Epigenetics in Inflammatory Breast Cancer: Biological Features and Therapeutic Perspectives
Abstract
1. Introduction
2. Clinical and Epidemiological Features
3. Molecular Profiling of Inflammatory Breast Carcinomas
4. Epigenetic of Mammary Gland Differentiation and Development
5. Epigenetic Alterations in Inflammatory and Non-Inflammatory Breast Cancer
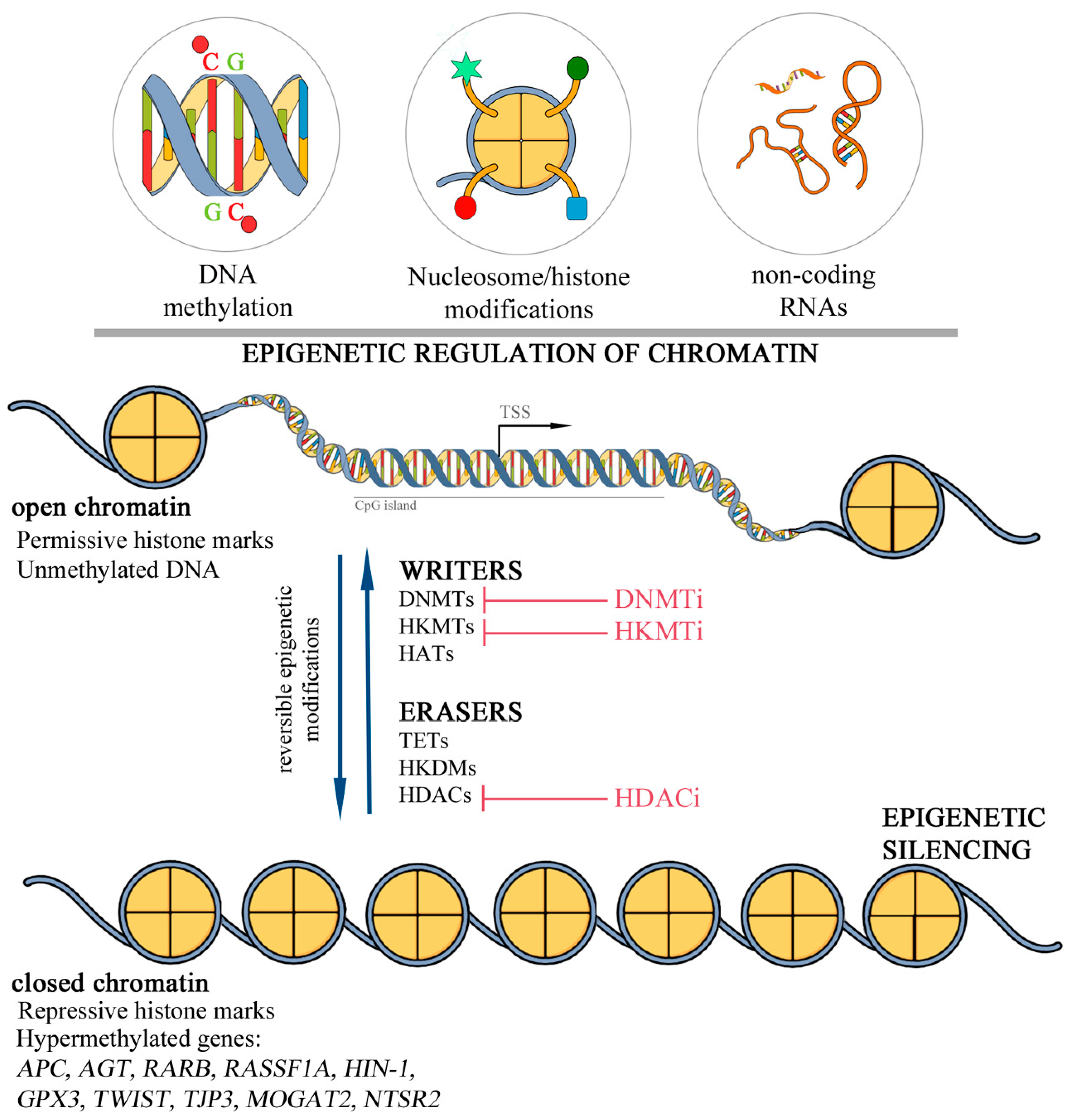
6. Evidence of Disrupted Epigenetic Program in the Inflammatory Breast Cancer Development
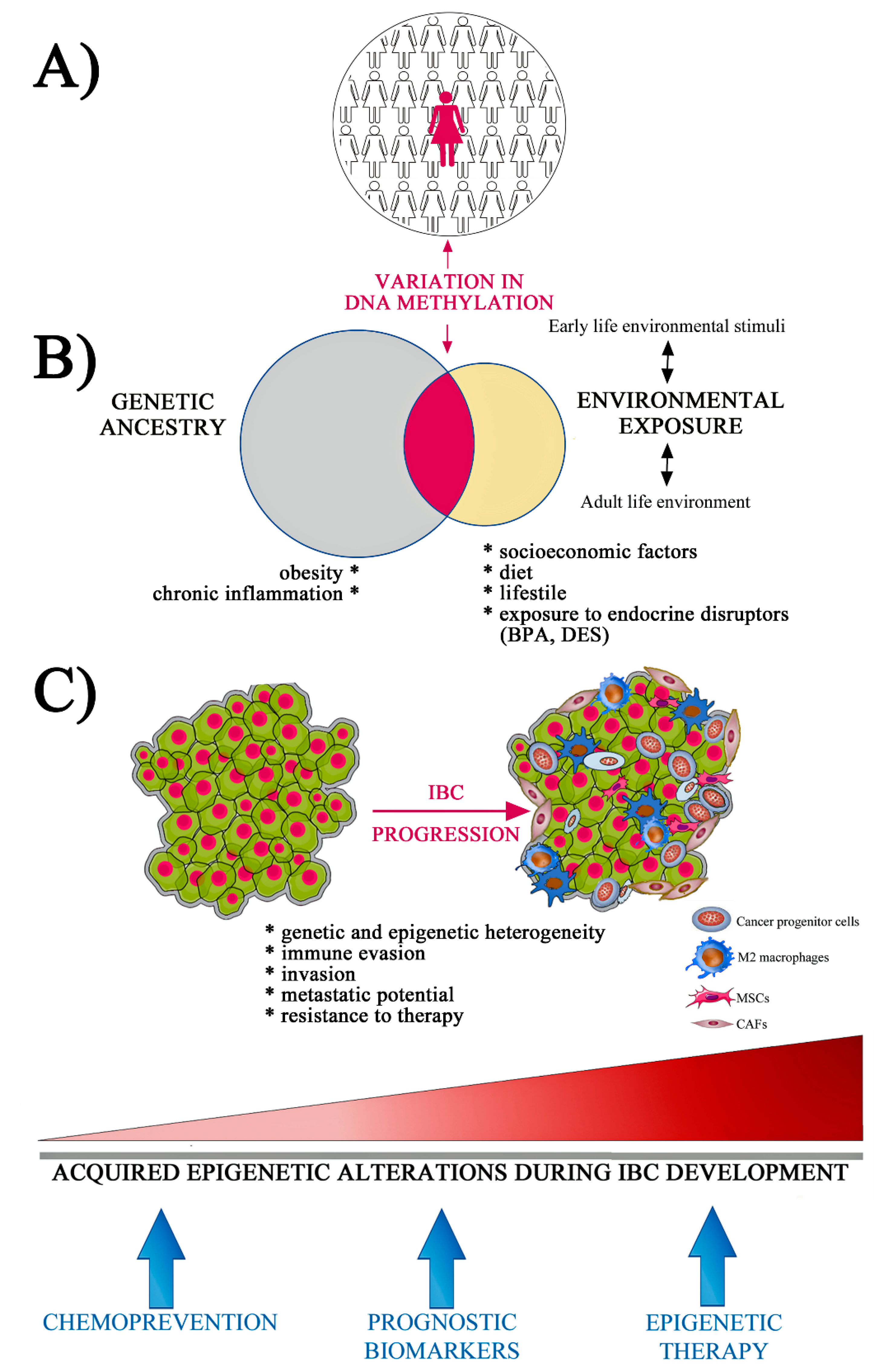
6.1. Racial Disparities and Epigenetic Alterations
6.2. Obesity and Body Mass Index (BMI)
6.3. Environmental Exposure to Endocrine Disruptors
6.4. Stemness and Microenvironment
6.5. Inflammatory Pathways
7. Epigenetic Therapy: The Inhibitors of Histone Deacetylases (HDACi)
8. Final Remarks and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer Statistics. CA Cancer J. Clin. 2017, 67, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Anderson, W.F.; Schairer, C.; Chen, B.E.; Hance, K.W.; Levine, P.H. Epidemiology of inflammatory breast cancer (IBC). Breast Dis. 2005–2006, 22, 9–23. [Google Scholar] [CrossRef] [PubMed]
- Aurit, S.J.; Devesa, S.S.; Soliman, A.S.; Schairer, C. Inflammatory and other breast cancer incidence rate trends by estrogen receptor status in the Surveillance, Epidemiology, and End Results database (2001–2015). Breast Cancer Res. Treat. 2019, 175, 755–764. [Google Scholar] [CrossRef] [PubMed]
- Hance, K.W.; Anderson, W.F.; Devesa, S.S.; Young, H.A.; Levine, P.H. Trends in inflammatory breast carcinoma incidence and survival: The Surveillance, Epidemiology, and End Results program at the National Cancer Institute. J. Natl. Cancer Inst. 2005, 97, 966–975. [Google Scholar] [CrossRef]
- Dawood, S.; Merajver, S.D.; Viens, P.; Vermeulen, P.B.; Swain, S.M.; Buchholz, T.A.; Dirix, L.Y.; Levine, P.H.; Lucci, A.; Krishnamurthy, S.; et al. International expert panel on inflammatory breast cancer: Consensus statement for standardized diagnosis and treatment. Ann. Oncol. 2005, 22, 515–523. [Google Scholar] [CrossRef]
- Lim, B.; Woodward, W.A.; Wang, X.; Reuben, J.M.; Ueno, N.T. Inflammatory breast cancer biology: The tumour microenvironment is key. Nat. Rev. Cancer 2018, 18, 485–499. [Google Scholar] [CrossRef]
- Fouad, T.M.; Kogawa, T.; Liu, D.D.; Shen, Y.; Masuda, H.; El-Zein, R.; Woodward, W.A.; Chavez-MacGregor, M.; Alvarez, R.H.; Arun, B.; et al. Overall survival differences between patients with inflammatory and noninflammatory breast cancer presenting with distant metastasis at diagnosis. Breast Cancer Res. Treat. 2015, 152, 407–416. [Google Scholar] [CrossRef]
- Kalish, J.M.; Jiang, C.; Bartolomei, M.S. Epigenetics and imprinting in human disease. Int. J. Dev. Biol. 2014, 58, 291–298. [Google Scholar] [CrossRef]
- Pasculli, B.; Barbano, R.; Parrella, P. Epigenetics of breast cancer: Biology and clinical implication in the era of precision medicine. Semin. Cancer Biol. 2018, 51, 22–35. [Google Scholar] [CrossRef]
- Esteller, M. Cancer epigenetics for the 21st century: What’s next? Genes Cancer 2011, 2, 604–606. [Google Scholar] [CrossRef]
- Cava, C.; Bertoli, G.; Castiglioni, I. Integrating genetics and epigenetics in breast cancer: Biological insights, experimental, computational methods and therapeutic potential. BMC Syst. Biol. 2015, 9, 62. [Google Scholar] [CrossRef] [PubMed]
- Widschwendter, M.; Apostolidou, S.; Raum, E.; Rothenbacher, D.; Fiegl, H.; Menon, U.; Stegmaier, C.; Jacobs, I.J.; Brenner, H. Epigenotyping in peripheral blood cell DNA and breast cancer risk: A proof of principle study. PLoS ONE 2008, 3, e2656. [Google Scholar] [CrossRef] [PubMed]
- Leygo, C.; Williams, M.; Jin, H.C.; Chan, M.W.Y.; Chu, W.K.; Grusch, M.; Cheng, Y.Y. DNA methylation as a noninvasive epigenetic biomarker for the detection of cancer. Dis. Markers 2017, 2017, 3726595. [Google Scholar] [CrossRef] [PubMed]
- Bhat, S.A.; Majid, S.; Wani, H.A.; Rashid, S. Diagnostic utility of epigenetics in Breast Cancer—A Review. Cancer Treat. Res. Commun. 2019, 19, 100125. [Google Scholar] [CrossRef]
- van Hoesel, A.Q.; Sato, Y.; Elashoff, D.A.; Turner, R.R.; Giuliano, A.E.; Shamonki, J.M.; Kuppen, P.J.; van de Velde, C.J.; Hoon, D.S. Assessment of DNA methylation status in early stages of breast cancer development. Br. J. Cancer 2013, 108, 2033–2038. [Google Scholar] [CrossRef]
- Martínez-Galán, J.; Torres-Torres, B.; Núñez, M.I.; López-Peñalver, J.; Del Moral, R.; Ruiz De Almodóvar, J.M.; Menjón, S.; Concha, A.; Chamorro, C.; Ríos, S.; et al. ESR1 gene promoter region methylation in free circulating DNA and its correlation with estrogen receptor protein expression in tumor tissue in breast cancer patients. BMC Cancer 2014, 14, 59. [Google Scholar] [CrossRef]
- Singla, H.; Ludhiadch, A.; Kaur, R.P.; Chander, H.; Kumar, V.; Munshi, A. Recent advances in HER2 positive breast cancer epigenetics: Susceptibility and therapeutic strategies. Eur. J. Med. Chem. 2017, 142, 316–327. [Google Scholar] [CrossRef]
- Palomeras, S.; Diaz-Lagares, Á.; Viñas, G.; Setien, F.; Ferreira, H.J.; Oliveras, G.; Crujeiras, A.B.; Hernández, A.; Lum, D.H.; Welm, A.L.; et al. Epigenetic silencing of TGFBI confers resistance to trastuzumab in human breast cancer. Breast Cancer Res. 2019, 21, 79. [Google Scholar] [CrossRef]
