Is T Cell Negative Selection a Learning Algorithm?
Abstract
1. Introduction
2. Results
2.1. Problem Definition and Model Design
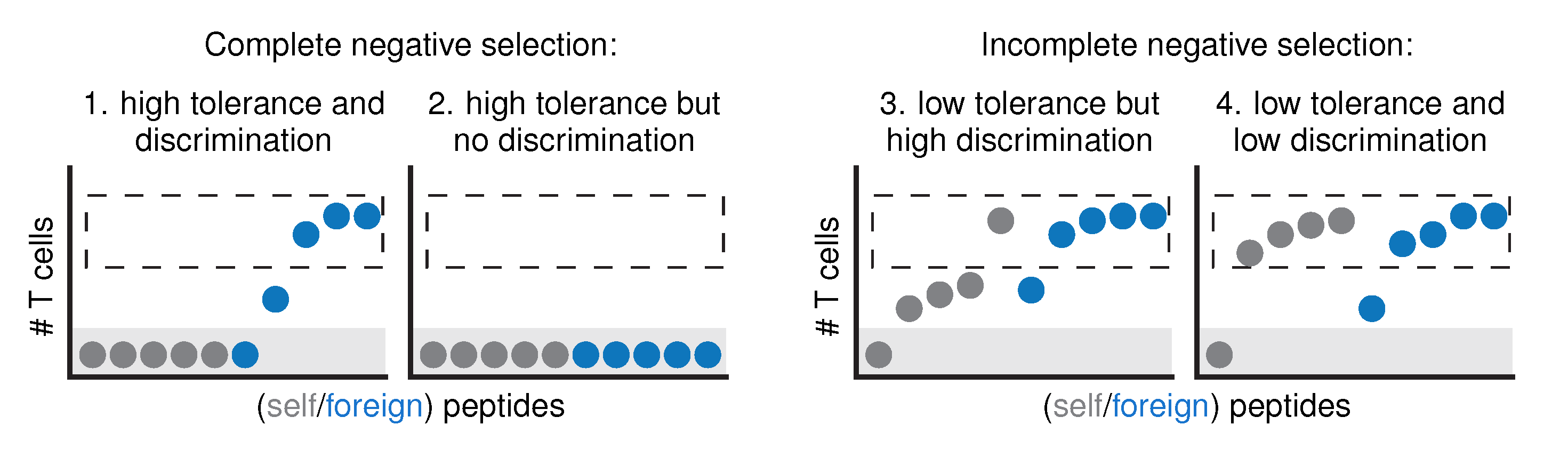
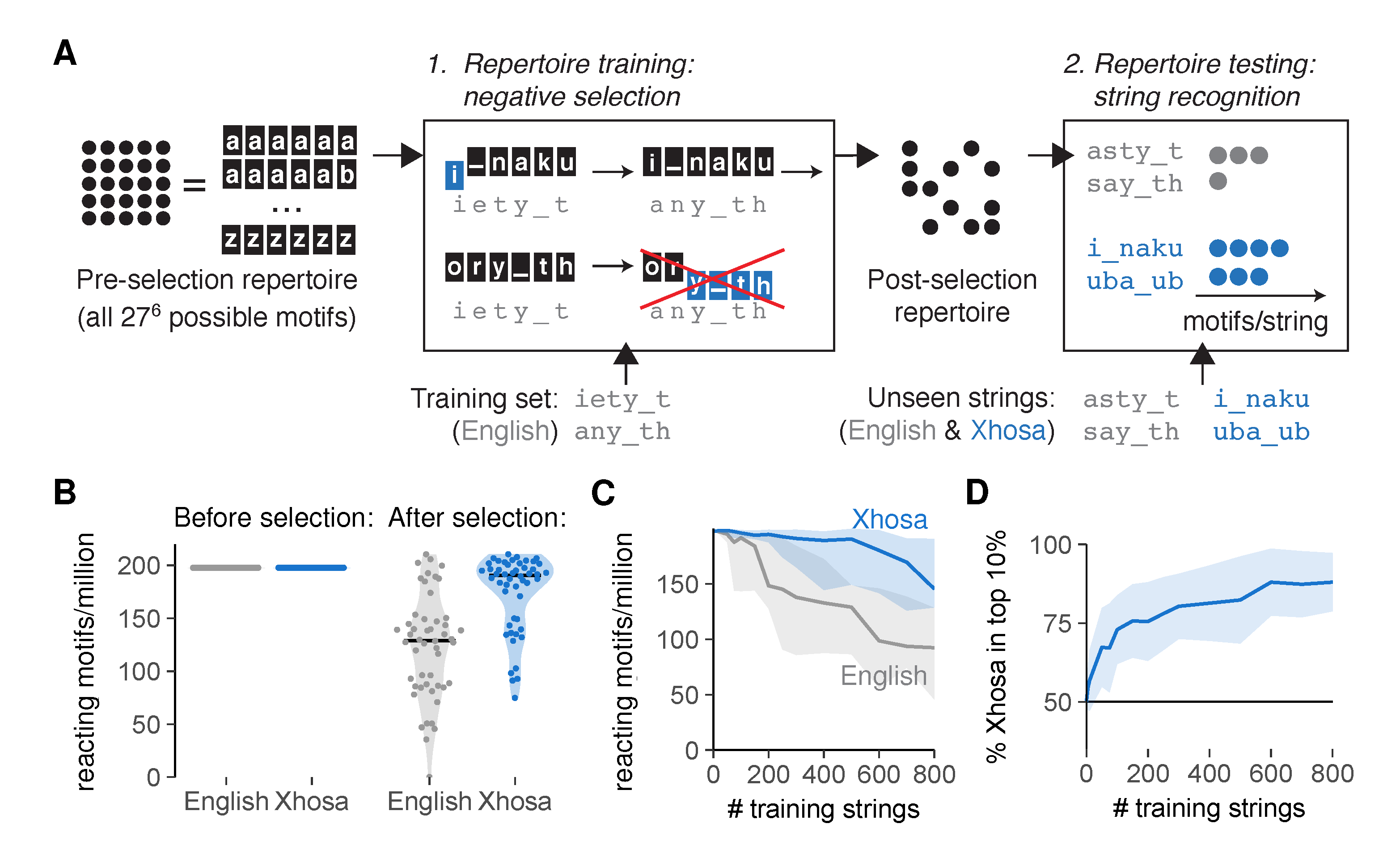
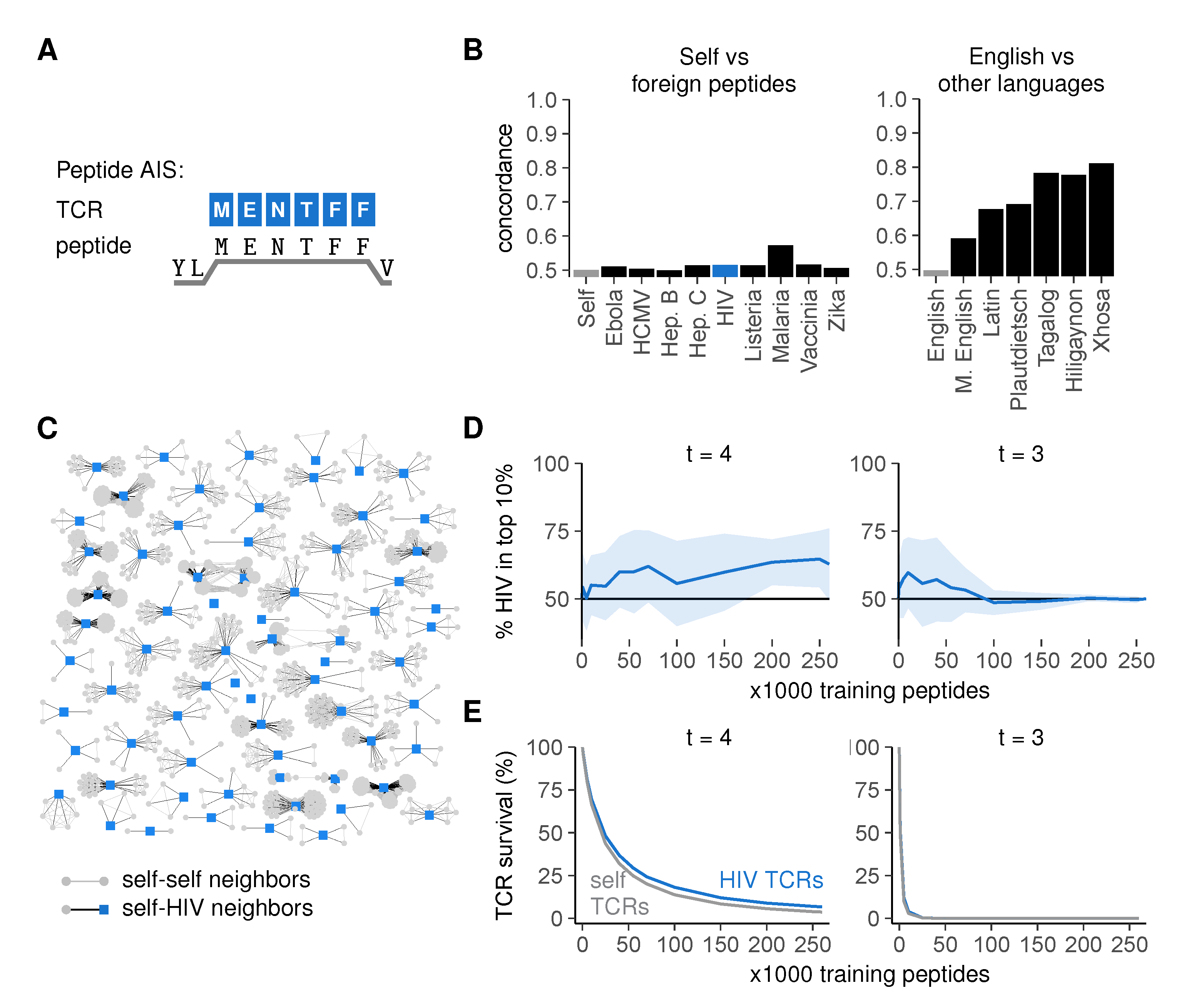
