Abstract
Wooden breast is a muscle disorder affecting modern commercial broiler chickens that causes a palpably firm pectoralis major muscle and severe reduction in meat quality. Most studies have focused on advanced stages of wooden breast apparent at market age, resulting in limited insights into the etiology and early pathogenesis of the myopathy. Therefore, the objective of this study was to identify early molecular signals in the wooden breast transcriptional cascade by performing gene expression analysis on the pectoralis major muscle of two-week-old birds that may later exhibit the wooden breast phenotype by market age at 7 weeks. Biopsy samples of the left pectoralis major muscle were collected from 101 birds at 14 days of age. Birds were subsequently raised to 7 weeks of age to allow sample selection based on the wooden breast phenotype at market age. RNA-sequencing was performed on 5 unaffected and 8 affected female chicken samples, selected based on wooden breast scores (0 to 4) assigned at necropsy where affected birds had scores of 2 or 3 (mildly or moderately affected) while unaffected birds had scores of 0 (no apparent gross lesions). Differential expression analysis identified 60 genes found to be significant at an FDR-adjusted p-value of 0.05. Of these, 26 were previously demonstrated to exhibit altered expression or genetic polymorphisms related to glucose tolerance or diabetes mellitus in mammals. Additionally, 9 genes have functions directly related to lipid metabolism and 11 genes are associated with adiposity traits such as intramuscular fat and body mass index. This study suggests that wooden breast disease is first and foremost a metabolic disorder characterized primarily by ectopic lipid accumulation in the pectoralis major.
1. Introduction
Wooden breast is one of several muscle abnormalities of modern commercial broiler chickens that causes substantial economic losses in the poultry industry due to its impact on meat quality. Emerging evidence suggests wooden breast may also be detrimental to bird welfare as affected chickens exhibit increased locomotor difficulties, decreased wing mobility, and higher mortality rates [1,2,3]. While the etiology of the myopathy is still poorly understood, many believe it to be a side-effect of improved management practices and selective breeding for performance traits due to increased susceptibility among broilers with high feed efficiency [4,5], breast muscle yield [4,6,7], and growth rate [8,9].
Macroscopic manifestations of the disorder include pale and hardened areas, subcutaneous and fascial edema, petechial hemorrhages, spongy areas with disintegrating myofiber bundles, and white fatty striations characteristic of white striping [2,10]. An early study of wooden breast characterized its microscopic presentation as polyphasic myodegeneration and necrosis with regeneration and interstitial connective tissue accumulation (fibrosis), primarily affecting the cranial end of the pectoralis major muscle [10]. However, it has since been demonstrated that venous inflammation (phlebitis) and perivascular lipid and inflammatory cell infiltration appear in the first week of age and precede other symptoms [2]. Differential gene expression analysis of the pectoralis major in 7-week-old broilers suggests that hypoxia, oxidative stress, fiber-type switching, and increased intracellular calcium may be important components of the myopathy [11]. In two- and three-week old birds, differentially expressed genes were mostly associated with increased inflammation, vascular disease, increased oxidative stress, extracellular matrix remodeling, dysregulation of carbohydrates and lipids, and impaired excitation-contraction coupling [12]. Metabolomic profiling is in agreement with these results and provides evidence of oxidative stress and dysregulated carbohydrate and lipid metabolism in affected birds at 7 weeks of age [13].
The objective of the present study was to better characterize the transcriptional anomalies that exist in the pectoralis major of two-week-old birds that later develop wooden breast by market age at 7 weeks. Only one other gene expression study has investigated early stages of wooden breast [12]. The current study serves as a continuation of that work, but makes two key changes. First, unlike the previous study that used only male birds, we included only female birds in the RNA-seq analysis. Second, birds in the affected group all possessed mild or moderate wooden breast phenotypes rather than severe symptoms, which allowed us to capture a clearer signal of the earliest transcriptomic perturbations associated with the myopathy.
2. Materials and Methods
2.1. Experimental Animals and Tissue Collection
The University of Delaware Institutional Animal Care and Use Committee approved the animal conditions and experimental procedures used in this scientific study under protocol number 48R-2015-0. For this experiment, 302 mixed male and female Cobb500 broilers were raised according to industry growing standards in two poultry houses from 1 day to 7 weeks of age. Birds were divided between two poultry houses due to capacity constraints, but both houses were maintained at the same environmental conditions. Chickens were provided with continuous free access to water and feed that met all nutritional recommendations for Cobb500 broilers. At 14 days of age, biopsy samples of the craniolateral area of the left pectoralis major muscle were collected in the same manner described by a previous study [12] from 101 birds randomly selected from both houses. After biopsy, all birds were grown out to 7 weeks of age, at which time they were euthanized by cervical dislocation. During necropsy, the pectoralis major muscles were evaluated for gross lesions and palpable firmness associated with wooden breast and each bird was assigned a wooden breast score using a 0-4 scale; 0-Normal indicates the bird had no macroscopic signs of the myopathy, 1-Very Mild indicates 1% or less of the breast muscle was affected, 2-Mild indicates between 1% and 10% of the breast muscle was affected, 3-Moderate indicates between 10% and 50% of the breast muscle was affected, and a score of 4-Severe indicates that more than 50% was affected. This scoring system is slightly different from the one previously used in our laboratory and separates unaffected and mildly and moderately affected chickens with higher resolution.
2.2. Sample Selection and RNA-Sequencing
Selection of samples for use in RNA-seq analysis was based on wooden breast scores assigned at necropsy at 7 weeks of age. A total of 6 unaffected and 8 affected birds were identified; affected birds had scores of 2 or 3 (mildly or moderately affected) while unaffected birds had scores of 0 (no apparent gross lesions). Only samples taken from female birds were used for RNA-seq. Total RNA was extracted from pectoralis major tissue samples using the mirVana miRNA Isolation Kit (Thermo Fisher Scientific, Waltham, MA, USA) according to the manufacturer’s protocol and stored at −80 °C until cDNA library preparation. Each RNA sample was quantified using the NanoDrop 1000 Spectrophotometer (Thermo Fisher Scientific) and quality was assessed with the Fragment Analyzer at the Delaware Biotechnology Institute (DBI). cDNA libraries were constructed using the ScriptSeq Complete Kit (Human/Mouse/Rat) (Illumina, San Diego, CA, USA) with the optional step of adding a user-defined barcode to the library. The 14 barcoded cDNA libraries were normalized and 10 µl of each sample were pooled in two tubes (7 samples in each pool). Pooled libraries were subsequently submitted to the DBI for paired-end 2 × 76-nucleotide sequencing on two lanes of a flow cell using the HiSeq 2500 Sequencing System (Illumina).
Raw sequencing reads were demultiplexed and then checked for quality using FastQC v0.11.7 [14]. All samples passed the quality check and were submitted to Trimmomatic v0.38 [15] to trim leading and trailing bases with quality below 20, remove reads with an average quality below 15, and remove reads that were shorter than 30 bases in length. Trimmed reads were then mapped to both Gallus_gallus-5.0 (Ensembl release 94) and GRCg6a (Ensembl release 95) chicken reference genomes using HISAT2 v2.1.0 [16] with concordant mapping required for both reads in each pair. Cuffdiff v2.2.1 [17] was used with the fragment bias correction option to identify differentially expressed genes between affected and unaffected birds. Genes were considered statistically significant if the FDR-adjusted p-value was ≤ 0.05. The use of two reference genome builds, the latter of which was released during the course of this study, provided validation of our results and allowed us to capture differentially expressed genes that may have borderline statistical significance due to assembly errors or bias. One sample (animal ID 424183) in the unaffected group displayed an extreme outlier expression pattern; it was therefore removed and differential expression analysis with Cuffdiff was repeated without this sample (5 unaffected vs. 8 affected). To compile the results generated from each reference genome, Ensembl Gene IDs from Gallus-gallus-5.0 were mapped to GRCg6a gene IDs using Ensembl’s ID History Converter; differentially expressed genes with annotation differences between the two reference genome releases were scrutinized for consistency. Pairwise correlation analysis and visualization of differentially expressed genes was conducted with the “stats” and “corrplot” packages [18] in R only using expression data generated with the GRCg6a reference genome build.
3. Results
An average of 19,616,353 paired-end sequence reads were generated per sample, which was reduced to an average of 19,609,044 paired-end reads after trimming. The average mapping rate per sample was 74.5% with the Gallus_gallus-5.0 reference genome build and 75.1% with GRCg6a. The total number of sequenced reads, trimmed reads, and mapped reads per sample can be found in Table S1.
There were 52 differentially expressed genes identified using the Gallus_gallus-5.0 reference genome build and 29 differentially expressed genes using the GRCg6a genome build. After accounting for changes in annotation of Ensembl Gene IDs between genome releases, a total of 60 genes were found to be differentially expressed between affected and unaffected groups across both analyses, with 18 differentially expressed genes overlapping between both Gallus_gallus-5.0 and GRCg6a. Three Ensembl Gene IDs from the earlier build were deprecated in GRCg6a and were excluded from further analysis. Of the 60 differentially expressed genes used for downstream analysis, 52 were up-regulated in affected birds and 8 were down-regulated in affected birds (Table 1).

