From Tank to Treatment: Modeling Melanoma in Zebrafish
Abstract
1. Melanoma in Humans
2. Melanocytes in Zebrafish
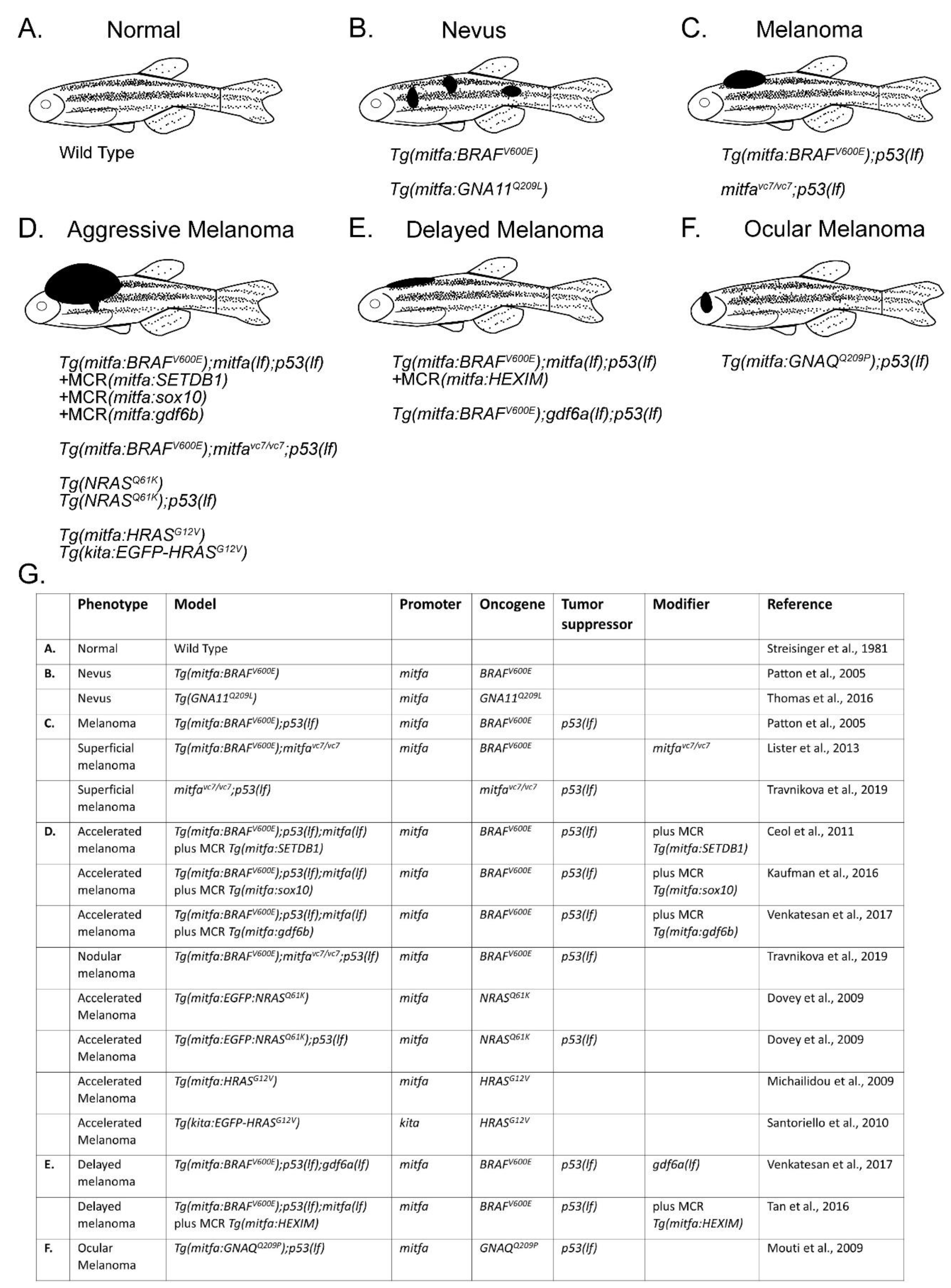
3. Modeling Melanoma Disease Drivers in Zebrafish
4. Using Zebrafish to Discover Genetic Modifiers of Melanoma
5. Intersection of Melanocyte Development and Melanoma in Zebrafish
6. Zebrafish Neural Crest Reactivation in Melanoma Initiation and Progression
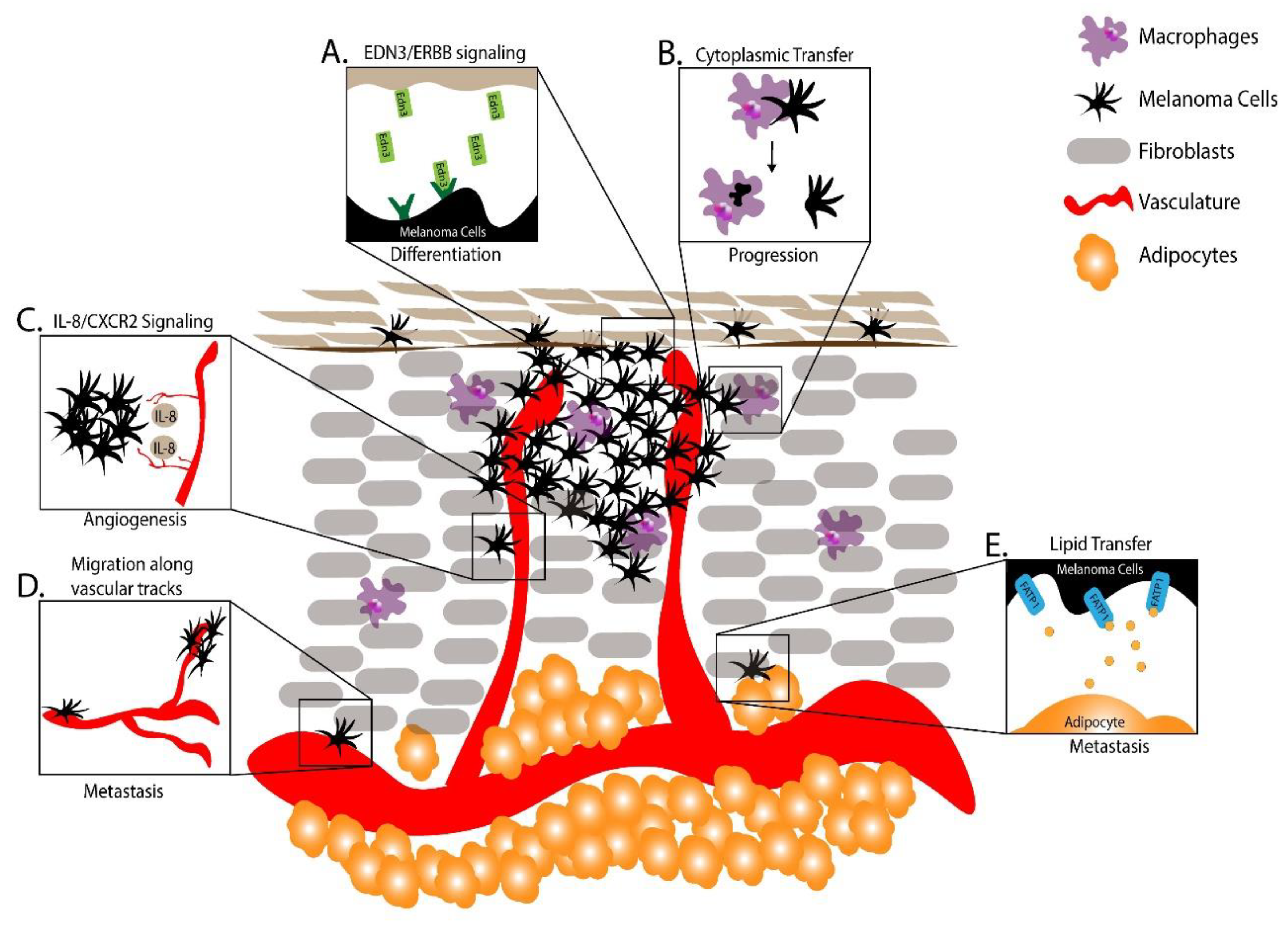
7. Utilizing Zebrafish to Understand the Melanoma Microenvironment
7.1. Using Zebrafish to Study Phenotype Switching in Melanoma
7.2. Using Zebrafish to Study Innate Immune Cells in Melanoma
7.3. Using Zebrafish to Study Vascular Cells in Melanoma
7.4. Using Zebrafish to Study Adipocytes in Melanoma
8. Future Directions
8.1. Understanding the Micro- and Macro-Environmental Factors that Encourage Disease Initiation, Progression, Metastasis and Relapse
8.2. Understanding the Role of Developmental Mechanisms in Melanoma Pathogenesis
8.3. Utilizing Zebrafish Xenografts to Model Patient Disease
8.4. Utilizing Zebrafish as a Screening Platform
9. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Cancer Facts and Figures 2020; American Cancer Society: Atlanta, GA, USA, 2020.
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef]
- Atkins, M.B.; Lotze, M.T.; Dutcher, J.P.; Fisher, R.I.; Weiss, G.; Margolin, K.; Abrams, J.; Sznol, M.; Parkinson, D.; Hawkins, M.; et al. High-dose recombinant interleukin 2 therapy for patients with metastatic melanoma: Analysis of 270 patients treated between 1985 and 1993. J. Clin. Oncol. 1999, 17, 2105–2116. [Google Scholar] [CrossRef]
- Hill, G.J., 2nd; Krementz, E.T.; Hill, H.Z. Dimethyl triazeno imidazole carboxamide and combination therapy for melanoma. IV. Late results after complete response to chemotherapy (Central Oncology Group protocols 7130, 7131, and 7131A). Cancer 1984, 53, 1299–1305. [Google Scholar] [CrossRef]
- Sosman, J.A.; Kim, K.B.; Schuchter, L.; Gonzalez, R.; Pavlick, A.C.; Weber, J.S.; McArthur, G.A.; Hutson, T.E.; Moschos, S.J.; Flaherty, K.T.; et al. Survival in BRAF V600-mutant advanced melanoma treated with vemurafenib. N. Engl. J. Med. 2012, 366, 707–714. [Google Scholar] [CrossRef] [PubMed]
- Robert, C.; Grob, J.J.; Stroyakovskiy, D.; Karaszewska, B.; Hauschild, A.; Levchenko, E.; Chiarion Sileni, V.; Schachter, J.; Garbe, C.; Bondarenko, I.; et al. Five-year outcomes with dabrafenib plus trametinib in metastatic melanoma. N. Engl. J. Med. 2019, 381, 626–636. [Google Scholar] [CrossRef] [PubMed]
- Larkin, J.; Chiarion-Sileni, V.; Gonzalez, R.; Grob, J.J.; Rutkowski, P.; Lao, C.D.; Cowey, C.L.; Schadendorf, D.; Wagstaff, J.; Dummer, R.; et al. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl. J. Med. 2019, 381, 1535–1546. [Google Scholar] [CrossRef] [PubMed]
- Eggermont, A.M.M.; Blank, C.U.; Mandala, M.; Long, G.V.; Atkinson, V.; Dalle, S.; Haydon, A.; Lichinitser, M.; Khattak, A.; Carlino, M.S.; et al. Adjuvant pembrolizumab versus placebo in resected stage III melanoma. N. Engl. J. Med. 2018, 378, 1789–1801. [Google Scholar] [CrossRef]
- Streisinger, G.; Walker, C.; Dower, N.; Knauber, D.; Singer, F. Production of clones of homozygous diploid zebra fish (Brachydanio rerio). Nature 1981, 291, 293–296. [Google Scholar] [CrossRef]
- Gomes, M.C.; Mostowy, S. The case for modeling human infection in zebrafish. Trends Microbiol. 2020, 28, 10–18. [Google Scholar] [CrossRef]
- Santoriello, C.; Anelli, V.; Alghisi, E.; Mione, M. Highly penetrant melanoma in a zebrafish model is independent of ErbB3b signaling. Pigment Cell Melanoma Res. 2012, 25, 287–289. [Google Scholar] [CrossRef]
- Langenau, D.M.; Traver, D.; Ferrando, A.A.; Kutok, J.L.; Aster, J.C.; Kanki, J.P.; Lin, S.; Prochownik, E.; Trede, N.S.; Zon, L.I.; et al. Myc-induced T cell leukemia in transgenic zebrafish. Science 2003, 299, 887–890. [Google Scholar] [CrossRef] [PubMed]
- Patton, E.E.; Widlund, H.R.; Kutok, J.L.; Kopani, K.R.; Amatruda, J.F.; Murphey, R.D.; Berghmans, S.; Mayhall, E.A.; Traver, D.; Fletcher, C.D.; et al. BRAF mutations are sufficient to promote nevi formation and cooperate with p53 in the genesis of melanoma. Curr. Biol. 2005, 15, 249–254. [Google Scholar] [CrossRef] [PubMed]
