Differential Gene Expression in Articular Cartilage and Subchondral Bone of Neonatal and Adult Horses
Abstract
1. Introduction
2. Materials and Methods
2.1. Tissue Collection and RNA Extraction
2.2. RNA Sequencing
2.3. Data Analysis
2.3.1. Alignment and Gene-Level Quantification
2.3.2. Filtering, Clustering and Surrogate Variable Analysis
2.3.3. Differential Expression Testing
2.3.4. Functional Annotation and Pathway Analysis
3. Results
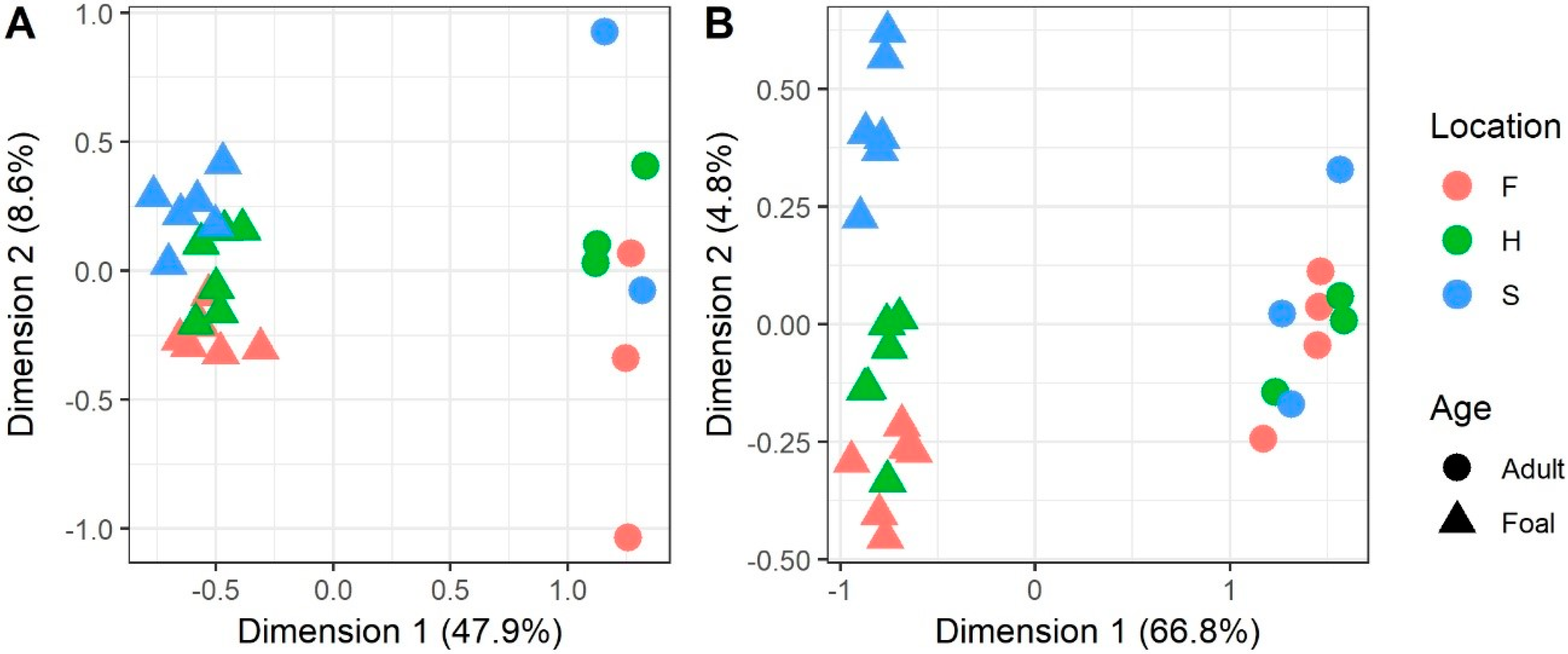
3.1. MDS Clustering
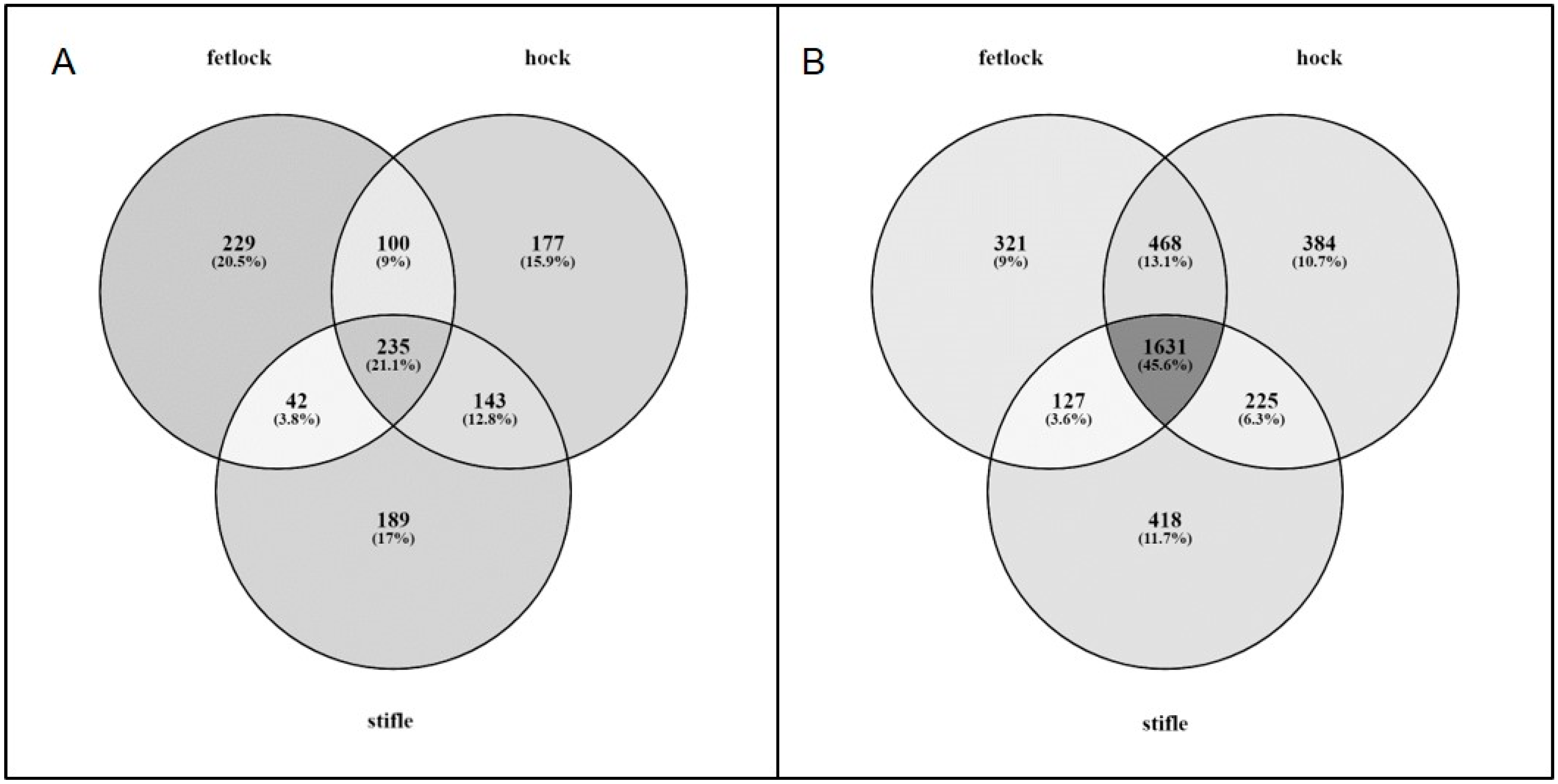
3.2. Differential Expression
3.2.1. Articular Cartilage
3.2.2. Subchondral Bone
3.3. Functional Annotation and Pathway Analysis
3.3.1. Articular Cartilage
3.3.2. Subchondral Bone
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
Appendix A.1. Quality Control
Appendix A.2. Alignment and Gene-Level Quantification
Appendix A.3. Normalization and Filtering
Appendix A.4. Clustering
Appendix A.5. Differential Expression Testing
References
- Karsenty, G. Transcriptional control of skeletogenesis. Annu. Rev. Genom. Hum. Genet. 2008, 9, 183–196. [Google Scholar] [CrossRef] [PubMed]
- Smeeton, J.; Askary, A.; Crump, J.G. Building and maintaining joints by exquisite local control of cell fate. Wiley Interdiscip. Rev. Dev. Biol. 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Mackie, E.J.; Tatarczuch, L.; Mirams, M. The skeleton: A multi-functional complex organ: The growth plate chondrocyte and endochondral ossification. J. Endocrinol. 2011, 211, 109–121. [Google Scholar] [CrossRef] [PubMed]
- Fretz, P.B.; Cymbaluk, N.F.; Pharr, J.W. Quantitative analysis of long-bone growth in the horse. Am. J. Vet. Res. 1984, 45, 1602–1609. [Google Scholar] [PubMed]
- Lawrence, L.A.; Ott, E.A.; Miller, G.J.; Poulos, P.W.; Piotrowski, G.; Asquith, R.L. The mechanical properties of equine third metacarpals as affected by age. J. Anim. Sci. 1994, 72, 2617–2623. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, S.; Furukawa, S.; Kitada, S.; Mori, M.; Saito, T.; Kawakami, K.; Belmonte, J.C.I.; Kawakami, Y.; Ito, Y.; Sato, T.; et al. Analysis of transcription factors expressed at the anterior mouse limb bud. PLoS ONE 2017, 12, e0175673. [Google Scholar] [CrossRef] [PubMed]
- Sensiate, L.A.; Marques-Souza, H. Bone growth as the main determinant of mouse digit tip regeneration after amputation. Sci. Rep. 2019, 9, 9720. [Google Scholar] [CrossRef] [PubMed]
- Hayes, A.J.; Mitchell, R.E.; Bashford, A.; Reynolds, S.; Caterson, B.; Hammond, C.L. Expression of glycosaminoglycan epitopes during zebrafish skeletogenesis. Dev. Dyn. 2013, 242, 778–789. [Google Scholar] [CrossRef]
- Enomoto-Iwamoto, M.; Kitagaki, J.; Koyama, E.; Tamamura, Y.; Wu, C.; Kanatani, N.; Koike, T.; Okada, H.; Komori, T.; Yoneda, T.; et al. The Wnt antagonist Frzb-1 regulates chondrocyte maturation and long bone development during limb skeletogenesis. Dev. Biol. 2002, 251, 142–156. [Google Scholar] [CrossRef]
- Hutchison, C.; Pilote, M.; Roy, S. The axolotl limb: A model for bone development, regeneration and fracture healing. Bone 2007, 40, 45–56. [Google Scholar] [CrossRef]
- Gao, J.; Li, X.; Zhang, Y.; Wang, H. Endochondral ossification in hindlimbs during bufo gargarizans metamorphosis: A model of studying skeletal development in vertebrates. Dev. Dyn. 2018, 247, 1121–1134. [Google Scholar] [CrossRef] [PubMed]
- Giffin, J.L.; Gaitor, D.; Franz-Odendaal, T.A. The Forgotten Skeletogenic Condensations: A Comparison of Early Skeletal Development Amongst Vertebrates. J. Dev. Biol. 2019, 7, 4. [Google Scholar] [CrossRef] [PubMed]
- Cosden, R.S.; Lattermann, C.; Romine, S.; Gao, J.; Voss, S.R.; MacLeod, J.N. Intrinsic repair of full-thickness articular cartilage defects in the axolotl salamander. Osteoarthr. Cartil. 2011, 19, 200–205. [Google Scholar] [CrossRef] [PubMed]
- Lydon, H.; Getgood, A.; Henson, F.M.D. Healing of Osteochondral Defects via Endochondral Ossification in an Ovine Model. Cartilage 2019, 10, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Witt, F.; Petersen, A.; Seidel, R.; Vetter, A.; Weinkamer, R.; Duda, G.N. Combined in vivo/in silico study of mechanobiological mechanisms during endochondral ossification in bone healing. Ann. Biomed. Eng. 2011, 39, 2531–2541. [Google Scholar] [CrossRef] [PubMed]
- McIlwraith, C.W.; Association, A.Q.H. Summary of panel findings. In Proceedings Panel on Developmental Orthopedic Disease, AQHA Developmental Orthopedic Symposium; McIlwraith, C.W., Ed.; American Quarter Horse Association: Amarillo, TX, USA, 1986; pp. 55–61. [Google Scholar]