Table 1.
Differentially expressed genes between wooden breast-affected pectoralis major muscle samples and unaffected samples at 2 weeks of age. Log2FC is calculated by log2(FPKMaffected/FPKMunaffected). Unknown gene names are indicated with a dash (-). Non-significant p-values (i.e., FDR-adjusted p-values > 0.05) are indicated as n.s.
Correlation analysis of differentially expressed genes revealed two major clusters with Pearson’s correlation coefficients greater than 0.8 for all gene pairs (see Figure 1). The first is a cluster of 8 genes, all of which were excluded from further analysis due to presumed skin contamination. These include serine palmitoyltransferase long-chain base subunit 3 (SPTLC3), desmoplakin (DSP), ELOVL fatty acid elongase 4 (ELOVL4), PERP2, TP53 apoptosis effector (PERP2), keratin 5 (KRT5), cell adhesion molecule L1 like (CHL1), lymphocyte antigen 6 complex, locus E-like (LY6CLEL), and transmembrane protein 254 (TMEM254). Several of these genes are known to be primarily expressed in the skin and a previous biopsy study using the same technique demonstrated that biopsy samples are prone to skin contamination [12]. Additionally, differential expression of these genes was driven by the same three samples, one unaffected and two affected, and expression in the remaining samples was relatively very low or approximately zero. The second cluster consisted of 9 protein-coding genes with demonstrated or putative involvement in lipid metabolism. Although no other clusters were apparent from correlation analysis, functional groupings of differentially expressed genes included muscle growth and function, calcium signaling, and endoplasmic reticulum (ER) stress response. We also found a substantial number of genes that are differentially expressed, implicated, or otherwise involved in metabolic syndrome in mammals, which is characterized primarily by diabetes, insulin resistance, obesity, elevated blood lipids, and high blood pressure.
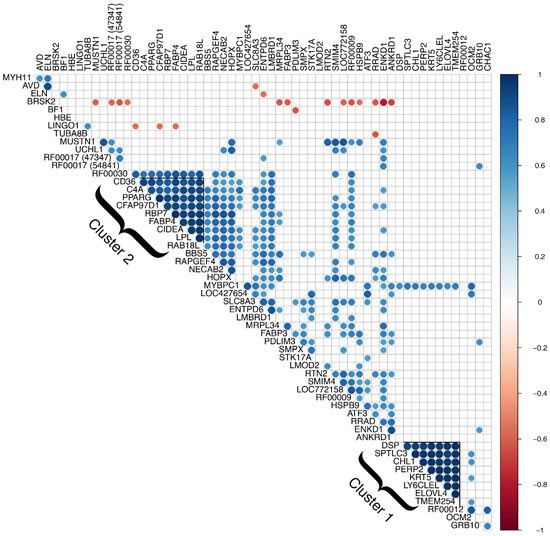
Figure 1.
Correlation analysis of differentially expressed genes. Genes with significantly correlated expression (p-value ≤ 0.05) are shown in blue (positive correlation) and red (negative correlation). Two major clusters of genes have Pearson’s correlation coefficients greater than 0.8 for all gene pairs. Cluster 1 consists of 8 genes, all of which were excluded from further analysis due to presumed skin contamination. Cluster 2 consists of 9 genes related to lipid metabolism or adiposity traits.
4. Discussion
Metabolic syndrome refers to a cluster of conditions, including obesity, high blood sugar, high serum triglycerides, low serum HDL (high-density lipoprotein) cholesterol, and high blood pressure, that put an individual at greater risk of developing type 2 diabetes and associated complications such as atherosclerosis, cardiomyopathy, non-alcoholic fatty liver disease, and diabetic nephropathy. Our data revealed a surprising number of differentially expressed genes implicated in or associated with metabolic syndrome in humans. Among the 60 differentially expressed genes identified in this study, 20 are previously reported to exhibit altered expression in relation to diabetes or a closely related metabolic condition and nine genes have been identified as candidate genes in association studies of glucose tolerance or diabetes mellitus (Table 2). One of these candidate genes, Bardet-Biedl syndrome 5 (BBS5), is associated with a rare ciliopathy that strongly predisposes individuals to diabetes and other metabolic complications: obesity and diabetes mellitus are actually considered diagnostic features of the disease [19,20]. Upon further examination, we found that many of the conditions surrounding metabolic syndrome in humans possessed important similarities to the wooden breast phenotype, namely inflammation, ectopic fat deposition, dysregulation of Ca2+ homeostasis, ER stress, oxidative stress, altered glucose metabolism, fibrosis, and hypertrophy.

Table 2.
Differentially expressed genes linked to diabetes and glucose tolerance. Of the 60 differentially expressed genes identified in this study, 26 are either proposed as candidate genes for glucose tolerance or diabetes mellitus or exhibit altered expression in relation to diabetes or a closely related metabolic condition.
4.1. Increased Expression of Genes Involved in Lipid Metabolism
The most important connection to metabolic syndrome in our results is the increased expression of genes involved in lipid metabolism in the pectoralis major of affected birds. Many of the differentially expressed genes from our analysis encode proteins with critical or rate-limiting functions in lipid metabolism and homeostasis such as lipoprotein triglyceride hydrolysis, fatty acid transport, and lipid droplet regulation. These genes include lipoprotein lipase (LPL), CD36 molecule (CD36), peroxisome proliferator-activated receptor gamma (PPARG), retinol-binding protein 7 (RBP7), fatty acid-binding protein 3 (FABP3), fatty acid-binding protein 4 (FABP4), cell death-inducing DFFA-like effector a (CIDEA), Ras-related protein Rab-18-B-like (RAB18L), and LMBR1 domain containing 1 (LMBRD1) [31,48,52,53,54,55,56,57,58,59,60]. An outstanding member of this group is LPL, which encodes lipoprotein lipase, a rate-limiting catalyst for hydrolysis of the triglycerides found in circulating lipoproteins, providing free fatty acids and monoglycerides for use by surrounding tissues [52,53]. LPL serves as a metabolic gatekeeper in terms of partitioning circulating lipids among tissues, with highest activity in tissues that oxidize or store large quantities of fatty acids [53]. Regulation of LPL expression occurs in a tissue-specific manner and alterations to LPL activity in one tissue can affect systemic nutrient partitioning by reducing substrate (lipid) availability to other tissues [53].
Several other genes, some of which are functionally uncharacterized or poorly understood regarding lipid metabolism, have expression or genetic polymorphisms correlated with adiposity traits such as body mass index, percent intramuscular fat, percent abdominal fat, or blood lipid levels. These include HOP homeobox (HOPX), myosin-binding protein-C, slow type (MYBPC1), Bardet-Biedl syndrome 5 (BBS5), growth factor receptor bound protein 10 (GRB10), CFAP97 domain containing 1 (CFAP97D1), hemoglobin subunit epsilon (HBE), ectonucleoside triphosphate diphosphohydrolase 6 (ENTPD6), complement C4A (Rodgers blood group) (C4A), mitochondrial ribosomal protein L34 (MRPL34), ATF3, and CHAC1 [20,61,62,63,64,65,66,67,68,69,70,71,72,73,74].
Notably, the present study identified a cluster of nine genes related to lipid metabolism that may represent a functional group for two main reasons. First, the genes in this cluster exhibit highly correlated expression (r > 0.8 for all gene pairs; see Figure 1). Second, this cluster includes the gene encoding the transcription factor PPARγ and several of its experimentally validated transcriptional targets (CD36, C4A, RBP7, FABP4, CIDEA, and LPL) [55,56,61,75,76]. As a master regulator of adipogenesis, PPARγ plays a crucial role in governing the distribution of lipid deposition in the body and the development of various metabolic conditions [77,78,79,80,81]. It is also one of the few established genes that has been associated with common forms of type 2 diabetes across multiple genome-wide association studies [43,44,45,46,47]. In skeletal muscle of broiler chickens and other meat-type animals, increased expression of PPARG and PPARγ target genes is frequently associated with higher intramuscular fat content [82,83,84,85].
Increased expression of genes related to lipid metabolism and fat deposition in the pectoralis major may signify the pathological overprovisioning of lipids to the muscle, consistent with histological characterization of early stages of wooden breast, in which lipid infiltration and accumulation was established as one of the first signs of disease even before wooden breast is grossly detectable [2]. The storage of excess lipids in tissues other than adipose tissue, that normally contain only small amounts of fat, is called ectopic lipid deposition and is linked to insulin resistance and metabolic dysfunction in mammals [86]. In fact, ectopic lipid deposition and the resulting lipotoxicity are considered to be a major feature of metabolic syndrome with the precise location of ectopic lipid accumulation dictating specific complications such as atherosclerosis, hepatic steatosis, and diabetic nephropathy [87]. This suggests that increased lipid deposition in the pectoralis major may be a major factor contributing to the wooden breast phenotype.