- Iyengar, S.; Kasheta, M.; Ceol, C.J. Poised regeneration of zebrafish melanocytes involves direct differentiation and concurrent replenishment of tissue-resident progenitor cells. Dev. Cell 2015, 33, 631–643. [Google Scholar] [CrossRef] [PubMed]
- Hirata, M.; Nakamura, K.; Kanemaru, T.; Shibata, Y.; Kondo, S. Pigment cell organization in the hypodermis of zebrafish. Dev. Dyn 2003, 227, 497–503. [Google Scholar] [CrossRef] [PubMed]
- Howe, K.; Clark, M.D.; Torroja, C.F.; Torrance, J.; Berthelot, C.; Muffato, M.; Collins, J.E.; Humphray, S.; McLaren, K.; Matthews, L.; et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature 2013, 496, 498–503. [Google Scholar] [CrossRef] [PubMed]
- Chakrabarti, S.; Streisinger, G.; Singer, F.; Walker, C. Frequency of gamma-ray induced specific locus and recessive lethal mutations in mature germ cells of the zebrafish, Brachydanio rerio. Genetics 1983, 103, 109–123. [Google Scholar]
- Kelsh, R.N.; Schmid, B.; Eisen, J.S. Genetic analysis of melanophore development in zebrafish embryos. Dev. Biol. 2000, 225, 277–293. [Google Scholar] [CrossRef]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF gene in human cancer. Nature 2002, 417, 949–954. [Google Scholar] [CrossRef]
- Pollock, P.M.; Harper, U.L.; Hansen, K.S.; Yudt, L.M.; Stark, M.; Robbins, C.M.; Moses, T.Y.; Hostetter, G.; Wagner, U.; Kakareka, J.; et al. High frequency of BRAF mutations in nevi. Nat. Genet. 2003, 33, 19–20. [Google Scholar] [CrossRef]
- Dovey, M.; White, R.M.; Zon, L.I. Oncogenic NRAS cooperates with p53 loss to generate melanoma in zebrafish. Zebrafish 2009, 6, 397–404. [Google Scholar] [CrossRef]
- Zhang, Y.; Xiong, Y.; Yarbrough, W.G. ARF Promotes MDM2 Degradation and Stabilizes p53: ARF-INK4a Locus Deletion Impairs Both the Rb and p53 Tumor Suppression Pathways. Cell 1998, 92, 725–734. [Google Scholar] [CrossRef]
- Zhuravleva, J.; Paggetti, J.; Martin, L.; Hammann, A.; Solary, E.; Bastie, J.N.; Delva, L. MOZ/TIF2-induced acute myeloid leukaemia in transgenic fish. Br. J. Haematol. 2008, 143, 378–382. [Google Scholar] [CrossRef] [PubMed]
- Bergamaschi, D.; Samuels, Y.; O’Neil, N.J.; Trigiante, G.; Crook, T.; Hsieh, J.K.; O’Connor, D.J.; Zhong, S.; Campargue, I.; Tomlinson, M.L.; et al. iASPP oncoprotein is a key inhibitor of p53 conserved from worm to human. Nat. Genet. 2003, 33, 162–167. [Google Scholar] [CrossRef] [PubMed]
- Gembarska, A.; Luciani, F.; Fedele, C.; Russell, E.A.; Dewaele, M.; Villar, S.; Zwolinska, A.; Haupt, S.; de Lange, J.; Yip, D.; et al. MDM4 is a key therapeutic target in cutaneous melanoma. Nat. Med. 2012, 18, 1239–1247. [Google Scholar] [CrossRef]
- Hodis, E.; Watson, I.R.; Kryukov, G.V.; Arold, S.T.; Imielinski, M.; Theurillat, J.P.; Nickerson, E.; Auclair, D.; Li, L.; Place, C.; et al. A landscape of driver mutations in melanoma. Cell 2012, 150, 251–263. [Google Scholar] [CrossRef]
- Santoriello, C.; Gennaro, E.; Anelli, V.; Distel, M.; Kelly, A.; Koster, R.W.; Hurlstone, A.; Mione, M. Kita driven expression of oncogenic HRAS leads to early onset and highly penetrant melanoma in zebrafish. PLoS ONE 2010, 5, e15170. [Google Scholar] [CrossRef]
- Michailidou, C.; Jones, M.; Walker, P.; Kamarashev, J.; Kelly, A.; Hurlstone, A.F. Dissecting the roles of Raf- and PI3K-signalling pathways in melanoma formation and progression in a zebrafish model. Dis. Model. Mech. 2009, 2, 399–411. [Google Scholar] [CrossRef]
- Anelli, V.; Santoriello, C.; Distel, M.; Koster, R.W.; Ciccarelli, F.D.; Mione, M. Global repression of cancer gene expression in a zebrafish model of melanoma is linked to epigenetic regulation. Zebrafish 2009, 6, 417–424. [Google Scholar] [CrossRef]
- Van Raamsdonk, C.D.; Bezrookove, V.; Green, G.; Bauer, J.; Gaugler, L.; O’Brien, J.M.; Simpson, E.M.; Barsh, G.S.; Bastian, B.C. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 2009, 457, 599–602. [Google Scholar] [CrossRef]