- van Uden, D.J.; van Laarhoven, H.W.; Westenberg, A.H.; de Wilt, J.H.; Blanken-Peeters, C.F. Inflammatory breast cancer: An overview. Crit. Rev. Oncol. Hematol. 2015, 93, 116–126. [Google Scholar] [CrossRef]
- Lehman, H.L.; Dashner, E.J.; Lucey, M.; Vermeulen, P.; Dirix, L.; Van Laere, S.; van Golen, K.L. Modeling and characterization of inflammatory breast cancer emboli grown in vitro. Int. J. Cancer 2013, 132, 2283–2294. [Google Scholar] [CrossRef]
- Woodward, W.A. Inflammatory breast cancer: Unique biological and therapeutic considerations. Lancet Oncol. 2015, 16, e568–e576. [Google Scholar] [CrossRef]
- Amin, M.B.; Edge, S.; Greene, F.; Byrd, D.R.; Brookland, R.K.; Washington, M.K.; Gershenwald, J.E.; Compton, C.C.; Hess, K.R.; Sullivan, D.C.; et al. AJCC Cancer Staging Manual—Breast Neoplasms, 8th ed.; Springer: New York, NY, USA, 2017; p. 1032. [Google Scholar]
- Soliman, A.S.; Kleer, C.G.; Mrad, K.; Karkouri, M.; Omar, S.; Khaled, H.M.; Benider, A.L.; Ayed, F.B.; Eissa, S.S.; Eissa, M.S.; et al. Inflammatory breast cancer in north Africa: Comparison of clinical and molecular epidemiologic characteristics of patients from Egypt, Tunisia, and Morocco. Breast Dis. 2011–2012, 33, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Yood, M.U.; Johnson, C.C.; Blount, A.; Abrams, J.; Wolman, E.; McCarthy, B.D.; Raju, U.; Nathanson, D.S.; Worsham, M.; Wolman, S.R. Race and differences in breast cancer survival in a managed care population. J. Natl. Cancer Inst. 1999, 91, 1487–1491. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Chang, S.; Buzdar, A.U.; Hursting, S.D. Inflammatory breast cancer and body mass index. J. Clin. Oncol. 1998, 16, 3731–3735. [Google Scholar] [CrossRef] [PubMed]
- Newman, L.A.; Kaljee, L.M. Health disparities and triple-negative breast cancer in African American women: A Review. JAMA Surg. 2017, 152, 485–493. [Google Scholar] [CrossRef]
- Yang, R.; Cheung, M.C.; Hurley, J.; Byrne, M.M.; Huang, Y.; Zimmers, T.A.; Koniaris, L.G. A comprehensive evaluation of outcomes for inflammatory breast cancer. Breast Cancer Res. Treat. 2009, 17, 631–641. [Google Scholar] [CrossRef]
- Schlichting, J.A.; Soliman, A.S.; Schairer, C.; Schottenfeld, D.; Merajver, S.D. Inflammatory and non-inflammatory breast cancer survival by socioeconomic position in the Surveillance, Epidemiology, and End Results database, 1990–2008. Breast Cancer Res. Treat. 2012, 134, 1257–1268. [Google Scholar] [CrossRef]
- Scott, L.; Mobley, L.R.; Il’yasova, D. Geospatial analysis of inflammatory breast cancer and associated community characteristics in the United States. Int. J. Environ. Res. Public Health 2017, 14, 404. [Google Scholar] [CrossRef]
- Fouad, T.M.; Ueno, N.T.; Yu, R.K.; Ensor, J.E.; Alvarez, R.H.; Krishnamurthy, S.; Lucci, A.; Reuben, J.M.; Yang, W.; Willey, J.S.; et al. Distinct epidemiological profiles associated with inflammatory breast cancer (IBC): A comprehensive analysis of the IBC registry at The University of Texas MD Anderson Cancer Center. PLoS ONE 2018, 13, e0204372. [Google Scholar] [CrossRef]
- Schairer, C.; Li, Y.; Frawley, P.; Graubard, B.I.; Wellman, R.D.; Buist, D.S.; Kerlikowske, K.; Onega, T.L.; Anderson, W.F.; Miglioretti, D.L. Risk factors for inflammatory breast cancer and other invasive breast cancers. J. Natl. Cancer Inst. 2013, 105, 1373–1384. [Google Scholar] [CrossRef]
- Chang, S.; Parker, S.L.; Pham, T.; Buzdar, A.U.; Hursting, S.D. Inflammatory breast carcinoma incidence and survival: The Surveillance, Epidemiology, and End Results program of the National Cancer Institute,1975–1992. Cancer 1998, 82, 2366–2372. [Google Scholar] [CrossRef]
- Atkinson, R.L.; El-Zein, R.; Valero, V.; Lucci, A.; Bevers, T.B.; Fouad, T.; Liao, W.; Ueno, N.T.; Woodward, W.A.; Brewster, A.M. Epidemiological risk factors associated with inflammatory breast cancer subtypes. Cancer Causes Control 2016, 27, 359–366. [Google Scholar] [CrossRef] [PubMed]
- Fouad, T.M.; Barrera, A.M.G.; Reuben, J.M.; Lucci, A.; Woodward, W.A.; Stauder, M.C.; Lim, B.; DeSnyder, S.M.; Arun, B.; Gildy, B.; et al. Inflammatory breast cancer: A proposed conceptual shift in the UICC-AJCC TNM staging system. Lancet Oncol. 2017, 18, e228–e232. [Google Scholar] [CrossRef]
- Cserni, G.; Charafe-Jauffret, E.; van Diest, P.J. Inflammatory breast cancer: The pathologists’ perspective. Eur. J. Surg. Oncol. 2018, 44, 1128–1134. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Gonzalez-Angulo, A.M.; Allen, P.K.; Yu, T.K.; Woodward, W.A.; Ueno, N.T.; Lucci, A.; Krishnamurthy, S.; Gong, Y.; Bondy, M.L.; et al. Triple-negative subtype predicts poor overall survival and high locoregional relapse in inflammatory breast cancer. Oncologist 2011, 16, 1675–1683. [Google Scholar] [CrossRef] [PubMed]
- Masuda, H.; Brewer, T.M.; Liu, D.D.; Iwamoto, T.; Shen, Y.; Hsu, L.; Willey, J.S.; Gonzalez-Angulo, A.M.; Chavez-MacGregor, M.; Fouad, T.M.; et al. Long-term treatment efficacy in primary inflammatory breast cancer by hormonal receptor- and HER2-defined subtypes. Ann. Oncol. 2014, 25, 384–391. [Google Scholar] [CrossRef] [PubMed]
- Rosenbluth, J.M.; Overmoyer, B.A. Inflammatory breast cancer: A separate entity. Curr. Oncol. Rep. 2019, 21, 86. [Google Scholar] [CrossRef]