- Lepeule, J.; Bareille, N.; Robert, C.; Ezanno, P.; Valette, J.P.; Jacquet, S.; Blanchard, G.; Denoix, J.M.; Seegers, H. Association of growth, feeding practices and exercise conditions with the prevalence of Developmental Orthopaedic Disease in limbs of French foals at weaning. Prev. Vet. Med. 2009, 89, 167–177. [Google Scholar] [CrossRef] [PubMed]
- Gabel, A.A.; Knight, D.A.; Reed, S.M. Comparison of incidence and severity of developmental orthopedic disease on 17 farms before and after adjustment of ration. Am. Assoc. Equine Pract. Proc. 1987, 33, 163–170. [Google Scholar]
- Philipsson, J.; Andreasson, E.; Sandgren, B.; Dalin, G.; Carlsten, J. Osteochondrosis in the tarsocrural joint and osteochondral fragments in the fetlock joints in Standardbred trotters. II. Heritability. Equine Vet. J. Suppl. 1993, 16, 38–41. [Google Scholar] [CrossRef]
- Van Grevenhof, E.M.; Schurink, A.; Ducro, B.J.; van Weeren, P.R.; Van Tartwijk, J.M.; Bijma, P.; van Arendonk, J.A. Genetic variables of various manifestations of osteochondrosis and their correlations between and within joints in Dutch warmblood horses. J. Anim. Sci. 2009, 87, 1906–1912. [Google Scholar] [CrossRef]
- McCoy, A.M.; Norton, E.M.; Kemper, A.M.; Beeson, S.K.; Mickelson, J.R.; McCue, M.E. SNP-based heritability and genetic architecture of tarsal osteochondrosis in North American Standardbred horses. Anim. Genet. 2019, 50, 78–81. [Google Scholar] [CrossRef]
- Baldridge, D.; Shchelochkov, O.; Kelley, B.; Lee, B. Signaling pathways in human skeletal dysplasias. Annu Rev. Genom. Hum. Genet. 2010, 11, 189–217. [Google Scholar] [CrossRef] [PubMed]
- Lefebvre, V.; Bhattaram, P. Vertebrate skeletogenesis. Curr. Top. Dev. Biol. 2010, 90, 291–317. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 2017, 14, 417–419. [Google Scholar] [CrossRef] [PubMed]
- Soneson, C.; Love, M.I.; Robinson, M.D. Differential analyses for RNA-seq: Transcript-level estimates improve gene-level inferences. F1000Research 2015, 4, 1521. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Lun, A.T.; Smyth, G.K. From reads to genes to pathways: Differential expression analysis of RNA-Seq experiments using Rsubread and the edgeR quasi-likelihood pipeline. F1000Research 2016, 5, 1438. [Google Scholar] [CrossRef] [PubMed]
- Leek, J.T.; Storey, J.D. Capturing heterogeneity in gene expression studies by surrogate variable analysis. PLoS Genet. 2007, 3, 1724–1735. [Google Scholar] [CrossRef] [PubMed]
- Leek, J.T.; Storey, J.D. A general framework for multiple testing dependence. Proc. Natl. Acad. Sci. USA 2008, 105, 18718–18723. [Google Scholar] [CrossRef]
- Law, C.W.; Chen, Y.; Shi, W.; Smyth, G.K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 2014, 15, R29. [Google Scholar] [CrossRef]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.H.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- McCarthy, D.J.; Smyth, G.K. Testing significance relative to a fold-change threshold is a TREAT. Bioinformatics 2009, 25, 765–771. [Google Scholar] [CrossRef] [PubMed]
- Huber, W.; Carey, V.J.; Gentleman, R.; Anders, S.; Carlson, M.; Carvalho, B.S.; Bravo, H.C.; Davis, S.; Gatto, L.; Girke, T.; et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 2015, 12, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Annotationhub: Client to Access Annotationhub Resources, R Package Version 2.16.1; Bioconductor 3.9: Buffalo, NY, USA, 2019.
- KEGGREST: Client-Side REST Access to KEGG, R Package Version 1.24.0; Bioconductor 3.9: Buffalo, NY, USA, 2019.
- Huerta-Cepas, J.; Szklarczyk, D.; Forslund, K.; Cook, H.; Heller, D.; Walter, M.C.; Rattei, T.; Mende, D.R.; Sunagawa, S.; Kuhn, M.; et al. eggNOG 4.5: A hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 2016, 44, D286–D293. [Google Scholar] [CrossRef] [PubMed]
- Thomas, P.D.; Campbell, M.J.; Kejariwal, A.; Mi, H.; Karlak, B.; Daverman, R.; Diemer, K.; Muruganujan, A.; Narechania, A. PANTHER: A library of protein families and subfamilies indexed by function. Genome Res. 2003, 13, 2129–2141. [Google Scholar] [CrossRef] [PubMed]
- Mi, H.; Dong, Q.; Muruganujan, A.; Gaudet, P.; Lewis, S.; Thomas, P.D. PANTHER version 7: Improved phylogenetic trees, orthologs and collaboration with the Gene Ontology Consortium. Nucleic Acids Res. 2010, 38, D204–D210. [Google Scholar] [CrossRef]
