Investigation of the Possible Role of RAD9 in Post-Diapaused Embryonic Development of the Brine Shrimp Artemia sinica
Abstract
1. Introduction
2. Results
2.1. Cloning and Bioinformatic Analysis of As–RAD9
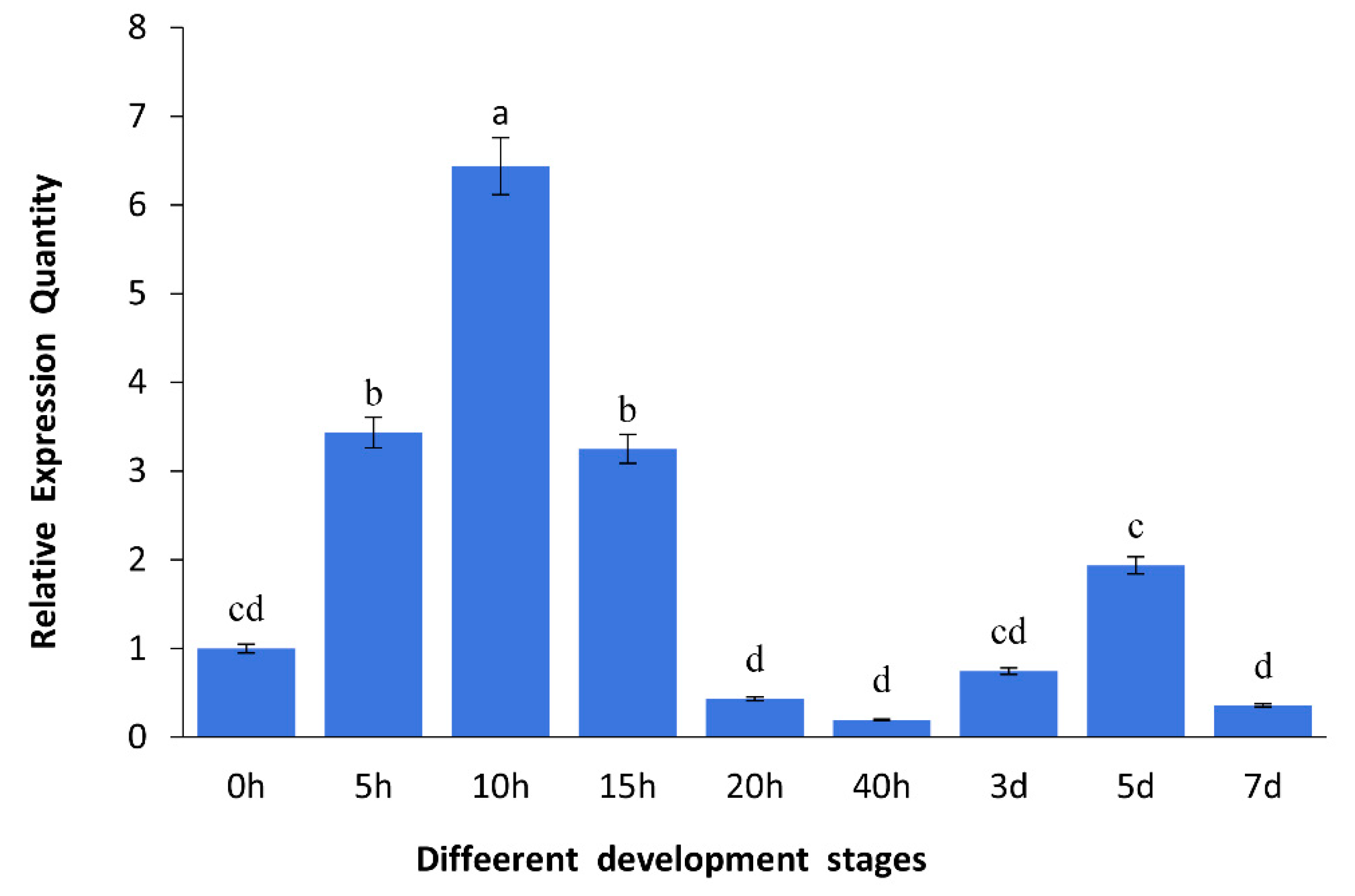
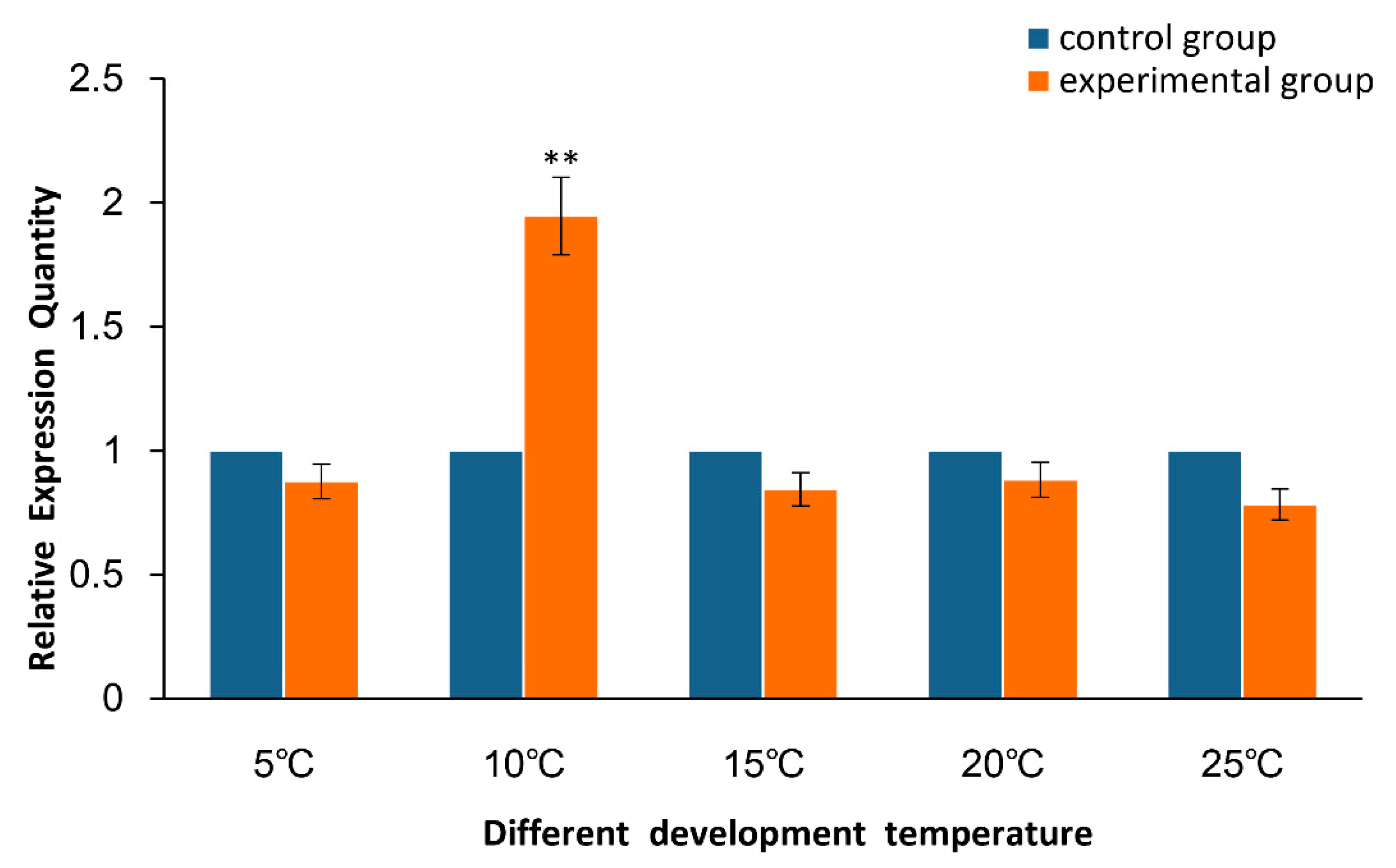
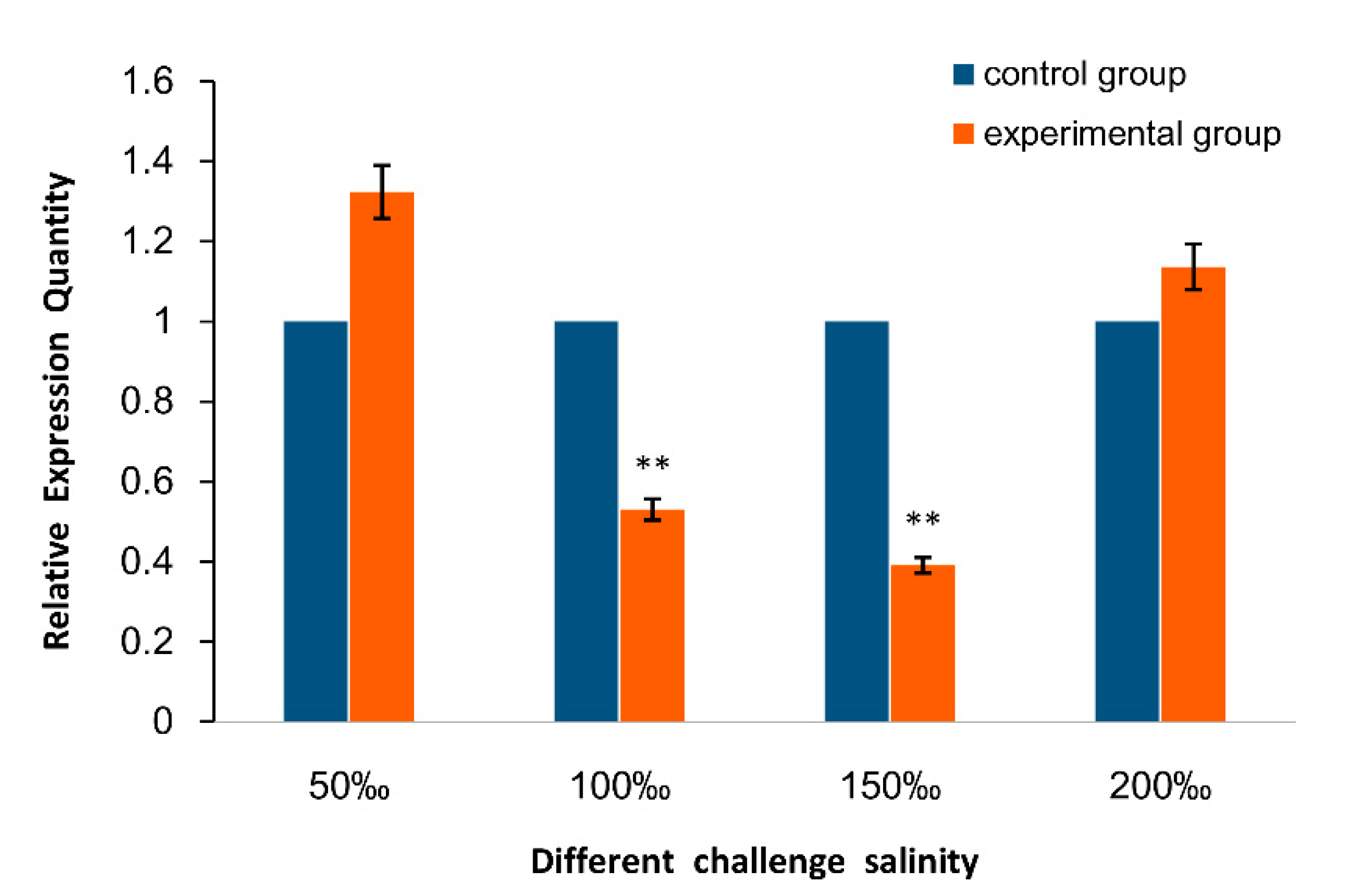
2.2. Expression of As–RAD9 by qPCR
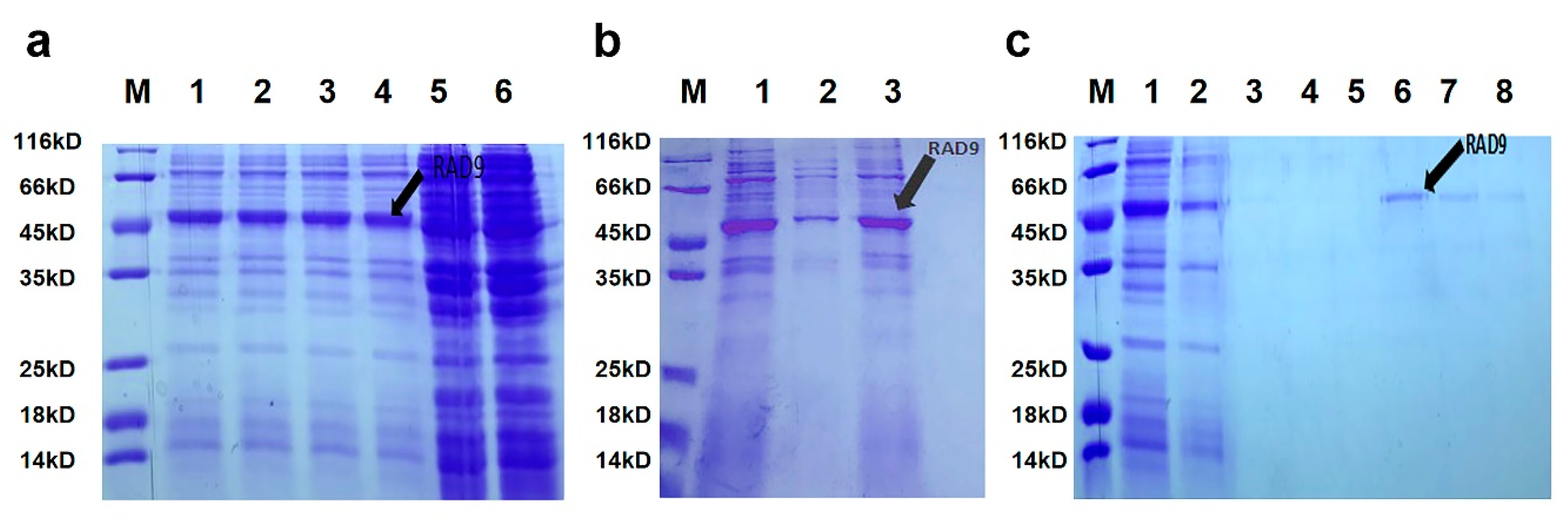
2.3. Purification and Expression of As–RAD9 Protein
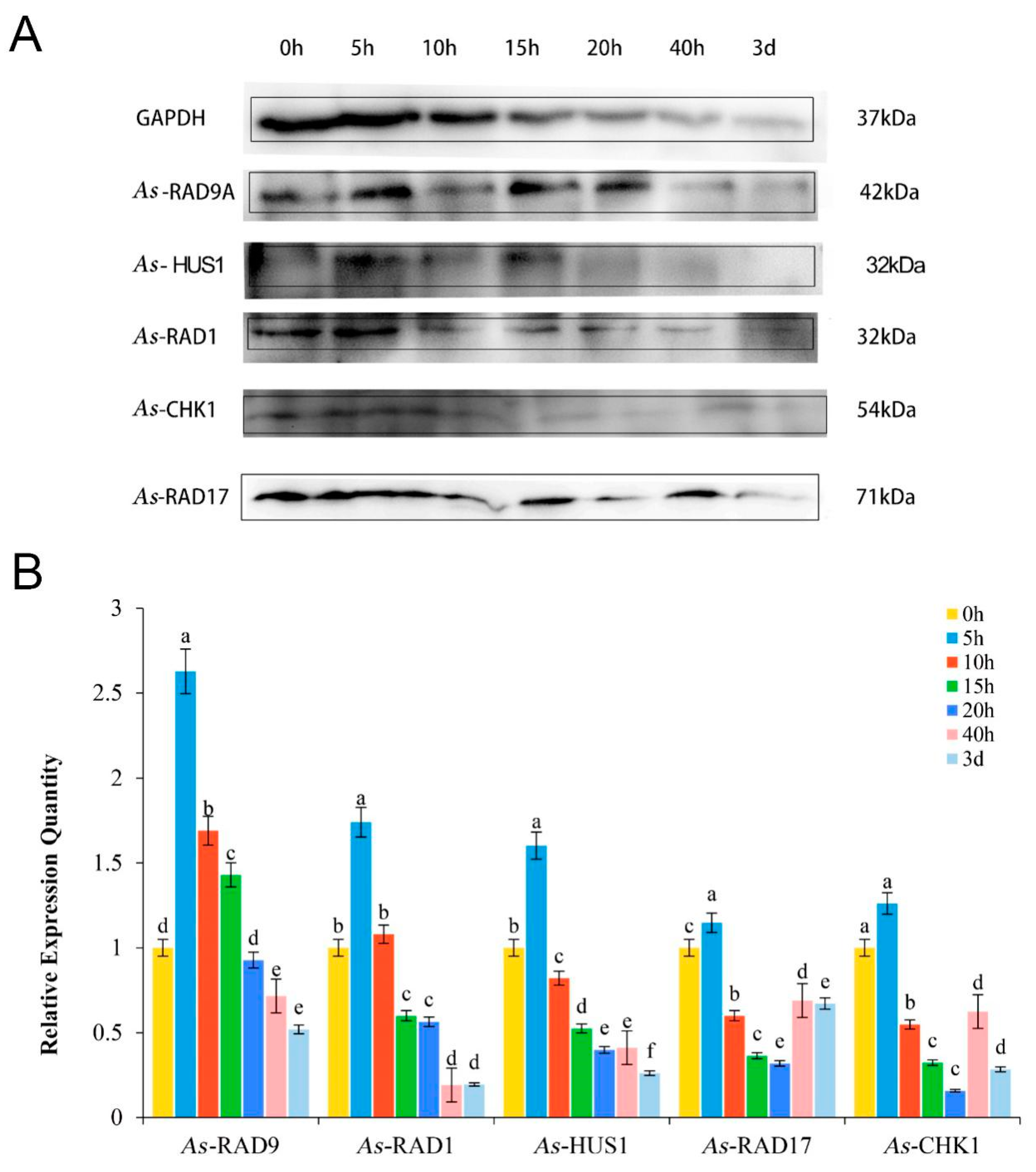
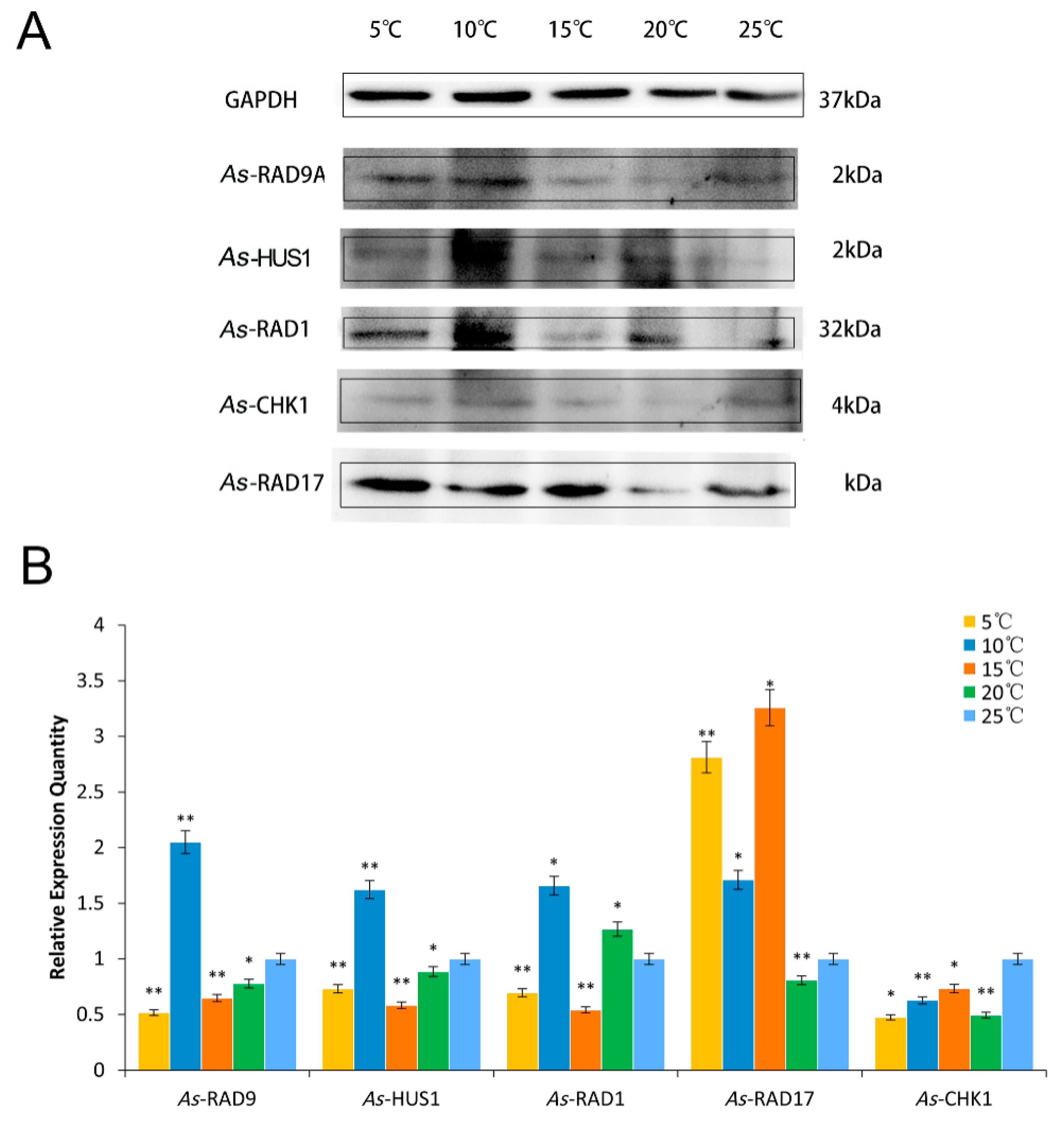
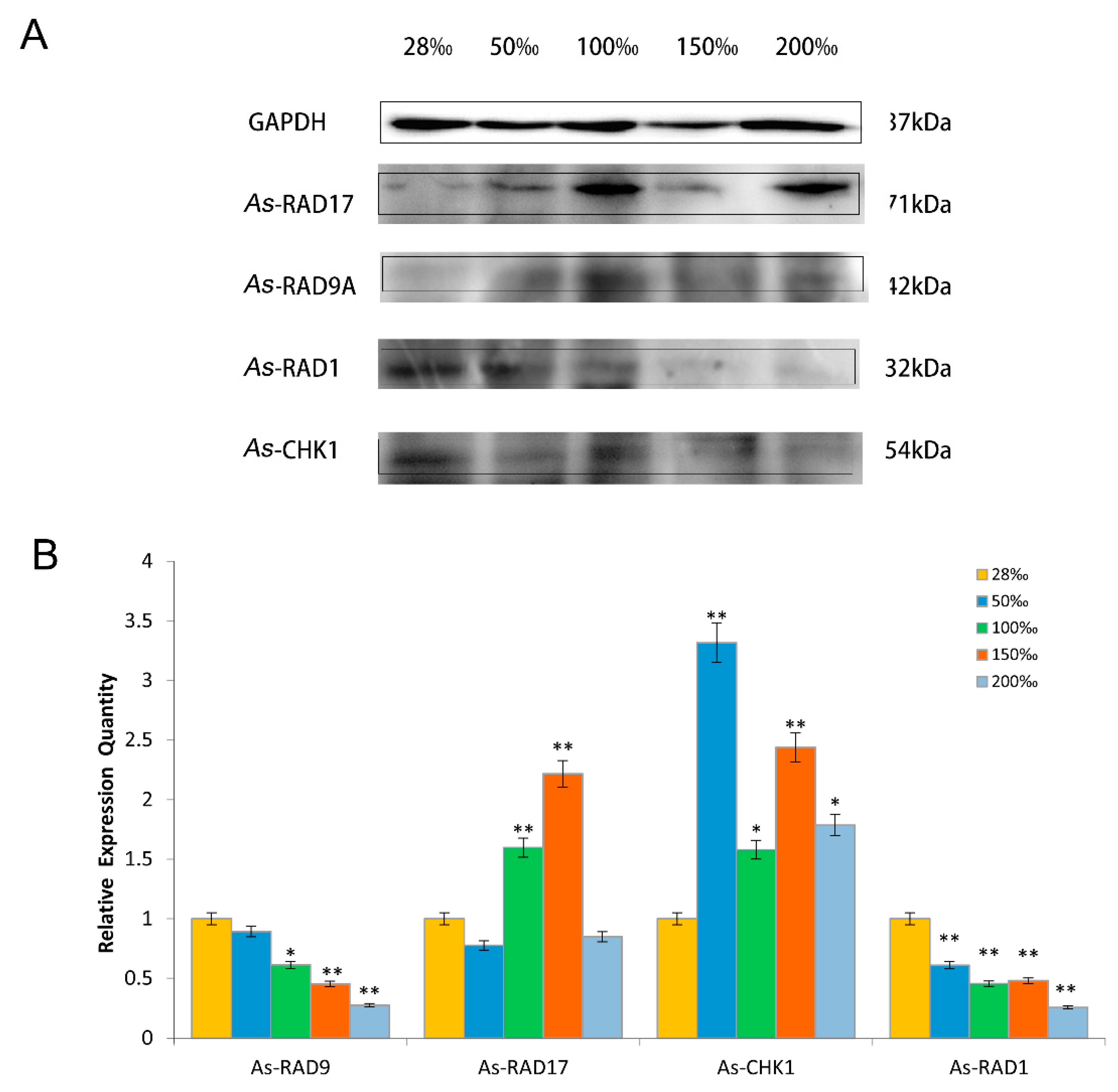
2.4. Expression Pattern of As–RAD9 Protein
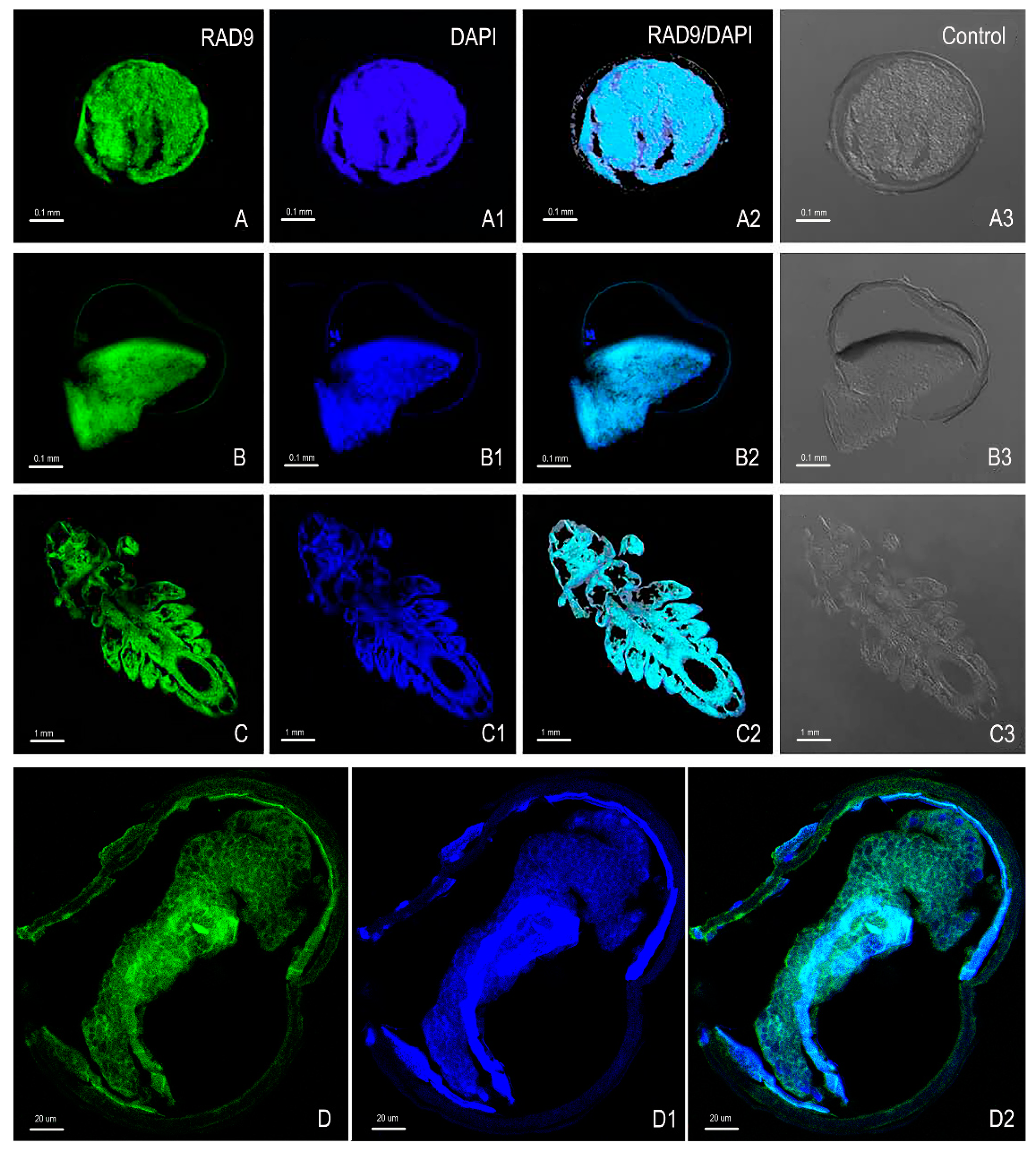
2.5. Immunofluorescence Analysis of As–RAD9
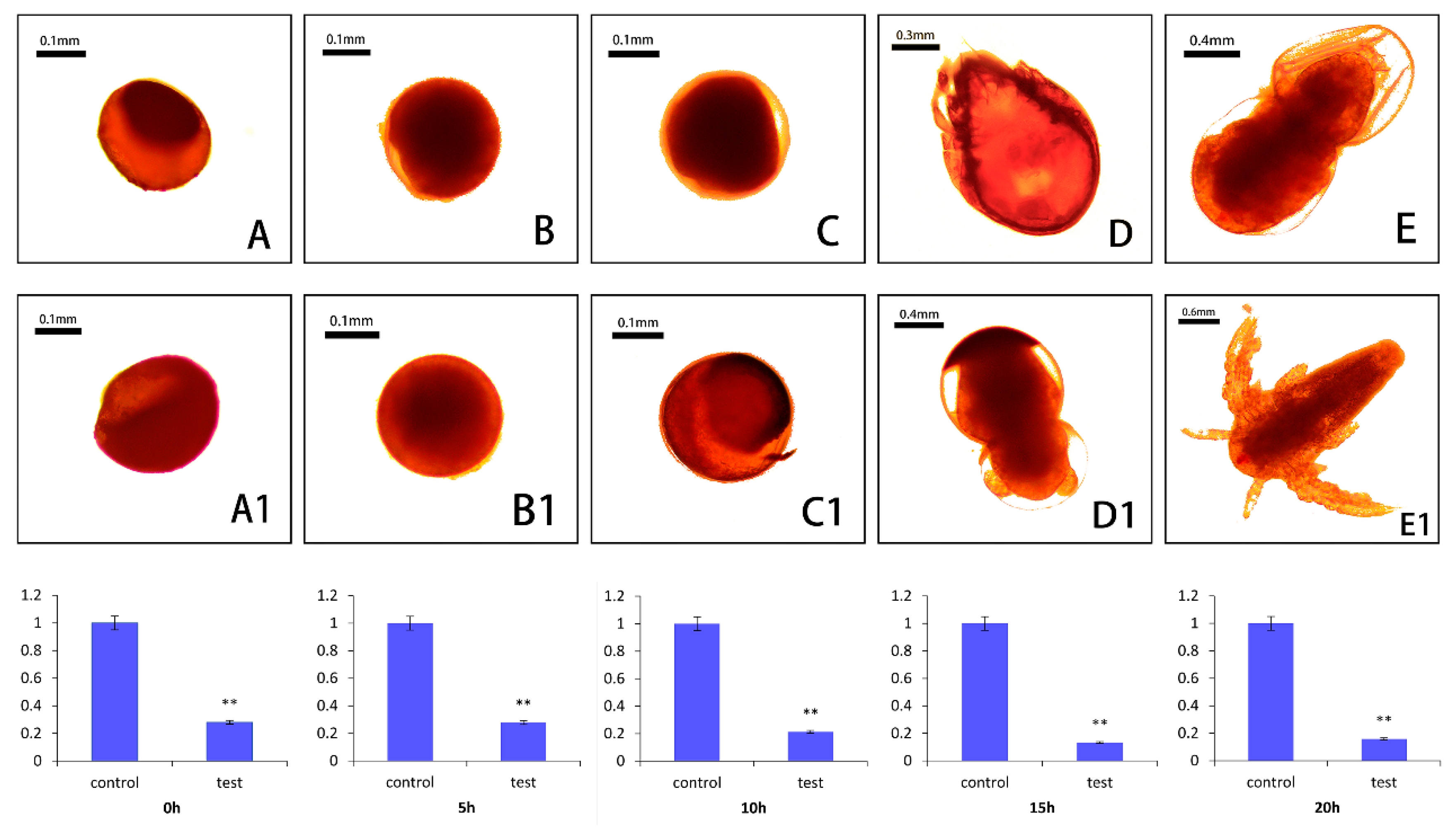
2.6. Small RNA Interference of As–RAD9
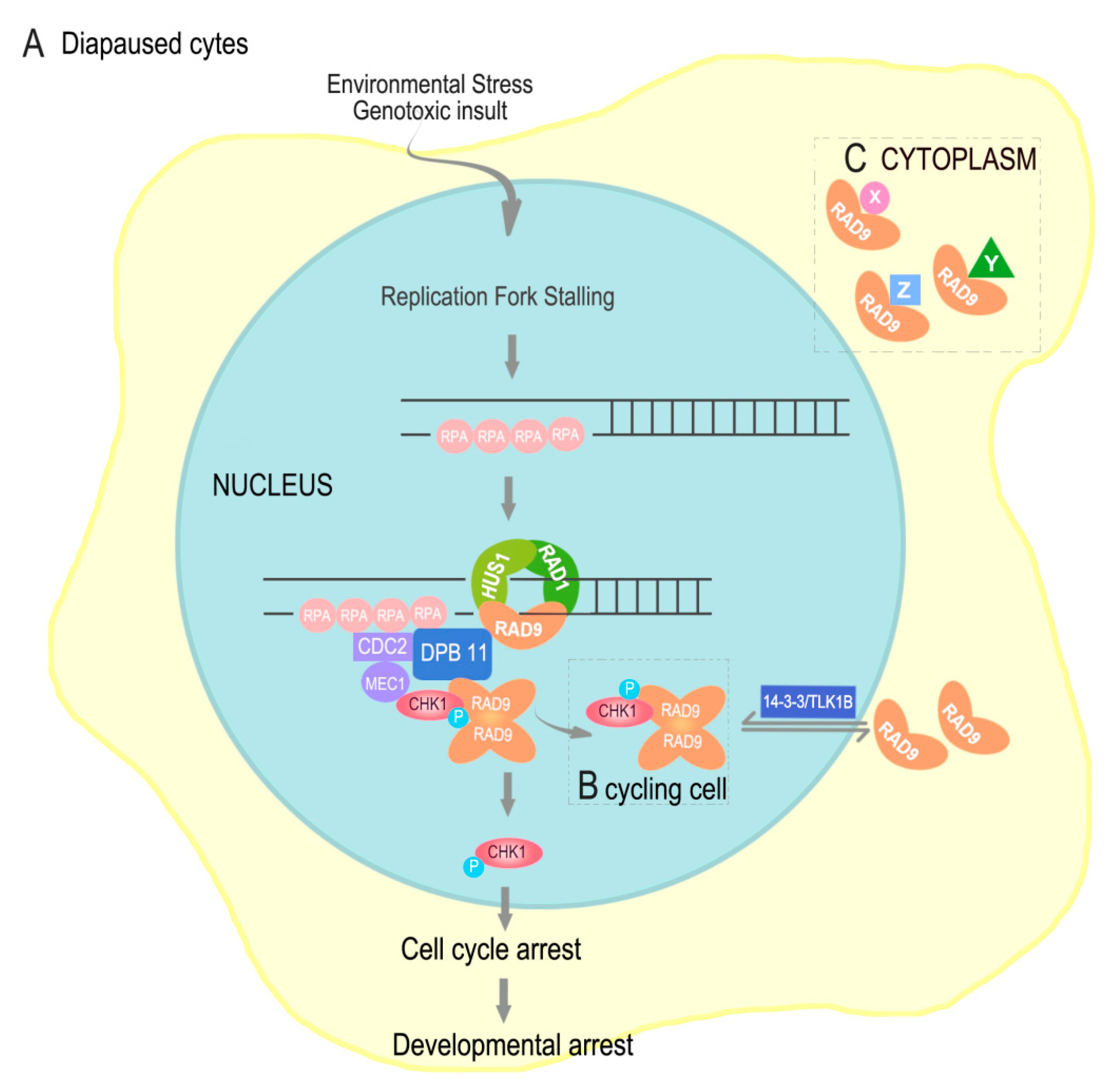
3. Discussion
4. Materials and Methods
4.1. Preparation of Animal
4.2. Cloning the Full-Length cDNA of As–RAD9
4.3. Bioinformatic and Biostatistics Analyses
4.4. Expression Pattern of As–RAD9 by qPCR
4.4.1. Expression of As–RAD9 in Early Embryo Development
4.4.2. Temperature and Salinity Stress Assays
4.5. Purification of Recombinant As–RAD9 Protein
4.6. Production of Polyclonal Antibodies
4.7. Western Blotting
4.8. Immunofluorescence (IF)
4.9. RNA Interference
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| As-rad9 | rad9 mRNA from Artemia sinica |
| As-RAD9 | RAD9 Cell cycle checkpoint control protein from Artemia sinica |
| As-RAD17 | RAD17 checkpoint clamp loader component protein from Artemia sinica |
| As-RAD1 | RAD1 checkpoint DNA exonuclease protein from Artemia sinica |
| As-CHK1 | serine/threonine protein kinase CHK1 from Artemia sinica |
| UTR | untranslated region |
| ATR | ataxia- telangiectasia |
| TOPBP1 | DNA topoisomerase II binding protein 1 |
| ATRIP | ATR interacting protein. |