- Bertucci, F.; Ueno, N.T.; Finetti, P.; Vermeulen, P.; Lucci, A.; Robertson, F.M.; Marsan, M.; Iwamoto, T.; Krishnamurthy, S.; Masuda, H.; et al. Gene expression profiles of inflammatory breast cancer: Correlation with response to neoadjuvant chemotherapy and metastasis-free survival. Ann. Oncol. 2014, 25, 358–365. [Google Scholar] [CrossRef]
- Van Laere, S.J.; Ueno, N.T.; Finetti, P.; Vermeulen, P.; Lucci, A.; Robertson, F.M.; Marsan, M.; Iwamoto, T.; Krishnamurthy, S.; Masuda, H.; et al. Uncovering the molecular secrets of inflammatory breast cancer biology: An integrated analysis of three distinct affymetrix gene expression datasets. Clin. Cancer Res. 2013, 19, 4685–4696. [Google Scholar] [CrossRef]
- Woodward, W.A.; Krishnamurthy, S.; Yamauchi, H.; El-Zein, R.; Ogura, D.; Kitadai, E.; Niwa, S.; Cristofanilli, M.; Vermeulen, P.; Dirix, L.; et al. Genomic and expression analysis of microdissected inflammatory breast cancer. Breast Cancer Res. Treat. 2013, 38, 761–772. [Google Scholar] [CrossRef]
- Bertucci, F.; Finetti, P.; Vermeulen, P.; Van Dam, P.; Dirix, L.; Birnbaum, D.; Viens, P.; Van Laere, S. Genomic profiling of inflammatory breast cancer: A review. Breast 2014, 23, 538–545. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.S.; Ali, S.M.; Wang, K.; Khaira, D.; Palma, N.A.; Chmielecki, J.; Palmer, G.A.; Morosini, D.; Elvin, J.A.; Fernandez, S.V.; et al. Comprehensive genomic profiling of inflammatory breast cancer cases reveals a high frequency of clinically relevant genomic alterations. Breast Cancer Res. Treat. 2015, 154, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Bekhouche, I.; Finetti, P.; Adelaïde, J.; Ferrari, A.; Tarpin, C.; Charafe-Jauffret, E.; Charpin, C.; Houvenaeghel, G.; Jacquemier, J.; Bidaut, G.; et al. High-resolution comparative genomic hybridization of inflammatory breast cancer and identification of candidate genes. PLoS ONE 2011, 6, e16950. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, N.; Lim, B.; Wang, Y.; Krishnamurthy, S.; Woodward, W.; Alvarez, R.H.; Lucci, A.; Valero, V.; Reuben, J.M.; Meric-Bernstam, F.; et al. Identification of frequent somatic mutations in inflammatory breast cancer. Breast Cancer Res. Treat. 2017, 163, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Vacher, S.; Boulai, A.; Bernard, V.; Baulande, S.; Bohec, M.; Bièche, I.; Lerebours, F.; Callens, C. Targeted next-generation sequencing identifies clinically relevant somatic mutations in a large cohort of inflammatory breast cancer. Breast Cancer Res. 2018, 20, 88. [Google Scholar] [CrossRef]
- Radford, E.J. An Introduction to epigenetic mechanisms. Prog. Mol. Biol. Transl. Sci. 2018, 158, 29–48. [Google Scholar]
- Arrowsmith, C.H.; Bountra, C.; Fish, P.V.; Lee, K.; Schapira, M. Epigenetic protein families: A new frontier for drug discovery. Nat. Rev. Drug. Discov. 2012, 11, 384–400. [Google Scholar] [CrossRef]
- Moris, N.; Pina, C.; Arias, A.M. Transition states and cell fate decisions in epigenetic landscapes. Nat. Rev. Genet. 2016, 17, 693–703. [Google Scholar] [CrossRef]
- Chung, C.Y.; Ma, Z.; Dravis, C.; Preissl, S.; Poirion, O.; Luna, G.; Hou, X.; Giraddi, R.R.; Ren, B.; Wahl, G.M. Single-cell chromatin analysis of mammary gland development reveals cell-state transcriptional regulators and lineage relationships. Cell Rep. 2019, 29, 495.e6–510.e6. [Google Scholar] [CrossRef]
- Holliday, H.; Baker, L.A.; Junankar, S.R.; Clark, S.J.; Swarbrick, A. Epigenomics of mammary gland development. Breast Cancer Res. 2018, 20, 100. [Google Scholar] [CrossRef]
- Inman, J.L.; Robertson, C.; Mott, J.D.; Bissell, M.J. Mammary gland development: Cell fate specification, stem cells and the microenvironment. Development 2015, 142, 1028–1042. [Google Scholar] [CrossRef] [PubMed]
- Huang, T.H.; Esteller, M. Chromatin remodeling in mammary gland differentiation and breast tumorigenesis. Cold Spring Harb. Perspect. Biol. 2010, 2, a004515. [Google Scholar] [CrossRef] [PubMed]
- Abdel-Hafiz, H.A.; Horwitz, K.B. Role of epigenetic modifications in luminal breast cancer. Epigenomics 2015, 7, 847–862. [Google Scholar] [CrossRef]
- Dravis, C.; Chung, C.Y.; Lytle, N.K.; Herrera-Valdez, J.; Luna, G.; Trejo, C.L.; Reya, T.; Wahl, G.M. Epigenetic and transcriptomic profiling of mammary gland development and tumor models disclose regulators of cell state plasticity. Cancer Cell 2018, 34, 466.e6–482.e6. [Google Scholar] [CrossRef] [PubMed]
- Gascard, P.; Bilenky, M.; Sigaroudinia, M.; Zhao, J.; Li, L.; Carles, A.; Delaney, A.; Tam, A.; Kamoh, B.; Cho, S.; et al. Epigenetic and transcriptional determinants of the human breast. Nat. Commun. 2015, 6, 6351. [Google Scholar] [CrossRef]
- Maruyama, R.; Choudhury, S.; Kowalczyk, A.; Bessarabova, M.; Beresford-Smith, B.; Conway, T.; Kaspi, A.; Wu, Z.; Nikolskaya, T.; Merino, V.F.; et al. Epigenetic regulation of cell type-specific expression patterns in the human mammary epithelium. PLoS Genet. 2011, 7, e1001369. [Google Scholar] [CrossRef]
- Bloushtain-Qimron, N.; Yao, J.; Shipitsin, M.; Maruyama, R.; Polyak, K. Epigenetic patterns of embryonic and adult stem cells. Cell Cycle 2009, 8, 809–817. [Google Scholar] [CrossRef]
- Lee, H.J.; Hinshelwood, R.A.; Bouras, T.; Gallego-Ortega, D.; Valdes-Mora, F.; Blazek, K.; Visvader, J.E.; Clark, S.J.; Ormandy, C.J. Lineage specific methylation of the Elf5 promoter in mammary epithelial cells. Stem Cells 2011, 29, 1611–1619. [Google Scholar] [CrossRef]