- Fabregat, A.; Jupe, S.; Matthews, L.; Sidiropoulos, K.; Gillespie, M.; Garapati, P.; Haw, R.; Jassal, B.; Korninger, F.; May, B.; et al. The Reactome Pathway Knowledgebase. Nucleic Acids Res. 2018, 46, D649–D655. [Google Scholar] [CrossRef]
- Staines, K.A.; Pollard, A.S.; McGonnell, I.M.; Farquharson, C.; Pitsillides, A.A. Cartilage to bone transitions in health and disease. J. Endocrinol. 2013, 219, R1–R12. [Google Scholar] [CrossRef]
- Pan, Z.; Zhang, X.; Shangguan, Y.; Hu, H.; Chen, L.; Wang, H. Suppressed osteoclast differentiation at the chondro-osseous junction mediates endochondral ossification retardation in long bones of Wistar fetal rats with prenatal ethanol exposure. Toxicol. Appl. Pharmacol. 2016, 305, 234–241. [Google Scholar] [CrossRef]
- Adhami, M.D.; Rashid, H.; Chen, H.; Clarke, J.C.; Yang, Y.; Javed, A. Loss of Runx2 in committed osteoblasts impairs postnatal skeletogenesis. J. Bone Miner. Res. 2015, 30, 71–82. [Google Scholar] [CrossRef] [PubMed]
- Glass, D.A., 2nd; Bialek, P.; Ahn, J.D.; Starbuck, M.; Patel, M.S.; Clevers, H.; Taketo, M.M.; Long, F.; McMahon, A.P.; Lang, R.A.; et al. Canonical Wnt signaling in differentiated osteoblasts controls osteoclast differentiation. Dev. Cell 2005, 8, 751–764. [Google Scholar] [CrossRef] [PubMed]
- Green, A.C.; Kocovski, P.; Jovic, T.; Walia, M.K.; Chandraratna, R.A.S.; Martin, T.J.; Baker, E.K.; Purton, L.E. Retinoic acid receptor signalling directly regulates osteoblast and adipocyte differentiation from mesenchymal progenitor cells. Exp. Cell Res. 2017, 350, 284–297. [Google Scholar] [CrossRef] [PubMed]
- Day, T.F.; Guo, X.; Garrett-Beal, L.; Yang, Y. Wnt/β-catenin signaling in mesenchymal progenitors controls osteoblast and chondrocyte differentiation during vertebrate skeletogenesis. Dev. Cell 2005, 8, 739–750. [Google Scholar] [CrossRef] [PubMed]
- Iwamoto, M.; Tamamura, Y.; Koyama, E.; Komori, T.; Takeshita, N.; Williams, J.A.; Nakamura, T.; Enomoto-Iwamoto, M.; Pacifici, M. Transcription factor ERG and joint and articular cartilage formation during mouse limb and spine skeletogenesis. Dev. Biol. 2007, 305, 40–51. [Google Scholar] [CrossRef] [PubMed]
- Teufel, S.; Hartmann, C. Wnt-signaling in skeletal development. Curr. Top. Dev. Biol. 2019, 133, 235–279. [Google Scholar] [CrossRef] [PubMed]
- Marcellini, S.; Henriquez, J.P.; Bertin, A. Control of osteogenesis by the canonical Wnt and BMP pathways in vivo: Cooperation and antagonism between the canonical Wnt and BMP pathways as cells differentiate from osteochondroprogenitors to osteoblasts and osteocytes. Bioessays 2012, 34, 953–962. [Google Scholar] [CrossRef]
- Chen, E.; Liu, G.; Zhou, X.; Zhang, W.; Wang, C.; Hu, D.; Xue, D.; Pan, Z. Concentration-dependent, dual roles of IL-10 in the osteogenesis of human BMSCs via P38/MAPK and NF-kappaB signaling pathways. FASEB J. 2018, 32, 4917–4929. [Google Scholar] [CrossRef]
- Hosseini-Farahabadi, S.; Geetha-Loganathan, P.; Fu, K.; Nimmagadda, S.; Yang, H.J.; Richman, J.M. Dual functions for WNT5A during cartilage development and in disease. Matrix Biol. 2013, 32, 252–264. [Google Scholar] [CrossRef]
- McCoy, A.M.; Toth, F.; Dolvik, N.I.; Ekman, S.; Ellermann, J.; Olstad, K.; Ytrehus, B.; Carlson, C.S. Articular osteochondrosis: A comparison of naturally-occurring human and animal disease. Osteoarthr. Cartil. 2013, 21, 1638–1647. [Google Scholar] [CrossRef]
- Kinsley, M.A.; Semevolos, S.A.; Duesterdieck-Zellmer, K.F. Wnt/β-catenin signaling of cartilage canal and osteochondral junction chondrocytes and full thickness cartilage in early equine osteochondrosis. J. Orthop. Res. 2015, 33, 1433–1438. [Google Scholar] [CrossRef] [PubMed]
- Semevolos, S.A.; Nixon, A.J.; Brower-Toland, B.D. Changes in molecular expression of aggrecan and collagen types I, II, and X, insulin-like growth factor-I, and transforming growth factor-beta1 in articular cartilage obtained from horses with naturally acquired osteochondrosis. Am. J. Vet. Res. 2001, 62, 1088–1094. [Google Scholar] [CrossRef] [PubMed]
- Semevolos, S.A.; Brower-Toland, B.D.; Bent, S.J.; Nixon, A.J. Parathyroid hormone-related peptide and indian hedgehog expression patterns in naturally acquired equine osteochondrosis. J. Orthop. Res. 2002, 20, 1290–1297. [Google Scholar] [CrossRef]
- Henson, F.M.; Schofield, P.N.; Jeffcott, L.B. Expression of transforming growth factor-β 1 in normal and dyschondroplastic articular growth cartilage of the young horse. Equine Vet. J. 1997, 29, 434–439. [Google Scholar] [CrossRef] [PubMed]