4.2. Endoplasmic Reticulum Stress and Dysregulation of Calcium Homeostasis
Results from the current study suggest that ER stress and dysregulation of calcium homeostasis are occurring in the pectoralis major muscle in the early stages of wooden breast. Evidence for ER stress is supported by the up-regulation of activating transcription factor 3 (ATF3), ChaC glutathione specific gamma-glutamylcyclotransferase 1 (CHAC1), and reticulon 2 (RTN2) and the down-regulation of BR serine/threonine kinase 2 (BRSK2). ATF3, CHAC1, and BRSK2 are part of the unfolded protein response [88,89], a highly conserved cellular stress response caused by an accumulation of unfolded or misfolded proteins in the ER [90]. An association between the unfolded protein response, lipid metabolism, dysregulation of calcium homeostasis, and metabolic syndrome has been clearly established, but the direction of causality is controversial [91,92,93,94]. The role of ATF3 in particular has been studied in the context of type 2 diabetes, non-alcoholic fatty liver disease, diabetic cardiomyopathy, atherosclerosis, and obesity, with some authors suggesting it may have both detrimental and beneficial functions related to insulin resistance, mitochondrial dysfunction, and inflammation in response to high-fat diets [22,67,95,96,97,98,99,100]. In arterial endothelial cells, ATF3 expression can be induced by exposure to high levels of triglyceride-rich lipoprotein lipolysis products [96], substantiating it as a link between high lipid metabolism and cellular stress response.
One of the most compelling links to metabolic syndrome and dysregulation of calcium homeostasis in our data is Ras-related glycolysis inhibitor and calcium channel regulator (RRAD), a gene encoding a small GTPase that binds directly to Ca2+ channel beta subunits to regulate intracellular Ca2+ signaling in muscle cells [101]. RRAD also has regulatory functions via its interaction with calmodulin [102]. This gene was originally named Ras-related associated with diabetes because it was identified via subtraction cloning as the only gene out of 4000 cDNA clones that was overexpressed in skeletal muscle of type 2 diabetic individuals compared to non-diabetic or type 1 diabetic individuals [26]. Overexpression of RRAD in cultured myocytes was found to reduce insulin-stimulated glucose uptake by 50–90%, which the authors speculated was due to a decrease in intrinsic activity of glucose transporter 4, the insulin-dependent glucose transporter [103]. An in vivo study of transgenic mice overexpressing RRAD in skeletal muscle found that high-fat feeding produced not only insulin resistance in transgenic mice, but also increased triglyceride metabolism compared to controls [104]. This suggests that RRAD may inhibit glycolysis independently from its action on glucose transporters via substrate competition with fatty acids [105]. Another regulator of intracellular Ca2+, RAPGEF4, up-regulated in the current study, is the direct target of some anti-diabetic drugs called sulfonylureas [49].
The role of Ca2+ in metabolic syndrome and diabetes is complex and not fully understood, but one theory suggests that the disruption of Ca2+ homeostasis is a feed-forward pathological cycle resulting from ER dysfunction during chronic exposure to excessive nutrients and energy [106]. The majority of intracellular Ca2+ pools are contained within the ER, known as the sarcoplasmic reticulum in muscle cells; however, high concentrations of fatty acids can mediate a substantial redistribution of ER luminal Ca2+ stores among subcompartments of the ER and from the ER to the cytosol, leading to ER stress and eventually cell death [107,108].
4.3. Increased Expression of Genes Related to Hypertrophy and Slow-Twitch Muscle
Several genes involved in myogenic differentiation and muscle hypertrophy are up-regulated in affected birds: musculoskeletal, embryonic nuclear protein 1 (MUSTN1), ankyrin repeat domain 1 (ANKRD1), and HOPX have roles in myotube formation, myofusion, and regulation of other myoblast differentiation genes [72,109,110,111,112,113], although studies of HOPX in chickens have found that it is most highly expressed in adipose tissue and has functions related to adipocyte differentiation [71,72]. Other up-regulated genes related to development and regeneration of the musculosketal system include PDZ and LIM domain 3 (PDLIM3), small muscle protein X-linked (SMPX), and leiomodin 2 (LMOD2). Two of these, SMPX and PDLIM3, encode Z-disc associated proteins with putative mechanosensory or stretch signaling roles in striated muscle [114,115]. Expression of MUSTN1, ANKRD1, PDLIM3, and SMPX can be induced by eccentric contraction exercises [116,117,118] or passive stretch [119], suggesting a role in muscle hypertrophy and repair [115]. Although some of these genes possess roles in muscle fiber regeneration, we do not believe that they are indicative of the regenerative process that characterizes later stages of wooden breast. Regeneration of new muscle fibers occurs in response to degeneration and necrosis and is not apparent microscopically until 3 weeks of age [2].
Up-regulation of genes related to hypertrophy in affected birds is in line with higher breast muscle yield in affected birds [4,6,7]. In fact, speculation on the cause of wooden breast and related muscle disorders has focused largely on impaired oxygen supply and buildup of metabolic waste resulting from sustained rapid growth of the pectoralis major [120,121,122,123]. However, up-regulation of genes involved in hypertrophy may also be part of the disease process, causing excessive growth of pectoralis major in affected chickens. Considering the metabolic and physiologic similarities between wooden breast and diabetes, it is possible that mechanisms underlying muscle hypertrophy in wooden breast are similar to those that cause hypertrophy of organs in diabetic complications. For example, diabetic cardiomyopathy, non-alcoholic fatty liver disease, and diabetic nephropathy can cause structural remodeling that includes hypertrophy of the heart, liver, and kidneys respectively [124,125,126].
Interestingly, our data showed up-regulation of several genes, such as MYBPC1 and SMPX [127,128], that are more closely associated with slow-twitch oxidative muscle rather than fast-twitch glycolytic muscle. For example, LMOD2 has been alternatively called cardiac leiomodin and its levels in cardiac muscle are directly linked to the length of actin-containing thin filaments due to competition for binding with tropomodulin-1 [129]. Overexpression of LMOD2 in the heart results in elongation of thin filaments and reduced cardiac function as proper thin filament length is necessary to generate contractile force [129,130]. Similarly, ANKRD1 was previously named cardiac ankyrin repeat protein (CARP) and has been proposed as a marker of cardiac hypertrophy due to its increased expression in three distinct models of cardiac hypertrophy in rats [131]. The lipid transporter FABP3, which is involved in the uptake, intracellular metabolism, and transport of long-chain fatty acids, is most abundantly expressed in slow-twitch skeletal and cardiac muscle of humans rather than fast-twitch skeletal muscle [66]. Up-regulation of these genes is consistent with previous reports of fiber-type switching in 7-week-old birds with wooden breast [11] and may suggest that the pectoralis major muscle of affected birds resembles cardiac or slow-twitch muscle at the transcriptional level.
4.4. Comparison with Prior Gene Expression Study of Early Stages of Wooden Breast
A considerable number of differentially expressed genes from the current study were previously identified by Papah et al. [12], who studied the early pathogenesis of wooden breast in male broilers. Of the 20 genes that were previously identified, the majority were found to be differentially expressed in 3-week-old birds rather than 2-week-old birds (Table 3). A key difference between these studies is the use of male birds in the previous study and female birds in the present experiment. Methodological discrepancies, such as broiler line and severity of disease in affected birds, prevent us from drawing conclusions from these two studies about sex-linked differences in global gene expression that might help explain the higher prevalence and severity of wooden breast among male birds compared to females [8]. However, it has been suggested that increased expression of genes related to fat metabolism and deposition in the pectoralis major of male broilers at 3 weeks of age may contribute to their increased susceptibility [132]. Incomplete dosage compensation in male broilers, which are homogametic and possess two Z chromosomes, may also play an important role [132]. The only Z-linked gene found to be differentially expressed in our results is LPL, which encodes a rate-limiting catalyst for lipoprotein triglyceride hydrolysis [52,53]. Up-regulation of LPL in the pectoralis major of affected birds may increase the rate that lipids are taken up by the muscle, making it a critical gene for ectopic lipid deposition.

Table 3.
Comparison of differentially expressed genes with previous study of wooden breast in male broilers. A total of 20 genes from the current study were also previously identified at early stages of wooden breast development in 2- and 3-week-old male broilers by Papah et al. [12].
