A Clonal NG2-Glia Cell Response in a Mouse Model of Multiple Sclerosis
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. StarTrack DNA Vectors
2.3. In Utero Electroporation (IUE)
2.4. EAE Induction
2.5. Immunohistochemistry
2.6. Imaging Acquisition and Data Analysis
3. Results
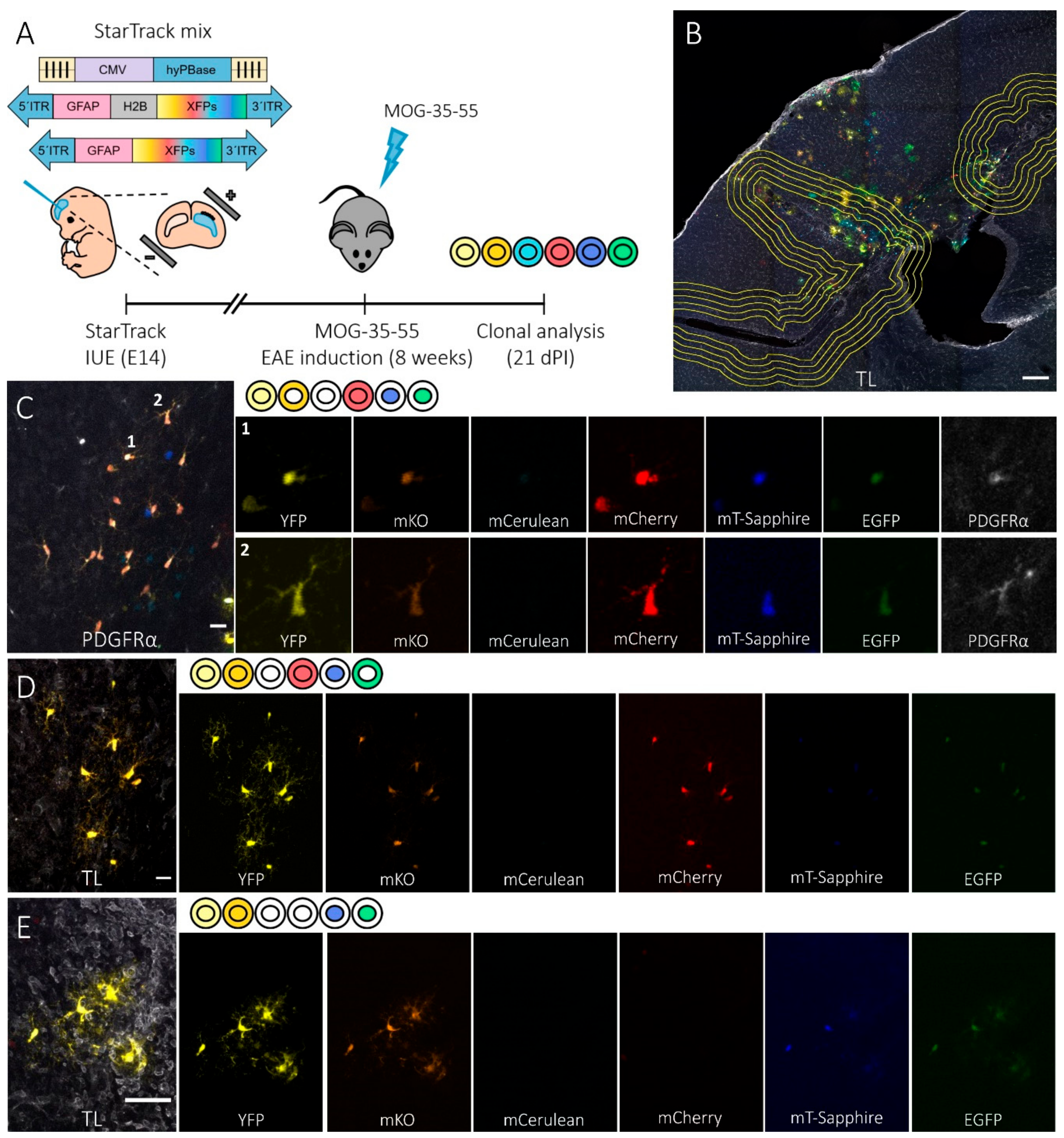
3.1. Spatial Distribution of the Cortical Progeny of NG2-Glia Derived from Single Embryonic Progenitors in EAE-Lesioned Mouse Brain
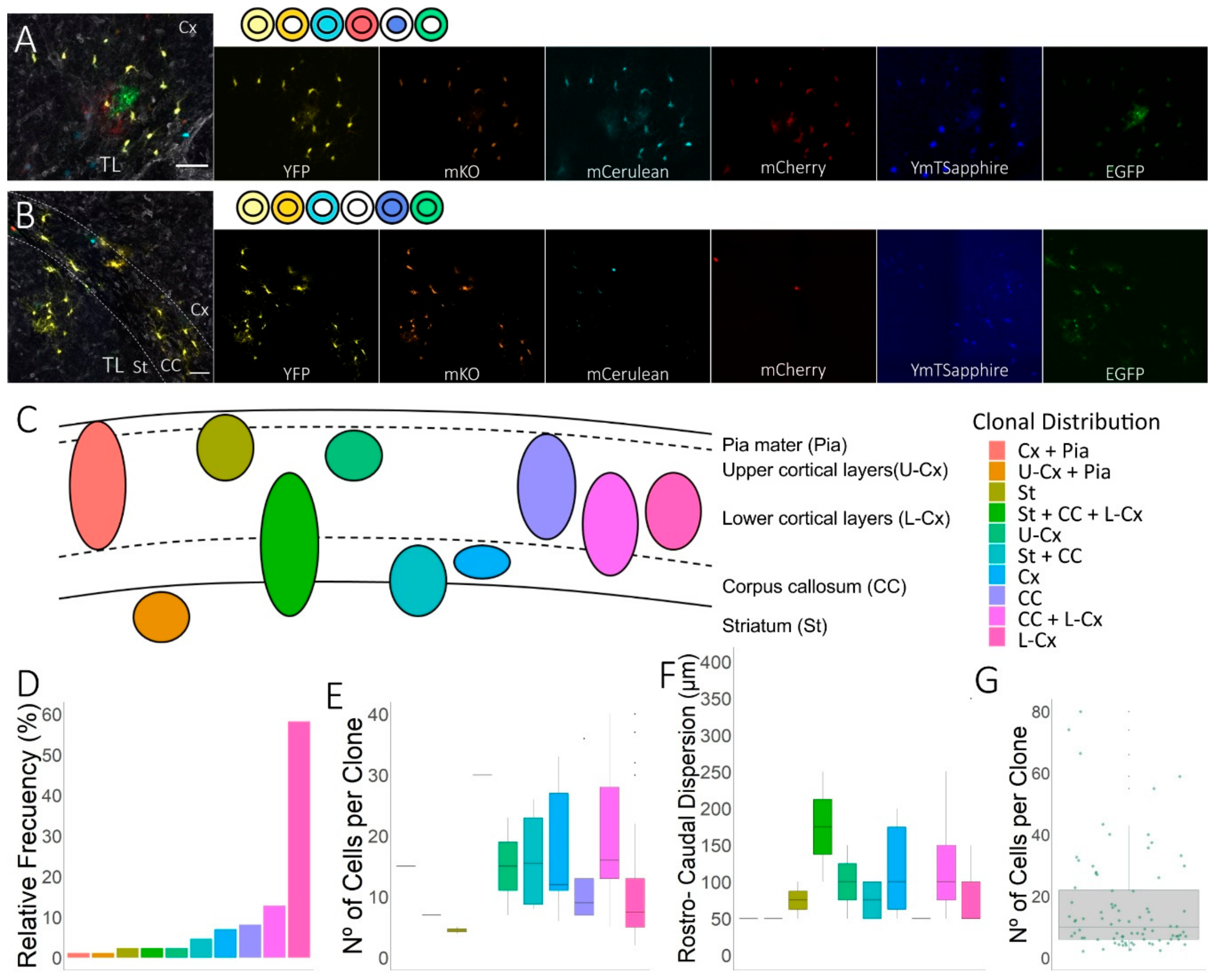
3.2. Clonal Distribution of the Cell Progeny from Individual Embryonic Pallial NG2-Glia Progenitors in EAE-Lesioned Brains
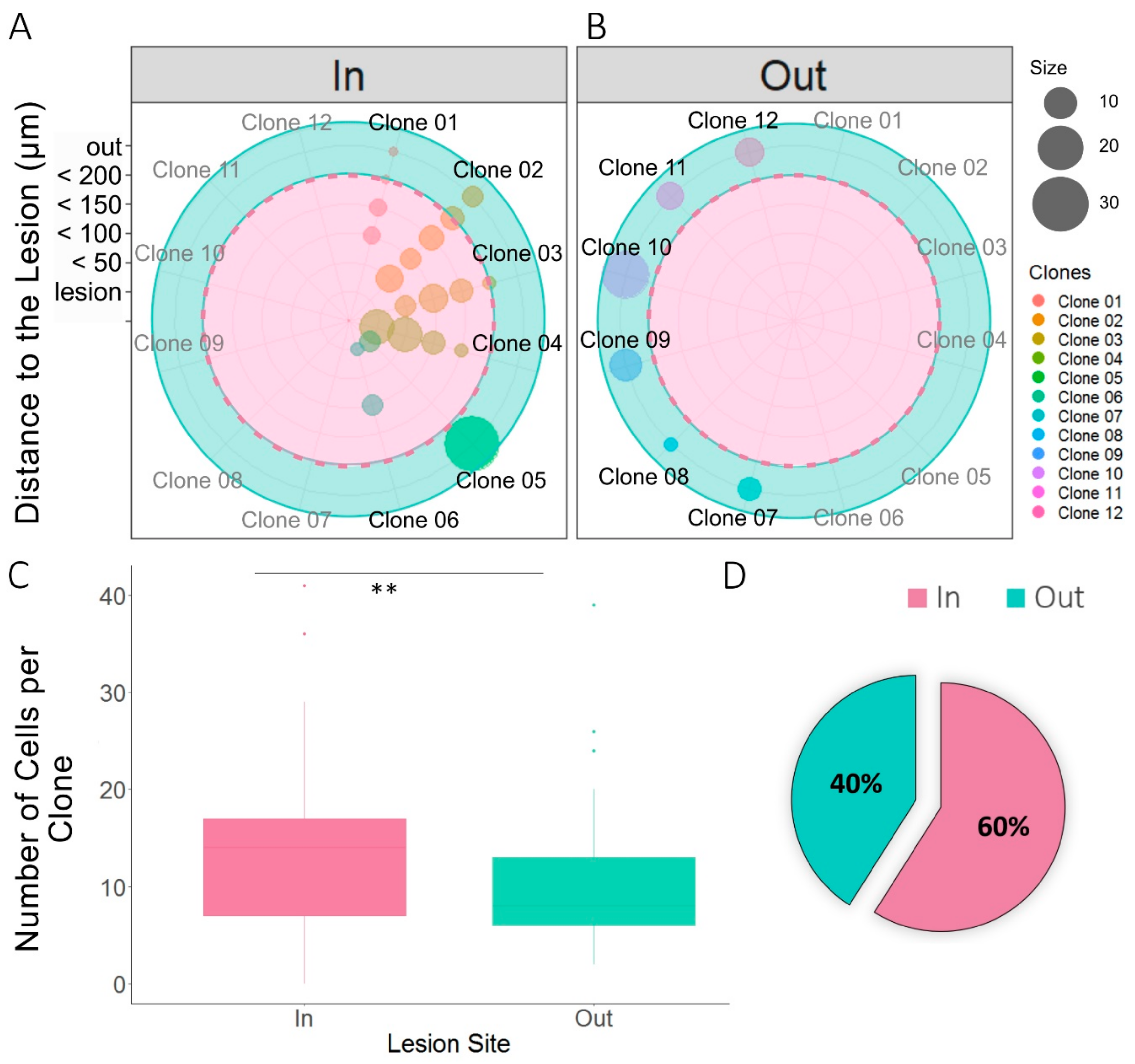
3.3. Relationship Between Sibling NG2-Glia and EAE Brain Lesions
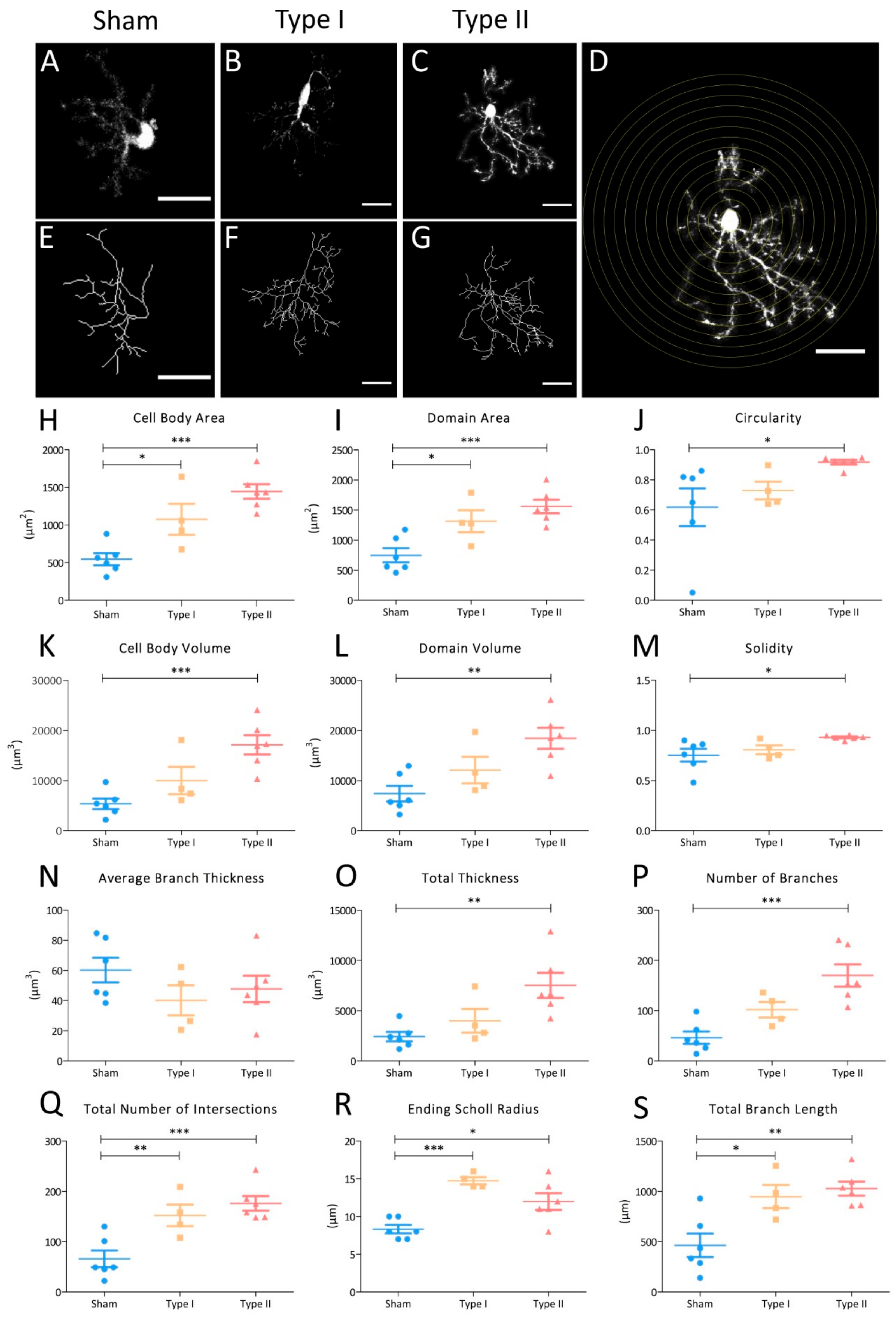
3.4. Morphometric Analyses of NG2-Glia in EAE Lesions
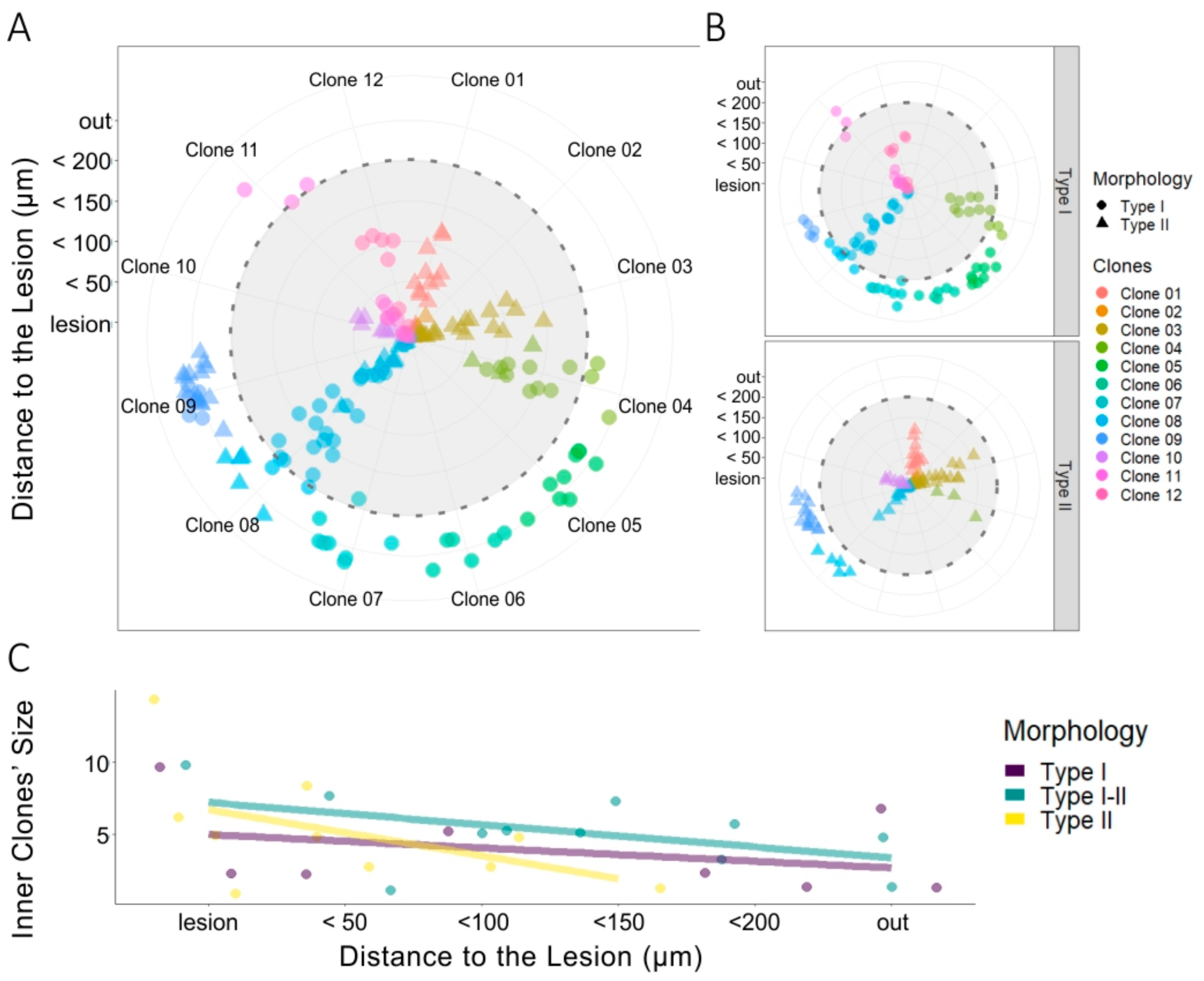
3.5. Clonal NG2-Glia Response to EAE Lesion
4. Discussion
4.1. Heterogeneity in the Clonal Distribution of NG2-Glia in a Mouse Model of MS
4.2. Heterogenic and Clonal NG2-Glia Responses to EAE Lesions
4.3. NG2-Glia Activate in Response to EAE Lesions, Undergoing a Progressive Morphological Change to Reactivity Characterized by an Increase in Their Size, Domain, and Arborization
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Lassmann, H. Cortical lesions in multiple sclerosis: Inflammation versus neurodegeneration. Brain 2012, 135, 2904–2905. [Google Scholar] [CrossRef] [PubMed]
- Prins, M.; Schul, E.; Geurts, J.; van der Valk, P.; Drukarch, B.; van Dam, A.M. Pathological differences between white and grey matter multiple sclerosis lesions. Ann. N. Y. Acad. Sci. 2015, 1351, 99–113. [Google Scholar] [CrossRef] [PubMed]