- Mirams, M.; Tatarczuch, L.; Ahmed, Y.A.; Pagel, C.N.; Jeffcott, L.B.; Davies, H.M.; Mackie, E.J. Altered gene expression in early osteochondrosis lesions. J. Orthop. Res. 2009, 27, 452–457. [Google Scholar] [CrossRef] [PubMed]
- Mirams, M.; Ayodele, B.A.; Tatarczuch, L.; Henson, F.M.; Pagel, C.N.; Mackie, E.J. Identification of novel osteochondrosis—Associated genes. J. Orthop. Res. 2016, 34, 404–411. [Google Scholar] [CrossRef] [PubMed]
- Austbo, L.; Roed, K.H.; Dolvik, N.I.; Skretting, G. Identification of differentially expressed genes associated with osteochondrosis in standardbred horses using RNA arbitrarily primed PCR. Anim. Biotechnol. 2010, 21, 135–139. [Google Scholar] [CrossRef] [PubMed]
- Serteyn, D.; Piquemal, D.; Vanderheyden, L.; Lejeune, J.P.; Verwilghen, D.; Sandersen, C. Gene expression profiling from leukocytes of horses affected by osteochondrosis. J. Orthop. Res. 2010, 28, 965–970. [Google Scholar] [CrossRef] [PubMed]
- Piepoli, T.; Mennuni, L.; Zerbi, S.; Lanza, M.; Rovati, L.C.; Caselli, G. Glutamate signaling in chondrocytes and the potential involvement of NMDA receptors in cell proliferation and inflammatory gene expression. Osteoarthr. Cartil. 2009, 17, 1076–1083. [Google Scholar] [CrossRef]
- Wang, L.; Hinoi, E.; Takemori, A.; Takarada, T.; Yoneda, Y. Abolition of chondral mineralization by group III metabotropic glutamate receptors expressed in rodent cartilage. Br. J. Pharmacol. 2005, 146, 732–743. [Google Scholar] [CrossRef]
- Wang, L.; Hinoi, E.; Takemori, A.; Yoneda, Y. Release of endogenous glutamate by AMPA receptors expressed in cultured rat costal chondrocytes. Biol. Pharm. Bull. 2005, 28, 990–993. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Grassel, S.G. The role of peripheral nerve fibers and their neurotransmitters in cartilage and bone physiology and pathophysiology. Arthritis Res. Ther. 2014, 16, 485. [Google Scholar] [CrossRef] [PubMed]
- Studle, C.; Occhetta, P.; Geier, F.; Mehrkens, A.; Barbero, A.; Martin, I. Challenges Toward the Identification of Predictive Markers for Human Mesenchymal Stromal Cells Chondrogenic Potential. Stem Cells Transl. Med. 2019, 8, 194–204. [Google Scholar] [CrossRef] [PubMed]
- Wen, Z.H.; Chang, Y.C.; Jean, Y.H. Excitatory amino acid glutamate: Role in peripheral nociceptive transduction and inflammation in experimental and clinical osteoarthritis. Osteoarthr. Cartil. 2015, 23, 2009–2016. [Google Scholar] [CrossRef] [PubMed]
- Xiao, W.; Wang, Y.; Pacios, S.; Li, S.; Graves, D.T. Cellular and Molecular Aspects of Bone Remodeling. Front. Oral Biol. 2016, 18, 9–16. [Google Scholar] [CrossRef] [PubMed]
- Herath, T.D.K.; Larbi, A.; Teoh, S.H.; Kirkpatrick, C.J.; Goh, B.T. Neutrophil-mediated enhancement of angiogenesis and osteogenesis in a novel triple cell co-culture model with endothelial cells and osteoblasts. J. Tissue Eng. Regen. Med. 2018, 12, e1221–e1236. [Google Scholar] [CrossRef] [PubMed]
- Ehrnthaller, C.; Huber-Lang, M.; Nilsson, P.; Bindl, R.; Redeker, S.; Recknagel, S.; Rapp, A.; Mollnes, T.; Amling, M.; Gebhard, F.; et al. Complement C3 and C5 deficiency affects fracture healing. PLoS ONE 2013, 8, e81341. [Google Scholar] [CrossRef] [PubMed]
- Huber-Lang, M.; Kovtun, A.; Ignatius, A. The role of complement in trauma and fracture healing. Semin. Immunol. 2013, 25, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, Y.; Yamasaki, S.; Oi, K.; Kuranobu, T.; Nojima, T.; Miyaki, S.; Ida, H.; Sugiyama, E. IL-1beta Enhances Wnt Signal by Inhibiting DKK1. Inflammation 2018, 41, 1945–1954. [Google Scholar] [CrossRef]
- Liu, S.; Li, Y.; Xia, L.; Shen, H.; Lu, J. IL-35 prevent bone loss through promotion of bone formation and angiogenesis in rheumatoid arthritis. Clin. Exp. Rheumatol. 2019, 37, 820–825. [Google Scholar]
- Moriwaki, S.; Into, T.; Suzuki, K.; Miyauchi, M.; Takata, T.; Shibayama, K.; Niida, S. γ-Glutamyltranspeptidase is an endogenous activator of Toll-like receptor 4-mediated osteoclastogenesis. Sci. Rep. 2016, 6, 35930. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.H.; Huang, D.; Li, Z.J.; Li, X.H.; Wang, X.; Yang, H.P.; Tian, S.P.; Mao, Y.; Liu, M.F.; Wang, Y.F.; et al. Toll-like receptor-4-dependence of the lipopolysaccharide-mediated inhibition of osteoblast differentiation. Genet. Mol. Res. 2016, 15. [Google Scholar] [CrossRef] [PubMed]