- Van Raamsdonk, C.D.; Griewank, K.G.; Crosby, M.B.; Garrido, M.C.; Vemula, S.; Wiesner, T.; Obenauf, A.C.; Wackernagel, W.; Green, G.; Bouvier, N.; et al. Mutations in GNA11 in uveal melanoma. N. Engl. J. Med. 2010, 363, 2191–2199. [Google Scholar] [CrossRef]
- Mouti, M.A.; Dee, C.; Coupland, S.E.; Hurlstone, A.F. Minimal contribution of ERK1/2-MAPK signalling towards the maintenance of oncogenic GNAQQ209P-driven uveal melanomas in zebrafish. Oncotarget 2016, 7, 39654–39670. [Google Scholar] [CrossRef] [PubMed]
- Berger, M.F.; Levin, J.Z.; Vijayendran, K.; Sivachenko, A.; Adiconis, X.; Maguire, J.; Johnson, L.A.; Robinson, J.; Verhaak, R.G.; Sougnez, C.; et al. Integrative analysis of the melanoma transcriptome. Genome. Res. 2010, 20, 413–427. [Google Scholar] [CrossRef] [PubMed]
- Yen, J.; White, R.M.; Wedge, D.C.; Van Loo, P.; de Ridder, J.; Capper, A.; Richardson, J.; Jones, D.; Raine, K.; Watson, I.R.; et al. The genetic heterogeneity and mutational burden of engineered melanomas in zebrafish models. Genome Biol. 2013, 14, R113. [Google Scholar] [CrossRef]
- Golan, T.; Parikh, R.; Jacob, E.; Vaknine, H.; Zemser-Werner, V.; Hershkovitz, D.; Malcov, H.; Leibou, S.; Reichman, H.; Sheinboim, D.; et al. Adipocytes sensitize melanoma cells to environmental TGF-beta cues by repressing the expression of miR-211. Sci. Signal 2019, 12, eaav6847. [Google Scholar] [CrossRef] [PubMed]
- Venkatesan, A.M.; Vyas, R.; Gramann, A.K.; Dresser, K.; Gujja, S.; Bhatnagar, S.; Chhangawala, S.; Gomes, C.B.F.; Xi, H.S.; Lian, C.G.; et al. Ligand-activated BMP signaling inhibits cell differentiation and death to promote melanoma. J. Clin. Investig. 2018, 128, 294–308. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.; Hoersch, S.; Amsterdam, A.; Whittaker, C.A.; Beert, E.; Catchen, J.M.; Farrington, S.; Postlethwait, J.H.; Legius, E.; Hopkins, N.; et al. Comparative oncogenomic analysis of copy number alterations in human and zebrafish tumors enables cancer driver discovery. PLoS Genet 2013, 9, e1003734. [Google Scholar] [CrossRef]
- Yeh, I.; von Deimling, A.; Bastian, B.C. Clonal BRAF mutations in melanocytic nevi and initiating role of BRAF in melanocytic neoplasia. J. Natl. Cancer Inst. 2013, 105, 917–919. [Google Scholar] [CrossRef]
- Shain, A.H.; Yeh, I.; Kovalyshyn, I.; Sriharan, A.; Talevich, E.; Gagnon, A.; Dummer, R.; North, J.; Pincus, L.; Ruben, B.; et al. The Genetic Evolution of Melanoma from Precursor Lesions. N. Engl. J. Med. 2015, 373, 1926–1936. [Google Scholar] [CrossRef]
- Berger, M.F.; Hodis, E.; Heffernan, T.P.; Deribe, Y.L.; Lawrence, M.S.; Protopopov, A.; Ivanova, E.; Watson, I.R.; Nickerson, E.; Ghosh, P.; et al. Melanoma genome sequencing reveals frequent PREX2 mutations. Nature 2012, 485, 502–506. [Google Scholar] [CrossRef]
- Kan, Z.; Jaiswal, B.S.; Stinson, J.; Janakiraman, V.; Bhatt, D.; Stern, H.M.; Yue, P.; Haverty, P.M.; Bourgon, R.; Zheng, J.; et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 2010, 466, 869–873. [Google Scholar] [CrossRef]
- Lawrence, M.S.; Stojanov, P.; Mermel, C.H.; Robinson, J.T.; Garraway, L.A.; Golub, T.R.; Meyerson, M.; Gabriel, S.B.; Lander, E.S.; Getz, G. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 2014, 505, 495–501. [Google Scholar] [CrossRef]
- Pleasance, E.D.; Cheetham, R.K.; Stephens, P.J.; McBride, D.J.; Humphray, S.J.; Greenman, C.D.; Varela, I.; Lin, M.L.; Ordonez, G.R.; Bignell, G.R.; et al. A comprehensive catalogue of somatic mutations from a human cancer genome. Nature 2010, 463, 191–196. [Google Scholar] [CrossRef] [PubMed]
- Thomas, A.C.; Zeng, Z.; Riviere, J.B.; O’Shaughnessy, R.; Al-Olabi, L.; St-Onge, J.; Atherton, D.J.; Aubert, H.; Bagazgoitia, L.; Barbarot, S.; et al. Mosaic Activating Mutations in GNA11 and GNAQ Are Associated with Phakomatosis Pigmentovascularis and Extensive Dermal Melanocytosis. J. Investig. Dermatol. 2016, 136, 770–778. [Google Scholar] [CrossRef] [PubMed]
- Lister, J.A.; Capper, A.; Zeng, Z.; Mathers, M.E.; Richardson, J.; Paranthaman, K.; Jackson, I.J.; Elizabeth Patton, E. A conditional zebrafish MITF mutation reveals MITF levels are critical for melanoma promotion vs. regression in vivo. J. Investig. Dermatol. 2014, 134, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Ceol, C.J.; Houvras, Y.; Jane-Valbuena, J.; Bilodeau, S.; Orlando, D.A.; Battisti, V.; Fritsch, L.; Lin, W.M.; Hollmann, T.J.; Ferre, F.; et al. The histone methyltransferase SETDB1 is recurrently amplified in melanoma and accelerates its onset. Nature 2011, 471, 513–517. [Google Scholar] [CrossRef] [PubMed]