- Bloushtain-Qimron, N.; Yao, J.; Snyder, E.L.; Shipitsin, M.; Campbell, L.L.; Mani, S.A.; Hu, M.; Chen, H.; Ustyansky, V.; Antosiewicz, J.E.; et al. Cell type-specific DNA methylation patterns in the human breast. Proc. Natl. Acad. Sci. USA 2008, 105, 14076–14081. [Google Scholar] [CrossRef]
- Kouros-Mehr, H.; Kim, J.W.; Bechis, S.K.; Werb, Z. GATA-3 and the regulation of the mammary luminal cell fate. Curr. Opin. Cell. Biol. 2008, 20, 164–170. [Google Scholar] [CrossRef]
- Pellacani, D.; Bilenky, M.; Kannan, N.; Heravi-Moussavi, A.; Knapp, D.J.H.F.; Gakkhar, S.; Moksa, M.; Carles, A.; Moore, R.; Mungall, A.J.; et al. Analysis of normal human mammary epigenomes reveals cell-specific active enhancer states and associated transcription factor networks. Cell Rep. 2016, 17, 2060–2074. [Google Scholar] [CrossRef] [PubMed]
- Lim, E.; Vaillant, F.; Wu, D.; Forrest, N.C.; Pal, B.; Hart, A.H.; Asselin-Labat, M.L.; Gyorki, D.E.; Ward, T.; Partanen, A.; et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat. Med. 2009, 15, 907–913. [Google Scholar] [CrossRef] [PubMed]
- Pal, B.; Bouras, T.; Shi, W.; Vaillant, F.; Sheridan, J.M.; Fu, N.; Breslin, K.; Jiang, K.; Ritchie, M.E.; Young, M.; et al. Global changes in the mammary epigenome are induced by hormonal cues and coordinated by Ezh2. Cell Rep. 2013, 3, 411–426. [Google Scholar] [CrossRef] [PubMed]
- Michalak, E.M.; Nacerddine, K.; Pietersen, A.; Beuger, V.; Pawlitzky, I.; Cornelissen-Steijger, P.; Wientjens, E.; Tanger, E.; Seibler, J.; van Lohuizen, M.; et al. Polycomb group gene Ezh2 regulates mammary gland morphogenesis and maintains the luminal progenitor pool. Stem Cells 2013, 31, 1910–1920. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, S.; Wu, Z.; Russnes, H.G.; Takagi, S.; Peluffo, G.; Vaske, C.; Zhao, X.; Moen Vollan, H.K.; Maruyama, R.; Ekram, M.B.; et al. JARID1B is a luminal lineage-driving oncogene in breast cancer. Cancer Cell 2014, 25, 762–777. [Google Scholar] [CrossRef] [PubMed]
- Kanwal, R.; Gupta, K.; Gupta, S. Cancer epigenetics: An introduction. Methods Mol. Biol. 2015, 1238, 3–25. [Google Scholar]
- Byler, S.; Goldgar, S.; Heerboth, S.; Leary, M.; Housman, G.; Moulton, K.; Sarkar, S. Genetic and epigenetic aspects of breast cancer progression and therapy. Anticancer Res. 2014, 34, 1071–1077. [Google Scholar]
- Constâncio, V.; Nunes, S.P.; Henrique, R.; Jerónimo, C. DNA methylation-based testing in liquid biopsies as detection and prognostic biomarkers for the four major cancer types. Cells 2020, 9, 624. [Google Scholar] [CrossRef]
- Li, S.; Rong, M.; Iacopetta, B. DNA hypermethylation in breast cancer and its association with clinicopathological features. Cancer Lett. 2006, 237, 272–280. [Google Scholar] [CrossRef]
- Downs, B.; Wang, S.M. Epigenetic changes in BRCA1-mutated familial breast cancer. Cancer Genet. 2015, 208, 237–240. [Google Scholar] [CrossRef][Green Version]
- Oh, T.G.; Wang, S.M.; Muscat, G.E. Therapeutic Implications of Epigenetic Signaling in Breast Cancer. Endocrinology 2017, 158, 431–447. [Google Scholar] [CrossRef] [PubMed]
- Veeck, J.; Ropero, S.; Setien, F.; Gonzalez-Suarez, E.; Osorio, A.; Benitez, J.; Herman, J.G.; Esteller, M. BRCA1 CpG island hypermethylation predicts sensitivity to poly(adenosine diphosphate)-ribose polymerase inhibitors. J. Clin. Oncol. 2010, 28, e565–e566. [Google Scholar] [CrossRef] [PubMed]
- Ter Brugge, P.; Kristel, P.; van der Burg, E.; Boon, U.; de Maaker, M.; Lips, E.; Mulder, L.; de Ruiter, J.; Moutinho, C.; Gevensleben, H.; et al. Mechanisms of therapy resistance in patient-derived xenograft models of BRCA1-deficient breast cancer. J. Natl. Cancer Inst. 2016, 5, 108. [Google Scholar] [CrossRef] [PubMed]
- Werner, R.J.; Kelly, A.D.; Issa, J.J. Epigenetics and precision oncology. Cancer J. 2017, 23, 262–269. [Google Scholar] [CrossRef] [PubMed]
- Van den Eynden, G.G.; Van Laere, S.J.; Van der Auwera, I.; Merajver, S.D.; Van Marck, E.A.; van Dam, P.; Vermeulen, P.B.; Dirix, L.Y.; van Golen, K.L. Overexpression of caveolin-1 and -2 in cell lines and in human samples of inflammatory breast cancer. Breast Cancer Res. Treat. 2006, 95, 219–228. [Google Scholar] [CrossRef]
- Van der Auwera, I.; Van Laere, S.J.; Van den Bosch, S.M.; Van den Eynden, G.G.; Trinh, B.X.; van Dam, P.A.; Colpaert, C.G.; van Engeland, M.; Van Marck, E.A.; Vermeulen, P.B.; et al. Aberrant methylation of the Adenomatous Polyposis Coli (APC) gene promoter is associated with the inflammatory breast cancer phenotype. Br. J. Cancer 2008, 99, 1735–1742. [Google Scholar] [CrossRef]
- Van der Auwera, I.; Bovie, C.; Svensson, C.; Limame, R.; Trinh, X.B.; van Dam, P.; Van Laere, S.J.; Van Marck, E.; Vermeulen, P.B.; Dirix, L.Y. Quantitative assessment of DNA hypermethylation in the inflammatory and non-inflammatory breast cancer phenotypes. Cancer Biol. Ther. 2009, 8, 2252–2259. [Google Scholar] [CrossRef][Green Version]
- Van der Auwera, I.; Yu, W.; Suo, L.; Van Neste, L.; van Dam, P.; Van Marck, E.A.; Pauwels, P.; Vermeulen, P.B.; Dirix, L.Y.; Van Laere, S.J. Array-based DNA methylation profiling for breast cancer subtype discrimination. PLoS ONE 2010, 5, e12616. [Google Scholar] [CrossRef]