Our results highlight the importance of increased lipid metabolism in governing susceptibility to wooden breast and establish high expression of lipid metabolism genes as a much earlier signature of the disease than was previously believed [12]. This is consistent with previous reports of metabolic perturbations in wooden breast [11,12,13], especially a recent study suggesting that the increased ability to direct alimentary resources, particularly fatty acids, to the pectoralis major muscle may underlie susceptibility to wooden breast [5]. While our findings reveal strong parallels between wooden breast in broilers and metabolic syndrome in humans, it is important to recognize that wooden breast is not associated with increased visceral adiposity [5,133] or elevated blood glucose levels [9]. Additionally, lipotoxicity in human skeletal muscle causes muscle atrophy [134] while wooden breast-affected birds have a significantly larger pectoralis major muscle compared to unaffected birds [6]. It is important to investigate how ectopic lipid accumulation in skeletal muscle can manifest with such disparate phenotypes in humans compared to broilers.
5. Conclusions
The findings of this study show that transcriptional changes associated with early stages of wooden breast disease in 2-week-old birds have significant overlap with genes that are dysregulated in metabolic syndrome in humans. Although the underlying causes of metabolic dysfunction possibly leading to pathological progression of wooden breast remain unknown, this study clearly demonstrates that early up-regulation of lipid metabolism in the pectoralis major is a key feature of the myopathy. Additionally, the finding that PPARγ and several of its transcriptional target genes are expressed higher in affected chickens provides critical insight into the early pathogenesis of wooden breast. Affected birds also show dysregulation of various genes involved in muscle growth and function as well as calcium signaling and ER stress. Additional research is needed to understand the mechanisms underlying the apparent metabolic dysfunction and to investigate the possible link between wooden breast and metabolic syndrome.
Supplementary Materials
The following are available online at https://www.mdpi.com/2073-4425/10/10/746/s1, Table S1: Sequencing and mapping statistics for RNA-Seq analysis of eight wooden breast-affected and six unaffected pectoralis major muscle samples.
Author Contributions
Conceptualization, B.A. and J.A.L.; methodology, B.A. and J.A.L.; software, J.A.L.; validation, B.A., J.A.L. and M.B.P.; formal analysis, J.A.L.; investigation, J.A.L. and M.B.P.; resources, B.A.; data curation, X.X.; writing—original draft preparation, J.A.L.; writing—review and editing, B.A. and M.B.P.; visualization, J.A.L.; supervision, B.A.; project administration, B.A.; funding acquisition, B.A.
Funding
This research was funded by the U.S. Department of Agriculture, Agriculture, and Food Research Initiative competitive grant number 2016-67015-25027.
Acknowledgments
The authors gratefully acknowledge the in-kind support by Cobb–Vantress Inc. in providing chicks and chicken feed for this experiment. We also greatly appreciate assistance with samples and data collection from many graduate and undergraduate students at the University of Delaware Department of Animal and Food Sciences. We would like to acknowledge the support from the University of Delaware Center for Bioinformatics and Computational Biology for utilization of their cluster BioMix, which was made possible through support from the Delaware INBRE (NIH GM103446), the state of Delaware and the Delaware Biotechnology Institute (DBI). RNA-sequencing services at DBI are greatly appreciated.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Gall, S.; Suyemoto, M.M.; Sather, H.M.L.; Sharpton, A.R.; Barnes, H.J.; Borst, L.B. Wooden breast in commercial broilers associated with mortality, dorsal recumbency, and pulmonary disease. Avian Dis. 2019, in press. [Google Scholar]
- Papah, M.B.; Brannick, E.M.; Schmidt, C.J.; Abasht, B. Evidence and role of phlebitis and lipid infiltration in the onset and pathogenesis of Wooden Breast Disease in modern broiler chickens. Avian Pathol. 2017, 46, 623–643. [Google Scholar] [CrossRef] [PubMed]
- Norring, M.; Valros, A.; Valaja, J.; Sihvo, H.; Immonen, K.; Puolanne, E. Wooden breast myopathy links with poorer gait in broiler chickens. Animal 2018, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Mutryn, M.F.; Fu, W.; Abasht, B. Incidence of Wooden Breast Disease and its correlation with broiler performance and ultimate pH of breast muscle. In Proceedings of the XXII European Symposium on Poultry Meat Quality, Nantes, France, 10–13 May 2015. [Google Scholar]
- Abasht, B.; Zhou, N.; Lee, W.R.; Zhuo, Z.; Peripolli, E. The metabolic characteristics of susceptibility to wooden breast disease in chickens with high feed efficiency. Poult. Sci. 2019, 98, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Dalle Zotte, A.; Cecchinato, M.; Quartesan, A.; Bradanovic, J.; Puolanne, E. How does “Wooden Breast” myodegradation affect poultry meat quality? In Proceedings of the 60th International Congress of Meat Science and Technology, Punta Del Este, Uruguay, 17–22 August 2014. [Google Scholar]
- Bailey, R.A.; Watson, K.A.; Bilgili, S.F.; Avendano, S. The genetic basis of pectoralis major myopathies in modern broiler chicken lines. Poult. Sci. 2015, 94, 2870–2879. [Google Scholar] [CrossRef] [PubMed]
- Trocino, A.; Piccirillo, A.; Birolo, M.; Radaelli, G.; Bertotto, D.; Filiou, E.; Petracci, M.; Xiccato, G. Effect of genotype, gender and feed restriction on growth, meat quality and the occurrence of white striping and wooden breast in broiler chickens. Poult. Sci. 2015, 94, 2996–3004. [Google Scholar] [CrossRef] [PubMed]
- Livingston, M.L.; Ferket, P.R.; Brake, J.; Livingston, K.A. Dietary amino acids under hypoxic conditions exacerbates muscle myopathies including wooden breast and white stripping. Poult. Sci. 2019, 98, 1517–1527. [Google Scholar] [CrossRef] [PubMed]
- Sihvo, H.K.; Immonen, K.; Puolanne, E. Myodegeneration with fibrosis and regeneration in the pectoralis major muscle of broilers. Vet. Pathol. 2014, 51, 619–623. [Google Scholar] [CrossRef] [PubMed]
- Mutryn, M.F.; Brannick, E.M.; Fu, W.; Lee, W.R.; Abasht, B. Characterization of a novel chicken muscle disorder through differential gene expression and pathway analysis using RNA-sequencing. BMC Genomics 2015, 16, 1–19. [Google Scholar] [CrossRef]
- Papah, M.B.; Brannick, E.M.; Schmidt, C.J.; Abasht, B. Gene expression profiling of the early pathogenesis of wooden breast disease in commercial broiler chickens using RNA-sequencing. PLoS ONE 2018, 13, e0207346. [Google Scholar] [CrossRef]
- Abasht, B.; Mutryn, M.F.; Michalek, R.D.; Lee, W.R. Oxidative stress and metabolic perturbations in wooden breast disorder in chickens. PLoS ONE 2016, 11, 1–16. [Google Scholar] [CrossRef] [PubMed]
- FastQC. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 20 February 2019).
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Hendrickson, D.G.; Sauvageau, M.; Goff, L.; Rinn, J.L.; Pachter, L. Differential analysis of gene regulation at transcript resolution with RNA-seq. Nat. Biotechnol. 2013, 31, 46–53. [Google Scholar] [CrossRef] [PubMed]
- Wei, T.; Simko, V. R package “corrplot”: Visualization of a correlation matrix (Version 0.84) 2017. Available online: https://cran.r-project.org/web/packages/corrplot/ (accessed on 5 March 2019).