| Dpb11 | DNA replication regulator DPB11 |
References
- Qin, T.; Zhao, X.; Luan, H.; Ba, H.; Yang, L.; Li, Z.; Hou, L.; Zou, X. Identification, expression pattern and functional characterization of As-MyD88 in bacteria challenge and during different developmental stages of Artemia sinica. Dev. Comp. Immunol. 2015, 50, 9–18. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Yao, F.; Qin, T.; Hou, L.; Zou, X. Identification, expression pattern and functional characterization of As-kip2 in diapause embryo restarting process of Artemia sinica. Gene 2017, 608, 28–40. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Yao, F.; Zhang, H.; Li, N.; Zou, X.; Sui, L.; Hou, L. The Potential Roles of the Apoptosis-Related Protein PDRG1 in Diapause Embryo Restarting of Artemia sinica. Int. J. Mol. Sci. 2018, 19, 126. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yao, F.; Liang, X.; Zhu, X.; Zheng, R.; Jia, B.; Hou, L.; Zou, X. Cloning and expression of retinoblastoma-binding protein 4 gene in embryo diapause termination and in response to salinity stress from brine shrimp Artemia sinica. Gene 2016, 591, 351–361. [Google Scholar] [CrossRef] [PubMed]
- MacRae, T.H. Stress tolerance during diapause and quiescence of the brine shrimp, Artemia. Cell Stress Chaperones 2016, 21, 9–18. [Google Scholar] [CrossRef]
- Yasutis, K.M.; Kozminski, K.G. Cell cycle checkpoint regulators reach a zillion. Cell Cycle 2013, 12, 1501–1509. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Siefert, J.C.; Clowdus, E.A.; Sansam, C.L. Cell cycle control in the early embryonic development of aquatic animal species. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 2015, 178, 8–15. [Google Scholar] [CrossRef]
- Wang, G.; Tong, X.; Weng, S.; Zhou, H. Multiple phosphorylation of Rad9 by CDK is required for DNA damage checkpoint activation. Cell Cycle 2012, 11, 3792–3800. [Google Scholar] [CrossRef]