2.2. An Artificial Immune System Discriminates Self from Foreign after Negative Selection
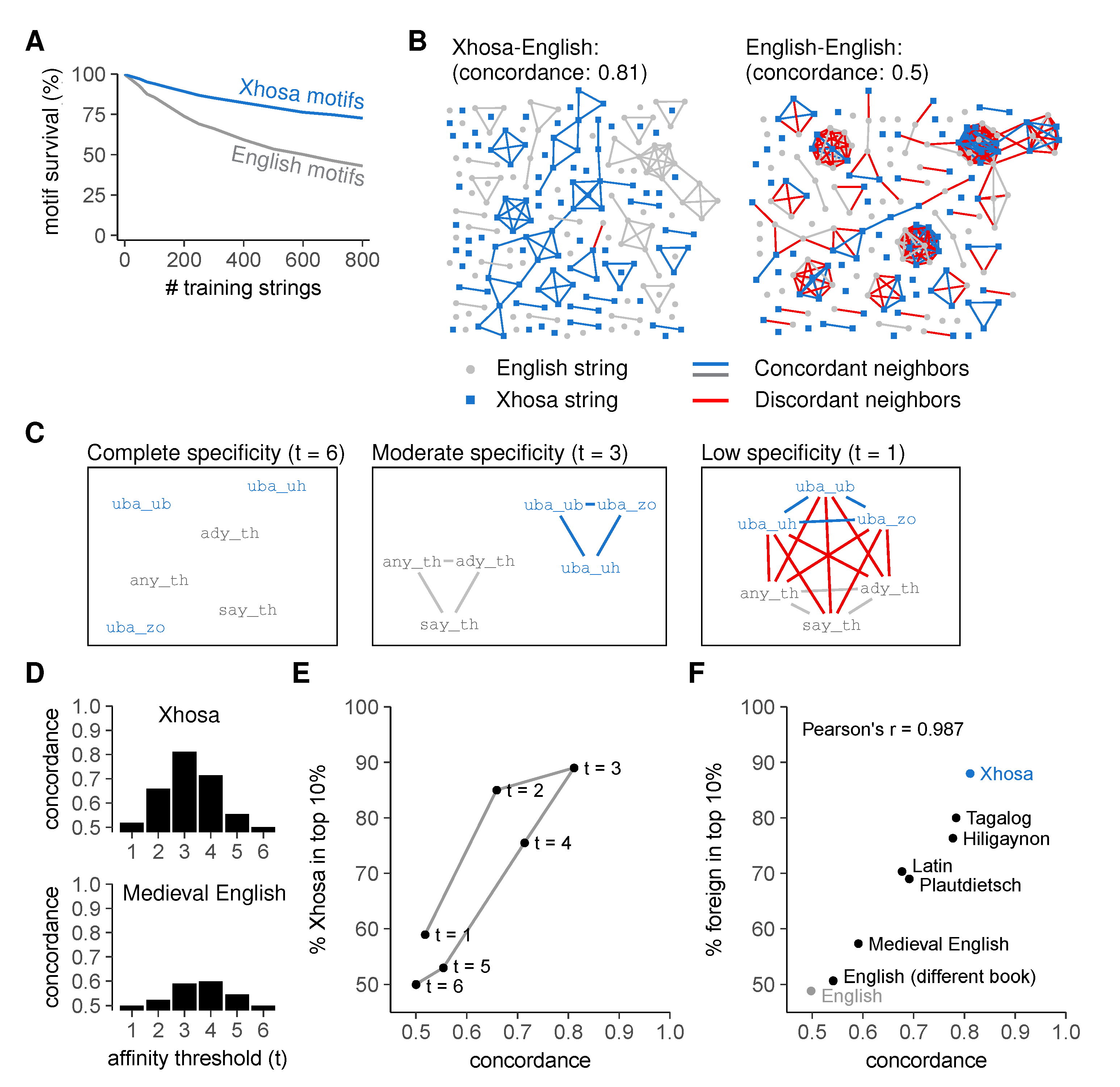
2.3. Discrimination Relies on Moderate Cross-Reactivity and Sequence Dissimilarity