- Rangkasenee, N.; Murani, E.; Schellander, K.; Cinar, M.U.; Ponsuksili, S.; Wimmers, K. Gene expression profiling of articular cartilage reveals functional pathways and networks of candidate genes for osteochondrosis in pigs. Physiol. Genom. 2013, 45, 856–865. [Google Scholar] [CrossRef] [PubMed]
- Rangkasenee, N.; Murani, E.; Brunner, R.M.; Schellander, K.; Cinar, M.U.; Luther, H.; Hofer, A.; Stoll, M.; Witten, A.; Ponsuksili, S.; et al. Genome-Wide Association Identifies TBX5 as Candidate Gene for Osteochondrosis Providing a Functional Link to Cartilage Perfusion as Initial Factor. Front. Genet. 2013, 4, 78. [Google Scholar] [CrossRef] [PubMed]
- Ewels, P.; Magnusson, M.; Lundin, S.; Kaller, M. MultiQC: Summarize analysis results for multiple tools and samples in a single report. Bioinformatics 2016, 32, 3047–3048. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef] [PubMed]
| Foal.F vs. Adult.F | Foal.H vs. Adult.H | Foal.S vs. Adult.S | ||
|---|---|---|---|---|
| Articular Cartilage | Down | 212 | 288 | 260 |
| Not Significant | 15,832 | 15,785 | 15,831 | |
| Up | 396 | 367 | 349 | |
| Subchondral Bone | Down | 1343 | 1360 | 1250 |
| Not Significant | 15,461 | 15,296 | 15,606 | |
| Up | 1205 | 1353 | 1153 |
| Genes in Reference List | Genes in Analyzed List | Fold Enrichment | Raw p-Value | FDR | |
|---|---|---|---|---|---|
| GO-Slim Biological Process | |||||
| extracellular matrix organization (GO:0030198) | 69 | 16 | 5.05 | 7.76 × 10−7 | 0.0002 |
| microtubule-based movement (GO:0007018) | 87 | 15 | 3.75 | 3.85 × 10−5 | 0.0041 |
| regulation of axonogenesis (GO:0050770) | 27 | 8 | 6.45 | 1.10 × 10−4 | 0.007 |
| chemical synaptic transmission (GO:0007268) | 330 | 33 | 2.18 | 8.10 × 10−5 | 0.0073 |
| postitive regulation of synaptic transmission (GO:0050806) | 24 | 7 | 6.35 | 3.22 × 10−4 | 0.017 |
| cell adhesion (GO:0007155) | 373 | 34 | 1.98 | 3.39 × 10−4 | 0.017 |
| Wnt signaling pathway (GO:0016055) | 98 | 14 | 3.11 | 3.98 × 10−4 | 0.018 |
| positive regulation of signal transduction (GO:0009967) | 112 | 15 | 2.91 | 4.65 × 10−4 | 0.018 |
| tissue development (GO:0009888) | 117 | 15 | 2.79 | 7.01 × 10−4 | 0.026 |
| angiogenesis (GO:0001525) | 30 | 7 | 5.08 | 1.00 × 10−3 | 0.035 |
| regulation of chemotaxis (GO:0050920) | 22 | 6 | 5.93 | 1.16 × 10−3 | 0.039 |
| axonogenesis (GO:0007409) | 98 | 13 | 2.89 | 1.18 × 10−3 | 0.039 |
| amino acid transport (GO:0007416) | 15 | 5 | 7.25 | 1.45 × 10−3 | 0.044 |
| Notch signaling pathway (GO:0007219) | 32 | 7 | 4.76 | 1.39 × 10−3 | 0.044 |
| dendritic spine organization (GO:0097061) | 3 | 3 | 21.76 | 1.53 × 10−3 | 0.045 |
| synapse assembly (GO:0007416) | 15 | 5 | 7.25 | 1.45 × 10−3 | 0.045 |
| negative regulation of neurogenesis (GO:0050768) | 24 | 6 | 5.44 | 1.70 × 10−3 | 0.048 |
| GO-Slim Molecular Function | |||||
| extracellular matrix structural component (GO:0005201) | 53 | 17 | 6.98 | 6.46 × 10−9 | 3.26 × 10−6 |
| microtubule binding (GO:0008017) | 123 | 17 | 3.01 | 1.43 × 10−4 | 0.01 |
| transmembrane receptor protein kinase activity (GO:0019199) | 62 | 10 | 3.51 | 1.16 × 10−3 | 0.031 |
| cation transmembrane transporter activity (GO:0008324) | 449 | 37 | 1.79 | 1.10 × 10−3 | 0.031 |
| metallopeptidase activity (GO:0008237) | 103 | 14 | 2.96 | 6.23 × 10−4 | 0.035 |
| calcium ion binding (GO:0005509) | 205 | 21 | 2.23 | 1.46 × 10−3 | 0.037 |
| cadherin binding (GO:0045296) | 43 | 8 | 4.05 | 1.60 × 10−3 | 0.039 |
| peptidase inhibitor activity (GO:0030414) | 117 | 14 | 2.6 | 1.88 × 10−3 | 0.043 |
| ATP-dependent microtubule motor activity, plus-end-directed (GO:0008574) | 25 | 6 | 5.22 | 2.03 × 10−3 | 0.045 |
| gated channel activity (GO:0022836) | 160 | 17 | 2.31 | 2.45 × 10−3 | 0.046 |
| GO-Slim Cellular Component | |||||
| collagen trimer (GO:0005581) | 10 | 6 | 13.05 | 3.88 × 10−5 | 0.0022 |
| microtubule associated complex (GO:0005875) | 58 | 12 | 4.5 | 4.89 × 10−5 | 0.0022 |
| extrinsic component of plasma membrane (GO:0019897) | 28 | 8 | 6.22 | 1.36 × 10−4 | 0.0041 |
| cell-cell junction (GO: 0005911) | 70 | 12 | 3.73 | 2.35 × 10−4 | 0.0066 |
| cation channel complex (GO:0034703) | 94 | 14 | 3.24 | 2.72 × 10−4 | 0.0072 |
| microtubule (GO:0005874) | 162 | 19 | 2.55 | 4.04 × 10−4 | 0.0095 |
| postsynaptic density (GO:0014069) | 46 | 9 | 4.26 | 6.06 × 10−4 | 0.013 |
| postsynaptic membrane (GO:0045211) | 58 | 10 | 3.75 | 7.35 × 10−4 | 0.015 |
| ionotropic glutamate receptor complex (GO:0008328) | 33 | 7 | 4.62 | 1.62 × 10−3 | 0.029 |
| extracellular space (GO:0005615) | 863 | 61 | 1.54 | 1.73 × 10−3 | 0.029 |