- Siede, W.; Friedberg, A.S.; Friedberg, E.C. RAD9-dependent G1 arrest defines a second checkpoint for damaged DNA in the cell cycle of Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 1993, 90, 7985–7989. [Google Scholar] [CrossRef]
- Hopkins, K.M.; Auerbach, W.; Wang, X.Y.; Hande, M.P.; Hang, H.; Wolgemuth, D.J.; Joyner, A.L.; Lieberman, H.B. Deletion of mouse rad9 causes abnormal cellular responses to DNA damage, genomic instability, and embryonic lethality. Mol. Cell. Biol. 2004, 24, 7235–7248. [Google Scholar] [CrossRef]
- Broustas, C.G.; Lieberman, H.B. Contributions of Rad9 to tumorigenesis. J. Cell. Biochem. 2012, 113, 742–751. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lieberman, H.B.; Bernstock, J.D.; Broustas, C.G.; Hopkins, K.M.; Leloup, C.; Zhu, A. The role of RAD9 in tumorigenesis. J. Mol. Cell Biol. 2011, 3, 39–43. [Google Scholar] [CrossRef] [PubMed]
- Cotta-Ramusino, C.; McDonald, E.R., 3rd; Hurov, K.; Sowa, M.E.; Harper, J.W.; Elledge, S.J. A DNA damage response screen identifies RHINO, a 9-1-1 and TopBP1 interacting protein required for ATR signaling. Science 2011, 332, 1313–1317. [Google Scholar] [CrossRef] [PubMed]
- Lindsey-Boltz, L.A.; Kemp, M.G.; Capp, C.; Sancar, A. RHINO forms a stoichiometric complex with the 9-1-1 checkpoint clamp and mediates ATR-Chk1 signaling. Cell Cycle 2015, 14, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Delacroix, S.; Wagner, J.M.; Kobayashi, M.; Yamamoto, K.; Karnitz, L.M. The Rad9-Hus1-Rad1 (9-1-1) clamp activates checkpoint signaling via TopBP1. Genes Dev. 2007, 21, 1472–1477. [Google Scholar] [CrossRef] [PubMed]
- Burrows, A.E.; Elledge, S.J. How ATR turns on: TopBP1 goes on ATRIP with ATR. Genes Dev. 2008, 22, 1416–1421. [Google Scholar] [CrossRef] [PubMed]
