Behavioral Changes in Stem-Cell Potency by HepG2-Exhausted Medium
Abstract
1. Introduction
2. Materials and Methods
2.1. WJ-MSC Isolation and Culture
2.2. HepG2
2.3. Treatment and Preparation of WJ-MSCs
2.4. RNA Extraction and Quantitative Polymerase Chain Reaction
2.5. RNA Extraction and Quantitative Polymerase Chain Reaction for miRNA
2.6. Exosome Experiments
2.7. Quantitative-PCR Analysis
2.8. Gene Expression: Statistical Analysis and Real-Time PCR Data Analysis
2.9. miRNA: Statistical Analysis and Real-Time PCR Data Analysis
2.10. Immunostaining
2.11. Alizarin Red Assay
3. Results
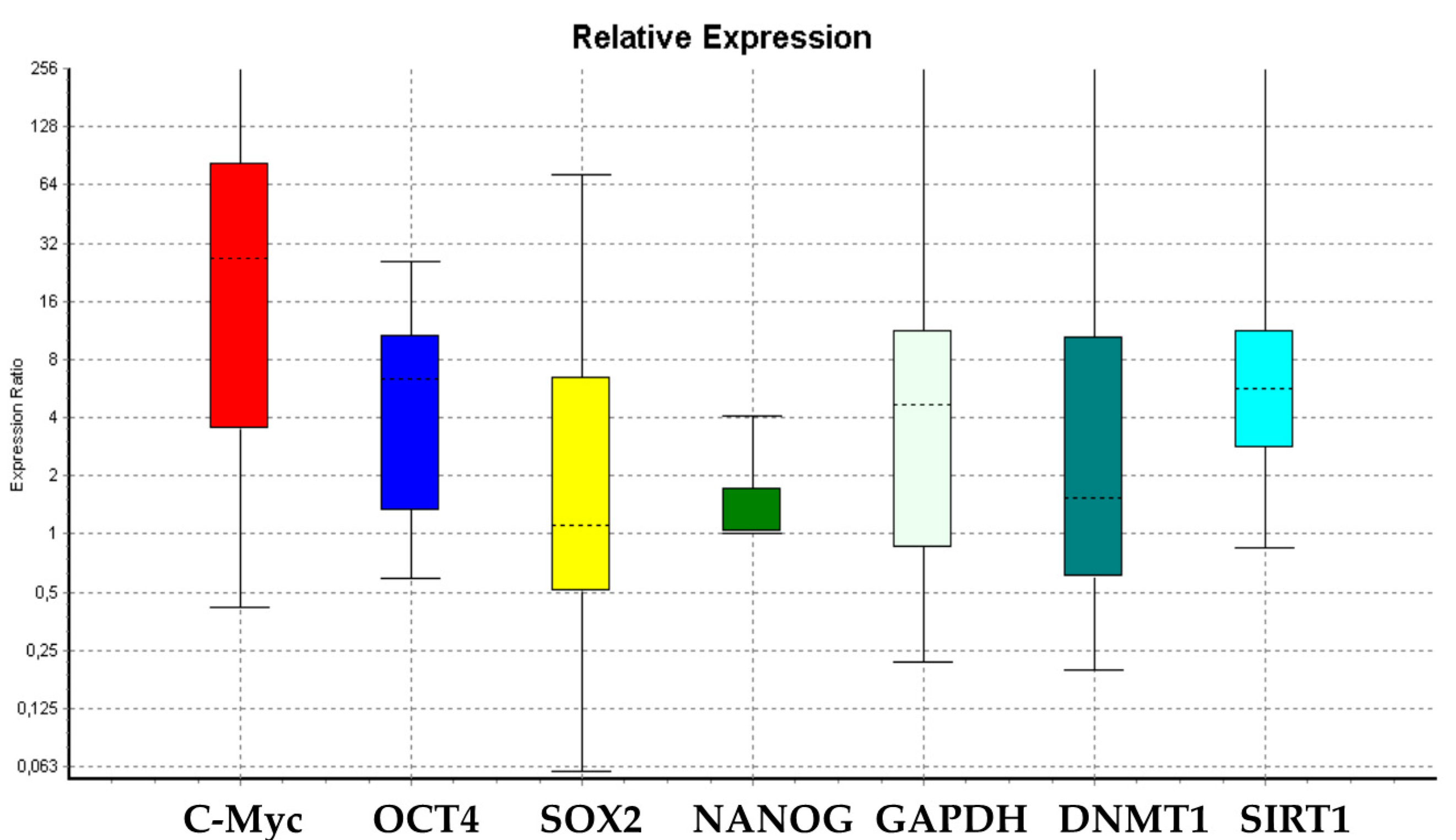
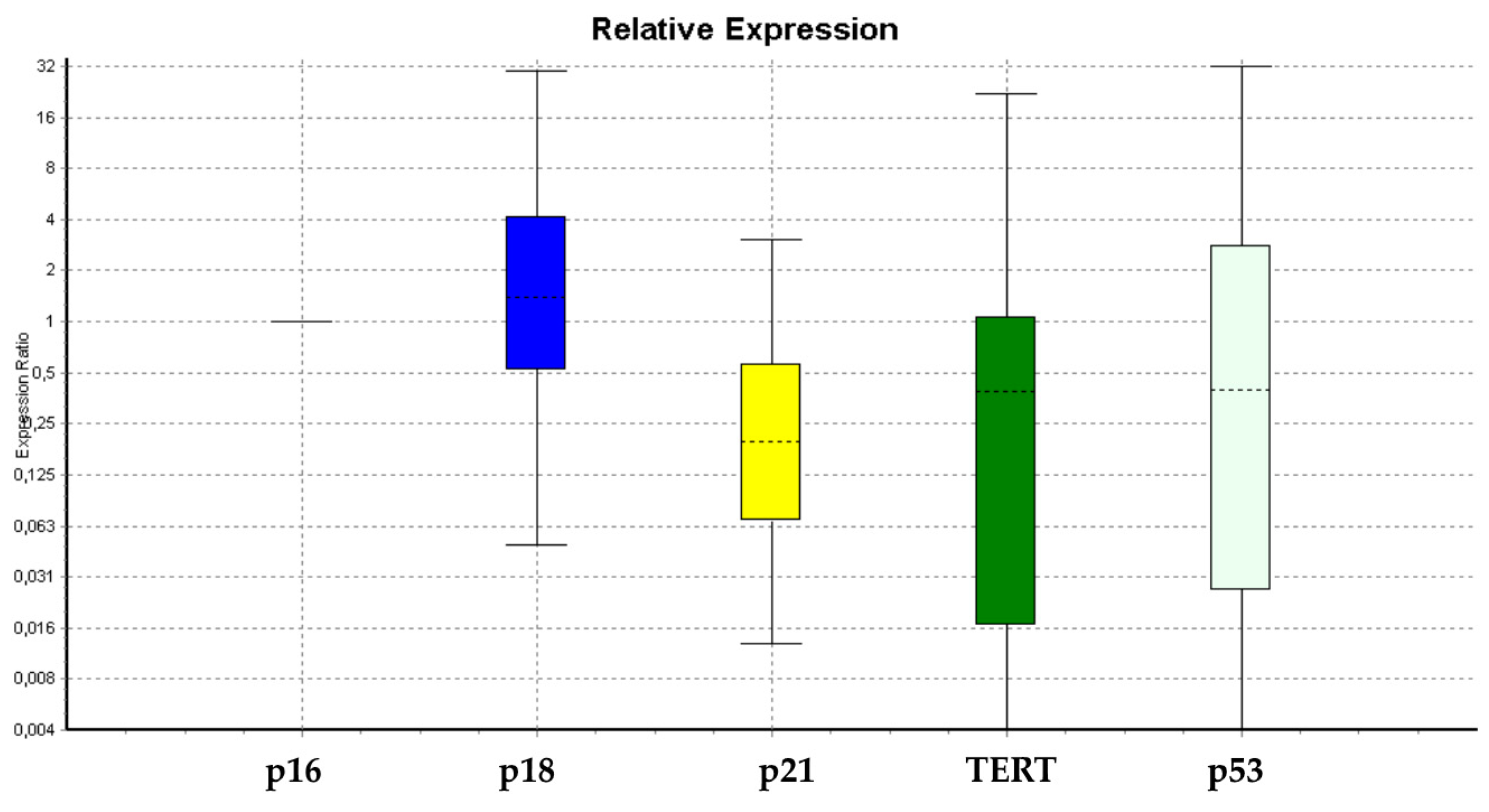
3.1. Gene Expression
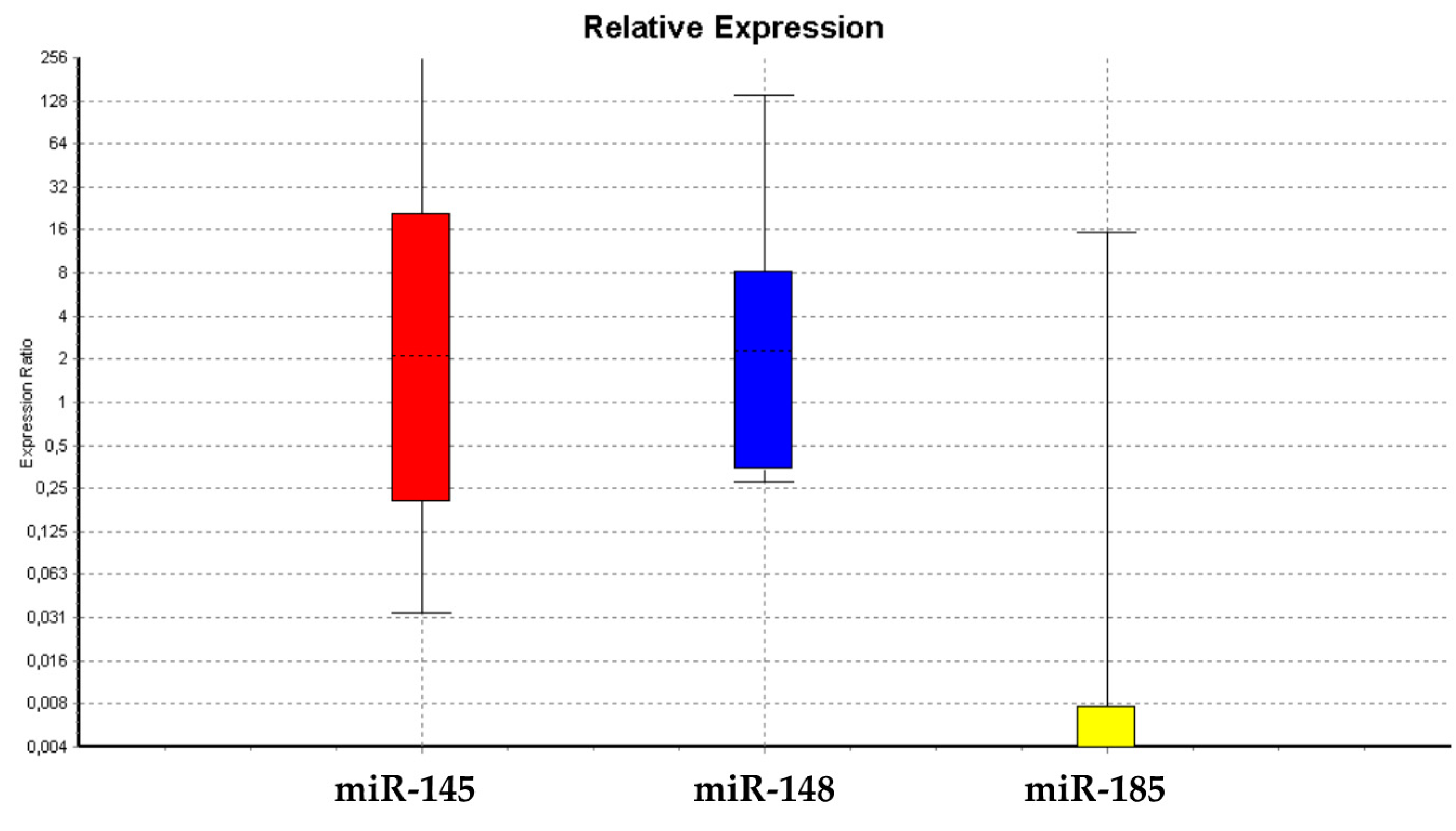
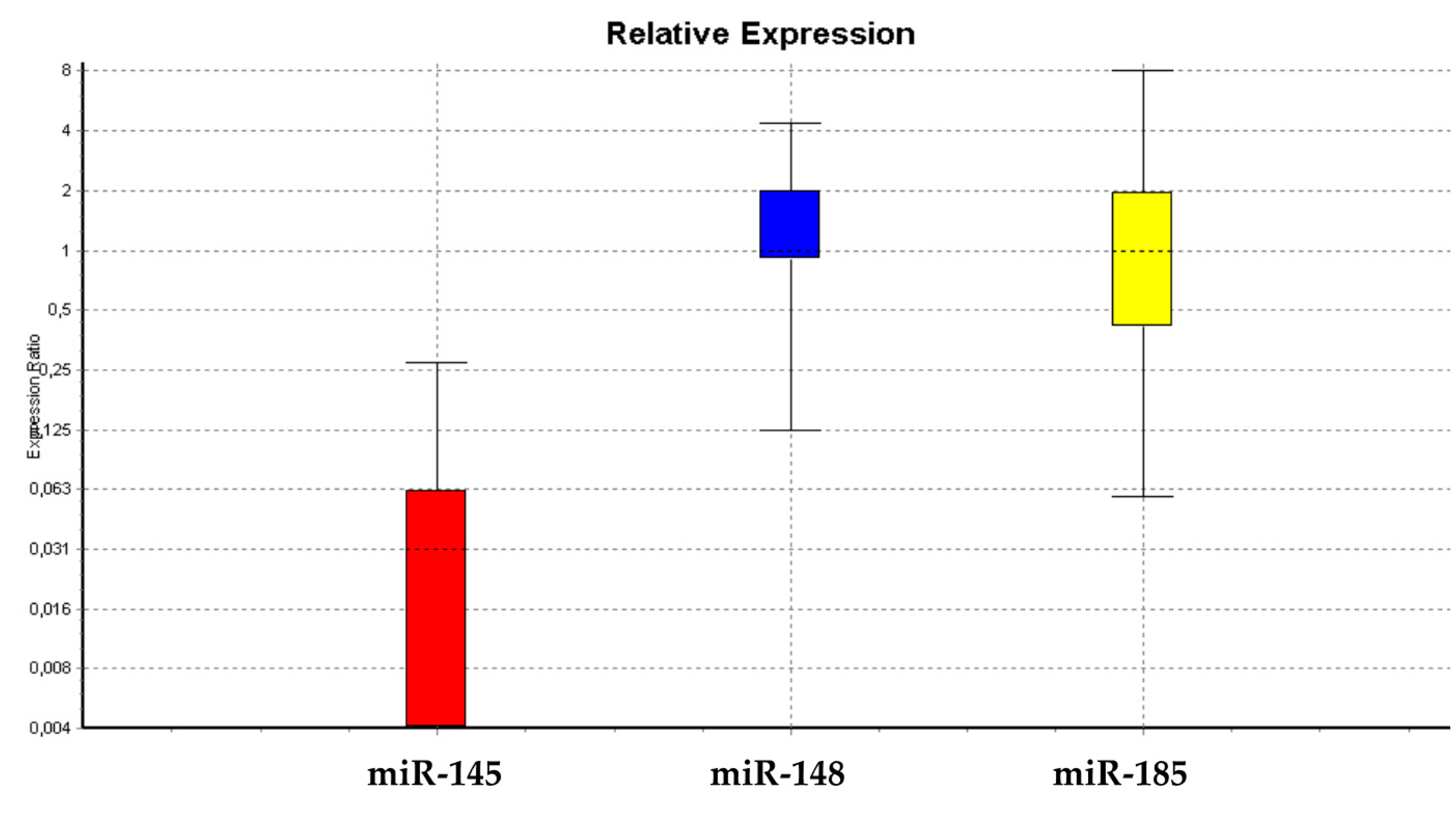
3.1.1. miRNA
3.1.2. Exosomes
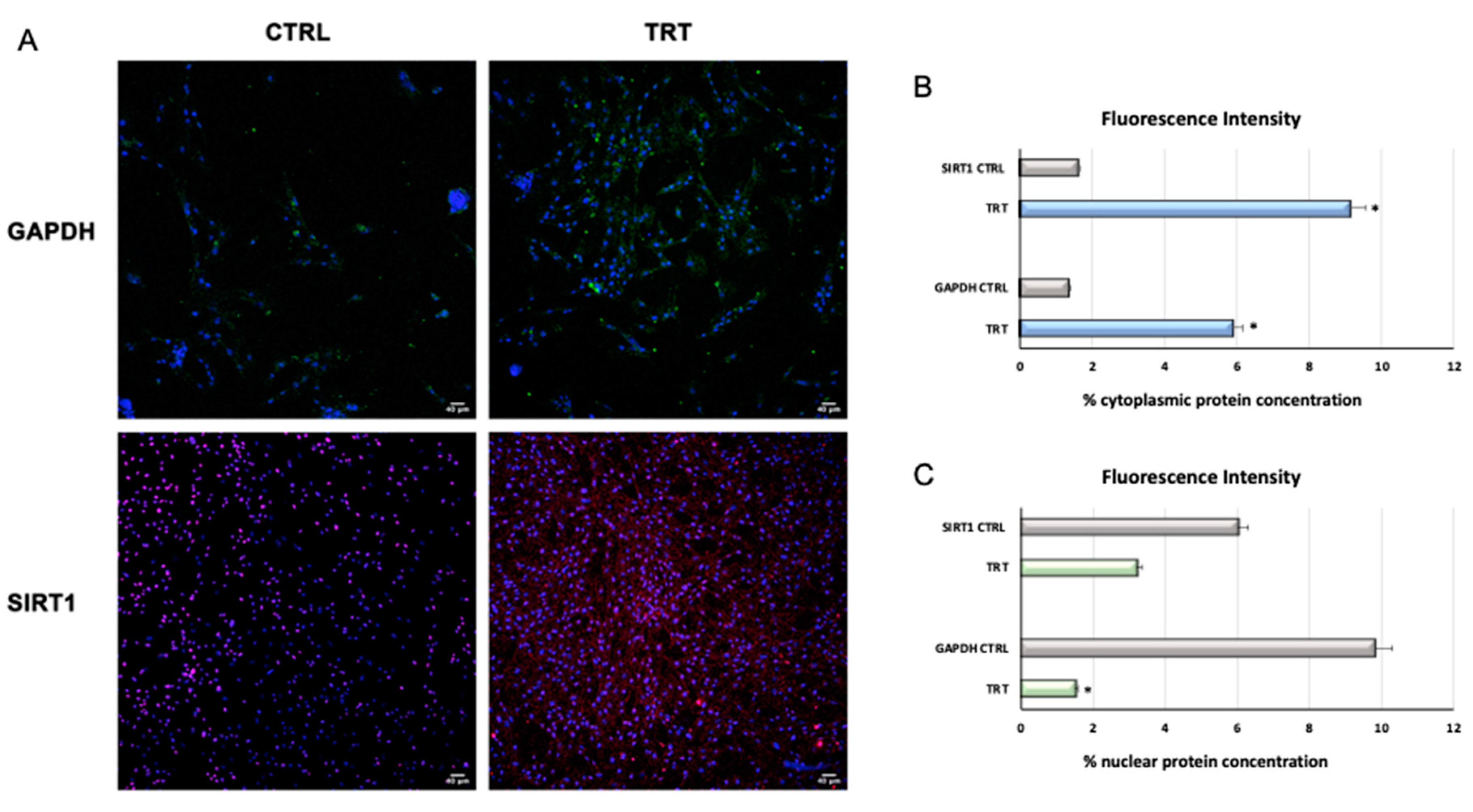
3.1.3. Immunostaining
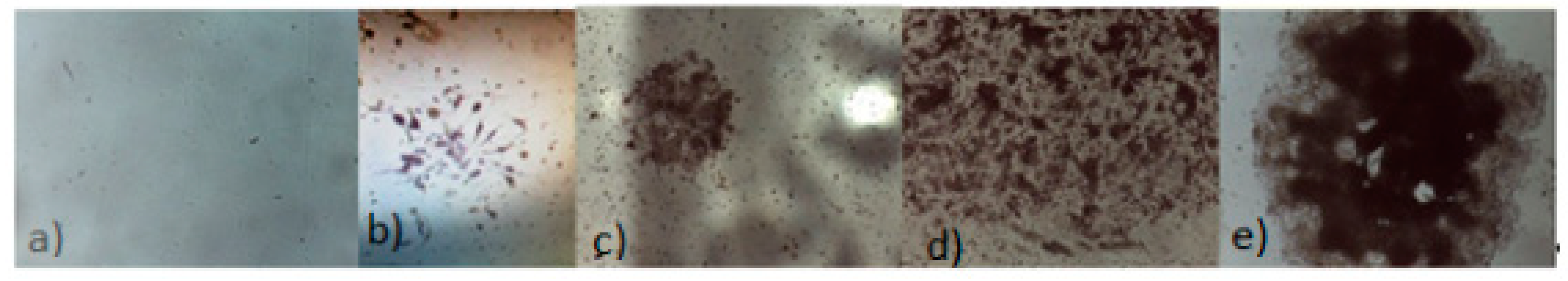
3.1.4. Alizarin Red Assay
3.1.5. Optical Microscope Analysis
4. Discussion
4.1. c-Myc’s Power
4.2. MicroRNA’s Power
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Maioli, M.; Basoli, V.; Santaniello, S.; Cruciani, S.; Delitala, A.P.; Pinna, R.; Milia, E.; Grillari-Voglauer, R.; Fontani, V.; Rinaldi, S.; et al. Osteogenesis from Dental Pulp Derived Stem Cells: A Novel Conditioned Medium Including Melatonin within a Mixture of Hyaluronic, Butyric, and Retinoic Acids. Stem Cells Int. 2016, 2016, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Rinaldi, S.; Maioli, M.; Santaniello, S.; Pigliaru, G.; Ventura, C.; Montella, A.; Sanna, R.; Bandiera, P.; Bagella, L.; Delitala, A.P.; et al. Amniotic fluid stem cells morph into a cardiovascular lineage: Analysis of a chemically induced cardiac and vascular commitment. Drug Des. Dev. Ther. 2013, 7, 1063–1073. [Google Scholar] [CrossRef][Green Version]