- Beales, P.L.; Elcioglu, N.; Woolf, A.S.; Parker, D.; Flinter, F.A. New criteria for improved diagnosis of Bardet-Biedl syndrome: Results of a population survey. J. Med. Genet. 1999, 36, 437–446. [Google Scholar] [PubMed]
- Forsythe, E.; Beales, P.L. Bardet-Biedl syndrome. Eur. J. Hum. Genet. 2013, 21, 8–13. [Google Scholar] [CrossRef] [PubMed]
- Matsuura, K.; Uesugi, N.; Hijiya, N.; Uchida, T.; Moriyama, M. Upregulated expression of cardiac ankyrin-repeated protein in renal podocytes is associated with proteinuria severity in lupus nephritis. Hum. Pathol. 2007, 38, 410–419. [Google Scholar] [CrossRef]
- Qi, L.; Saberi, M.; Zmuda, E.; Wang, Y.; Altarejos, J.; Zhang, X.; Dentin, R.; Hedrick, S.; Bandyopadhyay, G.; Hai, T.; et al. Adipocyte CREB promotes insulin resistance in obesity. Cell Metab. 2009, 9, 277–286. [Google Scholar] [CrossRef]
- Faustman, D.; Li, X.; Lin, H.Y.; Fu, Y.; Eisenbarth, G.; Avruch, J.; Guo, J. Linkage of faulty major histocompatibility complex class I to autoimmune diabetes. Science 1991, 254, 1756–1761. [Google Scholar] [CrossRef]
- Caplen, N.J.; Patel, A.; Millward, A.; Campbell, R.D.; Ratanachaiyavong, S.; Wong, F.S.; Demaine, A.G. Complement C4 and heat shock protein 70 (HSP70) genotypes and type I diabetes mellitus. Immunogenetics 1990, 32, 427–430. [Google Scholar] [CrossRef]
- Lappas, M. Lower circulating levels of complement split proteins C3a and C4a in maternal plasma of women with gestational diabetes mellitus. Diabet. Med. 2011, 28, 906–911. [Google Scholar] [CrossRef] [PubMed]
- Reynet, C.; Kahn, C.R. Rad: A member of the Ras family overexpressed in muscle of type II diabetic humans. Science 1993, 262, 1441. [Google Scholar] [CrossRef] [PubMed]
- Tan, A.L.M.; Langley, S.R.; Tan, C.F.; Chai, J.F.; Khoo, C.M.; Leow, M.K.S.; Khoo, E.Y.H.; Moreno-Moral, A.; Pravenec, M.; Rotival, M.; et al. Ethnicity-specific skeletal muscle transcriptional signatures and their relevance to insulin resistance in Singapore. J. Clin. Endocrinol. Metab. 2018, 104, 465–486. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Xu, L.; Ye, J.; Li, D.; Wang, W.; Li, X.; Wu, L.; Wang, H.; Guan, F.; Li, P. Cidea promotes hepatic steatosis by sensing dietary fatty acids. Hepatology 2012, 56, 95–107. [Google Scholar] [CrossRef] [PubMed]
- Rich, S.S.; Goodarzi, M.O.; Palmer, N.D.; Langefeld, C.D.; Ziegler, J.; Haffner, S.M.; Bryer-Ash, M.; Norris, J.M.; Taylor, K.D.; Haritunians, T.; et al. A genome-wide association scan for acute insulin response to glucose in Hispanic-Americans: The Insulin Resistance Atherosclerosis Family Study (IRAS FS). Diabetologia 2009, 52, 1326–1333. [Google Scholar] [CrossRef]
- Husi, H.; Van Agtmael, T.; Mullen, W.; Bahlmann, F.H.; Schanstra, J.P.; Vlahou, A.; Delles, C.; Perco, P.; Mischak, H. Proteome-based systems biology analysis of the diabetic mouse aorta reveals major changes in fatty acid biosynthesis as potential hallmark in diabetes mellitus-associated vascular disease. Circ. Cardiovasc. Genet. 2014, 7, 161–170. [Google Scholar] [CrossRef]
- Furuhashi, M.; Saitoh, S.; Shimamoto, K.; Miura, T. Fatty acid-binding protein 4 (FABP4): Pathophysiological insights and potent clinical biomarker of metabolic and cardiovascular diseases. Clin. Med. Insights Cardiol. 2014, 2014, 23–33. [Google Scholar] [CrossRef]
- Westerbacka, J.; Kolak, M.; Kiviluoto, T.; Arkkila, P.; Sire, J.; Hamsten, A.; Fisher, R.M.; Yki-ja, H. Genes involved in fatty acid partitioning and binding, inflammation are overexpressed in the human fatty liver of insulin-resistant subjects. Diabetes 2007, 56, 2759–2765. [Google Scholar] [CrossRef]
- Lim, M.A.; Riedel, H.; Liu, F. Grb10: More than a simple adaptor protein. Front. Biosci. 2007, 9, 387–403. [Google Scholar] [CrossRef]
- Yang, S.; Deng, H.; Zhang, Q.; Xie, J.; Zeng, H.; Jin, X.; Ling, Z.; Shan, Q.; Liu, M.; Ma, Y.; et al. Amelioration of diabetic mouse nephropathy by catalpol correlates with down-regulation of Grb10 adn activation of insulin-like growth factor 1/insulin-like growth factor 1 receptor signaling. PLoS ONE 2016, 11, e0151857. [Google Scholar]
- Abdullah, N.; Abdul Murad, N.A.; Attia, J.; Oldmeadow, C.; Mohd Haniff, E.A.; Syafruddin, S.E.; Abd Jalal, N.; Ismail, N.; Ishak, M.; Jamal, R.; et al. Characterizing the genetic risk for Type 2 diabetes in a Malaysian multi-ethnic cohort. Diabet. Med. 2015, 32, 1377–1384. [Google Scholar] [CrossRef] [PubMed]
- Morris, A.D.P.; Voight, B.F.; Teslovich, T.M.; Ferreira, T.; Segrè, A.V.; Steinthorsdottir, V.; Strawbridge, R.J.; Khan, H.; Grallert, H.; Mahajan, A.; et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 2012, 44, 981–990. [Google Scholar] [PubMed]
- Kang, H.P.; Yang, X.; Chen, R.; Zhang, B.; Corona, E.; Schadt, E.E.; Butte, A.J. Integration of disease-specific single nucleotide polymorphisms, Expression quantitative trait loci and coexpression networks reveal novel candidate genes for type 2 diabetes. Diabetologia 2012, 55, 2205–2213. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Howard, B.V. Lipoprotein metabolism in diabetes mellitus. J. Lipid Res. 1987, 28, 613–628. [Google Scholar]
- Yang, T.; Pang, C.P.; Tsang, M.W.; Lam, C.W.; Poon, P.M.K.; Chan, L.Y.S.; Wu, X.Q.; Tomlinson, B.; Baum, L. Pathogenic mutations of the lipoprotein lipase gene in Chinese patients with hypertriglyceridemic type 2 diabetes. Hum. Mutat. 2003, 21, 453. [Google Scholar] [CrossRef] [PubMed]
- Park, P.J.; Kong, S.W.; Tebaldi, T.; Lai, W.R.; Kasif, S.; Kohane, I.S. Integration of heterogeneous expression data sets extends the role of the retinol pathway in diabetes and insulin resistance. Bioinformatics 2009, 25, 3121–3127. [Google Scholar] [CrossRef]
- Moreno-Viedma, V.; Amor, M.; Sarabi, A.; Bilban, M.; Staffler, G.; Zeyda, M.; Stulnig, T.M. Common dysregulated pathways in obese adipose tissue and atherosclerosis. Cardiovasc. Diabetol. 2016, 15, 1–12. [Google Scholar] [CrossRef]
- Park, K.S.; Ciaraldi, T.P.; Abrams-carter, L.; Mudaliar, S.; Nikoulina, S.E.; Henry, R.R. PPAR-y gene expression is elevated in skeletal muscle of obese and type II diabetic subjects. Diabetes 1997, 46, 1230–1234. [Google Scholar] [CrossRef]
- Zeggini, E.; Scott, L.J.; Saxena, R.; Voight, B.F. Meta-analysis of genome-wide association data and large-scale replication identifies additional susceptibility loci for type 2 diabetes. Nat. Genet. 2008, 40, 638–645. [Google Scholar] [CrossRef] [PubMed]
- Scott, L.J.; Mohlke, K.L.; Bonnycastle, L.L.; Willer, C.J.; Li, Y.; Duren, W.L.; Erdos, M.R.; Stringham, H.M.; Chines, P.S.; Jackson, A.U.; et al. A genome-wide association study of type 2 diabetes in finns detects multiple susceptibility variants. Science 2007, 316, 1341–1345. [Google Scholar] [CrossRef] [PubMed]
- Sanghera, D.K.; Blackett, P.R. Type 2 diabetes genetics: Beyond GWAS. J. Diabetes Metab. 2012, 3, 6948. [Google Scholar] [CrossRef] [PubMed]
- Sladek, R.; Rocheleau, G.; Rung, J.; Dina, C.; Shen, L.; Serre, D.; Boutin, P.; Vincent, D.; Belisle, A.; Hadjadj, S.; et al. A genome-wide association study identifies novel risk loci for type 2 diabetes. Nature 2007, 445, 881–885. [Google Scholar] [CrossRef] [PubMed]