- Mohamed, M.M.; Sabet, S.; Peng, D.F.; Nouh, M.A.; El-Shinawi, M.; El-Rifai, W. Promoter hypermethylation and suppression of glutathione peroxidase 3 are associated with inflammatory breast carcinogenesis. Oxid. Med. Cell Longev. 2014, 2014, 787195. [Google Scholar] [CrossRef]
- Debeb, B.G.; Gong, Y.; Atkinson, R.L.; Sneige, N.; Huo, L.; Gonzalez-Angulo, A.M.; Hung, M.C.; Valero, V.; Ueno, N.T.; Woodward, W.A. EZH2 expression correlates with locoregional recurrence after radiation in inflammatory breast cancer. J. Exp. Clin. Cancer Res. 2014, 33, 58. [Google Scholar] [CrossRef][Green Version]
- Ding, Q.; Wang, Y.; Zuo, Z.; Gong, Y.; Krishnamurthy, S.; Li, C.W.; Lai, Y.J.; Wei, W.; Wang, J.; Manyam, G.C.; et al. Decreased expression of microRNA-26b in locally advanced and inflammatory breast cancer. Hum. Pathol. 2018, 77, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Kelly, A.D.; Issa, J.J. The promise of epigenetic therapy: Reprogramming the cancer epigenome. Curr. Opin. Genet. Dev. 2017, 42, 68–77. [Google Scholar] [CrossRef] [PubMed]
- St Laurent, G.; Wahlestedt, C.; Kapranov, P. The landscape of long noncoding RNA classification. Trends Genet. 2015, 31, 239–251. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.J.; Tan, X.L.; Guo, L. The long non-coding RNA DANCR regulates the inflammatory phenotype of breast cancer cells and promotes breast cancer progression via EZH2-dependent suppression of SOCS3 transcription. Mol. Oncol. 2020, 14, 309–328. [Google Scholar] [CrossRef]
- Ladd-Acosta, C.; Fallin, M.D. The role of epigenetics in genetic and environmental epidemiology. Epigenomics 2016, 8, 271–283. [Google Scholar] [CrossRef]
- Ahmad, A.; Azim, S.; Zubair, H.; Khan, M.A.; Singh, S.; Carter, J.E.; Rocconi, R.P.; Singh, A.P. Epigenetic basis of cancer health disparities: Looking beyond genetic differences. Biochim. Biophys. Acta Rev. Cancer 2017, 1868, 16–28. [Google Scholar] [CrossRef]
- Ambrosone, C.B.; Young, A.C.; Sucheston, L.E.; Wang, D.; Yan, L.; Liu, S.; Tang, L.; Hu, Q.; Freudenheim, J.L.; Shields, P.G.; et al. Genome-wide methylation patterns provide insight into differences in breast tumor biology between American women of African and European ancestry. Oncotarget 2014, 5, 237–248. [Google Scholar] [CrossRef]
- Williams, D.R.; Mohammed, S.A.; Shields, A.E. Understanding and effectively addressing breast cancer in African American women: Unpacking the social context. Cancer 2016, 122, 2138–2149. [Google Scholar] [CrossRef]
- Song, M.A.; Brasky, T.M.; Marian, C.; Weng, D.Y.; Taslim, C.; Dumitrescu, R.G.; Llanos, A.A.; Freudenheim, J.L.; Shields, P.G. Racial differences in genome-wide methylation profiling and gene expression in breast tissues from healthy women. Epigenetics 2015, 10, 1177–1187. [Google Scholar] [CrossRef]
- Galanter, J.M.; Gignoux, C.R.; Oh, S.S.; Torgerson, D.; Pino-Yanes, M.; Thakur, N.; Eng, C.; Hu, D.; Huntsman, S.; Farber, H.J.; et al. Differential methylation between ethnic sub-groups reflects the effect of genetic ancestry and environmental exposures. eLife 2017, 6, e20532. [Google Scholar] [CrossRef]
- Ligibel, J.A.; Alfano, C.M.; Courneya, K.S.; Demark-Wahnefried, W.; Burger, R.A.; Chlebowski, R.T.; Fabian, C.J.; Gucalp, A.; Hershman, D.L.; Hudson, M.M.; et al. American Society of Clinical Oncology position statement on obesity and cancer. J. Clin. Oncol. 2014, 32, 3568–3574. [Google Scholar] [CrossRef]
- Gillman, A.S.; Gardiner, C.K.; Koljack, C.E.; Bryan, A.D. Body mass index, diet, and exercise: Testing possible linkages to breast cancer risk via DNA methylation. Breast Cancer Res. Treat. 2018, 168, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Font-Burgada, J.; Sun, B.; Karin, M. Obesity and Cancer: The oil that feeds the flame. Cell Metab. 2016, 23, 48–62. [Google Scholar] [CrossRef] [PubMed]
- Woods, J.A.; Vieira, V.J.; Keylock, K.T. Exercise, inflammation, and innate immunity. Immunol. Allergy Clin. North Am. 2009, 29, 381–393. [Google Scholar] [CrossRef] [PubMed]
- Calder, P.C.; Ahluwalia, N.; Brouns, F.; Buetler, T.; Clement, K.; Cunningham, K.; Esposito, K.; Jönsson, L.S.; Kolb, H.; Lansink, M.; et al. Dietary factors and low-grade inflammation in relation to overweight and obesity. Br. J. Nutr. 2011, 106, S5–S78. [Google Scholar] [CrossRef]
- Xu, X.; Su, S.; Barnes, V.A.; De Miguel, C.; Pollock, J.; Ownby, D.; Shi, H.; Zhu, H.; Snieder, H.; Wang, X. A genome-wide methylation study on obesity: Differential variability and differential methylation. Epigenetics 2013, 8, 522–533. [Google Scholar] [CrossRef]
- Rönn, T.; Volkov, P.; Gillberg, L.; Kokosar, M.; Perfilyev, A.; Jacobsen, A.L.; Jørgensen, S.W.; Brøns, C.; Jansson, P.A.; Eriksson, K.F.; et al. Impact of age, BMI and HbA1c levels on the genome-wide DNA methylation and mRNA expression patterns in human adipose tissue and identification of epigenetic biomarkers in blood. Hum. Mol. Genet. 2015, 24, 3792–3813. [Google Scholar] [CrossRef]
- Hair, B.Y.; Troester, M.A.; Edmiston, S.N.; Parrish, E.A.; Robinson, W.R.; Wu, M.C.; Olshan, A.F.; Swift-Scanlan, T.; Conway, K. Body mass index is associated with gene methylation inestrogen receptor-positive breast tumors. Cancer Epidemiol. Biomarkers Prev. 2015, 24, 580–586. [Google Scholar] [CrossRef]