- Plaks, V.; Kong, N.; Werb, Z. The cancer stem cell niche: How essential is the niche in regulating stemness of tumor cells? Cell Stem Cell 2015, 16, 225–338. [Google Scholar] [CrossRef] [PubMed]
- Montanucci, P.; Pescara, T.; Greco, A.; Leonardi, G.; Marini, L.; Basta, G.; Calafiore, R. Co-Microencapsulation of human umbilical cord-derived mesenchymal stem and pancreatic islet-derived insulin producing cells: A new experimental approach for the cell therapy of Type 1 Diabetes Mellitus (T1D). Diabetes/Metab. Res. Rev. 2020, e3372. [Google Scholar] [CrossRef]
- Wright, A.; Snyder, L.J.; Knights, K.; He, H.; Springer, N.L.; Lillich, J.; Weiss, M. A Protocol for the Isolation, Culture, and Cryopreservation of Umbilical Cord-Derived Canine Mesenchymal Stromal Cells: Role of Cell Attachment in Long-Term Maintenance. Stem Cells Dev. 2020, 29, 695–713. [Google Scholar] [CrossRef]
- Qu, W.; Wang, Z.; Hare, J.M.; Bu, G.; Mallea, J.M.; Pascual, J.M.; Caplan, A.; Kurtzberg, J.; Zubair, A.; Kubrova, E.; et al. Cell-based therapy to reduce mortality from COVID -19: Systematic review and meta-analysis of human studies on acute respiratory distress syndrome. Stem Cells Transl. Med. 2020, 10. [Google Scholar] [CrossRef]
- Chen, J.; Hu, C.; Chen, L.; Tang, L.; Zhu, Y.; Xu, X.; Chen, L.; Gao, H.; Lu, X.; Yu, L.; et al. Clinical study of mesenchymal stem cell treating acute respiratory distress syndrome induced by epidemic influenza a (H7N9) infection, a hint for COVID-19 treatment. Engineering 2020. [Google Scholar] [CrossRef]