| basement membrane (GO:0005604) | 4 | 3 | 16.32 | 2.59 × 10−3 | 0.036 |
| Pathway | Genes in Reference List | Genes in Analyzed List | Fold Enrichment | Raw p-Value | FDR |
|---|---|---|---|---|---|
| PANTHER | |||||
| Collagen chain trimerization (R-HSA-8948216) | 44 | 14 | 6.92 | 1.52 × 10−7 | 4.15 × 10−5 |
| Collagen biosynthesis and modifying enzymes (R-HSA-1650814) | 67 | 18 | 5.85 | 2.38 × 10−8 | 8.68 × 10−6 |
| Collagen formation (R-HSA-1474290) | 89 | 21 | 5.13 | 1.17 × 10−8 | 8.53 × 10−6 |
| ECM organization (R-HSA-1474244) | 299 | 54 | 3.93 | 1.02 × 10−15 | 2.25 × 10−12 |
| NCAM1 interactions (R-HSA-8948216) | 42 | 13 | 6.73 | 5.48 × 10−7 | 0.0001 |
| NCAM signaling for neurite outgrowth (R-HSA-375165) | 60 | 14 | 5.08 | 3.54 × 10−6 | 0.00055 |
| Axon guidance (R-HSA-422475) | 550 | 52 | 2.06 | 3.97 × 10−6 | 0.00058 |
| Collagen degradation (R-HSA-1442490) | 64 | 16 | 5.44 | 3.27 × 10−7 | 7.16 × 10−5 |
| Degradation of the ECM (R-HSA-1474228) | 140 | 26 | 4.04 | 1.73 × 10−8 | 9.48 × 10−6 |
| Assembly of collagen fibrils and other multimeric structures (R-HSA-2022090) | 60 | 15 | 5.44 | 7.60 × 10−7 | 0.00013 |
| Integrin cell surface interactions (R-HSA-216083) | 84 | 20 | 5.18 | 2.28 × 10−8 | 9.99 × 10−6 |
| ECM proteoglycans (R-HSA-3000178) | 76 | 17 | 4.87 | 5.46 × 10−7 | 0.00011 |
| Kinesins (R-HSA-983189) | 61 | 13 | 4.64 | 1.82 × 10−5 | 0.0025 |
| Hemostasis (R-HSA-109582) | 671 | 54 | 1.75 | 1.99 × 10−4 | 0.021 |
| Elastic fibril formation (R-HSA-1566948) | 45 | 9 | 4.35 | 5.27 × 10−4 | 0.046 |
| Neurotransmitter receptors and postsynaptic signal transmission (R-HSA-112314) | 148 | 19 | 2.79 | 1.43 × 10−4 | 0.017 |
| Transmission across chemical synapses (R-HSA-112315) | 218 | 26 | 2.59 | 4.57 × 10−5 | 0.0056 |
| Neuronal system (R-HSA-112316) | 361 | 42 | 2.53 | 2.29 × 10−7 | 5.57 × 10−5 |
| Reactome | |||||
| ECM organization (R-HSA-1474244) | 329 | 54 | NA | 1.26 × 10−8 | 1.49 × 10−5 |
| Collagen chain trimerization (R-HSA-8948216) | 44 | 14 | 3.95 × 10−6 | 0.0017 | |
| Integrin cell surface interactions (R-HSA-216083) | 86 | 20 | 4.40 × 10−6 | 0.0017 | |
| Collagen biosynthesis and modifying enzymes (R-HSA-1650814) | 76 | 18 | 1.02 × 10−5 | 0.003 | |
| NCAM1 interactions (R-HSA-419037) | 44 | 13 | 1.86 × 10−5 | 0.0041 | |
| Collagen formation (R-HSA-1474290) | 104 | 21 | 2.07 × 10−5 | 0.0041 | |
| Degradation of the ECM (R-HSA-1474228) | 148 | 26 | 2.45 × 10−5 | 0.0041 | |
| Collagen degradation (R-HSA-1442490) | 69 | 16 | 4.00 × 10−5 | 0.0059 | |
| ECM proteoglycans (R-HSA-3000178) | 79 | 17 | 5.75 × 10−5 | 0.0075 | |
| Assembly of collagen fibrils and other multimeric structures (R-HSA-2022090) | 67 | 15 | 1.00 × 10−4 | 0.012 | |
| NCAM signaling for neurite out-growth (R-HSA-375165) | 69 | 14 | 4.46 × 10−4 | 0.048 | |
| Gene Family | Gene Symbol(s) | ECM Organization | Collagen Chain Trimerization | Collagen Biosynthesis and Modifying Enzymes | Collagen Formation | Degradation of the ECM | Collagen Degradation | ECM Proteoglycans | Assembly of Collagen Fibrils and Other Multimeric Structures | Integrin Cell Surface Interactions | NCAM1 Interactions | NCAM Signaling for Neurite Outgrowth |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Upregulated in foals | ||||||||||||
| ADAM metallopeptidase domain | ADAM12 ADAM19 | x | ||||||||||
| ADAM metallopeptidase with thrombospondin | ADAMTS2 ADAMTS3 | x | x | x | ||||||||
| ADAMTS8 | x | x | ||||||||||
| Calcium voltage-gated channel subunit | CACNA1G CACNA1H | x | x | |||||||||
| Calpain | CAPN6 | x | x | |||||||||
| Collagen | COL2A1 COL4A1 COL9A1 COL9A2 COL9A3 | x | x | x | x | x | x | x | x | x | x | x |
| COL8A1 | x | x | x | x | x | x | x | x | ||||
| COL11A1 COL11A2 | x | x | x | x | x | x | ||||||
| COL16A1 | x | x | x | x | x | x | x | |||||
| COL18A1 | x | x | x | x | x | x | x | x | ||||
| COL21A1 | x | x | x | x | ||||||||
| Elastin | ELN | x | x | |||||||||
| Immunoglobulin superfamily | F11R | x | x | |||||||||
| NCAM1 | x | x | x | x | ||||||||
| Fibulin | FBLN2 | x | ||||||||||
| Fibrillin | FBN2 | x | x | |||||||||
| Hyaluronan and proteoglycan link protein | HAPLN1 | x | x | |||||||||
| Heparain sulfate proteoglycan | HSPG2 | x | x | x | x | |||||||
| Integrin subunit | ITGA10 | x | x | |||||||||
| VEGF receptor | KDR | x | x | |||||||||
| Lysyl oxidase | LOXL2 | x | x | |||||||||
| Matrilin | MATN1 MATN3 | x | x | |||||||||
| Microfibrillar associated protein | MFAP2 | x | ||||||||||