- Kaufman, C.K.; Mosimann, C.; Fan, Z.P.; Yang, S.; Thomas, A.J.; Ablain, J.; Tan, J.L.; Fogley, R.D.; van Rooijen, E.; Hagedorn, E.J.; et al. A zebrafish melanoma model reveals emergence of neural crest identity during melanoma initiation. Science 2016, 351, aad2197. [Google Scholar] [CrossRef]
- Travnickova, J.; Wojciechowska, S.; Khamseh, A.; Gautier, P.; Brown, D.V.; Lefevre, T.; Brombin, A.; Ewing, A.; Capper, A.; Spitzer, M.; et al. Zebrafish MITF-Low Melanoma Subtype Models Reveal Transcriptional Subclusters and MITF-Independent Residual Disease. Cancer Res. 2019, 79, 5769–5784. [Google Scholar] [CrossRef]
- Tan, J.L.; Fogley, R.D.; Flynn, R.A.; Ablain, J.; Yang, S.; Saint-Andre, V.; Fan, Z.P.; Do, B.T.; Laga, A.C.; Fujinaga, K.; et al. Stress from Nucleotide Depletion Activates the Transcriptional Regulator HEXIM1 to Suppress Melanoma. Mol. Cell 2016, 62, 34–46. [Google Scholar] [CrossRef]
- Lin, W.M.; Baker, A.C.; Beroukhim, R.; Winckler, W.; Feng, W.; Marmion, J.M.; Laine, E.; Greulich, H.; Tseng, H.; Gates, C.; et al. Modeling genomic diversity and tumor dependency in malignant melanoma. Cancer Res. 2008, 68, 664–673. [Google Scholar] [CrossRef]
- Curtin, J.A.; Fridlyand, J.; Kageshita, T.; Patel, H.N.; Busam, K.J.; Kutzner, H.; Cho, K.H.; Aiba, S.; Brocker, E.B.; LeBoit, P.E.; et al. Distinct sets of genetic alterations in melanoma. N. Engl. J. Med. 2005, 353, 2135–2147. [Google Scholar] [CrossRef]
- Ablain, J.; Xu, M.; Rothschild, H.; Jordan, R.C.; Mito, J.K.; Daniels, B.H.; Bell, C.F.; Joseph, N.M.; Wu, H.; Bastian, B.C.; et al. Human tumor genomics and zebrafish modeling identify SPRED1 loss as a driver of mucosal melanoma. Science 2018, 362, 1055–1060. [Google Scholar] [CrossRef] [PubMed]
- Ablain, J.; Durand, E.M.; Yang, S.; Zhou, Y.; Zon, L.I. A CRISPR/Cas9 vector system for tissue-specific gene disruption in zebrafish. Dev. Cell 2015, 32, 756–764. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, D.R.; Yu, J.; Shanker, K.; Deshpande, N.; Varambally, R.; Ghosh, D.; Barrette, T.; Pandey, A.; Chinnaiyan, A.M. Oncomine: A cancer microarray database and integrated data-mining platform. Neoplasia 2004, 6, 1–6. [Google Scholar] [CrossRef]
- Contreras, X.; Barboric, M.; Lenasi, T.; Peterlin, B.M. HMBA releases P-TEFb from HEXIM1 and 7SK snRNA via PI3K/Akt and activates HIV transcription. PLoS Pathog. 2007, 3, 1459–1469. [Google Scholar] [CrossRef]
- Scahill, C.M.; Digby, Z.; Sealy, I.M.; Wojciechowska, S.; White, R.J.; Collins, J.E.; Stemple, D.L.; Bartke, T.; Mathers, M.E.; Patton, E.E.; et al. Loss of the chromatin modifier Kdm2aa causes BrafV600E-independent spontaneous melanoma in zebrafish. PLoS Genet 2017, 13, e1006959. [Google Scholar] [CrossRef]
- Salhi, A.; Farhadian, J.A.; Giles, K.M.; Vega-Saenz de Miera, E.; Silva, I.P.; Bourque, C.; Yeh, K.; Chhangawala, S.; Wang, J.; Ye, F.; et al. RSK1 activation promotes invasion in nodular melanoma. Am. J. Pathol. 2015, 185, 704–716. [Google Scholar] [CrossRef]
- Wu, M.; Hemesath, T.J.; Takemoto, C.M.; Horstmann, M.A.; Wells, A.G.; Price, E.R.; Fisher, D.Z.; Fisher, D.E. c-Kit triggers dual phosphorylations, which couple activation and degradation of the essential melanocyte factor Mi. Gene. Develop. 2000, 14, 301–312. [Google Scholar]
- Salhi, A.; Jordan, A.C.; Bochaca, I.I.; Izsak, A.; Darvishian, F.; Houvras, Y.; Giles, K.M.; Osman, I. Oxidative Phosphorylation Promotes Primary Melanoma Invasion. Am. J. Pathol. 2020, 190, 1108–1117. [Google Scholar] [CrossRef]
- Haq, R.; Shoag, J.; Andreu-Perez, P.; Yokoyama, S.; Edelman, H.; Rowe, G.C.; Frederick, D.T.; Hurley, A.D.; Nellore, A.; Kung, A.L.; et al. Oncogenic BRAF regulates oxidative metabolism via PGC1alpha and MITF. Cancer Cell 2013, 23, 302–315. [Google Scholar] [CrossRef]