- Zhang, Y.; Ding, J.; Ren, S.; Wang, W.; Yang, Y.; Li, S.; Meng, M.; Wu, T.; Liu, D.; Tian, S.; et al. Intravenous infusion of human umbilical cord Wharton’s jelly-derived mesenchymal stem cells as a potential treatment for patients with COVID-19 pneumonia. Stem Cell Res. Ther. 2020, 11, 1–6. [Google Scholar] [CrossRef]
- De Gregorio, C.; Contador, D.; Díaz, D.; Cárcamo, C.; Santapau, D.; Lobos-Gonzalez, L.; Acosta, C.; Campero, M.; Carpio, D.; Gabriele, C.; et al. Human adipose-derived mesenchymal stem cell-conditioned medium ameliorates polyneuropathy and foot ulceration in diabetic BKS db/db mice. Stem Cell Res. Ther. 2020, 11, 1–21. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of pluripotent stem cellsfrom mouse embryonic and adult fibroblast cultures by definedfactors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef]
- Warren, L.; Manos, P.D.; Ahfeldt, T.; Loh, Y.-H.; Li, H.; Lau, F.; Ebina, W.; Mandal, P.; Smith, Z.D.; Meissner, A.; et al. Highly efficientreprogramming to pluripotency and directed differentiation ofhuman cells with synthetic modified mRNA. Cell Stem Cell 2010, 7, 618–630. [Google Scholar] [CrossRef] [PubMed]
- Rais, Y.; Zviran, A.; Geula, S.; Gafni, O.; Chomsky, E.; Viukov, S.; Mansour, A.A.; Caspi, I.; Krupalnik, V.; Zerbib, M.; et al. Deterministic directreprogramming of somatic cells to pluripotency. Nature 2013, 502, 65–70. [Google Scholar] [CrossRef]
- Polo, J.M.; Anderssen, E.; Walsh, R.M.; Schwarz, B.A.; Nefzger, C.M.; Lim, S.M.; Borkent, M.; Apostolou, E.; Alaei, S.; Cloutier, J.; et al. A molecular roadmapof reprogramming somatic cells into iPS cells. Cell 2012, 151, 1617–1632. