scRNA-seq Profiling of Human Testes Reveals the Presence of the ACE2 Receptor, A Target for SARS-CoV-2 Infection in Spermatogonia, Leydig and Sertoli Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Sources
2.2. scRNA-Seq Data Processing
2.3. Single-Cell Transcriptomes to Identify Cell Types
2.4. Identification of Differential Expression Genes
2.5. Gene Function Analysis
3. Results
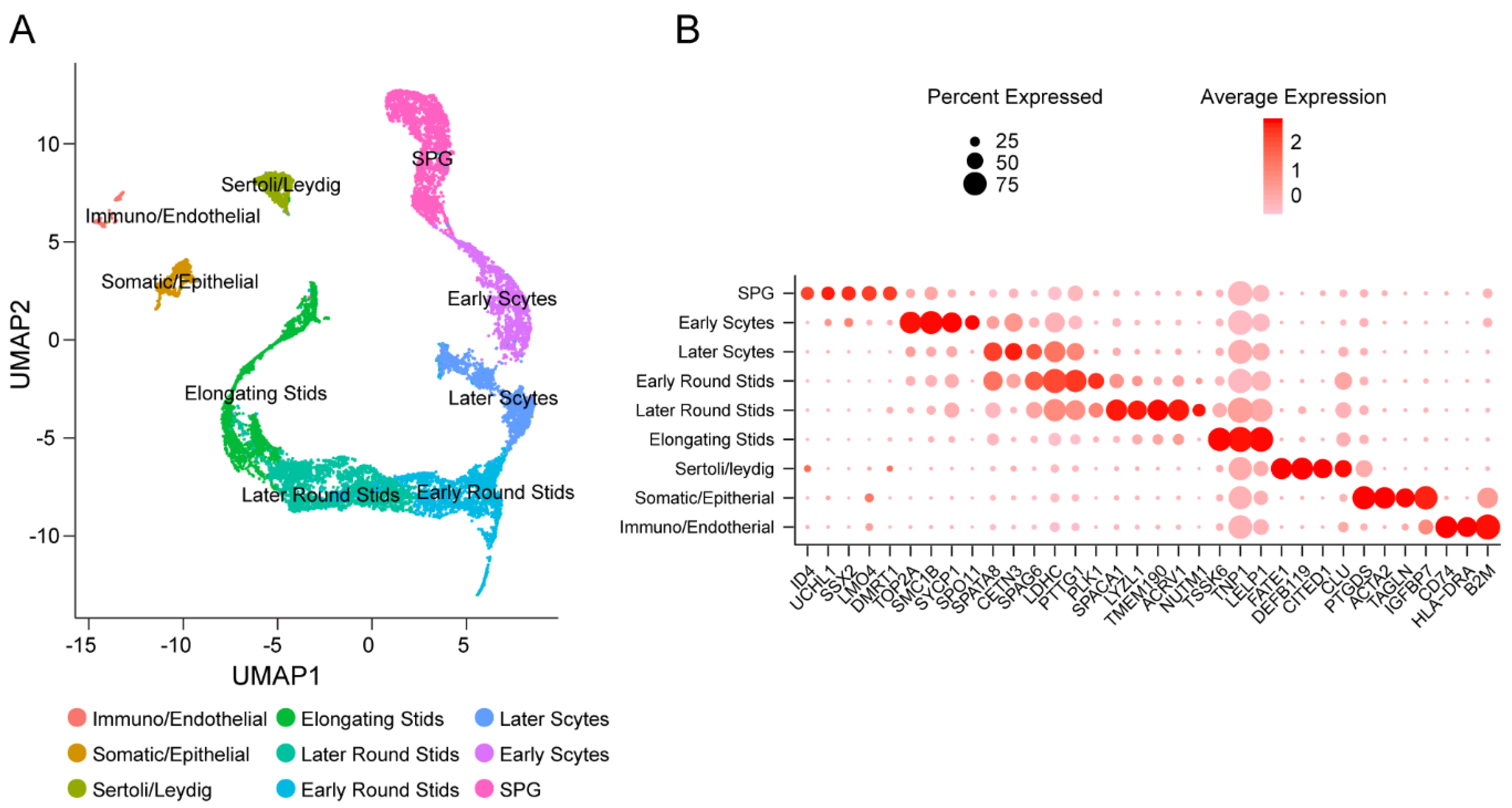
3.1. Identification of Cell Types in Adult Human Testes
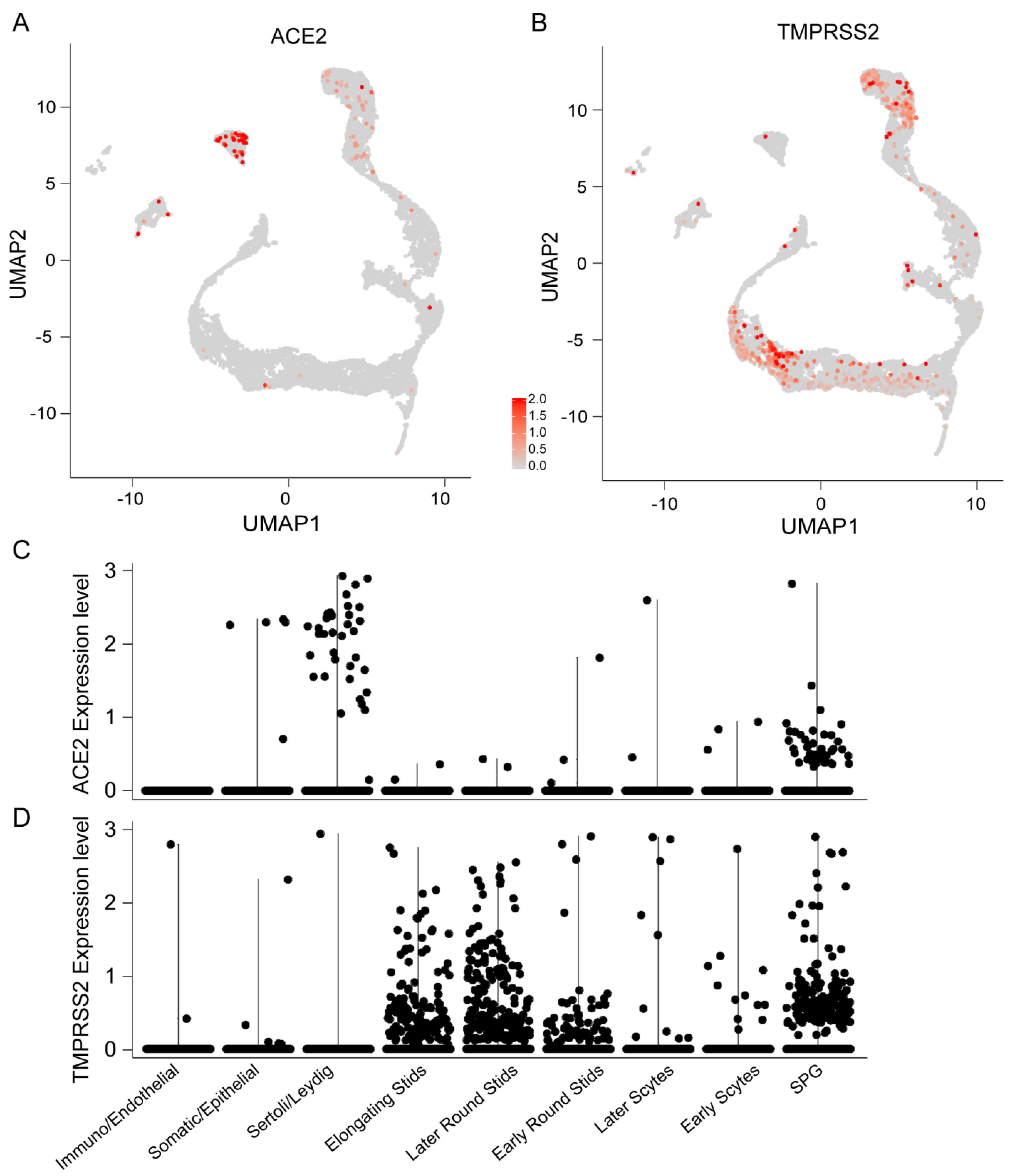
3.2. Cell-Specific Expression of ACE2
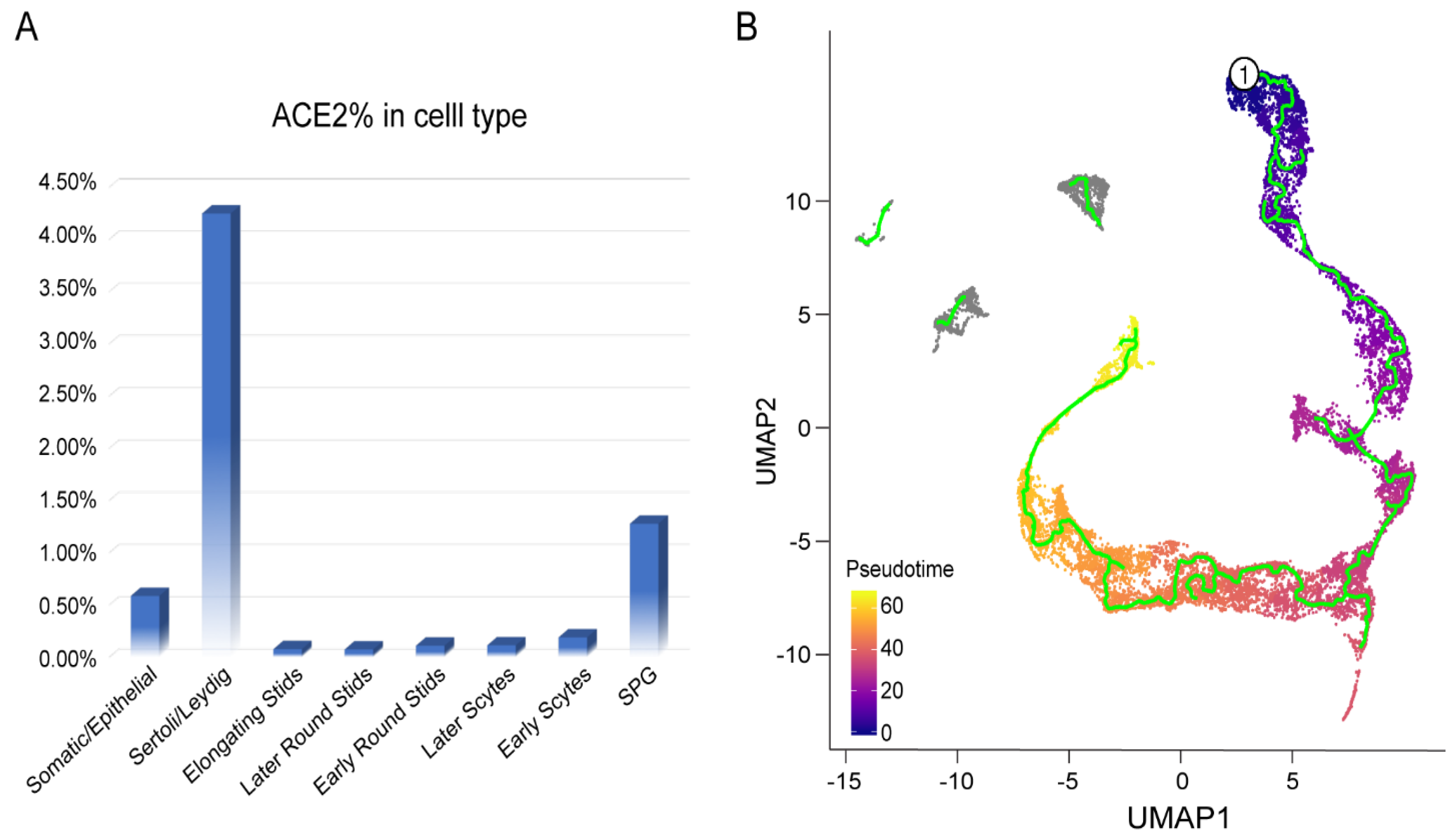
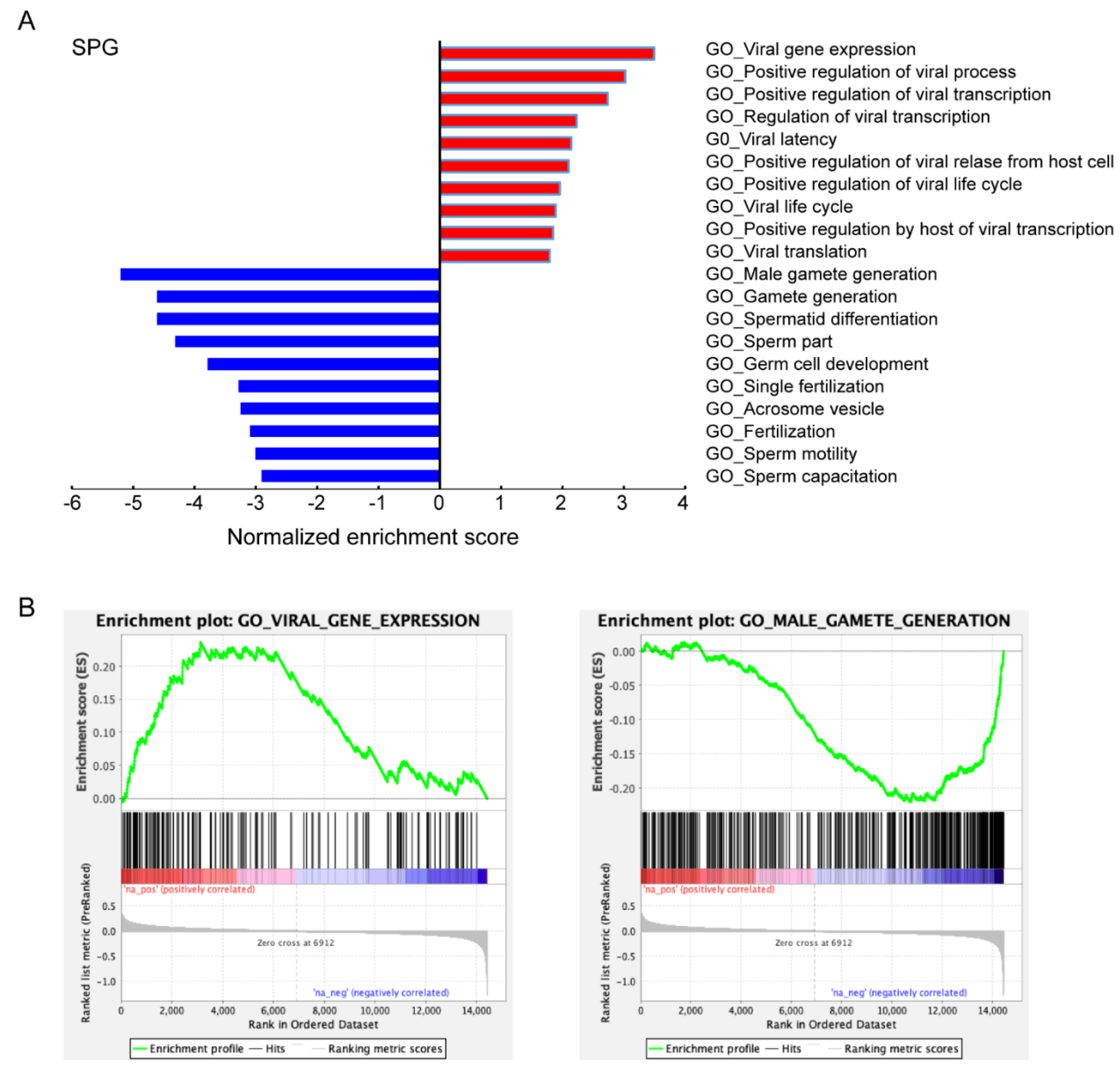
3.3. Characteristics of ACE2-Positive Cells in Human Testis
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Chen, N.; Zhou, M.; Dong, X.; Qu, J.; Gong, F.; Han, Y.; Qiu, Y.; Wang, J.; Liu, Y.; Wei, Y.; et al. Epidemiological and Clinical Characteristics of 99 Cases of 2019 Novel Coronavirus Pneumonia in Wuhan, China: A Descriptive Study. Lancet 2020, 395, 507–513. [Google Scholar] [CrossRef]