2.4. Sequence Similarity Hampers Discrimination between Self- and Foreign Peptides
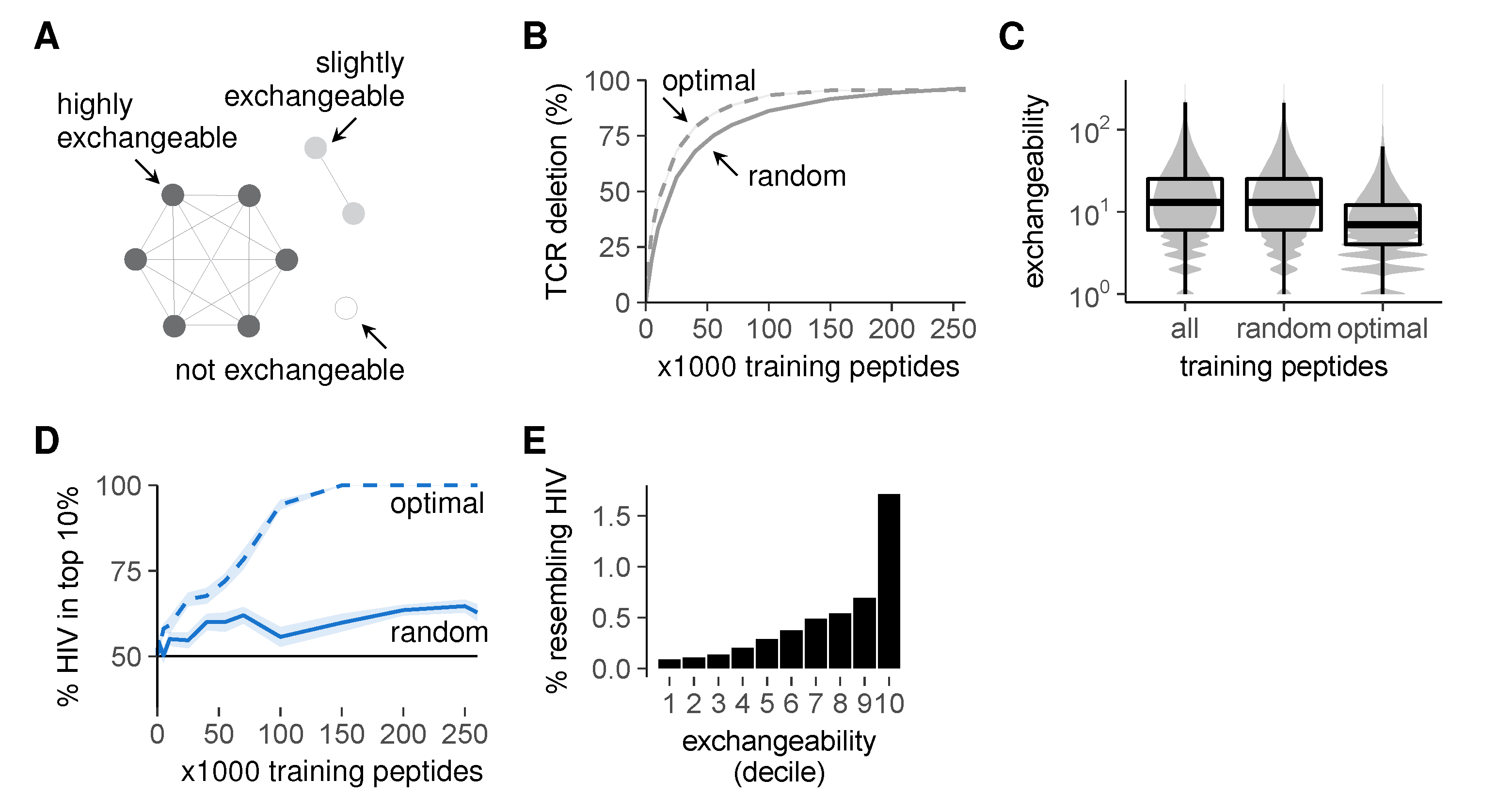
2.5. Selection on Non-Random Peptides Greatly Improves Self-Foreign Discrimination
3. Materials and Methods
3.1. Data and Code Availability
3.2. Simulation of Negative Selection
- Generation of an unbiased TCR repertoire containing all possible motifs of length 6. For details, see Repertoire model of negative selection (Appendix A.2).