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Papagiannakopoulos, T.; Pan, G.; Thomson, J.A.; Kosik, K.S. MicroRNA-145 rgulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009, 137, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Sommer, C.A.; Stadtfeld, M.; Murphy, G.J.; Hochedlinger, K.; Kotton, D.N.; Mostoslavsky, G. Induced pluripotent stem cell generation using a single lentiviral stem cell cassette. Stem Cells 2009, 27, 543–549. [Google Scholar] [CrossRef] [PubMed]
- Beer, S.; Zetterberg, A.; Ihrie, R.A.; McTaggart, R.A.; Yang, Q.; Bradon, N.; Arvanitis, C.; Attardi, L.D.; Feng, S.; Ruebner, B.; et al. Developmental Context Determines Latency of MYC-Induced Tumorigenesis. PLoS Boil. 2004, 2, e332. [Google Scholar] [CrossRef]
- Stine, Z.E.; Walton, Z.E.; Altman, B.J.; Hsieh, A.L.; Dang, C.V. MYC, Metabolism, and Cancer. Cancer Discov. 2015, 5, 1024–1039. [Google Scholar] [CrossRef]
- Conacci-Sorrell, M.; McFerrin, L.; Eisenman, R.N. An overview of MYC and its interactome. Cold Spring Harb. Perspect. Med. 2014, 4, a014357. [Google Scholar] [CrossRef]
- Dang, C.V. MYC on the Path to Cancer. Cell 2012, 149, 22–35. [Google Scholar] [CrossRef]
- Lin, C.-P.; Liu, C.-R.; Lee, C.-N.; Chan, T.-S.; Liu, H.E. Targeting c-Myc as a novel approach for hepatocellular carcinoma. World J. Hepatol. 2010, 2, 16–20. [Google Scholar] [CrossRef]
- Korc, M. Beyond Kras: MYC Rules in Pancreatic Cancer. Cell. Mol. Gastroenterol. Hepatol. 2018, 6, 223–224. [Google Scholar] [CrossRef] [PubMed]
- Gabay, M.; Li, Y.; Felsher, D.W. MYC Activation Is a Hallmark of Cancer Initiation and Maintenance. Cold Spring Harb. Perspect. Med. 2014, 4, a014241. [Google Scholar] [CrossRef] [PubMed]