- Chan, J.F.; Yuan, S.; Kok, K.H.; To, K.K.; Chu, H.; Yang, J.; Xing, F.; Liu, J.; Yip, C.C.; Poon, R.W.; et al. A Familial Cluster of Pneumonia Associated with the 2019 Novel Coronavirus Indicating Person-to-Person Transmission: A Study of a Family Cluster. Lancet 2020, 395, 514–523. [Google Scholar] [CrossRef]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical Features of Patients Infected with 2019 Novel Coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Li, W.; Moore, M.J.; Vasilieva, N.; Sui, J.; Wong, S.K.; Berne, M.A.; Somasundaran, M.; Sullivan, J.L.; Luzuriaga, K.; Greenough, T.C.; et al. Angiotensin-Converting Enzyme 2 Is a Functional Receptor for the Sars Coronavirus. Nature 2003, 426, 450–454. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. Sars-Cov-2 Cell Entry Depends on Ace2 and Tmprss2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Zhao, Z.; Wang, Y.; Zhou, Y.; Ma, Y.; Zuo, W. Single-Cell Rna Expression Profiling of Ace2, the Putative Receptor of Wuhan 2019-Ncov. BioRxiv 2020. [Google Scholar] [CrossRef]
- Yan, R.; Zhang, Y.; Li, Y.; Xia, L.; Guo, Y.; Zhou, Q. Structural Basis for the Recognition of the Sars-Cov-2 by Full-Length Human Ace2. Science 2020, 367, 1444–1448. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.Y.; et al. Structural and Functional Basis of Sars-Cov-2 Entry by Using Human Ace2. Cell 2020. [Google Scholar] [CrossRef]