- Stumvoll, M.; Häring, H. The peroxisome proliferator–activated receptor-gamma2 Pro12Ala polymorphism. Diabetes 2002, 51, 2341–2347. [Google Scholar] [CrossRef] [PubMed]
- Pulido, M.R.; Diaz-Ruiz, A.; Jiménez-Gómez, Y.; Garcia-Navarro, S.; Gracia-Navarro, F.; Tinahones, F.; López-Miranda, J.; Frühbeck, G.; Vázquez-Martínez, R.; Malagón, M.M. Rab18 dynamics in adipocytes in relation to lipogenesis, lipolysis and obesity. PLoS ONE 2011, 6, e22931. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.-L.; Katoh, M.; Shibasaki, T.; Minami, K.; Sunaga, Y.; Takahashi, H.; Yokoi, N.; Iwasaki, M.; Miki, T.; Susumu, S. The cAMP sensor Epac2 is a direct target of antidiabetic sulfonylurea drugs. Science 2009, 325, 607–610. [Google Scholar] [CrossRef] [PubMed]
- Nie, J.; DuBois, D.C.; Xue, B.; Jusko, W.J.; Almon, R.R. Effects of high-fat feeding on skeletal muscle gene expression in diabetic Goto-Kakizaki rats. Gene Regul. Syst. Biol. 2017, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Luo, W.; Sun, Y.; Qiao, Y.; Zhang, L.; Zhao, Z.; Lv, S. Wnt/β-catenin signaling mediated-UCH-L1 expression in podocytes of diabetic nephropathy. Int. J. Mol. Sci. 2016, 17, 1404. [Google Scholar] [CrossRef] [PubMed]
- Mead, J.R.; Irvine, S.A.; Ramji, D.P. Lipoprotein lipase: Structure, function, regulation, and role in disease. J. Mol. Med. 2002, 80, 753–769. [Google Scholar] [CrossRef]
- Wang, H.; Eckel, R.H. Lipoprotein lipase: From gene to obesity. Am. J. Physiol. Metab. 2009, 297, E271–E288. [Google Scholar] [CrossRef]
- Bonen, A.; Parolin, M.L.; Steinberg, G.R.; Calles-Escandon, J.; Tandon, N.N.; Glatz, J.F.C.; Luiken, J.J.F.P.; Heigenhauser, G.J.F.; Dyck, D.J. Triacylglycerol accumulation in human obesity and type 2 diabetes is associated with increased rates of skeletal muscle fatty acid transport and increased sarcolemmal FAT/CD36. FASEB J. 2004, 18, 1144–1146. [Google Scholar] [CrossRef]
- Zizola, C.F.; Schwartz, G.J.; Vogel, S. Cellular retinol-binding protein type III is a PPARγ target gene and plays a role in lipid metabolism. Am. J. Physiol. Metab. 2008, 295, E1358–E1368. [Google Scholar] [CrossRef] [PubMed]
- Puri, V.; Ranjit, S.; Konda, S.; Nicoloro, S.M.C.; Straubhaar, J.; Chawla, A.; Chouinard, M.; Lin, C.; Burkart, A.; Corvera, S.; et al. Cidea is associated with lipid droplets and insulin sensitivity in humans. Proc. Natl. Acad. Sci. USA 2008, 105, 7833–7838. [Google Scholar] [CrossRef] [PubMed]
- Martin, S.; Parton, R.G. Characterization of Rab18, a lipid droplet-associated Small GTPase. Methods Enzymol. 2008, 438, 109–129. [Google Scholar]
- Green, C.R.; Wallace, M.; Divakaruni, A.S.; Phillips, S.A.; Murphy, A.N.; Ciaraldi, T.P.; Metallo, C.M. Branched-chain amino acid catabolism fuels adipocyte differentiation and lipogenesis. Nat. Chem. Biol. 2016, 12, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Rutsch, F.; Gailus, S.; Suormala, T.; Fowler, B. LMBRD1: The gene for the cblF defect of vitamin B12 metabolism. J. Inherit. Metab. Dis. 2011, 34, 121–126. [Google Scholar] [CrossRef]
- Zerega, B.; Camardella, L.; Cermelli, S.; Sala, R.; Cancedda, R.; Descalzi Cancedda, F. Avidin expression during chick chondrocyte and myoblast development in vitro and in vivo: regulation of cell proliferation. J. Cell Sci. 2001, 114, 1473–1482. [Google Scholar] [PubMed]
- Szatmari, I.; Töröcsik, D.; Agostini, M.; Nagy, T.; Gurnell, M.; Barta, E.; Chatterjee, K.; Nagy, L. PPARγ regulates the function of human dendritic cells primarily by altering lipid metabolism. Blood 2007, 110, 3271–3280. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, E.M.; Karssen, L.C.; Deelen, J.; Isaacs, A.; Medina-Gomez, C.; Mbarek, H.; Kanterakis, A.; Trompet, S.; Postmus, I.; Verweij, N.; et al. Genome of the Netherlands population-specific imputations identify an ABCA6 variant associated with cholesterol levels. Nat. Commun. 2015, 6. [Google Scholar] [CrossRef]
- Kilpeläinen, T.O.; Bentley, A.R.; Noordam, R.; Sung, Y.J.; Schwander, K.; Winkler, T.W.; Jakupović, H.; Chasman, D.I.; Manning, A.; Ntalla, I.; et al. Multi-ancestry study of blood lipid levels identifies four loci interacting with physical activity. Nat. Commun. 2019, 10, 376. [Google Scholar] [CrossRef]
- Willer, C.J.; Schmidt, E.M.; Sengupta, S.; Peloso, G.M.; Gustafsson, S.; Kanoni, S.; Ganna, A.; Chen, J.; Buchkovich, M.L.; Mora, S.; et al. Discovery and refinement of loci associated with lipid levels. Nat. Genet. 2013, 45, 1274–1285. [Google Scholar]
- Surakka, I.; Horikoshi, M.; Mägi, R.; Sarin, A.-P.; Mahajan, A.; Lagou, V.; Marullo, L.; Ferreira, T.; Miraglio, B.; Timonen, S.; et al. The impact of low-frequency and rare variants on lipid levels. Nat. Genet. 2015, 47, 589–597. [Google Scholar] [CrossRef]
- Heuckeroth, R.O.; Birkenmeier, E.H.; Levin, M.S.; Gordon, J.I. Analysis of the tissue-specific expression, developmental regulation, and linkage relationships of a rodent gene encoding heart fatty acid binding protein. J. Biol. Chem. 1987, 262, 9709–9717. [Google Scholar] [PubMed]
- Jang, M.K.; Son, Y.; Jung, M.H. ATF3 plays a role in adipocyte hypoxia-mediated mitochondria dysfunction in obesity. Biochem. Biophys. Res. Commun. 2013, 431, 421–427. [Google Scholar] [CrossRef] [PubMed]
- Puig-Oliveras, A.; Ramayo-Caldas, Y.; Corominas, J.; Estellé, J.; Pérez-Montarelo, D.; Hudson, N.J.; Casellas, J.; Ballester, J.M.F.M. Differences in muscle transcriptome among pigs phenotypically extreme for fatty acid composition. PLoS ONE 2014, 9, e99720. [Google Scholar] [CrossRef] [PubMed]
- Resnyk, C.W.; Carré, W.; Wang, X.; Porter, T.E.; Simon, J.; Le Bihan-Duval, E.; Duclos, M.J.; Aggrey, S.E.; Cogburn, L.A. Transcriptional analysis of abdominal fat in chickens divergently selected on bodyweight at two ages reveals novel mechanisms controlling adiposity: Validating visceral adipose tissue as a dynamic endocrine and metabolic organ. BMC Genomics 2017, 18, 1–31. [Google Scholar] [CrossRef] [PubMed]
- Turcot, V.; Lu, Y.; Highland, H.M.; Schurmann, C.; Justice, A.E.; Fine, R.S.; Bradfield, J.P.; Esko, T.; Giri, A.; Graff, M.; et al. Protein-altering variants associated with body mass index implicate pathways that control energy intake and expenditure in obesity. Nat. Genet. 2018, 50, 26–35. [Google Scholar] [CrossRef] [PubMed]
- Hong-yan, S.H.I.; Qi, H.E.; Min, C.; Ying-ning, S.U.N.; Hui, L.I.; Ning, W. Effect of HOPX gene overexpression on chicken preadipocyte proliferation. 2015, 48, 1624–1631. [Google Scholar]
- Zhang, K.; Zhang, Z.W.; Wang, W.S.; Yan, X.H.; Li, H.; Wang, N. Cloning and expression of chicken HOPX gene. J. Northeast Agric. Univ. 2012, 43, 46–53. [Google Scholar]
- Chen, Z.; Zhao, T.-J.; Li, J.; Gao, Y.-S.; Meng, F.-G.; Yan, Y.-B.; Zhou, H.-M. Slow skeletal muscle myosin-binding protein-C (MyBPC1) mediates recruitment of muscle-type creatine kinase (CK) to myosin. Biochem. J. 2011, 436, 437–445. [Google Scholar] [CrossRef]
- Liu, M.; Bai, J.; He, S.; Villarreal, R.; Hu, D.; Zhang, C.; Yang, X.; Liang, H.; Slaga, T.J.; Yu, Y.; et al. Grb10 promotes lipolysis and thermogenesis by phosphorylation-dependent feedback inhibition of mTORC1. Cell Metab. 2014, 19, 967–980. [Google Scholar] [CrossRef]