- Perry, J.A.; Kornbluth, S. Cdc25 and Wee1: Analogous opposites? Cell Div. 2007, 2, 12. [Google Scholar] [CrossRef] [PubMed]
- Ueda, S.; Takeishi, Y.; Ohashi, E.; Tsurimoto, T. Two serine phosphorylation sites in the C-terminus of Rad9 are critical for 9-1-1 binding to TopBP1 and activation of the DNA damage checkpoint response in HeLa cells. Genes Cells 2012, 17, 807–816. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef] [PubMed]
- Dang, T.; Bao, S.; Wang, X.F. Human Rad9 is required for the activation of S-phase checkpoint and the maintenance of chromosomal stability. Genes Cells 2005, 10, 287–295. [Google Scholar] [CrossRef]
- Wysocki, R.; Javaheri, A.; Allard, S.; Sha, F.; Cote, J.; Kron, S.J. Role of Dot1-dependent histone H3 methylation in G1 and S phase DNA damage checkpoint functions of Rad9. Mol. Cell Biol. 2005, 25, 8430–8443. [Google Scholar] [CrossRef] [PubMed]
- Kelly, R.; Davey, S.K. Tousled-like kinase-dependent phosphorylation of Rad9 plays a role in cell cycle progression and G2/M checkpoint exit. PLoS ONE 2013, 8, e85859. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Bai, L.; Gong, Y.; Xie, W.; Hang, H.; Jiang, T. Structure and functional implications of the human rad9-hus1-rad1 cell cycle checkpoint complex. J. Biol. Chem. 2009, 284, 20457–20461. [Google Scholar] [CrossRef] [PubMed]
- Sohn, S.Y.; Cho, Y. Crystal structure of the human rad9-hus1-rad1 clamp. J. Mol. Biol. 2009, 390, 490–502. [Google Scholar] [CrossRef] [PubMed]
- Awate, S.; De Benedetti, A. TLK1B mediated phosphorylation of Rad9 regulates its nuclear/cytoplasmic localization and cell cycle checkpoint. BMC Mol. Biol. 2016, 17, 3. [Google Scholar] [CrossRef] [PubMed]
- Hustedt, N.; Gasser, S.M.; Shimada, K. Replication checkpoint: Tuning and coordination of replication forks in s phase. Genes 2013, 4, 388–434. [Google Scholar] [CrossRef]