- Chau, T.N.; Lee, K.C.; Yao, H.; Tsang, T.Y.; Chow, T.C.; Yeung, Y.C.; Choi, K.W.; Tso, Y.K.; Lau, T.; Lai, S.T.; et al. Sars-Associated Viral Hepatitis Caused by a Novel Coronavirus: Report of Three Cases. Hepatology 2004, 39, 302–310. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Kim, Y.J.; Chung, E.H.; Kim, D.W.; Jeong, I.; Kim, Y.; Yun, M.R.; Kim, S.S.; Kim, G.; Joh, J.S. The Clinical and Virological Features of the First Imported Case Causing Mers-Cov Outbreak in South Korea, 2015. BMC Infect. Dis. 2017, 17, 498. [Google Scholar] [CrossRef] [PubMed]
- Chai, X.; Hu, L.; Zhang, Y.; Han, W.; Lu, Z.; Ke, A.; Zhou, J.; Shi, G.; Fang, N.; Fan, J.; et al. Specific Ace2 Expression in Cholangiocytes May Cause Liver Damage after 2019-Ncov Infection. BioRxiv 2020. [Google Scholar] [CrossRef]
- Zhang, H.; Kang, Z.; Gong, H.; Xu, D.; Wang, J.; Li, Z.; Cui, X.; Xiao, J.; Meng, T.; Zhou, W.; et al. The Digestive System Is a Potential Route of 2019-Ncov Infection: A Bioinformatics Analysis Based on Single-Cell Transcriptomes. BioRxiv 2020. [Google Scholar] [CrossRef]