- Lefterova, M.I.; Zhang, Y.; Steger, D.J.; Schupp, M.; Schug, J.; Cristancho, A.; Feng, D.; Zhuo, D.; Stoeckert, C.J.; Liu, X.S.; et al. PPARγ and C/EBP factors orchestrate adipocyte biology via adjacent binding on a genome-wide scale. Genes Dev. 2008, 22, 2941–2952. [Google Scholar] [CrossRef]
- Laplante, M.; Sell, H.; MacNaul, K.L.; Richard, D.; Berger, J.P.; Deshaies, Y. PPAR-gamma activation mediates adipose depot-specific effects on gene expression and lipoprotein lipase activity. Diabetes 2003, 52, 291–299. [Google Scholar] [CrossRef] [PubMed]
- Semple, R.K.; Chatterjee, V.K.K.; Rahilly, S.O. PPARγ and human metabolic disease. J. Clin. Invest. 2006, 116, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Phua, W.W.T.; Wong, M.X.Y.; Liao, Z.; Tan, N.S. An apparent functional consequence in skeletal muscle physiology via peroxisome proliferator-activated receptors. Int. J. Mol. Sci. 2018, 19, 1425. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.X. PPARs: Diverse regulators in energy metabolism and metabolic diseases. Cell Res. 2010, 20, 124–137. [Google Scholar] [CrossRef] [PubMed]
- Mitch, W.E.; Medina, R.; Grieber, S.; May, R.C.; England, B.K.; Price, S.R.; Bailey, J.L.; Goldberg, A.L. Metabolic acidosis stimulates muscle protein degradation by activating the adenosine triphosphate-dependent pathway involving ubiquitin and proteasomes. J. Clin. Invest. 1994, 93, 2127–2133. [Google Scholar] [CrossRef] [PubMed]
- Medina-Gomez, G.; Gray, S.; Vidal-Puig, A. Adipogenesis and lipotoxicity: Role of peroxisome proliferator-activated receptor γ (PPARγ) and PPARγ coactivator-1 (PGC1). Public Health Nutr. 2007, 10, 1132–1137. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.H.; Byrne, K.A.; Reverter, A.; Harper, G.S.; Taniguchi, M.; McWilliam, S.M.; Mannen, H.; Oyama, K.; Lehnert, S.A. Transcriptional profiling of skeletal muscle tissue from two breeds of cattle. Mamm. Genome 2005, 16, 201–210. [Google Scholar] [CrossRef] [PubMed]
- Jeong, J.; Kwon, E.G.; Im, S.K.; Seo, K.S.; Baik, M. Expression of fat deposition and fat removal genes is associated with intramuscular fat content in longissimus dorsi muscle of Korean cattle steers. J. Anim. Sci. 2012, 90, 2044–2053. [Google Scholar] [CrossRef]
- Cui, H.; Zheng, M.; Zhao, G.; Liu, R.; Wen, J. Identification of differentially expressed genes and pathways for intramuscular fat metabolism between breast and thigh tissues of chickens. BMC Genomics 2018, 19, 1–9. [Google Scholar] [CrossRef]
- Liu, L.; Cui, H.; Fu, R.; Zheng, M.; Liu, R.; Zhao, G.; Wen, J. The regulation of IMF deposition in pectoralis major of fast- and slow- growing chickens at hatching. J. Anim. Sci. Biotechnol. 2017, 8, 1–8. [Google Scholar] [CrossRef]
- Tumova, J.; Andel, M.; Trnka, J. Excess of free fatty acids as a cause of metabolic dysfunction in skeletal muscle. Physiol. Res. 2016, 65, 193–207. [Google Scholar] [PubMed]
- Rasouli, N.; Molavi, B.; Elbein, S.C.; Kern, P.A. Ectopic fat accumulation and metabolic syndrome. Diabetes Obes. Metab. 2007, 9, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Mungrue, I.N.; Pagnon, J.; Kohannim, O.; Gargalovic, P.S.; Lusis, A.J. CHAC1/MGC4504 is a novel proapoptotic component of the unfolded protein response, downstream of the ATF4-ATF3-CHOP cascade. J. Immunol. 2009, 182, 466–476. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wan, B.; Li, D.; Zhou, J.; Li, R.; Bai, M.; Chen, F.; Yu, L. BRSK2 is regulated by ER stress in protein level and involved in ER stress-induced apoptosis. Biochem. Biophys. Res. Commun. 2012, 423, 813–818. [Google Scholar] [CrossRef] [PubMed]
- Schröder, M.; Kaufman, R.J. ER stress and the unfolded protein response. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2005, 569, 29–63. [Google Scholar] [CrossRef] [PubMed]
- Volmer, R.; Ron, D. Lipid-dependent regulation of the unfolded protein response. Curr. Opin. Cell Biol. 2015, 33, 67–73. [Google Scholar] [CrossRef]
- Eizirik, D.L.; Cardozo, A.K.; Cnop, M. The role for endoplasmic reticulum stress in diabetes mellitus. Endocr. Rev. 2008, 29, 42–61. [Google Scholar] [CrossRef]
- Basseri, S.; Austin, R.C. Endoplasmic reticulum stress and lipid metabolism: Mechanisms and therapeutic potential. Biochem. Res. Int. 2012, 2012. [Google Scholar] [CrossRef]
- Cnop, M.; Foufelle, F.; Velloso, L.A. Endoplasmic reticulum stress, obesity and diabetes. Trends Mol. Med. 2012, 18, 59–68. [Google Scholar] [CrossRef]
- Zmuda, E.J.; Qi, L.; Zhu, M.X.; Mirmira, R.G.; Montminy, M.R.; Hai, T. The roles of ATF3, an adaptive-response gene, in high-fat-diet-induced diabetes and pancreatic β-cell dysfunction. Mol. Endocrinol. 2010, 24, 1423–1433. [Google Scholar] [CrossRef]
- Aung, H.H.; Lame, M.W.; Gohil, K.; An, C.I.; Wilson, D.W.; Rutledge, J.C. Induction of ATF3 gene network by triglyceride-rich lipoprotein lipolysis products increases vascular apoptosis and inflammation. Arterioscler. Thromb. Vasc. Biol. 2013, 33, 2088–2096. [Google Scholar] [CrossRef] [PubMed]
- Kalfon, R.; Koren, L.; Aviram, S.; Schwartz, O.; Hai, T.; Aronheim, A. ATF3 expression in cardiomyocytes preserves homeostasis in the heart and controls peripheral glucose tolerance. Cardiovasc. Res. 2017, 113, 134–146. [Google Scholar] [CrossRef]
- Kim, J.Y.; Park, K.J.; Hwang, J.Y.; Kim, G.H.; Lee, D.Y.; Lee, Y.J.; Song, E.H.; Yoo, M.G.; Kim, B.J.; Suh, Y.H.; et al. Activating transcription factor 3 is a target molecule linking hepatic steatosis to impaired glucose homeostasis. J. Hepatol. 2017, 67, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Song, N.J.; Bahn, G.; Chang, S.H.; Yun, U.J.; Ku, J.M.; Jo, D.G.; Park, K.W. Atf3 induction is a therapeutic target for obesity and metabolic diseases. Biochem. Biophys. Res. Commun. 2018, 504, 903–908. [Google Scholar] [CrossRef] [PubMed]
- Hyun, B.K.; Kong, M.; Tae, M.K.; Young, H.S.; Kim, W.H.; Joo, H.L.; Ji, H.S.; Myeong, H.J. NFATc4 and ATF3 negatively regulate adiponectin gene expression in 3T3-L1 adipocytes. Diabetes 2006, 55, 1342–1352. [Google Scholar]
- Finlin, B.S.; Crump, S.M.; Satin, J.; Andres, D.A. Regulation of voltage-gated calcium channel activity by the Rem and Rad GTPases. Proc. Natl. Acad. Sci. USA 2003, 100, 14469–14474. [Google Scholar] [CrossRef] [PubMed]
- Moyers, J.S.; Bilan, P.J.; Zhu, J.; Kahn, C.R. Rad and Rad-related GTPases interact with calmodulin and calmodulin- dependent protein kinase II. J. Biol. Chem. 1997, 272, 11832–11839. [Google Scholar] [CrossRef]
- Moyers, J.S.; Bilan, P.J.; Reynet, C.; Ronald Kahn, C. Overexpression of Rad inhibits glucose uptake in cultured muscle and fat cells. J. Biol. Chem. 1996, 271, 23111–23116. [Google Scholar] [CrossRef]
- Ilany, J.; Bilan, P.J.; Kapur, S.; Caldwell, J.S.; Patti, M.-E.; Marette, A.; Kahn, C.R. Overexpression of Rad in muscle worsens diet-induced insulin resistance and glucose intolerance and lowers plasma triglyceride level. Proc. Natl. Acad. Sci. USA 2006, 103, 4481–4486. [Google Scholar] [CrossRef]
- Randle, P.J.; Priestman, D.A.; Mistry, S.C.; Halsall, A. Glucose fatty acid interactions and the regulation of glucose disposal. J. Cell. Biochem. 1994, 55, 1–11. [Google Scholar] [CrossRef]