- Selection of a training set of either n English strings or n self peptides. See Sequences (Appendix A.1) for details on the sequences used, and Training set selection (Appendix A.3) for details on the manners in which training sets are sampled. The training set selection method was random unless mentioned otherwise in the figure legend. The value of n can also be found in the figure legend.
- Negative selection of TCRs on the training set. All TCR motifs that match any of the training sequences in at least t adjacent positions are removed from the repertoire. Unless mentioned otherwise, negative selection was performed with an affinity threshold t = 3 for strings and t = 4 for peptides (see figure legends). All TCRs that remain make up the post-selection repertoire. For details on computational methods, see Repertoire model of negative selection (Appendix A.2).
- Analysis of the recognition of test sequences by the post-selection repertoire. Test sets always consist of “unseen” sequences that were not part of the training set used for negative selection. See figure legends for details on the number and source of the test sequences used. See Post-selection repertoire analysis (Appendix A.5) for details on specific analysis metrics used.
3.3. Supporting Methods
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| AA | Amino acid |
| AIS | Artificial immune system |
| ANN | Artificial neural network |
| HCMV | Human cytomegalovirus |
| HIV | Human immunodeficiency virus |
| MHC | Major histocompatibility complex |
| SD | Standard deviation |
| SEM | Standard error of the mean |
| TCR | T cell receptor |
Appendix A. Supplementary Methods
Appendix A.1. Sequences
Strings
Peptides
| Organism | Proteome Details | Proteins | ID | Download Date (d/m/y) | Unique 6-mers (#) |
|---|---|---|---|---|---|
| Ebola virus | Mayinga, Zaire, 1976 | 9 | UP000007209 | 27/09/2017 | 140 |
| Human cyto- megalovirus (HCMV) | Human herpesvirus 5 AD169 Isolate Unknown X17403 | 190 | UP000008991 | 27/09/2017 | 2090 |
| Hepatitis B virus | Genotype D subtype ayw (isolate France/Tiollais/1979) | 7 | UP000007930 | 27/09/2017 | 65 |
| Hepatitis C virus | H77 isolate Unknown AF009606 | 2 | UP000000518 | 27/09/2017 | 112 |
| Human immuno- deficiency virus (HIV) | Type 1 group M subtype B (isolate HXB2) | 9 | UP000002241 | 27/09/2017 | 69 |
| Vaccinia virus | Strain Copenhagen | 257 | UP000008269 | 27/09/2017 | 1955 |
| Zika virus | MR 766 Isolate Unknown AY632535 | 1 | UP000054557 | 27/09/2017 | 118 |
| Listeria monocytogenes | serovar 1/2a (strain ATCC BAA-679/EGD-e ) | 2844 | UP000000817 | 27/09/2017 | 31,251 |
| Plasmodium ovale (Malaria) | Wallikeri | 8636 | UP000078550 | 27/09/2017 | 89,408 |
| Homo sapiens (human) | - | 20,230 | UP000005640 | 01/06/2017 | 263,216 |
Appendix A.2. Repertoire Model of Negative Selection
Appendix A.3. Training Set Selection
Optimal Training Peptide Selection
- List the self-reactive TCR motifs that still remain in the repertoire;
- Select the self peptide that deletes the most of these remaining self-reactive TCRs. If multiple self peptides delete an equal number of remaining TCRs, we pick only those self peptides that do not overlap in the TCRs they delete.