- Delgado-Cruzata, L.; Zhang, W.; McDonald, J.A.; Tsai, W.Y.; Valdovinos, C.; Falci, L.; Wang, Q.; Crew, K.D.; Santella, R.M.; Hershman, D.L.; et al. Dietary modifications, weight loss, and changes in metabolic markers affect global DNA methylation in Hispanic, African American, and Afro-Caribbean breast cancer survivors. J. Nutr. 2015, 145, 783–790. [Google Scholar] [CrossRef]
- Donovan, M.G.; Wren, S.N.; Cenker, M.; Selmin, O.I.; Romagnolo, D.F. Dietary fat and obesity as modulators of breast cancer risk: Focus on DNA methylation. Br. J. Pharmacol. 2020, 177, 1331–1350. [Google Scholar] [CrossRef]
- Kanherkar, R.R.; Bhatia-Dey, N.; Csoka, A.B. Epigenetics across the human lifespan. Front. Cell Dev. Biol. 2014, 2, 49. [Google Scholar] [CrossRef]
- Diamanti-Kandarakis, E.; Bourguignon, J.P.; Giudice, L.C.; Hauser, R.; Prins, G.S.; Soto, A.M.; Zoeller, R.T.; Gore, A.C. Endocrine-disrupting chemicals: An Endocrine Society scientific statement. Endocr. Rev. 2009, 30, 293–342. [Google Scholar] [CrossRef]
- Anway, M.D.; Skinner, M.K. Epigenetic transgenerational actions of endocrine disruptors. Endocrinology 2006, 147, S43–S49. [Google Scholar] [CrossRef]
- Muñoz-de-Toro, M.; Markey, C.M.; Wadia, P.R.; Luque, E.H.; Rubin, B.S.; Sonnenschein, C.; Soto, A.M. Perinatal exposure to bisphenol-A alters peripubertal mammary gland development in mice. Endocrinology 2005, 146, 4138–4147. [Google Scholar] [CrossRef]
- Murata, M.; Kang, J.H. Bisphenol A (BPA) and cell signaling pathways. Biotechnol. Adv. 2018, 36, 311–327. [Google Scholar] [CrossRef]
- Doherty, L.F.; Bromer, J.G.; Zhou, Y.; Aldad, T.S.; Taylor, H.S. In utero exposure to diethylstilbestrol (DES) or bisphenol-A (BPA) increases EZH2 expression in the mammary gland: An epigenetic mechanism linking endocrine disruptors to breast cancer. Horm. Cancer 2010, 3, 146–155. [Google Scholar] [CrossRef] [PubMed]
- Dolinoy, D.C.; Huang, D.; Jirtle, R.L. Maternal nutrient supplementation counteracts bisphenol A-induced DNA hypomethylation in early development. Proc. Natl. Acad. Sci. USA 2007, 104, 13056–13061. [Google Scholar] [CrossRef]
- Huang, R.P.; Liu, Z.H.; Yin, H.; Dang, Z.; Wu, P.X.; Zhu, N.W.; Lin, Z. Bisphenol A concentrations in human urine, human intakes across six continents, and annual trends of average intakes in adult and child populations worldwide: A thorough literature review. Sci. Total Environ. 2018, 626, 971–981. [Google Scholar] [CrossRef] [PubMed]
- Skinner, M.K. Environmental epigenetic transgenerational inheritance and somatic epigenetic mitotic stability. Epigenetics 2011, 6, 838–842. [Google Scholar] [PubMed]
- Zama, A.M.; Uzumcu, M. Fetal and neonatal exposure to the endocrine disruptor methoxychlor causes epigenetic alterations in adult ovarian genes. Endocrinology 2009, 150, 4681–4691. [Google Scholar] [CrossRef] [PubMed]
- Armenti, A.E.; Zama, A.M.; Passantino, L.; Uzumcu, M. Developmental methoxychlor exposure affects multiple reproductive parameters and ovarian folliculogenesis and gene expression in adult rats. Toxicol. Appl. Pharmacol. 2008, 233, 286–296. [Google Scholar] [CrossRef]
- Galliot, B.; Ghila, L. Cell plasticity in homeostasis and regeneration. Mol. Reprod. Dev. 2010, 77, 837–855. [Google Scholar] [CrossRef]
- Chaffer, C.L.; Marjanovic, N.D.; Lee, T.; Bell, G.; Kleer, C.G.; Reinhardt, F.; D’Alessio, A.C.; Young, R.A.; Weinberg, R.A. Poised chromatin at the ZEB1 promoter enables breast cancer cell plasticity and enhances tumorigenicity. Cell 2013, 154, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Yager, J.D.; Davidson, N.E. Estrogen carcinogenesis in breast cancer. N. Engl. J. Med. 2006, 354, 270–282. [Google Scholar] [CrossRef] [PubMed]
- Fenton, S.E. Endocrine-disrupting compounds and mammary gland development: Early exposure and later life consequences. Endocrinology 2006, 147, S18–S24. [Google Scholar] [CrossRef] [PubMed]
- Cheng, A.S.; Culhane, A.C.; Chan, M.W.; Venkataramu, C.R.; Ehrich, M.; Nasir, A.; Rodriguez, B.A.; Liu, J.; Yan, P.S.; Quackenbush, J.; et al. Epithelial progeny of estrogen-exposed breast progenitor cells display a cancer-like methylome. Cancer Res. 2008, 68, 1786–1796. [Google Scholar] [CrossRef] [PubMed]
- Hsu, P.Y.; Deatherage, D.E.; Rodriguez, B.A.; Liyanarachchi, S.; Weng, Y.I.; Zuo, T.; Liu, J.; Cheng, A.S.; Huang, T.H. Xenoestrogen-induced epigenetic repression of microRNA-9-3 in breast epithelial cells. Cancer Res. 2009, 69, 5936–5945. [Google Scholar] [CrossRef]
- Jensen, E.V.; Jordan, V.C. The estrogen receptor: A model for molecular medicine. Clin. Cancer Res. 2003, 9, 1980–1989. [Google Scholar]
- Patterson, A.R.; Mo, X.; Shapiro, A.; Wernke, K.E.; Archer, T.K.; Burd, C.J. Sustained reprogramming of the estrogen response after chronic exposure to endocrine disruptors. Mol. Endocrinol. 2015, 29, 384–395. [Google Scholar] [CrossRef]
- Miranda, T.B.; Voss, T.C.; Sung, M.H.; Baek, S.; John, S.; Hawkins, M.; Grøntved, L.; Schiltz, R.L.; Hager, G.L. Reprogramming the chromatin landscape: Interplay of the estrogen and glucocorticoid receptors at the genomic level. Cancer Res. 2016, 73, 5130–5139. [Google Scholar] [CrossRef]
- Hoffman, J.A.; Trotter, K.W.; Ward, J.M.; Archer, T.K. BRG1 governs glucocorticoid receptor interactions with chromatin and pioneer factors across the genome. eLife 2018, 24, E35073. [Google Scholar] [CrossRef] [PubMed]