- Hand, S.C.; Denlinger, D.L.; Podrabsky, J.E.; Roy, R. Mechanisms of animal diapause: Recent developments from nematodes, crustaceans, insects, and fish. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2016, 310, 1193–1211. [Google Scholar] [CrossRef]
- Li, N.; Yao, F.; Huang, H.; Zhang, H.; Zhang, W.; Zou, X.; Sui, L.; Hou, L. The potential role of Annexin 3 in diapause embryo restart of Artemia sinica and in response to stress of low temperature. Mol. Reprod. Dev. 2019, 86, 530–542. [Google Scholar] [CrossRef]
- Zhao, W.; Yao, F.; Zhang, M.; Jing, T.; Zhang, S.; Hou, L.; Zou, X. The Potential Roles of the G1LEA and G3LEA Proteins in Early Embryo Development and in Response to Low Temperature and High Salinity in Artemia sinica. PLoS ONE 2016, 11, e0162272. [Google Scholar] [CrossRef]
- Xu, X.; Shi, R.; Zheng, L.; Guo, Z.; Wang, L.; Zhou, M.; Zhao, Y.; Tian, B.; Truong, K.; Chen, Y.; et al. SUMO-1 modification of FEN1 facilitates its interaction with Rad9-Rad1-Hus1 to counteract DNA replication stress. J. Mol. Cell Biol. 2018, 10, 460–474. [Google Scholar] [CrossRef]
- Liu, S.; Song, N.; Zou, L. The conserved C terminus of Claspin interacts with Rad9 and promotes rapid activation of Chk1. Cell Cycle 2012, 11, 2711–2716. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zhang, G.; Ma, F.; Li, L.; Li, J.; Li, P.; Zeng, S.; Sun, H.; Li, E. Palbociclib triggers apoptosis in bladder cancer cells by Cdk2-induced Rad9-mediated reorganization of the Bak.Bcl-xl complex. Biochem. Pharmacol. 2019, 163, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Lieberman, H.B.; Panigrahi, S.K.; Hopkins, K.M.; Wang, L.; Broustas, C.G. p53 and RAD9, the DNA Damage Response, and Regulation of Transcription Networks. Radiat. Res. 2017, 187, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Ghandhi, S.A.; Ponnaiya, B.; Panigrahi, S.K.; Hopkins, K.M.; Cui, Q.; Hei, T.K.; Amundson, S.A.; Lieberman, H.B. RAD9 deficiency enhances radiation induced bystander DNA damage and transcriptomal response. Radiat. Oncol. 2014, 9, 206. [Google Scholar] [CrossRef] [PubMed]