- Arand, J.; Spieler, D.; Karius, T.; Branco, M.R.; Meilinger, D.; Meissner, A.; Jenuwein, T.; Xu, G.; Leonhardt, H.; Wolf, V.; et al. In Vivo Control of CpG and Non-CpG DNA Methylation by DNA Methyltransferases. PLoS Genet. 2012, 8, e1002750. [Google Scholar] [CrossRef] [PubMed]
- Ehrlich, M. DNA methylation in cancer: Too much, but also too little. Oncogene 2002, 21, 5400–5413. [Google Scholar] [CrossRef]
- Garzon, R.; Liu, S.; Fabbri, M.; Liu, Z.; Heaphy, C.E.; Callegari, E.; Schwind, S.; Pang, J.; Yu, J.; Muthusamy, N.; et al. MicroRNA-29binduces global DNA hypomethylation and tumor suppressor gene reexpression in acute myeloid leukemia by targeting directly DNMT3A and3B and indirectly DNMT1. Blood 2009, 113, 6411–6418. [Google Scholar] [CrossRef]
- Mudbhary, R.; Hoshida, Y.; Chernyavskaya, Y.; Jacob, V.; Villanueva, A.; Fiel, M.I.; Chen, X.; Kojima, K.; Thung, S.; Bronson, R.T.; et al. UHRF1 overexpression drives DNA hypomethylation and hepatocellular carcinoma. Cancer Cell 2014, 25, 196–209. [Google Scholar] [CrossRef]
- Schöler, H.R.; Balling, R.; Hatzopoulos, A.; Suzuki, N.; Gruss, P. Octamer binding proteins confer transcriptional activity in early mouse embryogenesis. EMBO J. 1989, 8, 2551–2557. [Google Scholar] [CrossRef]
- Youngilyeom, Y.L.; Ha, H.S.; Balling, R.; Schöler, H.R.; Artzt, K. Structure, expression and chromosomal location of the Oct-4 gene. Mech. Dev. 1991, 35, 171–179. [Google Scholar] [CrossRef]
- Boiani, M.; Schöler, H.R. Regulatory networks in embryo-derived pluripotent stem cells. Nat. Rev. Mol. Cell Boil. 2005, 6, 872–881. [Google Scholar] [CrossRef]
- Nichols, J.; Zevnik, B.; Anastassiadis, K.; Niwa, H.; Klewe-Nebenius, D.; Chambers, I.; Schöler, H.R.; Smith, A. Formation of Pluripotent Stem Cells in the Mammalian Embryo Depends on the POU Transcription Factor Oct4. Cell 1998, 95, 379–391. [Google Scholar] [CrossRef]