- Takaku, M.; Grimm, S.A.; Shimbo, T.; Perera, L.; Menafra, R.; Stunnenberg, H.G.; Archer, T.K.; Machida, S.; Kurumizaka, H.; Wade, P.A. GATA3-dependent cellular reprogramming requires activation-domain dependent recruitment of a chromatin remodeler. Genome Biol. 2016, 17, 36. [Google Scholar] [CrossRef]
- Swinstead, E.E.; Paakinaho, V.; Hager, G.L. Chromatin reprogramming in breast cancer. Endocr. Relat. Cancer 2018, 25, R385–R404. [Google Scholar] [CrossRef]
- Herlofsen, S.R.; Bryne, J.C.; Høiby, T.; Wang, L.; Issner, R.; Zhang, X.; Coyne, M.J.; Boyle, P.; Gu, H.; Meza-Zepeda, L.A.; et al. Genome-wide map of quantified epigenetic changes during in vitro chondrogenic differentiation of primary human mesenchymal stem cells. BMC Genomics 2013, 14, 105. [Google Scholar] [CrossRef]
- Li, L.; Tang, P.; Li, S.; Qin, X.; Yang, H.; Wu, C.; Liu, Y. Notch signaling pathway networks in cancer metastasis: A new target for cancer therapy. Med. Oncol. 2017, 34, 180. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Ye, Y.; Zou, X.; Jones, S.; Yearsley, K.; Shetuni, B.; Tellez, J.; Barsky, S.H. The lymphovascular embolus of inflammatory breast cancer exhibits a Notch 3 addiction. Oncogene 2011, 30, 287–300. [Google Scholar] [CrossRef] [PubMed]
- Yoo, E.J.; Chung, J.J.; Choe, S.S.; Kim, K.H.; Kim, J.B.J. Down-regulation of histone deacetylases stimulates adipocyte differentiation. Biol. Chem. 2006, 281, 6608–6615. [Google Scholar] [CrossRef] [PubMed]
- Topalian, S.L.; Weiner, G.J.; Pardoll, D.M. Cancer immunotherapy comes of age. J. Clin. Oncol. 2011, 29, 4828–4836. [Google Scholar] [CrossRef]
- Jolly, M.K.; Boareto, M.; Debeb, B.G.; Aceto, N.; Farach-Carson, M.C.; Woodward, W.A.; Levine, H. Inflammatory breast cancer: A model for investigating cluster-based dissemination. NPJ Breast Cancer 2017, 3, 21. [Google Scholar] [CrossRef]
- Shah, J.N.; Shao, G.; Hei, T.K.; Zhao, Y. Methylation screening of the TGFBI promoter in human lung and prostate cancer by methylation-specific PCR. BMC Cancer 2008, 8, 284. [Google Scholar] [CrossRef]
- Yasmin, R.; Siraj, S.; Hassan, A.; Khan, A.R.; Abbasi, R.; Ahmad, N. Epigenetic regulation of inflammatory cytokines and associated genes in human malignancies. Mediators Inflamm. 2015, 2015, 201703. [Google Scholar] [CrossRef] [PubMed]
- Kumari, N.; Dwarakanath, B.S.; Das, A.; Bhatt, A.N. Role of interleukin-6 in cancer progression and therapeutic resistance. Tumour Biol. 2016, 37, 11553–11572. [Google Scholar] [CrossRef] [PubMed]
- Wolfe, A.R.; Trenton, N.J.; Debeb, B.G.; Larson, R.; Ruffell, B.; Chu, K.; Hittelman, W.; Diehl, M.; Reuben, J.M.; Ueno, N.T.; et al. Mesenchymal stem cells and macrophages interact through IL-6 to promote inflammatory breast cancer in pre-clinical models. Oncotarget 2016, 7, 82482–82492. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.C.; Lin, J.H.; Hsu, T.W.; Su, K.; Li, A.F.; Hsu, H.S.; Hung, S.C. IL-6 enriched lung cancer stem-like cell population by inhibition of cell cycle regulators via DNMT1 upregulation. Int. J. Cancer 2015, 136, 547–559. [Google Scholar] [PubMed]
- Itoh, H.; Kadomatsu, T.; Tanoue, H.; Yugami, M.; Miyata, K.; Endo, M.; Morinaga, J.; Kobayashi, E.; Miyamoto, T.; Kurahashi, R.; et al. TET2-dependent IL-6 induction mediated by the tumor microenvironment promotes tumor metastasis in osteosarcoma. Oncogene 2018, 37, 2903–2920. [Google Scholar] [CrossRef]
- Huang, L.; Hu, B.; Ni, J.; Wu, J.; Jiang, W.; Chen, C.; Yang, L.; Zeng, Y.; Wan, R.; Hu, G.; et al. Transcriptional repression of SOCS3 mediated by IL-6/STAT3 signaling via DNMT1 promotes pancreatic cancer growth and metastasis. J. Exp. Clin. Cancer Res. 2016, 35, 27. [Google Scholar] [CrossRef]
- D’Anello, L.; Sansone, P.; Storci, G.; Mitrugno, V.; D’Uva, G.; Chieco, P.; Bonafé, M. Epigenetic control of the basal-like gene expression profile via Interleukin-6 in breast cancer cells. Mol. Cancer 2010, 9, 300. [Google Scholar] [CrossRef]
- Shih, R.H.; Wang, C.Y.; Yang, C.M. NF-kappaB signaling pathways in neurological inflammation: A mini review. Front. Mol. Neurosci. 2015, 8, 77. [Google Scholar] [CrossRef]
- Drygin, D.; Ho, C.B.; Omori, M.; Bliesath, J.; Proffitt, C.; Rice, R.; Siddiqui-Jain, A.; O’Brien, S.; Padgett, C.; Lim, J.K.; et al. Protein kinase CK2 modulates IL-6 expression in inflammatory breast cancer. Biochem. Biophys. Res. Commun. 2011, 415, 163–167. [Google Scholar] [CrossRef]
- Lu, T.; Stark, G.R. NF-κB: Regulation by methylation. Cancer Res. 2015, 75, 3692–3695. [Google Scholar] [CrossRef]
- Morel, D.; Jeffery, D.; Aspeslagh, S.; Almouzni, G.; Postel-Vinay, S. Combining epigenetic drugs with other therapies for solid tumours—past lessons and future promise. Nat. Rev. Clin. Oncol. 2020, 17, 91–107. [Google Scholar] [CrossRef]
- Miranda Furtado, C.L.; Dos Santos Luciano, M.C.; Silva Santos, R.D.; Furtado, G.P.; Moraes, M.O.; Pessoa, C. Epidrugs: Targeting epigenetic marks in cancer treatment. Epigenetics 2019, 14, 1164–1176. [Google Scholar] [CrossRef]
- Toh, T.B.; Lim, J.J.; Chow, E.K. Epigenetics in cancer stem cells. Mol. Cancer 2017, 16, 29. [Google Scholar] [CrossRef]
- Parbin, S.; Kar, S.; Shilpi, A.; Sengupta, D.; Deb, M.; Rath, S.K.; Patra, S.K. Histone deacetylases: A saga of perturbed acetylation homeostasis in cancer. J. Histochem. Cytochem. 2014, 62, 11–33. [Google Scholar] [CrossRef]