Biased Training Peptide Selection
Appendix A.4. Sequence Analysis
String Graphs
Peptide Graphs
Concordance
AA Enrichment
Exchangeability
Appendix A.5. Post-Selection Repertoire Analysis
Sequence Recognition
Self-Foreign Discrimination
Affinity Distribution
TCR Survival/Deletion
Appendix A.6. Statistical Analysis
References
- Cooper, M.D.; Alder, M.N. The Evolution of Adaptive Immune Systems. Cell 2006, 124, 815–822. [Google Scholar] [CrossRef] [PubMed]
- Flajnik, M.F.; Kasahara, M. Origin and evolution of the adaptive immune system: Genetic events and selective pressures. Nat. Rev. Genet. 2009, 11, nrg2703. [Google Scholar] [CrossRef] [PubMed]
- Qi, Q.; Liu, Y.; Cheng, Y.; Glanville, J.; Zhang, D.; Lee, J.Y.; Olshen, R.A.; Weyand, C.M.; Boyd, S.D.; Goronzy, J.J. Diversity and clonal selection in the human T-cell repertoire. Proc. Natl. Acad. Sci. USA 2014, 111, 13139–13144. [Google Scholar] [CrossRef]
- Davis, M.M.; Bjorkman, P.J. T-cell antigen receptor genes and T-cell recognition. Nature 1988, 334, 334395a0. [Google Scholar] [CrossRef] [PubMed]
- Zarnitsyna, V.; Evavold, B.; Schoettle, L.; Blattman, J.; Antia, R. Estimating the Diversity, Completeness, and Cross-Reactivity of the T Cell Repertoire. Front. Immunol. 2013, 4. [Google Scholar] [CrossRef]
- Silverstein, A.M. Autoimmunity versus horror autotoxicus: The struggle for recognition. Nat. Immunol. 2001, 2, ni0401. [Google Scholar] [CrossRef]
- Detours, V.; Mehr, R.; Perelson, A.S. Deriving Quantitative Constraints on T Cell Selection from Data on the Mature T Cell Repertoire. J. Immunol. 2000, 164, 121–128. [Google Scholar] [CrossRef]
- Müller, V.; Bonhoeffer, S. Quantitative constraints on the scope of negative selection. Trends Immunol. 2003, 24, 132–135. [Google Scholar] [CrossRef]
- Vrisekoop, N.; Monteiro, J.; Mandl, J.; Germain, R. Revisiting Thymic Positive Selection and the Mature T Cell Repertoire for Antigen. Immunity 2014, 41, 181–190. [Google Scholar] [CrossRef]
- Yu, W.; Jiang, N.; Ebert, P.R.; Kidd, B.; Müller, S.; Lund, P.; Juang, J.; Adachi, K.; Tse, T.; Birnbaum, M.; et al. Clonal Deletion Prunes but Does Not Eliminate Self-Specific αβCD8+ T Lymphocytes. Immunity 2015, 42, 929–941. [Google Scholar] [CrossRef]
- Legoux, F.P.; Lim, J.B.; Cauley, A.W.; Dikiy, S.; Ertelt, J.; Mariani, T.J.; Sparwasser, T.; Way, S.S.; Moon, J.J. CD4+ T Cell Tolerance to Tissue-Restricted Self Antigens Is Mediated by Antigen-Specific Regulatory T Cells Rather than Deletion. Immunity 2015, 43, 896–908. [Google Scholar] [CrossRef] [PubMed]
- Davis, M. Not-So-Negative Selection. Immunity 2015, 43, 833–835. [Google Scholar] [CrossRef] [PubMed]
- Calis, J.J.A.; Boer, R.J.D.; Kesmir, C. Degenerate T-cell Recognition of Peptides on MHC Molecules Creates Large Holes in the T-cell Repertoire. PLoS Comput. Biol. 2012, 8, e1002412. [Google Scholar] [CrossRef] [PubMed]
- Gold, E.M. Language identification in the limit. Inf. Control. 1967, 10, 447–474. [Google Scholar] [CrossRef]
- McClelland, J.L.; McNaughton, B.L.; O’Reilly, R.C. Why there are complementary learning systems in the hippocampus and neocortex: Insights from the successes and failures of connectionist models of learning and memory. Psychol. Rev. 1995, 102, 419–457. [Google Scholar] [CrossRef]
- Forrest, S.; Hofmeyr, S.A.; Somayaji, A. Computer Immunology. Commun. ACM 1997, 40, 88–96. [Google Scholar] [CrossRef]