- Scholer, H.R. Octamania: The POU factors in murine development. Trends Genet. 1991, 7, 323–329. [Google Scholar] [CrossRef]
- Pera, M.F. Defining pluripotency. Nat. Methods 2010, 7, 885–887. [Google Scholar] [CrossRef] [PubMed]
- Gafni, O.; Weinberger, L.; Mansour, A.A.; Manor, Y.S.; Chomsky, E.; Ben-Yosef, D.; Kalma, Y.; Viukov, S.; Maza, I.; Zviran, A.; et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 2013, 504, 282–286. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Go, Y.; Kang, I.; Han, Y.; Kim, J. Oct-4 controls cell-cycle progression of embryonic stem cells. Biochem. J. 2010, 426, 171–181. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Yuan, Z.; Ling, H.; Fukasawa, K.; Robertson, K.D.; Olashaw, N.; Koomen, J.; Chen, J.; Lane, W.S.; Seto, E. SIRT1 Deacetylates the DNA Methyltransferase 1 (DNMT1) Protein and Alters Its Activities. Mol. Cell. Boil. 2011, 31, 4720–4734. [Google Scholar] [CrossRef]
- Ferhi, S.; Santaniello, S.; Zerizer, S.; Cruciani, S.; Fadda, A.; Sanna, D.; Dore, A.; Maioli, M.; D’Hallewin, G. Total Phenols from Grape Leaves Counteract Cell Proliferation and Modulate Apoptosis-Related Gene Expression in MCF-7 and HepG2 Human Cancer Cell Lines. Molecules 2019, 24, 612. [Google Scholar] [CrossRef]
- Bai, Y.; Xue, Y.; Xie, X.; Yu, T.; Zhu, Y.; Ge, Q.; Lu, Z. The RNA expression signature of the HepG2 cell line as determined by the integrated analysis of miRNA and mRNA expression profiles. Gene 2014, 548, 91–100. [Google Scholar] [CrossRef]
- Cellosaurus Hep-G2 (CVCL_0027)33, 04-Apr-2012. Available online: https://web.expasy.org/cellosaurus/CVCL_0027 (accessed on 2 July 2020).