- Van Munster, C.E.P.; Jonkman, L.E.; Weinstein, H.C.; Uitdehaag, B.M.; Geurts, J.J. Gray matter damage in multiple sclerosis: Impact on clinical symptoms. Neuroscience 2015, 303, 446–461. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, A.; Chang, A.; Trapp, B.D. NG2+ glial cells: A novel glial cell population in the adult brain. J. Neuropathol. Exp. Neurol. 1999, 58, 1113–1124. [Google Scholar] [CrossRef]
- Hill, R.A.; Nishiyama, A. NG2 cells (polydendrocytes): Listeners to the neural network with diverse properties. Glia 2014, 62, 1195–1210. [Google Scholar] [CrossRef]
- Dimou, L.; Gallo, V. NG2-glia and their functions in the central nervous system. Glia 2015, 63, 1429–1451. [Google Scholar] [CrossRef]
- Orduz, D.; Maldonado, P.P.; Balia, M.; Vélez-Fort, M.; de Sars, V.; Yanagawa, Y.; Emiliani, V.; Angulo, M.C. Interneurons and oligodendrocyte progenitors form a structured synaptic network in the developing neocortex. Elife 2015, 2015, 1–53. [Google Scholar] [CrossRef]
- Nakano, M.; Tamura, Y.; Yamato, M.; Kume, S.; Eguchi, A.; Takata, K.; Watanabe, Y.; Kataoka, Y. NG2 glial cells regulate neuroimmunological responses to maintain neuronal function and survival. Sci. Rep. 2017, 7, 42041. [Google Scholar] [CrossRef]
- Xiao, L.; Ohayon, D.; McKenzie, I.A.; Sinclair-Wilson, A.; Wright, J.L.; Fudge, A.D.; Emery, B.; Li, H.; Richardson, W.D. Rapid production of new oligodendrocytes is required in the earliest stages of motor-skill learning. Nat. Neurosci. 2016, 19, 1210–1217. [Google Scholar] [CrossRef]
- Hill, R.A.; Li, A.M.; Grutzendler, J. Lifelong cortical myelin plasticity and age-related degeneration in the live mammalian brain. Nat. Neurosci. 2018, 21, 683–695. [Google Scholar] [CrossRef]
- Hughes, E.G.; Orthmann-Murphy, J.L.; Langseth, A.J.; Bergles, D.E. Myelin remodeling through experience-dependent oligodendrogenesis in the adult somatosensory cortex. Nat. Neurosci. 2018, 21, 696–706. [Google Scholar] [CrossRef] [PubMed]