- Arruda, A.P.; Hotamisligil, G.S. Calcium homeostasis and organelle function in the pathogenesis of obesity and diabetes. Cell Metab. 2015, 22, 381–397. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Wang, D.; Gentile, C.L.; Pagliassotti, M.J. Reduced endoplasmic reticulum luminal calcium links saturated fatty acid-mediated endoplasmic reticulum stress and cell death in liver cells. Mol. Cell. Biochem. 2009, 331, 31–40. [Google Scholar] [CrossRef] [PubMed]
- Rys-Sikora, K.E.; Gill, D.L. Fatty acid-mediated calcium sequestration within intracellular calcium pools. J. Biol. Chem. 1998, 273, 32627–32635. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kee, H.J.; Kim, J.R.; Nam, K.I.; Hye, Y.P.; Shin, S.; Jeong, C.K.; Shimono, Y.; Takahashi, M.; Myung, H.J.; Kim, N.; et al. Enhancer of polycomb1, a novel homeodomain only protein-binding partner, induces skeletal muscle differentiation. J. Biol. Chem. 2007, 282, 7700–7709. [Google Scholar] [CrossRef] [PubMed]
- Shin, C.H.; Liu, Z.P.; Passier, R.; Zhang, C.L.; Wang, D.Z.; Harris, T.M.; Yamagishi, H.; Richardson, J.A.; Childs, G.; Olson, E.N. Modulation of cardiac growth and development by HOP, an unusual homeodomain protein. Cell 2002, 110, 725–735. [Google Scholar] [CrossRef]
- Kojic, S.; Nestorovic, A.; Rakicevic, L.; Belgrano, A.; Stankovic, M.; Divac, A.; Faulkner, G. A novel role for cardiac ankyrin repeat protein Ankrd1/CARP as a co-activator of the p53 tumor suppressor protein. Arch. Biochem. Biophys. 2010, 502, 60–67. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Zhang, Y.; Ma, G.; Zhao, X.; Chen, Y.; Zhu, D. Identification of gene expression modifications in myostatin-stimulated myoblasts. Biochem. Biophys. Res. Commun. 2005, 326, 660–666. [Google Scholar] [CrossRef]
- Ma, G.; Wang, H.; Gu, X.; Li, W.; Zhang, X.; Cui, L.; Li, Y.; Zhang, Y.; Zhao, B.; Li, K. CARP, a myostatin-downregulated gene in CFM cells, is a novel essential positive regulator of myogenesis. Int. J. Biol. Sci. 2014, 10, 309–320. [Google Scholar] [CrossRef]
- Bagnall, R.D.; Yeates, L.; Semsarian, C. Analysis of the Z-disc genes PDLIM3 and MYPN in Patients with Hypertrophic Cardiomyopathy. Int. J. Cardiol. 2010, 145, 601–602. [Google Scholar] [CrossRef] [PubMed]
- Eftestøl, E.; Norman Alver, T.; Gundersen, K.; Bruusgaard, J.C. Overexpression of SMPX in adult skeletal muscle does not change skeletal muscle fiber type or size. PLoS ONE 2014, 9, e99232. [Google Scholar] [CrossRef]
- Kostek, M.C.; Chen, Y.-W.; Cuthbertson, D.J.; Shi, R.; Fedele, M.J.; Esser, K.A.; Rennie, M.J. Gene expression responses over 24 h to lengthening and shortening contractions in human muscle: major changes in CSRP3, MUSTN1, SIX1, and FBXO32. Physiol. Genomics 2007, 31, 42–52. [Google Scholar] [CrossRef] [PubMed]
- Barash, I.A.; Mathew, L.; Ryan, A.F.; Chen, J.; Lieber, R.L. Rapid muscle-specific gene expression changes after a single bout of eccentric contractions in the mouse. Am. J. Physiol. Physiol. 2004, 286, C355–C364. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.W.; Nader, G.A.; Baar, K.R.; Fedele, M.J.; Hoffman, E.P.; Esser, K.A. Response of rat muscle to acute resistance exercise defined by transcriptional and translational profiling. J. Physiol. 2002, 545, 27–41. [Google Scholar] [CrossRef] [PubMed]
- Kemp, T.J.; Sadusky, T.J.; Simon, M.; Brown, R.; Eastwood, M.; Sassoon, D.A.; Coulton, G.R. Identification of a novel stretch-responsive skeletal muscle gene (Smpx). Genomics 2001, 72, 260–271. [Google Scholar] [CrossRef] [PubMed]
- MacRae, V.E.; Mahon, M.; Gilpin, S.; Sandercock, D.A.; Mitchell, M.A. Skeletal muscle fibre growth and growth associated myopathy in the domestic chicken (Gallus domesticus). Br. Poult. Sci. 2006, 47, 264–272. [Google Scholar] [CrossRef] [PubMed]
- Mudalal, S.; Lorenzi, M.; Soglia, F.; Cavani, C.; Petracci, M. Implications of white striping and wooden breast abnormalities on quality traits of raw and marinated chicken meat. Animal 2015, 9, 728–734. [Google Scholar] [CrossRef] [PubMed]
- Kuttappan, V.A.; Shivaprasad, H.L.; Shaw, D.P.; Valentine, B.A.; Hargis, B.M.; Clark, F.D.; McKee, S.R.; Owens, C.M. Pathological changes associated with white striping in broiler breast muscles. Poult. Sci. 2013, 92, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Lilburn, M.S.; Griffin, J.R.; Wick, M. From muscle to food: Oxidative challenges and developmental anomalies in poultry breast muscle. Poult. Sci. 2018, 98, 4255–4260. [Google Scholar] [CrossRef] [PubMed]
- Satriano, J. Kidney growth, hypertrophy and the unifying mechanism of diabetic complications. Amino Acids 2007, 33, 331–339. [Google Scholar] [CrossRef]
- Sharma, V.; McNeill, J.H. Diabetic cardiomapathy: Where are we 40 years later? Can. J. Cardiol. 2006, 22, 305–308. [Google Scholar] [CrossRef]
- Liang, W.; Menke, A.L.; Driessen, A.; Koek, G.H.; Lindeman, J.H.; Stoop, R.; Havekes, L.M.; Kleemann, R.; Van Den Hoek, A.M. Establishment of a general NAFLD scoring system for rodent models and comparison to human liver pathology. PLoS ONE 2014, 9, 1–17. [Google Scholar] [CrossRef]
- Jingting, S.; Qin, X.; Yanju, S.; Ming, Z.; Yunjie, T.; Gaige, J.; Zhongwei, S.; Jianmin, Z. Oxidative and glycolytic skeletal muscles show marked differences in gene expression profile in Chinese Qingyuan partridge chickens. PLoS ONE 2017, 12, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Palmer, S.; Groves, N.; Schindeler, A.; Yeoh, T.; Biben, C.; Wang, C.-C.; Sparrow, D.B.; Barnett, L.; Jenkins, N.A.; Copeland, N.G.; et al. The Small Muscle-specific Protein Csl Modifies Cell Shape and Promotes Myocyte Fusion in an. J. Cell Biol. 2001, 153, 985. [Google Scholar] [CrossRef] [PubMed]
- Tsukada, T.; Pappas, C.T.; Moroz, N.; Antin, P.B.; Kostyukova, A.S.; Gregorio, C.C. Leiomodin-2 is an antagonist of tropomodulin-1 at the pointed end of the thin filaments in cardiac muscle. J. Cell Sci. 2010, 123, 3136–3145. [Google Scholar] [CrossRef] [PubMed]
- Pappas, C.T.; Farman, G.P.; Mayfield, R.M.; Konhilas, J.P.; Gregorio, C.C. Cardiac-specific knockout of Lmod2 results in a severe reduction in myofilament force production and rapid cardiac failure. J. Mol. Cell. Cardiol. 2018, 122, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Aihara, Y.; Kurabayashi, M.; Saito, Y.; Ohyama, Y.; Tanaka, T.; Takeda, S.; Tomaru, K.; Sekiguchi, K.; Arai, M.; Nakamura, T.; et al. Cardiac ankyrin repeat protein is a novel marker of cardiac hypertrophy: Role of M-CAT element within the promoter. Hypertension 2000, 36, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Brothers, B.; Zhuo, Z.; Papah, M.B.; Abasht, B. RNA-seq analysis reveals spatial and sex differences in pectoralis major muscle of broiler chickens contributing to difference in susceptibility to wooden breast disease. Front. Physiol. 2019, 10, 764. [Google Scholar] [CrossRef]
- Zhuo, Z.; Lamont, S.J.; Lee, W.R.; Abasht, B. RNA-seq analysis of abdominal fat reveals differences between modern commercial broiler chickens with high and low feed efficiencies. PLoS ONE 2015, 10, e0135810. [Google Scholar] [CrossRef]
- Tamilarasan, K.P.; Temmel, H.; Das, S.K.; Al Zoughbi, W.; Schauer, S.; Vesely, P.W.; Hoefler, G. Skeletal muscle damage and impaired regeneration due to LPL-mediated lipotoxicity. Cell Death Dis. 2012, 3, e354–e358. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).