- Franko, A.; Hartwig, S.; Kotzka, J.; Ruoß, M.; Nussler, A.K.; Königsrainer, A.; Häring, H.-U.; Lehr, S.; Peter, A. Identification of the Secreted Proteins Originated from Primary Human Hepatocytes and HepG2 Cells. Nutrients 2019, 11, 1795. [Google Scholar] [CrossRef]
- Chen, J.; Chen, L.; Zern, M.A.; Theise, N.D.; Diehl, A.M.; Liu, P.; Duan, Y. The diversity and plasticity of adult hepatic progenitor cells and their niche. Liver Int. 2017, 37, 1260–1271. [Google Scholar] [CrossRef]
- Hang, W.; Feng, Y.; Sang, Z.; Yang, Y.; Zhu, Y.; Huang, Q.; Xi, X. Downregulation of miR-145-5p in cancer cells and their derived exosomes may contribute to the development of ovarian cancer by targeting CT. Int. J. Mol. Med. 2018, 43, 256–266. [Google Scholar] [CrossRef]
- Lei, Z.; Shi, H.; Li, W.; Yu, D.; Shen, F.; Yu, X.; Lu, D.; Sun, C.; Liao, K. miR-185 inhibits non-small cell lung cancer cell proliferation and invasion through targeting of SOX9 and regulation of Wnt signaling. Mol. Med. Rep. 2017, 17, 1742–1752. [Google Scholar] [CrossRef]
- Takahashi, Y.; Forrest, A.R.R.; Maeno, E.; Hashimoto, T.; Daub, C.O.; Yasuda, J. MiR-107 and MiR-185 Can Induce Cell Cycle Arrest in Human Non Small Cell Lung Cancer Cell Lines. PLoS ONE 2009, 4, e6677. [Google Scholar] [CrossRef] [PubMed]
- Santaniello, S.; Cruciani, S.; Basoli, V.; Balzano, F.; Bellu, E.; Garroni, G.; Ginesu, G.C.; Cossu, M.L.; Facchin, F.; Delitala, A.P.; et al. Melatonin and Vitamin D Orchestrate Adipose Derived Stem Cell Fate by Modulating Epigenetic Regulatory Genes. Int. J. Med. Sci. 2018, 15, 1631–1639. [Google Scholar] [CrossRef] [PubMed]
- Balzano, F.; Campesi, I.; Cruciani, S.; Garroni, G.; Bellu, E.; Giudici, S.D.; Angius, A.; Oggiano, A.; Rallo, V.; Capobianco, G.; et al. Epigenetics, Stem Cells, and Autophagy: Exploring a Path Involving miRNA. Int. J. Mol. Sci. 2019, 20, 5091. [Google Scholar] [CrossRef] [PubMed]
- Basoli, V.; Santaniello, S.; Cruciani, S.; Ginesu, G.C.; Cossu, M.L.; Delitala, A.P.; Serra, P.A.; Ventura, C.; Maioli, M. Melatonin and Vitamin D Interfere with the Adipogenic Fate of Adipose-Derived Stem Cells. Int. J. Mol. Sci. 2017, 18, 981. [Google Scholar] [CrossRef] [PubMed]
- Rinaldi, S.; Maioli, M.; Santaniello, S.; Castagna, A.; Pigliaru, G.; Gualini, S.; Margotti, M.L.; Carta, A.; Fontani, V.; Ventura, C. Regenerative treatment using a radioelectric asymmetric conveyor as a novel tool in antiaging medicine: An in vitro beta-galactosidase study. Clin. Interv. Aging 2012, 7, 191–194. [Google Scholar] [CrossRef]
- Balzano, F.; Bellu, E.; Basoli, V.; Dei Giudici, S.; Santaniello, S.; Cruciani, S.; Facchin, F.; Oggiano, A.; Capobianco, G.; Dessole, F.; et al. Lessons from human umbilical cord: Gender differences in stem cells from Wharton’s jelly. Eur. J. Obstet. Gynecol. Reprod. Boil. 2019, 234, 143–148. [Google Scholar] [CrossRef]
- Balzano, F.; Deiana, M.; Giudici, S.D.; Oggiano, A.; Baralla, A.; Pasella, S.; Mannu, A.; Pescatori, M.; Porcu, B.; Fanciulli, G.; et al. miRNA Stability in Frozen Plasma Samples. Molecules 2015, 20, 19030–19040. [Google Scholar] [CrossRef]
- Maioli, M.; Rinaldi, S.; Pigliaru, G.; Santaniello, S.; Basoli, V.; Castagna, A.; Fontani, V.; Ventura, C. REAC technology and hyaluron synthase 2, an interesting network to slow down stem cell senescence. Sci. Rep. 2016, 6, 28682. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST©) forgroup-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 2002, 30, 9. [Google Scholar] [CrossRef]
- Jin, Q.; Yan, T.; Ge, X.; Sun, C.; Shi, X.; Zhai, Q. Cytoplasm-localized SIRT1 enhances apoptosis. J. Cell. Physiol. 2007, 213, 88–97. [Google Scholar] [CrossRef] [PubMed]
- Faller, Y.D.A.D.V. Transcription Regulation by Class III Histone Deacetylases (HDACs)—Sirtuins. Transl. Oncogenomics 2008, 1, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Tanno, M.; Sakamoto, J.; Miura, T.; Shimamoto, K.; Horio, Y. Nucleocytoplasmic Shuttling of the NAD+-dependent Histone Deacetylase SIRT1. J. Boil. Chem. 2006, 282, 6823–6832. [Google Scholar] [CrossRef]