- Van Tilborg, E.; de Theije, C.G.M.; van Hal, M.; Wagenaar, N.; de Vries, L.S.; Benders, M.J.; Rowitch, D.H.; Nijboer, C.H. Origin and dynamics of oligodendrocytes in the developing brain: Implications for perinatal white matter injury. Glia 2018, 66, 221–238. [Google Scholar] [CrossRef] [PubMed]
- García-Marqués, J.; Núñez-Llaves, R.; López-Mascaraque, L. NG2-glia from pallial progenitors produce the largest clonal clusters of the brain: Time frame of clonal generation in cortex and olfactory bulb. J. Neurosci. 2014, 34, 13–2305. [Google Scholar] [CrossRef] [PubMed]
- Boda, E.; Di Maria, S.; Rosa, P.; Taylor, V.; Abbracchio, M.P.; Buffo, A. Early phenotypic asymmetry of sister oligodendrocyte progenitor cells after mitosis and its modulation by aging and extrinsic factors. Glia 2015, 63, 271–286. [Google Scholar] [CrossRef] [PubMed]
- Keirstead, H.S.; Levine, J.M.; Blakemore, W.F. Response of the oligodendrocyte progenitor cell population (Defined by NG2 labelling) to demyelination of the adult spinal cord. Glia 1998, 22, 161–170. [Google Scholar] [CrossRef]
- McTigue, D.M.; Wei, P.; Stokes, B.T. Proliferation of NG2-positive cells and altered oligodendrocyte numbers in the contused rat spinal cord. J. Neurosci. 2001, 21, 3392–3400. [Google Scholar] [CrossRef]
- Jin, X.; Riew, T.-R.; Kim, H.L.; Choi, J.-H.; Lee, M.-Y. Morphological characterization of NG2 glia and their association with neuroglial cells in the 3-nitropropionic acid-lesioned striatum of rat. Sci. Rep. 2018, 8, 5942. [Google Scholar] [CrossRef]
- Martín-López, E.; García-Marques, J.; Núñez-Llaves, R.; López-Mascaraque, L. Clonal astrocytic response to cortical injury. PLoS ONE 2013, 8, e74039. [Google Scholar] [CrossRef]
- Bribian, A.; Pérez-Cerdá, F.; Matute, C.; López-Mascaraque, L. Clonal Glial Response in a Multiple Sclerosis Mouse Model. Front. Cell. Neurosci. 2018, 12, 375. [Google Scholar] [CrossRef]