- Azad, N.; Zahnow, C.A.; Rudin, C.M.; Baylin, S.B. The future of epigenetic therapy in solid tumours-lessons from the past. Nat. Rev. Clin. Oncol. 2013, 10, 256–266. [Google Scholar] [CrossRef]
- Topper, M.J.; Vaz, M.; Marrone, K.A.; Brahmer, J.R.; Baylin, S.B. The emerging role of epigenetic therapeutics in immuno-oncology. Nat. Rev. Clin. Oncol. 2020, 17, 75–90. [Google Scholar] [CrossRef]
- Seto, E.; Yoshida, M. Erasers of histone acetylation: The histone deacetylase enzymes. Cold Spring Harb. Perspect. Biol. 2014, 6, a018713. [Google Scholar] [CrossRef]
- Chatterjee, N.; Wang, W.L.; Conklin, T.; Chittur, S.; Tenniswood, M. Histone deacetylase inhibitors modulate miRNA and mRNA expression, block metaphase, and induce apoptosis in inflammatory breast cancer cells. Cancer Biol. Ther. 2013, 14, 658–671. [Google Scholar] [CrossRef]
- Hollestelle, A.; Nagel, J.H.; Smid, M.; Lam, S.; Elstrodt, F.; Wasielewski, M.; Ng, S.S.; French, P.J.; Peeters, J.K.; Rozendaal, M.J.; et al. Distinct gene mutation profiles among luminal-type and basal-type breast cancer cell lines. Breast Cancer Res. Treat. 2010, 121, 53–64. [Google Scholar] [CrossRef]
- Putcha, P.; Yu, J.; Rodriguez-Barrueco, R.; Saucedo-Cuevas, L.; Villagrasa, P.; Murga-Penas, E.; Quayle, S.N.; Yang, M.; Castro, V.; Llobet-Navas, D.; et al. HDAC6 activity is a non-oncogene addiction hub for inflammatory breast cancers. Breast Cancer Res. 2015, 17, 149. [Google Scholar] [CrossRef]
- Lee, J.; Bartholomeusz, C.; Mansour, O.; Humphries, J.; Hortobagyi, G.N.; Ordentlich, P.; Ueno, N.T. A class I histone deacetylase inhibitor, entinostat, enhances lapatinib efficacy in HER2-overexpressing breast cancer cells through FOXO3-mediated Bim1 expression. Breast Cancer Res. Treat. 2014, 146, 259–272. [Google Scholar] [CrossRef] [PubMed]
- Debeb, B.G.; Lacerda, L.; Larson, R.; Wolfe, A.R.; Krishnamurthy, S.; Reuben, J.M.; Ueno, N.T.; Gilcrease, M.; Woodward, W.A. Histone deacetylase inhibitor-induced cancer stem cells exhibit high pentose phosphate pathway metabolism. Oncotarget 2016, 7, 28329–28339. [Google Scholar] [CrossRef] [PubMed]
- Torres-Adorno, A.M.; Lee, J.; Kogawa, T.; Ordentlich, P.; Tripathy, D.; Lim, B.; Ueno, N.T. Histone deacetylase inhibitor enhances the efficacy of MEK inhibitor through NOXA-Mediated MCL1 degradation in triple-negative and inflammatory breast cancer. Clin. Cancer Res. 2017, 23, 4780–4792. [Google Scholar] [CrossRef] [PubMed]
- Italiano, A.; Soria, J.C.; Toulmonde, M.; Michot, J.M.; Lucchesi, C.; Varga, A.; Coindre, J.M.; Blakemore, S.J.; Clawson, A.; Suttle, B.; et al. Tazemetostat, an EZH2 inhibitor, in relapsed or refractory B-cell non-Hodgkin lymphoma and advanced solid tumours: A first-in-human, open-label, phase 1 study. Lancet Oncol. 2018, 19, 649–659. [Google Scholar] [CrossRef]
- Easwaran, H.; Tsai, H.C.; Baylin, S.B. Cancer epigenetics: Tumor heterogeneity, plasticity of stem-like states, and drug resistance. Mol. Cell 2014, 54, 716–727. [Google Scholar] [CrossRef]
- Mio, C.; Bulotta, S.; Russo, D.; Damante, G. Reading cancer: Chromatin readers as druggable targets for cancer treatment. Cancers 2019, 11, 61. [Google Scholar] [CrossRef]
- Zhu, H.; Wei, T.; Cai, Y.; Jin, J. Small molecules targeting the specific domains of histone-mark readers in cancer therapy. Molecules 2020, 25, 578. [Google Scholar] [CrossRef]
- Khandekar, D.; Tiriveedhi, V. Role of BET inhibitors in triple negative breast cancers. Cancers 2020, 12, 784. [Google Scholar] [CrossRef]
| Target Gene(s) | Strategy | Sample Size | Main Findings | Refs. |
|---|---|---|---|---|
| APC | Quantitative Methylation-Specific PCR | 21 IBC/ 35 Non-IBC | High methylation levels in IBC versus non-IBC | [77,78] |
| APC DAPK HIN-1 RASSF1A RARB TWIST1 | Quantitative Methylation-Specific PCR using a six-gene panel | 19 IBC/ 81 Non-IBC | Differential methylation of RARB in IBC RARB and APC methylation frequency were significantly increased in IBC | [78] |
| Array-based DNA methylation profiling | Illumina Infinium Human Methylation 27 Beadchip | 19 IBC/ 43 non-IBC | Four-gene based signature (TJP3, MOGAT2, NTSR2, and AGT) differentiates IBC from non-IBC | [79] |
| GPX3 | qualitative Methylation-Specific PCR | 20 IBC/ 20 non-IBC | High promoter hypermethylation of GPX3 gene in IBC versus non-IBC | [80] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Faldoni, F.L.C.; Rainho, C.A.; Rogatto, S.R. Epigenetics in Inflammatory Breast Cancer: Biological Features and Therapeutic Perspectives. Cells 2020, 9, 1164. https://doi.org/10.3390/cells9051164
Faldoni FLC, Rainho CA, Rogatto SR. Epigenetics in Inflammatory Breast Cancer: Biological Features and Therapeutic Perspectives. Cells. 2020; 9(5):1164. https://doi.org/10.3390/cells9051164
Chicago/Turabian StyleFaldoni, Flavia Lima Costa, Cláudia Aparecida Rainho, and Silvia Regina Rogatto. 2020. "Epigenetics in Inflammatory Breast Cancer: Biological Features and Therapeutic Perspectives" Cells 9, no. 5: 1164. https://doi.org/10.3390/cells9051164
APA StyleFaldoni, F. L. C., Rainho, C. A., & Rogatto, S. R. (2020). Epigenetics in Inflammatory Breast Cancer: Biological Features and Therapeutic Perspectives. Cells, 9(5), 1164. https://doi.org/10.3390/cells9051164