- Yuan, J.; Minter-Dykhouse, K.; Lou, Z. A c-Myc–SIRT1 feedback loop regulates cell growth and transformation. J. Cell Boil. 2009, 185, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Liang, X.-H.; Chen, L.-X.; Bao, S.-M.; Yan, Z. SIRT1 is highly expressed in brain metastasis tissues of non-small cell lung cancer (NSCLC) and in positive regulation of NSCLC cell migration. Int. J. Clin. Exp. Pathol. 2013, 6, 2357–2365. [Google Scholar]
- Roth, M.; Chen, W.Y. Sorting out functions of sirtuins in cancer. Oncogene 2013, 33, 1609–1620. [Google Scholar] [CrossRef]
- Joo, H.-Y.; Woo, S.R.; Shen, Y.-N.; Yun, M.Y.; Shin, H.-J.; Park, E.-R.; Kim, S.-H.; Park, J.-E.; Ju, Y.-J.; Hong, S.H.; et al. SIRT1 interacts with and protects glyceraldehyde-3-phosphate dehydrogenase (GAPDH) from nuclear translocation: Implications for cell survival after irradiation. Biochem. Biophys. Res. Commun. 2012, 424, 681–686. [Google Scholar] [CrossRef]
- Schmitz, H.-D. Reversible nuclear translocation of glyceraldehyde-3-phosphate dehydrogenase upon serum depletion. Eur. J. Cell Boil. 2001, 80, 419–427. [Google Scholar] [CrossRef]
- Zhou, N.; Lin, X.; Dong, W. SIRT1 alleviates senescence of degenerative human intervertebral disc cartilage endo-plate cells via the p53/p21 pathway. Sci. Rep. 2016, 2, 26–28. [Google Scholar] [CrossRef]
- Fujino, T.; Yokokawa, R.; Oshima, T.; Hayakawa, M. SIRT1 knockdown up-regulates p53 and p21/Cip1 expression in renal adenocarcinoma cells but not in normal renal-derived cells in a deacetylase-independent manner. J. Toxicol. Sci. 2018, 43, 711–715. [Google Scholar] [CrossRef]
- Wandzioch, E.; Zaret, K.S. Dynamic Signaling Network for the Specification of Embryonic Pancreas and Liver Progenitors. Science 2009, 324, 1707–1710. [Google Scholar] [CrossRef] [PubMed]
- Abbas, T.; Dutta, A. p21 in cancer: Intricate networks and multiple activities. Nat. Rev. Cancer 2009, 9, 400–414. [Google Scholar] [CrossRef] [PubMed]


| Primer Name | Forward | Reverse |
|---|---|---|
| P16 | CAACGCACCGCCTAGTTACGG | AACTTCGTCCTCCAGAGTCGC |
| P19 | GCCTTCGGCTGACTGGCTGG | TCGTCCTCCAGAGTCGCCCG |
| P21 | CAAAGGCCCGCTCTACATCTT | AGGAACCTCCATTCACCCGA |
| P53 | TGGCCTTGAAACCACCTTTT | AACTACCAACCCACCAGCCAA |
| Oct-4 | GAGGAGTCCCAGGACATCAA | CATCGGCCTGTGTATATCCC |
| SOX2 | CCGTTCATGTAGGTCTGCGAGCTG | CAACGGCAGCTACAGCATGATGC |
| NANOG | CATGAGTGTGGATCCAGCT | CCTGAATAAGCAGATCCAT |
| c-Myc | GGACGACGAGACCTTCATCAA | GCACCGAGTCGTAGTCGAG |
| GAPDH | GAGTCAACGGATTTGGTCGTGA | CTCCTTGGGCCGCGCATCAT |
| DNMT1 | CGTCCGAGCGTCACACA | GAGCCTTTGCCATTCTTCGC |
| SIRT1 | CATTTCCATGGCGCTGAGG | TGCTGGTGGAACAATTCCTGT |
| Accession ID Number | Symbol | Sequence |
|---|---|---|
| MIMAT0000437 | hsa-miR-145-5p | GUCCAGUUUUCCCAGGAAUCCCU |
| MIMAT0000243 | hsa-miR-148a-3p | UCAGUGCACUACAGAACUUUGU |
| MIMAT0004611 | hsa-miR-185-3p | AGGGGCUGGCUUUCCUCUGGUC |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Balzano, F.; Garroni, G.; Cruciani, S.; Bellu, E.; Dei Giudici, S.; Oggiano, A.; Capobianco, G.; Dessole, S.; Ventura, C.; Maioli, M. Behavioral Changes in Stem-Cell Potency by HepG2-Exhausted Medium. Cells 2020, 9, 1890. https://doi.org/10.3390/cells9081890
Balzano F, Garroni G, Cruciani S, Bellu E, Dei Giudici S, Oggiano A, Capobianco G, Dessole S, Ventura C, Maioli M. Behavioral Changes in Stem-Cell Potency by HepG2-Exhausted Medium. Cells. 2020; 9(8):1890. https://doi.org/10.3390/cells9081890
Chicago/Turabian StyleBalzano, Francesca, Giuseppe Garroni, Sara Cruciani, Emanuela Bellu, Silvia Dei Giudici, Annalisa Oggiano, Giampiero Capobianco, Salvatore Dessole, Carlo Ventura, and Margherita Maioli. 2020. "Behavioral Changes in Stem-Cell Potency by HepG2-Exhausted Medium" Cells 9, no. 8: 1890. https://doi.org/10.3390/cells9081890
APA StyleBalzano, F., Garroni, G., Cruciani, S., Bellu, E., Dei Giudici, S., Oggiano, A., Capobianco, G., Dessole, S., Ventura, C., & Maioli, M. (2020). Behavioral Changes in Stem-Cell Potency by HepG2-Exhausted Medium. Cells, 9(8), 1890. https://doi.org/10.3390/cells9081890