- Damashek, M. Gauging Similarity with n-Grams: Language-Independent Categorization of Text. Science 1995, 267, 843–848. [Google Scholar] [CrossRef]
- Jenkins, M.K.; Moon, J.J. The Role of Naive T Cell Precursor Frequency and Recruitment in Dictating Immune Response Magnitude. J. Immunol. 2012, 188, 4135–4140. [Google Scholar] [CrossRef]
- Martinez, R.J.; Evavold, B.D. Lower Affinity T Cells are Critical Components and Active Participants of the Immune Response. Front. Immunol. 2015, 6, 468. [Google Scholar] [CrossRef]
- Castro, L.D.; Timmis, J. Artificial Immune Systems: A New Computational Intelligence Approach; Springer Science & Business Media: London, UK, 2002. [Google Scholar]
- Percus, J.K.; Percus, O.E.; Perelson, A.S. Predicting the size of the T-cell receptor and antibody combining region from consideration of efficient self-nonself discrimination. Proc. Natl. Acad. Sci. USA 1993, 90, 1691–1695. [Google Scholar] [CrossRef]
- Elberfeld, M.; Textor, J. Negative selection algorithms on strings with efficient training and linear-time classification. Theor. Comput. Sci. 2011, 412, 534–542. [Google Scholar] [CrossRef]
- Frankild, S.; Boer, R.J.D.; Lund, O.; Nielsen, M.; Kesmir, C. Amino Acid Similarity Accounts for T Cell Cross-Reactivity and for “Holes” in the T Cell Repertoire. PLoS ONE 2008, 3, e1831. [Google Scholar] [CrossRef] [PubMed]
- Košmrlj, A.; Jha, A.K.; Huseby, E.S.; Kardar, M.; Chakraborty, A.K. How the thymus designs antigen-specific and self-tolerant T cell receptor sequences. Proc. Natl. Acad. Sci. USA 2008, 105, 16671–16676. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Chakraborty, A.K.; Kardar, M. How nonuniform contact profiles of T cell receptors modulate thymic selection outcomes. Phys. Rev. E 2018, 97, 032413. [Google Scholar] [CrossRef] [PubMed]
- Birnbaum, M.E.; Mendoza, J.L.; Sethi, D.K.; Dong, S.; Glanville, J.; Dobbins, J.; Ozkan, E.; Davis, M.M.; Wucherpfennig, K.W.; Garcia, K.C. Deconstructing the peptide-MHC specificity of T cell recognition. Cell 2014, 157, 1073–1087. [Google Scholar] [CrossRef]
- Nelson, R.W.; Beisang, D.; Tubo, N.J.; Dileepan, T.; Wiesner, D.L.; Nielsen, K.; Wüthrich, M.; Klein, B.S.; Kotov, D.I.; Spanier, J.A.; et al. T cell receptor cross-reactivity between similar foreign and self peptides influences naive cell population size and autoimmunity. Immunity 2015, 42, 95–107. [Google Scholar] [CrossRef]
- Riley, T.P.; Hellman, L.M.; Gee, M.H.; Mendoza, J.L.; Alonso, J.A.; Foley, K.C.; Nishimura, M.I.; Vander Kooi, C.W.; Garcia, K.C.; Baker, B.M. T cell receptor cross-reactivity expanded by dramatic peptide-MHC adaptability. Nat. Chem. Biol. 2018, 14, 934–942. [Google Scholar] [CrossRef]
- Dash, P.; Fiore-Gartland, A.J.; Hertz, T.; Wang, G.C.; Sharma, S.; Souquette, A.; Crawford, J.C.; Clemens, E.B.; Nguyen, T.H.O.; Kedzierska, K.; et al. Quantifiable predictive features define epitope-specific T cell receptor repertoires. Nature 2017, 547, 89–93. [Google Scholar] [CrossRef]
- Glanville, J.; Huang, H.; Nau, A.; Hatton, O.; Wagar, L.E.; Rubelt, F.; Ji, X.; Han, A.; Krams, S.M.; Pettus, C.; et al. Identifying specificity groups in the T cell receptor repertoire. Nature 2017, 547, 94–98. [Google Scholar] [CrossRef]