- Damasceno, J.D.; Obonaga, R.; Santos, E.V.; Scott, A.; McCulloch, R.; Tosi, L.R. Functional compartmentalization of Rad9 and Hus1 reveals diverse assembly of the 9-1-1 complex components during the DNA damage response in Leishmania. Mol. Microbiol. 2016, 101, 1054–1068. [Google Scholar] [CrossRef] [PubMed]
- Abreu, C.M.; Kumar, R.; Hamilton, D.; Dawdy, A.W.; Creavin, K.; Eivers, S.; Finn, K.; Balsbaugh, J.L.; O’Connor, R.; Kiely, P.A.; et al. Site-specific phosphorylation of the DNA damage response mediator rad9 by cyclin-dependent kinases regulates activation of checkpoint kinase 1. PLoS Genet. 2013, 9, e1003310. [Google Scholar] [CrossRef]
- Zhang, S.; Yao, F.; Jing, T.; Zhang, M.; Zhao, W.; Zou, X.; Sui, L.; Hou, L. Cloning, expression pattern, and potential role of apoptosis inhibitor 5 in the termination of embryonic diapause and early embryo development of Artemia sinica. Gene 2017, 628, 170–179. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, E.W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol 1990, 215, 403–410. [Google Scholar] [CrossRef]
- Rombel, I.T.; Sykes, K.F.; Rayner, S.; Johnston, S.A. ORF-FINDER: A vector for high-throughput gene identification. Gene 2002, 282, 33–41. [Google Scholar] [CrossRef]
- Sigrist, C.J.; Cerutti, L.; Hulo, N.; Gattiker, A.; Falquet, L.; Pagni, M.; Bairoch, A.; Bucher, P. PROSITE: A documented database using patterns and profiles as motif descriptors. Brief. Bioinform. 2002, 3, 265–274. [Google Scholar] [CrossRef]