- Vazquez, F.; Lim, J.H.; Chim, H.; Bhalla, K.; Girnun, G.; Pierce, K.; Clish, C.B.; Granter, S.R.; Widlund, H.R.; Spiegelman, B.M.; et al. PGC1alpha expression defines a subset of human melanoma tumors with increased mitochondrial capacity and resistance to oxidative stress. Cancer Cell 2013, 23, 287–301. [Google Scholar] [CrossRef]
- Henderson, F.; Johnston, H.R.; Badrock, A.P.; Jones, E.A.; Forster, D.; Nagaraju, R.T.; Evangelou, C.; Kamarashev, J.; Green, M.; Fairclough, M.; et al. Enhanced Fatty Acid Scavenging and Glycerophospholipid Metabolism Accompany Melanocyte Neoplasia Progression in Zebrafish. Cancer Res. 2019, 79, 2136–2151. [Google Scholar] [CrossRef] [PubMed]
- Balaban, S.; Lee, L.S.; Schreuder, M.; Hoy, A.J. Obesity and cancer progression: Is there a role of fatty acid metabolism? Biomed. Res. Int. 2015, 2015, 274585. [Google Scholar] [CrossRef] [PubMed]
- Hodgkinson, C.A.; Moore, K.J.; Nakayama, A.; Steingrímsson, E.; Copeland, N.G.; Jenkins, N.A.; Arnheiter, H. Mutations at the mouse microphthalmia locus are associated with defects in a gene encoding a novel basic-helix-loop-helix-zipper protein. Cell 1993, 74, 395–404. [Google Scholar] [CrossRef]
- Krakowsky, J.M.; Boissy, R.E.; Neumann, J.C.; Lingrel, J.B. A DNA insertional mutation results in microphthalmia in transgenic mice. Transgenic Res. 1993, 2, 14–20. [Google Scholar] [CrossRef]
- Geissler, E.N.; Ryan, M.A.; Housman, D.E. The dominant-white spotting (W) locus of the mouse encodes the c-kit proto-oncogene. Cell 1988, 55, 185–192. [Google Scholar] [CrossRef]
- Garraway, L.A.; Widlund, H.R.; Rubin, M.A.; Getz, G.; Berger, A.J.; Ramaswamy, S.; Beroukhim, R.; Milner, D.A.; Granter, S.R.; Du, J.; et al. Integrative genomic analyses identify MITF as a lineage survival oncogene amplified in malignant melanoma. Nature 2005, 436, 117–122. [Google Scholar] [CrossRef]
- Curtin, J.A.; Busam, K.; Pinkel, D.; Bastian, B.C. Somatic activation of KIT in distinct subtypes of melanoma. J. Clin. Oncol. 2006, 24, 4340–4346. [Google Scholar] [CrossRef]
- Beadling, C.; Jacobson-Dunlop, E.; Hodi, F.S.; Le, C.; Warrick, A.; Patterson, J.; Town, A.; Harlow, A.; Cruz, F., 3rd; Azar, S.; et al. KIT gene mutations and copy number in melanoma subtypes. Clin. Cancer Res. 2008, 14, 6821–6828. [Google Scholar] [CrossRef]
- Levy, C.; Khaled, M.; Fisher, D.E. MITF: Master regulator of melanocyte development and melanoma oncogene. Trends Mol. Med. 2006, 12, 406–414. [Google Scholar] [CrossRef]
- Steingrimsson, E.; Moore, K.J.; Lamoreux, M.L.; Ferre-D’Amare, A.R.; Burley, S.K.; Zimring, D.C.; Skow, L.C.; Hodgkinson, C.A.; Arnheiter, H.; Copeland, N.G.; et al. Molecular basis of mouse microphthalmia (mi) mutations helps explain their developmental and phenotypic consequences. Nat. Genet. 1994, 8, 256–263. [Google Scholar] [CrossRef]
- Tassabehji, M.; Newton, V.E.; Read, A.P. Waardenburg syndrome type 2 caused by mutations in the human microphthalmia (MITF) gene. Nat. Genet. 1994, 8, 251–255. [Google Scholar] [CrossRef] [PubMed]
- Lister, J.A.; Robertson, C.P.; Lepage, T.; Johnson, S.L.; Raible, D.W. nacre encodes a zebrafish microphthalmia-related protein that regulates neural-crest-derived pigment cell fate. Development 1999, 126, 3757–3767. [Google Scholar] [PubMed]
- Hertwig, P. Neue Mutationen und Koppelungsgruppen bei der Hausmaus. Zeitschrift für Induktive Abstammungs- und Vererbungslehre 1942, 80, 220–246. [Google Scholar] [CrossRef]
- Hoek, K.S.; Schlegel, N.C.; Brafford, P.; Sucker, A.; Ugurel, S.; Kumar, R.; Weber, B.L.; Nathanson, K.L.; Phillips, D.J.; Herlyn, M.; et al. Metastatic potential of melanomas defined by specific gene expression profiles with no BRAF signature. Pigment Cell Res. 2006, 19, 290–302. [Google Scholar] [CrossRef]
- Muller, J.; Krijgsman, O.; Tsoi, J.; Robert, L.; Hugo, W.; Song, C.; Kong, X.; Possik, P.A.; Cornelissen-Steijger, P.D.; Geukes Foppen, M.H.; et al. Low MITF/AXL ratio predicts early resistance to multiple targeted drugs in melanoma. Nat. Commun. 2014, 5, 5712. [Google Scholar] [CrossRef]
- Zeng, Z.; Johnson, S.L.; Lister, J.A.; Patton, E.E. Temperature-sensitive splicing of mitfa by an intron mutation in zebrafish. Pigment Cell Melanoma Res. 2015, 28, 229–232. [Google Scholar] [CrossRef]