- Okada, S.; Hara, M.; Kobayakawa, K.; Matsumoto, Y.; Nakashima, Y. Astrocyte reactivity and astrogliosis after spinal cord injury. Neurosci. Res. 2018, 126, 39–43. [Google Scholar] [CrossRef]
- Hughes, E.G.; Kang, S.H.; Fukaya, M.; Bergles, D.E. Oligodendrocyte progenitors balance growth with self-repulsion to achieve homeostasis in the adult brain. Nat. Neurosci. 2013, 16, 668–676. [Google Scholar] [CrossRef] [PubMed]
- Valny, M.; Honsa, P.; Waloschkova, E.; Matuskova, H.; Kriska, J.; Kirdajova, D.; Androvic, P.; Valihrach, L.; Kubista, M.; Anderova, M. A single-cell analysis reveals multiple roles of oligodendroglial lineage cells during post-ischemic regeneration. Glia 2018, 66, 1068–1081. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, Y.; Tozuka, Y.; Takata, T.; Shimazu, N.; Matsumura, N.; Ohta, A.; Hisatsune, T. Excitatory GABAergic activation of cortical dividing glial cells. Cereb. Cortex 2009, 19, 2181–2195. [Google Scholar] [CrossRef] [PubMed]
- Kang, Z.; Wang, C.; Zepp, J.; Wu, L.; Sun, K.; Zhao, J.; Chandrasekharan, U.; DiCorleto, P.E.; Trapp, B.D.; Ransohoff, R.M.; et al. Act1 mediates IL-17-induced EAE pathogenesis selectively in NG2+ glial cells. Nat. Neurosci. 2013, 16, 8–1401. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.H.; Miyamoto, N.; Hayakawa, K.; Pham, L.D.D.; Maki, T.; Ayata, C.; Kim, K.W.; Lo, E.H.; Arai, K. Oligodendrocyte precursors induce early blood-brain barrier opening after white matter injury. J. Clin. Invest. 2013, 123, 782–786. [Google Scholar] [CrossRef]
- Rodriguez, J.P.; Coulter, M.; Miotke, J.; Meyer, R.L.; Takemaru, K.I.; Levine, J.M. Abrogation of β-Catenin signaling in oligodendrocyte precursor cells reduces glial scarring and promotes axon regeneration after CNS injury. J. Neurosci. 2014, 34, 10285–10297. [Google Scholar] [CrossRef]