- Letunic, I.; Bork, P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018, 46, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R.; Gasteiger, E.; Bairoch, A.; Sanchez, J.C.; Williams, K.L.; Appel, R.D.; Hochstrasser, D.F. Protein identification and analysis tools in the ExPASy server. Methods Mol. Biol. 1999, 112, 531–552. [Google Scholar] [PubMed]
- Bannai, H.; Tamada, Y.; Maruyama, O.; Nakai, K.; Miyano, S. Extensive feature detection of N-terminal protein sorting signals. Bioinformatics 2002, 18, 298–305. [Google Scholar] [CrossRef] [PubMed]
- Hofmann, K.; Stoffel, W. TMbase-A database of membrane spanning proteins segments. Biol. Chem. Hoppe Seyler 1993, 374, 166. [Google Scholar]
- Zhang, M.; Yao, F.; Luan, H.; Zhao, W.; Jing, T.; Zhang, S.; Hou, L.; Zou, X. APC/C(CDC20) and APC/C play pivotal roles in the process of embryonic development in Artemia sinica. Sci. Rep. 2016, 6, 39047. [Google Scholar] [CrossRef] [PubMed]
| Name | Position | Context a | Score b | Name | Position | Context a | Score b |
|---|---|---|---|---|---|---|---|
| Ser | 2 | ---ASHLPY | 0.002 | Thr | 27 | GLLQTATKQ | 0.025 |
| 41 | PVVESKWRT | 0.350 | 29 | LQTATKQAE | 0.038 | ||
| 59 | WNFISGIQY | 0.004 | 45 | SKWRTLEAF | 0.100 | ||
| 68 | LAGLSTLPG | 0.007 | 69 | AGLSTLPGN | 0.028 | ||
| 78 | PAIASLMAF | 0.003 | 83 | LMAFTASIT | 0.067 | ||
| 85 | AFTASITSP | 0.0079 | 87 | TASITSPLT | 0.046 | ||
| 88 | ASITSPLTT | 0.662 | 91 | TSPLTTQHT | 0.096 | ||
| 113 | LAPPSAASA | 0.032 | 92 | SPLTTQHTL | 0.192 | ||
| 116 | PSAASAFVG | 0.114 | 95 | TTQHTLLFN | 0.005 | ||
| 130 | AAVGSIGLG | 0.030 | 165 | EMPSTEDLV | 0.547 | ||
| 159 | FKVMSGEMP | 0.956 | 221 | HVSPTHYVP | 0.212 | ||
| 164 | GEMPSTEDL | 0.996 | 234 | AARVTQILS | 0.174 | ||
| 177 | PAILSPGAL | 0.069 | 241 | LSSLTITQL | 0.037 | ||
| 213 | IAFASRGNH | 0.046 | 243 | SLTITQLLK | 0.015 | ||
| 219 | GNHVSPTHY | 0.294 | 259 | EDCSTPC-- | 0.382 | ||
| 227 | YVPESDAAA | 0.305 | Tyr | 6 | SHLPYIEQG | 0.683 | |