- Giebel, L.B.; Spritz, R.A. Mutation of the KIT (mast/stem cell growth factor receptor) protooncogene in human piebaldism. Proc. Natl. Acad. Sci. USA 1991, 88, 8696–8699. [Google Scholar] [CrossRef]
- Williams, D.E.; Lyman, S.D. Characterization of the gene-product of the Steel locus. Prog. Growth Factor Res. 1991, 3, 235–242. [Google Scholar] [CrossRef]
- Parichy, D.M.; Rawls, J.F.; Pratt, S.J.; Whitfield, T.T.; Johnson, S.L. Zebrafish sparse corresponds to an orthologue of c-kit and is required for the morphogenesis of a subpopulation of melanocytes, but is not essential for hematopoiesis or primordial germ cell development. Development 1999, 126, 3425–3436. [Google Scholar]
- Hultman, K.A.; Bahary, N.; Zon, L.I.; Johnson, S.L. Gene Duplication of the zebrafish kit ligand and partitioning of melanocyte development functions to kit ligand a. PLoS Genet 2007, 3, e17. [Google Scholar] [CrossRef]
- Ryu, B.; Kim, D.S.; Deluca, A.M.; Alani, R.M. Comprehensive expression profiling of tumor cell lines identifies molecular signatures of melanoma progression. PLoS ONE 2007, 2, e594. [Google Scholar] [CrossRef] [PubMed]
- Natali, P.G.; Nicotra, M.R.; Winkler, A.B.; Cavaliere, R.; Bigotti, A.; Ullrich, A. Progression of human cutaneous melanoma is associated with loss of expression of c-kit proto-oncogene receptor. Int. J. Cancer 1992, 52, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Neiswender, J.V.; Kortum, R.L.; Bourque, C.; Kasheta, M.; Zon, L.I.; Morrison, D.K.; Ceol, C.J. KIT Suppresses BRAF(V600E)-Mutant Melanoma by Attenuating Oncogenic RAS/MAPK Signaling. Cancer Res 2017, 77, 5820–5830. [Google Scholar] [CrossRef] [PubMed]
- Bittner, M.; Meltzer, P.; Chen, Y.; Jiang, Y.; Seftor, E.; Hendrix, M.; Radmacher, M.; Simon, R.; Yakhini, Z.; Ben-Dor, A.; et al. Molecular classification of cutaneous malignant melanoma by gene expression profiling. Nature 2000, 406, 536–540. [Google Scholar] [CrossRef]
- Carr, K.M.; Bittner, M.; Trent, J.M. Gene-expression profiling in human cutaneous melanoma. Oncogene 2003, 22, 3076–3080. [Google Scholar] [CrossRef]
- Hoek, K.; Rimm, D.L.; Williams, K.R.; Zhao, H.; Ariyan, S.; Lin, A.; Kluger, H.M.; Berger, A.J.; Cheng, E.; Trombetta, E.S.; et al. Expression profiling reveals novel pathways in the transformation of melanocytes to melanomas. Cancer Res. 2004, 64, 5270–5282. [Google Scholar] [CrossRef]
- Topczewska, J.M.; Postovit, L.M.; Margaryan, N.V.; Sam, A.; Hess, A.R.; Wheaton, W.W.; Nickoloff, B.J.; Topczewski, J.; Hendrix, M.J. Embryonic and tumorigenic pathways converge via Nodal signaling: Role in melanoma aggressiveness. Nat. Med. 2006, 12, 925–932. [Google Scholar] [CrossRef]
- Lee, L.M.; Seftor, E.A.; Bonde, G.; Cornell, R.A.; Hendrix, M.J. The fate of human malignant melanoma cells transplanted into zebrafish embryos: Assessment of migration and cell division in the absence of tumor formation. Dev. Dyn. 2005, 233, 1560–1570. [Google Scholar] [CrossRef]
- Kulesa, P.M.; Kasemeier-Kulesa, J.C.; Teddy, J.M.; Margaryan, N.V.; Seftor, E.A.; Seftor, R.E.; Hendrix, M.J. Reprogramming metastatic melanoma cells to assume a neural crest cell-like phenotype in an embryonic microenvironment. Proc. Natl. Acad. Sci. USA 2006, 103, 3752–3757. [Google Scholar] [CrossRef]
- White, R.M.; Cech, J.; Ratanasirintrawoot, S.; Lin, C.Y.; Rahl, P.B.; Burke, C.J.; Langdon, E.; Tomlinson, M.L.; Mosher, J.; Kaufman, C.; et al. DHODH modulates transcriptional elongation in the neural crest and melanoma. Nature 2011, 471, 518–522. [Google Scholar] [CrossRef]
- Santoriello, C.; Sporrij, A.; Yang, S.; Flynn, R.A.; Henriques, T.; Dorjsuren, B.; Custo Greig, E.; McCall, W.; Stanhope, M.E.; Fazio, M.; et al. RNA helicase DDX21 mediates nucleotide stress responses in neural crest and melanoma cells. Nat. Cell Biol. 2020, 22, 372–379. [Google Scholar] [CrossRef]
- Hanson, K.; Robinson, S.D.; Al-Yousuf, K.; Hendry, A.E.; Sexton, D.W.; Sherwood, V.; Wheeler, G.N. The anti-rheumatic drug, leflunomide, synergizes with MEK inhibition to suppress melanoma growth. Oncotarget 2018, 9, 3815–3829. [Google Scholar] [CrossRef]
