,
10]. The more open chromatin composition of these bands is proposed to be more susceptible to breakage than more condensed regions of chromosomes, although it has been suggested that cytogeneticists may be biased towards placing rearrangements within G-negative bands due to the contrasting nature of light and dark bands. Nevertheless, studies of R-banded chromosomes indicate that most rearrangements are identified in R-positive (G-negative) bands [
24,
25]. Therefore, our findings provide more evidences that sustain the notion that euchromatic chromosome regions are more susceptible to breakage and subsequent rearrangement.
The presence of common fragile sites within G-negative bands was found to be associated with higher translocation frequency. In contrast the presence of common fragile sites in G-positive bands appeared to have no influence on translocation frequency. Common fragile sites in the pig karyotype have been previously noted to overlap with the cytogenetic positions of translocation breakpoints [
6,
16,
26,
27]. Our results are generally consistent with those found in humans, as cytogenetic bands with fragile sites in the human genome were found to translocate more frequently than bands without fragile sites [
9,
28,
29,
30]. These results suggest that the presence of common fragile sites on G-negative bands may influence rearrangement by promoting breakage on more open chromatin regions which already appear more susceptible to rearrangement by their nature.
Considering the features associated with breakpoint number, we determined that the chromatin density (e.g., type of banding) followed by length of bands, and the presence of common fragile sites were found to significantly contribute to a regression model of translocation breakpoint number. Only chromatin density contributed to a model of translocation frequency, however, which is in line with the presence of common fragile sites only influencing translocation frequency when present in G-negative bands. Chromatin density and physical chromosome length have previously been shown to influence breakpoint number in the human genome [
10,
22], however, little work has been done to demonstrate if these factors influence translocation frequency in the human genome. Although it is apparent that length, chromatin density, and the presence of fragile sites play a role in influencing breakpoint number, together at most they explain approximately a third of the variation present amongst bands. We may speculate then that more specific chromosome features, unique to each band, may contribute more specifically to the promotion of translocation events.
We determined that translocation breakpoints appeared to be nonrandomly distributed across cytogenetic bands, and that factors such as length, G-banding, and the presence of fragile sites were all related to breakpoint number. We thus sought to propose hotspots for chromosome rearrangement in the pig genome based on these characteristics. Starting with the average number of breakpoints per band, 2.4, and average translocation frequency, 0.29 per Mb, we proposed that any band that had at least five breakpoints and/or a translocation frequency of 0.58 per Mb or higher to be hotspots for rearrangement in the pig genome. In total, nineteen bands based on number of breakpoints (
Table 10), and fifteen bands based on translocation frequency (
Table 11) were proposed as hotspots for rearrangement. Six bands were shared between the lists. These bands are derived from a variety of chromosomes and positions. All bands are in G-negative regions, and twelve had a common fragile site. Notably, many bands from shorter chromosomes with high translocation frequencies feature prominently amongst our proposed hotspot bands. For instance, chromosome 12, with just seventeen breakpoints has three bands amongst our hotspots, indicating the specific bands on this chromosome that appear to drive the high translocation frequency of this chromosome. In total, thirty bands with varied characteristics were proposed as hotspots for rearrangement in the pig genome.
For the first time, all known porcine reciprocal translocations were analyzed, revealing the chromosomes and cytogenetic bands with the highest number of breakpoints and highest translocation frequencies. The cytogenetic bands with the highest number of breakpoints and highest translocation frequencies were selected as potential hotspots for rearrangement in the pig karyotype. This work can serve as a foundation for future endeavors to assist breeders in the selection of pigs that are less susceptible to chromosome rearrangement, ultimately increasing efficiency and profitability for breeders. The comparison of chromosome rearrangements in the pig and human genomes suggests that the pig may serve as an appropriate biomedical model to study chromosome rearrangements. Although less than 200 porcine translocations were considered, this is the most complete analysis of chromosome rearrangements in the pig and it provides a baseline for future considerations of porcine chromosome rearrangements. Given the analyses concerning chromosome features easily mappable to the pig karyotype, future considerations should be given to the specific molecular and genomic landscape present on cytogenetic bands, in order to characterize which factors promote rearrangement, and explain why specific cytogenetic bands may have more breakpoints or higher translocation frequencies than other bands.