- Liu, J.; Dietz, K.; Hodes, G.E.; Russo, S.J.; Casaccia, P. Widespread transcriptional alternations in oligodendrocytes in the adult mouse brain following chronic stress. Dev. Neurobiol. 2018, 78, 152–162. [Google Scholar] [CrossRef]
- Kessaris, N.; Fogarty, M.; Iannarelli, P.; Grist, M.; Wegner, M.; Richardson, W.D. Competing waves of oligodendrocytes in the forebrain and postnatal elimination of an embryonic lineage. Nat. Neurosci. 2006, 9, 173–179. [Google Scholar] [CrossRef]
- Winkler, C.C.; Yabut, O.R.; Fregoso, S.P.; Gomez, H.G.; Dwyer, B.E.; Pleasure, S.J.; Franco, S.J. The Dorsal Wave of Neocortical Oligodendrogenesis Begins Embryonically and Requires Multiple Sources of Sonic Hedgehog. J. Neurosci. 2018, 38, 5237–5250. [Google Scholar] [CrossRef]
- Tong, C.K.; Fuentealba, L.C.; Shah, J.K.; Lindquist, R.A.; Ihrie, R.A.; Guinto, C.D.; Rodas-Rodriguez, J.L.; Alvarez-Buylla, A. A dorsal SHH-dependent domain in the V-SVZ produces large numbers of oligodendroglial lineage cells in the postnatal brain. Stem Cell Rep. 2015, 5, 461–470. [Google Scholar] [CrossRef]
- García-Marqués, J.; López-Mascaraque, L. Clonal Identity Determines Astrocyte Cortical Heterogeneity. Cereb. Cortex 2013, 23(6), 1463–1475. [Google Scholar]
- Matute, C.; Torre, I.; Pérez-Cerdá, F.; Pérez-Samartín, A.; Alberdi, E.; Etxebarria, E.; Arranz, A.M.; Ravid, R.; Rodríguez-Antigüedad, A.; Sánchez-Gómez, M.; et al. P2X(7) receptor blockade prevents ATP excitotoxicity in oligodendrocytes and ameliorates experimental autoimmune encephalomyelitis. J. Neurosci. 2007, 27, 33–9525. [Google Scholar] [CrossRef] [PubMed]
- Pampliega, O.; Domercq, M.; Soria, F.N.; Villoslada, P.; Rodríguez-Antigüedad, A.; Matute, C. Increased expression of cystine/glutamate antiporter in multiple sclerosis. J. Neuroinflammation 2011, 8, 63. [Google Scholar] [CrossRef] [PubMed]