- McConnell, A.M.; Mito, J.K.; Ablain, J.; Dang, M.; Formichella, L.; Fisher, D.E.; Zon, L.I. Neural crest state activation in NRAS driven melanoma, but not in NRAS-driven melanocyte expansion. Dev. Biol. 2019, 449, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Ciarlo, C.; Kaufman, C.K.; Kinikoglu, B.; Michael, J.; Yang, S.; D′Amato, C.; Blokzijl-Franke, S.; den Hertog, J.; Schlaeger, T.M.; Zhou, Y.; et al. A chemical screen in zebrafish embryonic cells establishes that Akt activation is required for neural crest development. Elife 2017, 6. [Google Scholar]
- White, R.M.; Sessa, A.; Burke, C.; Bowman, T.; LeBlanc, J.; Ceol, C.; Bourque, C.; Dovey, M.; Goessling, W.; Burns, C.E.; et al. Transparent adult zebrafish as a tool for in vivo transplantation analysis. Cell Stem Cell 2008, 2, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.S.; Heilmann, S.; Kansler, E.R.; Zhang, Y.; Zimmer, M.; Ratnakumar, K.; Bowman, R.L.; Simon-Vermot, T.; Fennell, M.; Garippa, R.; et al. Microenvironment-derived factors driving metastatic plasticity in melanoma. Nat. Commun. 2017, 8, 14343. [Google Scholar] [CrossRef] [PubMed]
- Heilmann, S.; Ratnakumar, K.; Langdon, E.; Kansler, E.; Kim, I.; Campbell, N.R.; Perry, E.; McMahon, A.; Kaufman, C.; van Rooijen, E.; et al. A Quantitative System for Studying Metastasis Using Transparent Zebrafish. Cancer Res. 2015, 75, 4272–4282. [Google Scholar] [CrossRef] [PubMed]
- Roh-Johnson, M.; Shah, A.N.; Stonick, J.A.; Poudel, K.R.; Kargl, J.; Yang, G.H.; di Martino, J.; Hernandez, R.E.; Gast, C.E.; Zarour, L.R.; et al. Macrophage-Dependent Cytoplasmic Transfer during Melanoma Invasion In Vivo. Dev. Cell 2017, 43, 549–562 e546. [Google Scholar] [CrossRef]
- Grivennikov, S.I.; Greten, F.R.; Karin, M. Immunity, inflammation, and cancer. Cell 2010, 140, 883–899. [Google Scholar] [CrossRef]
- Gomez-Abenza, E.; Ibanez-Molero, S.; Garcia-Moreno, D.; Fuentes, I.; Zon, L.I.; Mione, M.C.; Cayuela, M.L.; Gabellini, C.; Mulero, V. Zebrafish modeling reveals that SPINT1 regulates the aggressiveness of skin cutaneous melanoma and its crosstalk with tumor immune microenvironment. J. Exp. Clin. Cancer Res. 2019, 38, 405. [Google Scholar] [CrossRef]
- Nicoli, S.; Presta, M. The zebrafish/tumor xenograft angiogenesis assay. Nat. Protoc. 2007, 2, 2918–2923. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Zhang, W.; Zhao, Y.; Yang, Y.; Luo, H.; Ji, G.; Dong, E.; Deng, H.; Lin, S.; Wei, Y.; et al. Endothelial Cords Promote Tumor Initial Growth prior to Vascular Function through a Paracrine Mechanism. Sci. Rep. 2016, 6, 19404. [Google Scholar] [CrossRef] [PubMed]
- Gabellini, C.; Gomez-Abenza, E.; Ibanez-Molero, S.; Tupone, M.G.; Perez-Oliva, A.B.; de Oliveira, S.; Del Bufalo, D.; Mulero, V. Interleukin 8 mediates bcl-xL-induced enhancement of human melanoma cell dissemination and angiogenesis in a zebrafish xenograft model. Int. J. Cancer 2018, 142, 584–596. [Google Scholar] [CrossRef] [PubMed]
- Fornabaio, G.; Barnhill, R.L.; Lugassy, C.; Bentolila, L.A.; Cassoux, N.; Roman-Roman, S.; Alsafadi, S.; Del Bene, F. Angiotropism and extravascular migratory metastasis in cutaneous and uveal melanoma progression in a zebrafish model. Sci. Rep. 2018, 8, 10448. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.; Di Martino, J.S.; Bowman, R.L.; Campbell, N.R.; Baksh, S.C.; Simon-Vermot, T.; Kim, I.S.; Haldeman, P.; Mondal, C.; Yong-Gonzales, V.; et al. Adipocyte-Derived Lipids Mediate Melanoma Progression via FATP Proteins. Cancer Discov. 2018, 8, 1006–1025. [Google Scholar] [CrossRef] [PubMed]
- Bennett, C.M.; Kanki, J.P.; Rhodes, J.; Liu, T.X.; Paw, B.H.; Kieran, M.W.; Langenau, D.M.; Delahaye-Brown, A.; Zon, L.I.; Fleming, M.D.; et al. Myelopoiesis in the zebrafish, Danio rerio. Blood 2001, 98, 643–651. [Google Scholar] [CrossRef]
- Willett, C.E.; Kawasaki, H.; Amemiya, C.T.; Lin, S.; Steiner, L.A. Ikaros expression as a marker for lymphoid progenitors during zebrafish development. Dev. Dyn. 2001, 222, 694–698. [Google Scholar] [CrossRef]