- Zou, X.; Chen, K.; Zou, J.; Han, P.; Hao, J.; Han, Z. The Single Cell Rna Seq Data Analysis on the Receptor Ace2 Expression Reveals the Potential Risk of Different Human Organs Vulnerable to Wuhan 2019 Ncov Infection. Front. Med. 2020, 12, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Qi, L.; Chi, X.; Yang, J.; Wei, X.; Gong, E.; Peh, S.; Gu, J. Orchitis: A Complication of Severe Acute Respiratory Syndrome (Sars). Biol. Reprod. 2006, 74, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Hermann, B.P.; Cheng, K.; Singh, A.; Roa-De La Cruz, L.; Mutoji, K.N.; Chen, I.C.; Gildersleeve, H.; Lehle, J.D.; Mayo, M.; Westernströer, B.; et al. The Mammalian Spermatogenesis Single-Cell Transcriptome, from Spermatogonial Stem Cells to Spermatids. Cell Rep. 2018, 25, 1650–1667. [Google Scholar] [CrossRef] [PubMed]
- de Wit, E.; van Doremalen, N.; Falzarano, D.; Munster, V.J. Sars and Mers: Recent Insights into Emerging Coronaviruses. Nat. Rev. Microbiol. 2016, 14, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.R.; Liu, Y.X. Regulation of Spermatogonial Stem Cell Self-Renewal and Spermatocyte Meiosis by Sertoli Cell Signaling. Reproduction 2015, 149, R159–R167. [Google Scholar] [CrossRef] [PubMed]