- Tavares, G.; Martins, M.; Correia, J.S.; Sardinha, V.M.; Guerra-Gomes, S.; das Neves, S.P.; Marques, F.; Sousa, N.; Oliveira, J.F. Employing an open-source tool to assess astrocyte tridimensional structure. Brain Struct. Funct. 2017, 222, 1989–1999. [Google Scholar] [CrossRef]
- Scholl, D.A. Dendritic organization in the neurons of the visual and motor cortices of the cat. J. Anat. 1953, 87, 387–406. [Google Scholar]
- Yamada, J.; Jinno, S. Novel objective classification of reactive microglia following hypoglossal axotomy using hierarchical cluster analysis. J. Comp. Neurol. 2013, 521, 1184–1201. [Google Scholar] [CrossRef]
- Figueres-Onãte, M.; Garciá-Marqués, J.; López-Mascaraque, L. UbC-StarTrack, a clonal method to target the entire progeny of individual progenitors. Sci. Rep. 2016, 6, 33896. [Google Scholar] [CrossRef]
- Butt, A.M.; Hamilton, N.; Hubbard, P.; Pugh, M.; Ibrahim, M. Synantocytes: The fifth element. J. Anat. 2005, 207, 695–706. [Google Scholar] [CrossRef]
- Khakh, B.S.; Sofroniew, M.V. Diversity of astrocyte functions and phenotypes in neural circuits. Nat. Neurosci. 2015, 18, 942–952. [Google Scholar] [CrossRef]
- Malatesta, P.; Hack, M.A.; Hartfuss, E.; Kettenmann, H.; Klinkert, W.; Kirchhoff, F.; Götz, M. Neuronal or glial progeny: Regional differences in radial glia fate. Neuron 2003, 37, 751–764. [Google Scholar] [CrossRef]
- Simon, C.; Götz, M.; Dimou, L. Progenitors in the adult cerebral cortex: Cell cycle properties and regulation by physiological stimuli and injury. Glia 2011, 59, 869–881. [Google Scholar] [CrossRef] [PubMed]
- Dimou, L.; Götz, M. Glial cells as progenitors and stem cells: New roles in the healthy and diseased brain. Physiol. Rev. 2014, 94, 709–737. [Google Scholar] [CrossRef] [PubMed]