| Matrix metalloproteinase | MMP2 MMP15 | x | x | x | ||||||||
| MMP9 | x | x | x | x | x | |||||||
| Nidogen | NID2 | x | ||||||||||
| Platelet and endothelial cell adhesion molecule | PECAM1 | x | x | |||||||||
| Serpin | SERPINH1 | x | x | x | ||||||||
| Secreted protein acidic and cysteine rich | SPARC | x | x | |||||||||
| Spectrin | SPTBN2 | x | ||||||||||
| Downregulated in foals | ||||||||||||
| Calpain | CAPN2 | x | x | |||||||||
| Transmembrane 4 superfamily | CD151 | x | x | x | ||||||||
| Collagen | COL4A3 | x | x | x | x | x | x | x | x | x | x | x |
| COL6A5 COL6A6 | x | x | x | x | x | x | x | x | x | x | x | |
| Small leucine-rich proteoglycan | DCN | x | x | x | ||||||||
| Integrin subunit | ITGA3 ITGAM | x | x | |||||||||
| ITGB5 | x | x | x | |||||||||
| latent TGFb binding protein | LTB2 LTB3 LTB4 | x | ||||||||||
| Procollagen C-endopeptidase enhancer | PCOLCE2 | x | x | x | ||||||||
| Prion protein | PRNP | x | x | |||||||||
| Signal peptide, CUB domain and EGF-like domain | SCUBE1 | x | x | |||||||||
| Secreted phosphoprotein | SPP1 | x | x | x | ||||||||
| Sialyltransferase | ST8SIA4 | x | x | |||||||||
| Tissue inhibitor of metalloproteinase | TIMP2 | x | x | |||||||||
| Tenascin | TNXB | x | x | |||||||||
| Genes in Reference List | Genes in Analyzed List | Fold Enrichment | Raw p-Value | FDR | |
|---|---|---|---|---|---|
| GO-Slim Biological Process | |||||
| mitotic nuclear division (GO:0140014) | 209 | 63 | 2.17 | 5.19 × 10−7 | 8.47 × 10−5 |
| positive regulation of cytosolic calcium ion concentration (GO:0007204) | 65 | 24 | 2.65 | 1.33 × 10−4 | 0.0096 |
| inflammatory response (GO:0006954) | 101 | 32 | 2.28 | 1.50 × 10−4 | 0.01 |
| intracellular signal transduction (GO:0035556) | 777 | 151 | 1.4 | 2.24 × 10−4 | 0.013 |
| extracellular matrix organization (GO:0030198) | 69 | 24 | 2.5 | 3.32 × 10−4 | 0.018 |
| tissue development (GO:0009888) | 117 | 34 | 2.09 | 3.93 × 10−4 | 0.02 |
| cytoskeleton organization (GO:0007010) | 226 | 55 | 1.75 | 4.47 × 10−4 | 0.021 |
| regulation of cell proliferation (GO:0042127) | 96 | 29 | 2.17 | 5.34 × 10−4 | 0.022 |
| transmembrane receptor protein tyrosine kinase signaling pathway (GO:0007169) | 197 | 49 | 1.79 | 5.51 × 10−4 | 0.022 |
| cell migration (GO:0016477) | 177 | 45 | 1.83 | 6.07 × 10−4 | 0.024 |
| neuron development (GO:0048666) | 172 | 44 | 1.84 | 7.42 × 10−4 | 0.028 |
| cell-cell adhesion (GO:0098609) | 104 | 30 | 2.07 | 8.37 × 10−4 | 0.03 |
| regulation of mitotic nuclear division (GO:0007088) | 26 | 12 | 3.32 | 1.37 × 10−3 | 0.044 |
| GO-Slim Molecular Function | |||||
| extracellular matrix structural component (GO:0005201) | 53 | 22 | 2.98 | 3.00 × 10−5 | 0.0034 |
| oxidoreductase activity (GO:0016491) | 527 | 111 | 1.51 | 1.08 × 10−4 | 0.0055 |
| cytokine receptor activity (GO:0004896) | 69 | 25 | 2.6 | 1.81 × 10−4 | 0.0076 |
| G-protein coupled peptide receptor activity (GO:0008528) | 76 | 25 | 2.36 | 5.99 × 10−4 | 0.014 |
| calcium ion binding (GO:0005509) | 205 | 51 | 1.79 | 4.92 × 10−4 | 0.014 |
| growth factor binding (GO:0019838) | 46 | 18 | 2.81 | 7.18 × 10−4 | 0.017 |
| actin filament binding (GO:0051015) | 65 | 21 | 2.32 | 1.49 × 10−3 | 0.029 |
| transmembrane receptor protein kinase activity (GO:0004714) | 60 | 20 | 2.39 | 1.60 × 10−3 | 0.03 |
| peptidase inhibitor activity (GO:0030414) | 117 | 32 | 1.96 | 1.48 × 10−3 | 0.03 |
| C-C chemokine binding (GO:0019957) | 24 | 11 | 3.29 | 2.25 × 10−3 | 0.038 |
| metallopeptidase activity (GO:0008237) | 103 | 28 | 1.95 | 2.91 × 10−3 | 0.043 |
| GO-Slim Cellular Component | |||||
| integral component of plasma membrane (GO:0005887) | 733 | 165 | 1.62 | 5.71 × 10−8 | 1.28 × 10−5 |
| collagen-containing ECM (GO:0062023) | 33 | 15 | 3.27 | 4.09 × 10−4 | 0.0087 |
| receptor complex (GO:0043235) | 177 | 45 | 1.83 | 6.07 × 10−4 | 0.011 |
| focal adhesion (GO:0005925) | 25 | 12 | 3.45 | 1.05 × 10−3 | 0.016 |
| condensed nuclear chromosome kinetochore (GO:0000778) | 9 | 7 | 5.59 | 1.69 × 10−3 | 0.022 |
| microtubule (GO:0005874) | 162 | 40 | 1.77 | 2.30 × 10−3 | 0.025 |
| extracellular space (GO:0005615) | 863 | 157 | 1.31 | 2.06 × 10−3 | 0.025 |
| cyclin-dependent protein kinase holoenzyme complex (GO:0000307) | 28 | 12 | 3.08 | 2.24 × 10−3 | 0.026 |
| spindle (GO:0005819) | 46 | 16 | 2.5 | 2.91 × 10−3 | 0.03 |
| cell surface (GO:0009986) | 322 | 67 | 1.49 | 3.75 × 10−3 | 0.038 |
| Pathway | Genes in Reference List | Genes in Analyzed List | Fold Enrichment | Raw p-Value | FDR |