- Zirkin, B.R.; Papadopoulos, V. Leydig Cells: Formation, Function, and Regulation. Biol. Reprod. 2018, 99, 101–111. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, Z.; Xu, X. scRNA-seq Profiling of Human Testes Reveals the Presence of the ACE2 Receptor, A Target for SARS-CoV-2 Infection in Spermatogonia, Leydig and Sertoli Cells. Cells 2020, 9, 920. https://doi.org/10.3390/cells9040920
Wang Z, Xu X. scRNA-seq Profiling of Human Testes Reveals the Presence of the ACE2 Receptor, A Target for SARS-CoV-2 Infection in Spermatogonia, Leydig and Sertoli Cells. Cells. 2020; 9(4):920. https://doi.org/10.3390/cells9040920
Chicago/Turabian StyleWang, Zhengpin, and Xiaojiang Xu. 2020. "scRNA-seq Profiling of Human Testes Reveals the Presence of the ACE2 Receptor, A Target for SARS-CoV-2 Infection in Spermatogonia, Leydig and Sertoli Cells" Cells 9, no. 4: 920. https://doi.org/10.3390/cells9040920
APA StyleWang, Z., & Xu, X. (2020). scRNA-seq Profiling of Human Testes Reveals the Presence of the ACE2 Receptor, A Target for SARS-CoV-2 Infection in Spermatogonia, Leydig and Sertoli Cells. Cells, 9(4), 920. https://doi.org/10.3390/cells9040920