- Hesp, Z.C.; Yoseph, R.Y.; Suzuki, R.; Jukkola, P.; Wilson, C.; Nishiyama, A.; McTigue, D.M. Proliferating NG2-Cell-Dependent Angiogenesis and Scar Formation Alter Axon Growth and Functional Recovery After Spinal Cord Injury in Mice. J. Neurosci. 2018, 38, 1366–1382. [Google Scholar] [CrossRef] [PubMed]
- Jäkel, S.; Dimou, L. Glial cells and their function in the adult brain: A journey through the history of their ablation. Front. Cell. Neurosci. 2017, 11, 24. [Google Scholar] [CrossRef]
- Bonfanti, E.; Gelosa, P.; Fumagalli, M.; Dimou, L.; Viganò, F.; Tremoli, E.; Cimino, M.; Sironi, L.; Abbracchio, M.P. The role of oligodendrocyte precursor cells expressing the GPR17 receptor in brain remodeling after stroke. Cell Death Dis. 2017, 8, e2871. [Google Scholar] [CrossRef]
- Boda, E.; Viganò, F.; Rosa, P.; Fumagalli, M.; Labat-Gest, V.; Tempia, F.; Abbracchio, M.P.; Dimou, L.; Buffo, A. The GPR17 receptor in NG2 expressing cells: Focus on in vivocell maturation and participation in acute trauma and chronic damage. Glia 2011, 59, 1958–1973. [Google Scholar] [CrossRef]
- Kirby, B.B.; Takada, N.; Latimer, A.J.; Shin, J.; Carney, T.J.; Kelsh, R.N.; Appel, B. In vivo time-lapse imaging shows dynamic oligodendrocyte progenitor behavior during zebrafish development. Nat. Neurosci. 2006, 9, 1506–1511. [Google Scholar] [CrossRef]
- Parolisi, R.; Boda, E. NG2 Glia: Novel Roles beyond Re-/Myelination. Neuroglia 2018, 1, 151–175. [Google Scholar] [CrossRef]
- Levine, J.M. Increased expression of the NG2 chondroitin-sulfate proteoglycan after brain injury. J. Neurosci. 1994, 14, 4716–4730. [Google Scholar] [CrossRef]
- Wight, T.N.; Kinsella, M.G.; Qwarnström, E.E. The role of proteoglycans in cell adhesion, migration and proliferation. Curr. Opin. Cell Biol. 1992, 4, 793–801. [Google Scholar] [CrossRef]
- Schäfer, M.K.E.; Tegeder, I. NG2/CSPG4 and progranulin in the posttraumatic glial scar. Matrix Biol. 2018, 68-69, 571–588. [Google Scholar]