|---|---|---|---|---|---|
| PANTHER | |||||
| Neutrophil degranulation (R-HSA-6798695) | 479 | 167 | 2.5 | 1.2 × 10−21 | 1.31 × 10−18 |
| Resolution of sister chromatid cohesion (R-HSA-2500257) | 122 | 50 | 2.94 | 2.69 × 10−9 | 4.54 × 10−7 |
| RHO GTPases activate formins (R-HSA-5663220) | 135 | 51 | 2.71 | 2.52 × 10−8 | 3.25 × 10−6 |
| Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal (R-HSA-141444) | 92 | 39 | 3.04 | 1.05 × 10−7 | 1.15 × 10−5 |
| Integrin cell surface interactions (R-HSA-216083) | 84 | 36 | 3.08 | 1.73 × 10−7 | 1.65 × 10−5 |
| ECM proteoglycans (R-HSA-3000178) | 76 | 31 | 2.93 | 3.60 × 10−6 | 2.46 × 10−4 |
| Deposition of new CENPA-containing nucleosomes at the centromere (R-HSA-606279) | 54 | 25 | 3.33 | 4.40 × 10−6 | 3.60 × 10−4 |
| Non-integrin membrane-ECM interactions (R-HSA-3000171) | 59 | 26 | 3.17 | 5.76 × 10−6 | 3.60 × 10−4 |
| Defective B3GALTL causes Peters-plus syndrome (PpS) (R-HSA-5083635) | 38 | 20 | 3.78 | 9.26 × 10−6 | 5.63 × 10−4 |
| O-glycosylation of TSR domain-containing proteins (R-HSA-5173214) | 39 | 20 | 3.68 | 1.24 × 10−5 | 7.35 × 10−4 |
| Collagen degradation (R-HSA-1442490) | 64 | 25 | 2.81 | 5.24 × 10−5 | 0.0026 |
| Assembly of collagen fibrils and other multimeric structures (R-HSA-2022090) | 60 | 24 | 2.87 | 6.26 × 10−5 | 0.0029 |
| Kinesins (R-HSA-983189) | 61 | 24 | 2.83 | 7.11 × 10−5 | 0.0033 |
| NCAM1 interactions (R-HSA-419037) | 42 | 19 | 3.25 | 7.67 × 10−5 | 0.033 |
| Molecules associated with elastic fibres (R-HSA-2129379) | 38 | 18 | 3.4 | 7.43 × 10−5 | 0.0033 |
| Separation of sister chromatids (R-HSA-2467813) | 186 | 50 | 1.93 | 8.50 × 10−5 | 0.0035 |
| GPVI-mediated activation cascade (R-HSA-114604) | 35 | 17 | 3.49 | 9.21 × 10−5 | 0.0037 |
| Platelet degranuation (R-HSA-114608) | 127 | 38 | 2.15 | 1.04 × 10−4 | 0.0039 |
| Cell surface interactions at the vascular wall (R-HSA-202733) | 198 | 50 | 1.81 | 3.96 × 10−4 | 0.012 |
| Laminin interactions (R-HSA-3000157) | 30 | 14 | 3.35 | 5.15 × 10−4 | 0.015 |
| Constituative signaling by aberrant PI3K in cancer (R-HSA-2219530) | 55 | 20 | 2.61 | 5.72 × 10−4 | 0.016 |
| Metabolism of folate and pterines (R-HSA-196757) | 16 | 10 | 4.49 | 6.05 × 10−4 | 0.016 |
| MyD88 deficiency (TLR2/4) (R-HSA-5602498) | 10 | 8 | 5.75 | 6.91 × 10−4 | 0.018 |
| Neuronal system (R-HSA-112316) | 361 | 78 | 1.55 | 6.78 × 10−4 | 0.018 |
| Signaling by interleukins (R-HSA-449147) | 449 | 93 | 1.49 | 6.88 × 10−4 | 0.018 |
| Reactome | |||||
| Neutrophil degranulation (R-HSA-6798695) | 480 | 164 | NA | 1.47 × 10−9 | 2.79 × 10−6 |
| RHO GTPase effectors (R-HSA-195258) | 326 | 105 | 1.49 × 10−5 | 0.014 | |
| ECM organization (R-HSA-1474244) | 329 | 105 | 2.13 × 10−5 | 0.014 | |
| Signaling by RHO GTPases (R-HSA-194315) | 457 | 136 | 3.79 × 10−5 | 0.018 | |
| Aplification of signal from the kinetochores (R-HSA-141424) | 94 | 39 | 6.82 × 10−5 | 0.022 | |
| Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal (R-HSA-141444) | 94 | 39 | 6.82 × 10−5 | 0.022 | |
| Resolution of sister chromatid cohesion (R-HSA-2500257) | 134 | 50 | 9.86 × 10−5 | 0.025 | |
| Integrin cell surface interactions (R-HSA-216083) | 86 | 36 | 1.07 × 10−4 | 0.025 | |
| Deposition of new CENPA-containing nucleosomes at the centromere (R-HSA-606279) | 54 | 25 | 2.71 × 10−4 | 0.052 | |
| Nucleosome assembly (R-HSA-774815) | 54 | 25 | 2.71 × 10−4 | 0.052 | |
| Defective B3GALTL causes Peters-plus syndrome (PpS) (R-HSA-5083635) | 39 | 20 | 3.04 × 10−4 | 0.052 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kemper, A.M.; Drnevich, J.; McCue, M.E.; McCoy, A.M. Differential Gene Expression in Articular Cartilage and Subchondral Bone of Neonatal and Adult Horses. Genes 2019, 10, 745. https://doi.org/10.3390/genes10100745
Kemper AM, Drnevich J, McCue ME, McCoy AM. Differential Gene Expression in Articular Cartilage and Subchondral Bone of Neonatal and Adult Horses. Genes. 2019; 10(10):745. https://doi.org/10.3390/genes10100745
Chicago/Turabian StyleKemper, Ann M., Jenny Drnevich, Molly E. McCue, and Annette M. McCoy. 2019. "Differential Gene Expression in Articular Cartilage and Subchondral Bone of Neonatal and Adult Horses" Genes 10, no. 10: 745. https://doi.org/10.3390/genes10100745
APA StyleKemper, A. M., Drnevich, J., McCue, M. E., & McCoy, A. M. (2019). Differential Gene Expression in Articular Cartilage and Subchondral Bone of Neonatal and Adult Horses. Genes, 10(10), 745. https://doi.org/10.3390/genes10100745