| 238 | TQILSSLTI | 0.003 | 63 | SGIQYLAGL | 0.127 | ||
| 239 | QILSSLTIT | 0.157 | 144 | ILAGYGAGV | 0.008 | ||
| 258 | NEDCSTPC- | 0.116 | 223 | SPTHYVPES | 0.714 |
| Primer | Sequence(5’–3’) |
|---|---|
| RAD9F | TGGTGATTACATTTACTTTG |
| RAD9R | CGGCACATCAACTACATCAC |
| 3’RAD9 | AGATTTGGCGTTGTGAGGTCTTAC |
| 5’RAD9 | GGAAGTAGAGCGGAAAACAGTCAG |
| RT-rad9F | CTAACCCGAAGTTGGATGCTCT |
| RT-rad9R | CAGATGGACTTGTTTGCTCGC |
| β-actinF | GTGTGACGATGATGTTGCGG |
| β-actinR | GCTGTCCTTTTGACCCATTCC |
| ORF-rad9F | CCGGAATTCATGGGGAGCGCAAGAATTTT |
| ORF-rad9R | ACGCGTCGACTTAATCTTCATCTGAATCAA |
| SiRNAA1 | CCTACGAGCAAGAAACAAT |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, H.; Chen, C.; Yao, F.; Li, X.; Wang, Y.; Shao, Y.; Wang, X.; Zhang, X.; Jiang, T.; Hou, L. Investigation of the Possible Role of RAD9 in Post-Diapaused Embryonic Development of the Brine Shrimp Artemia sinica. Genes 2019, 10, 768. https://doi.org/10.3390/genes10100768
Huang H, Chen C, Yao F, Li X, Wang Y, Shao Y, Wang X, Zhang X, Jiang T, Hou L. Investigation of the Possible Role of RAD9 in Post-Diapaused Embryonic Development of the Brine Shrimp Artemia sinica. Genes. 2019; 10(10):768. https://doi.org/10.3390/genes10100768
Chicago/Turabian StyleHuang, Huifang, Ce Chen, Feng Yao, Xiuling Li, Yanan Wang, Yuting Shao, Xinyao Wang, Xingzheng Zhang, Tao Jiang, and Lin Hou. 2019. "Investigation of the Possible Role of RAD9 in Post-Diapaused Embryonic Development of the Brine Shrimp Artemia sinica" Genes 10, no. 10: 768. https://doi.org/10.3390/genes10100768
APA StyleHuang, H., Chen, C., Yao, F., Li, X., Wang, Y., Shao, Y., Wang, X., Zhang, X., Jiang, T., & Hou, L. (2019). Investigation of the Possible Role of RAD9 in Post-Diapaused Embryonic Development of the Brine Shrimp Artemia sinica. Genes, 10(10), 768. https://doi.org/10.3390/genes10100768