- Heinrich, C.; Bergami, M.; Gascón, S.; Lepier, A.; Viganò, F.; Dimou, L.; Sutor, B.; Berninger, B.; Götz, M. Sox2-mediated conversion of NG2 glia into induced neurons in the injured adult cerebral cortex. Stem Cell Rep. 2014, 3, 1000–1014. [Google Scholar] [CrossRef] [PubMed]
- Pereira, M.; Birtele, M.; Shrigley, S.; Benitez, J.A.; Hedlund, E.; Parmar, M.; Ottosson, D.R. Direct Reprogramming of Resident NG2 Glia into Neurons with Properties of Fast-Spiking Parvalbumin-Containing Interneurons. Stem Cell Rep. 2017, 9, 742–751. [Google Scholar] [CrossRef] [PubMed]
- Yavarpour-Bali, H.; Ghasemi-Kasman, M.; Shojaei, A. Direct reprogramming of terminally differentiated cells into neurons: A novel and promising strategy for Alzheimer’s disease treatment. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2020, 98, 109820. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Barriola, S.; Pérez-Cerdá, F.; Matute, C.; Bribián, A.; López-Mascaraque, L. A Clonal NG2-Glia Cell Response in a Mouse Model of Multiple Sclerosis. Cells 2020, 9, 1279. https://doi.org/10.3390/cells9051279
Barriola S, Pérez-Cerdá F, Matute C, Bribián A, López-Mascaraque L. A Clonal NG2-Glia Cell Response in a Mouse Model of Multiple Sclerosis. Cells. 2020; 9(5):1279. https://doi.org/10.3390/cells9051279
Chicago/Turabian StyleBarriola, Sonsoles, Fernando Pérez-Cerdá, Carlos Matute, Ana Bribián, and Laura López-Mascaraque. 2020. "A Clonal NG2-Glia Cell Response in a Mouse Model of Multiple Sclerosis" Cells 9, no. 5: 1279. https://doi.org/10.3390/cells9051279
APA StyleBarriola, S., Pérez-Cerdá, F., Matute, C., Bribián, A., & López-Mascaraque, L. (2020). A Clonal NG2-Glia Cell Response in a Mouse Model of Multiple Sclerosis. Cells, 9(5), 1279. https://doi.org/10.3390/cells9051279