- Willett, C.E.; Zapata, A.G.; Hopkins, N.; Steiner, L.A. Expression of zebrafish rag genes during early development identifies the thymus. Dev. Biol. 1997, 182, 331–341. [Google Scholar] [CrossRef]
- Greenhalgh, P.; Steiner, L.A. Recombination activating gene 1 (Rag1) in zebrafish and shark. Immunogenetics 1995, 41, 54–55. [Google Scholar] [CrossRef]
- Grigura, V.; Barbier, M.; Zarov, A.P.; Kaufman, C.K. Feeding amount significantly alters overt tumor onset rate in a zebrafish melanoma model. Biol. Open 2018, 7, bio030726. [Google Scholar] [CrossRef]
- Gramann, A.K.; Venkatesan, A.M.; Guerin, M.; Ceol, C.J. Regulation of zebrafish melanocyte development by ligand-dependent BMP signaling. Elife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Stoletov, K.; Montel, V.; Lester, R.D.; Gonias, S.L.; Klemke, R. High-resolution imaging of the dynamic tumor cell vascular interface in transparent zebrafish. Proc. Natl. Acad. Sci. USA 2007, 104, 17406–17411. [Google Scholar] [CrossRef] [PubMed]
- Haldi, M.; Ton, C.; Seng, W.L.; McGrath, P. Human melanoma cells transplanted into zebrafish proliferate, migrate, produce melanin, form masses and stimulate angiogenesis in zebrafish. Angiogenesis 2006, 9, 139–151. [Google Scholar] [CrossRef] [PubMed]
- Lieschke, G.J.; Trede, N.S. Fish immunology. Curr. Biol. 2009, 19, R678–R682. [Google Scholar] [CrossRef] [PubMed]
- Traver, D.; Winzeler, A.; Stern, H.M.; Mayhall, E.A.; Langenau, D.M.; Kutok, J.L.; Look, A.T.; Zon, L.I. Effects of lethal irradiation in zebrafish and rescue by hematopoietic cell transplantation. Blood 2004, 104, 1298–1305. [Google Scholar] [CrossRef]
- Yan, C.; Brunson, D.C.; Tang, Q.; Do, D.; Iftimia, N.A.; Moore, J.C.; Hayes, M.N.; Welker, A.M.; Garcia, E.G.; Dubash, T.D.; et al. Visualizing Engrafted Human Cancer and Therapy Responses in Immunodeficient Zebrafish. Cell 2019, 177, 1903–1914 e1914. [Google Scholar] [CrossRef]
- Tang, Q.; Abdelfattah, N.S.; Blackburn, J.S.; Moore, J.C.; Martinez, S.A.; Moore, F.E.; Lobbardi, R.; Tenente, I.M.; Ignatius, M.S.; Berman, J.N.; et al. Optimized cell transplantation using adult rag2 mutant zebrafish. Nat. Methods 2014, 11, 821–824. [Google Scholar] [CrossRef]
- Moore, J.C.; Tang, Q.; Yordan, N.T.; Moore, F.E.; Garcia, E.G.; Lobbardi, R.; Ramakrishnan, A.; Marvin, D.L.; Anselmo, A.; Sadreyev, R.I.; et al. Single-cell imaging of normal and malignant cell engraftment into optically clear prkdc-null SCID zebrafish. J. Exp. Med. 2016, 213, 2575–2589. [Google Scholar] [CrossRef]
- Peterson, R.T.; Link, B.A.; Dowling, J.E.; Schreiber, S.L. Small molecule developmental screens reveal the logic and timing of vertebrate development. Proc. Natl. Acad. Sci. USA 2000, 97, 12965–12969. [Google Scholar] [CrossRef]
- Precazzini, F.; Pancher, M.; Gatto, P.; Tushe, A.; Adami, V.; Anelli, V.; Mione, M.C. Automated in vivo screen in zebrafish identifies Clotrimazole as targeting a metabolic vulnerability in a melanoma model. Dev. Biol. 2020, 457, 215–225. [Google Scholar] [CrossRef]
- Peterson, R.T.; Shaw, S.Y.; Peterson, T.A.; Milan, D.J.; Zhong, T.P.; Schreiber, S.L.; MacRae, C.A.; Fishman, M.C. Chemical suppression of a genetic mutation in a zebrafish model of aortic coarctation. Nat. Biotechnol. 2004, 22, 595–599. [Google Scholar] [CrossRef] [PubMed]
- Yu, P.B.; Hong, C.C.; Sachidanandan, C.; Babitt, J.L.; Deng, D.Y.; Hoyng, S.A.; Lin, H.Y.; Bloch, K.D.; Peterson, R.T. Dorsomorphin inhibits BMP signals required for embryogenesis and iron metabolism. Nat. Chem. Biol. 2008, 4, 33–41. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Frantz, W.T.; Ceol, C.J. From Tank to Treatment: Modeling Melanoma in Zebrafish. Cells 2020, 9, 1289. https://doi.org/10.3390/cells9051289
Frantz WT, Ceol CJ. From Tank to Treatment: Modeling Melanoma in Zebrafish. Cells. 2020; 9(5):1289. https://doi.org/10.3390/cells9051289
Chicago/Turabian StyleFrantz, William Tyler, and Craig J Ceol. 2020. "From Tank to Treatment: Modeling Melanoma in Zebrafish" Cells 9, no. 5: 1289. https://doi.org/10.3390/cells9051289
APA StyleFrantz, W. T., & Ceol, C. J. (2020). From Tank to Treatment: Modeling Melanoma in Zebrafish. Cells, 9(5), 1289. https://doi.org/10.3390/cells9051289