- Dunning, T. Statistical Identification of Language; Technical Report; New Mexico State University: Las Cruces, NM, USA, 1994. [Google Scholar]
- Ishizuka, J.; Grebe, K.; Shenderov, E.; Peters, B.; Chen, Q.; Peng, Y.; Wang, L.; Dong, T.; Pasquetto, V.; Oseroff, C.; et al. Quantitating T Cell Cross-Reactivity for Unrelated Peptide Antigens. J. Immunol. 2009, 183, 4337–4345. [Google Scholar] [CrossRef]
- Blattman, J.N.; Antia, R.; Sourdive, D.J.D.; Wang, X.; Kaech, S.M.; Murali-Krishna, K.; Altman, J.D.; Ahmed, R. Estimating the Precursor Frequency of Naive Antigen-specific CD8 T Cells. J. Exp. Med. 2002, 195, 657–664. [Google Scholar] [CrossRef] [PubMed]
- Alanio, C.; Lemaitre, F.; Law, H.K.W.; Hasan, M.; Albert, M.L. Enumeration of human antigen– specific naive CD8+ T cells reveals conserved precursor frequencies. Blood 2010, 115, 3718–3725. [Google Scholar] [CrossRef] [PubMed]
- Legoux, F.; Debeaupuis, E.; Echasserieau, K.; Salle, H.D.L.; Saulquin, X.; Bonneville, M. Impact of TCR Reactivity and HLA Phenotype on Naive CD8 T Cell Frequency in Humans. J. Immunol. 2010, 184, 6731–6738. [Google Scholar] [CrossRef] [PubMed]
- Schmidt, J.; Neumann-Haefelin, C.; Altay, T.; Gostick, E.; Price, D.A.; Lohmann, V.; Blum, H.E.; Thimme, R. Immunodominance of HLA-A2-Restricted Hepatitis C Virus-Specific CD8+ T Cell Responses Is Linked to Naïve-Precursor Frequency. J. Virol. 2011, 85, 5232–5236. [Google Scholar] [CrossRef]
- Hoof, I.; Peters, B.; Sidney, J.; Pedersen, L.E.; Sette, A.; Lund, O.; Buus, S.; Nielsen, M. NetMHCpan, a method for MHC class I binding prediction beyond humans. Immunogenetics 2009, 61, 1. [Google Scholar] [CrossRef]
- Sewell, A.K. Why must T cells be cross-reactive? Nat. Rev. Immunol. 2012, 12, nri3279. [Google Scholar] [CrossRef]
- Yates, A.J. Theories and quantification of thymic selection. Front. Immunol. 2014, 5, 13. [Google Scholar] [CrossRef]
- Butler, T.C.; Kardar, M.; Chakraborty, A.K. Quorum sensing allows T cells to discriminate between self and nonself. Proc. Natl. Acad. Sci. USA 2013, 110, 11833–11838. [Google Scholar] [CrossRef]
- Voisinne, G.; Nixon, B.G.; Melbinger, A.; Gasteiger, G.; Vergassola, M.; Altan-Bonnet, G. T Cells Integrate Local and Global Cues to Discriminate between Structurally Similar Antigens. Cell Rep. 2015, 11, 1208–1219. [Google Scholar] [CrossRef]
- Klein, L.; Kyewski, B.; Allen, P.M.; Hogquist, K.A. Positive and negative selection of the T cell repertoire: What thymocytes see (and don’t see). Nat. Rev. Immunol. 2014, 14, nri3667. [Google Scholar] [CrossRef]
- Nitta, T.; Murata, S.; Sasaki, K.; Fujii, H.; Ripen, A.M.; Ishimaru, N.; Koyasu, S.; Tanaka, K.; Takahama, Y. Thymoproteasome Shapes Immunocompetent Repertoire of CD8+ T Cells. Immunity 2010, 32, 29–40. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, K.; Takada, K.; Ohte, Y.; Kondo, H.; Sorimachi, H.; Tanaka, K.; Takahama, Y.; Murata, S. Thymoproteasomes produce unique peptide motifs for positive selection of CD8+ T cells. Nat. Commun. 2015, 6, ncomms8484. [Google Scholar] [CrossRef] [PubMed]
- Adamopoulou, E.; Tenzer, S.; Hillen, N.; Klug, P.; Rota, I.A.; Tietz, S.; Gebhardt, M.; Stevanovic, S.; Schild, H.; Tolosa, E.; et al. Exploring the MHC-peptide matrix of central tolerance in the human thymus. Nat. Commun. 2013, 4. [Google Scholar] [CrossRef]
- Schuster, H.; Shao, W.; Weiss, T.; Pedrioli, P.G.; Roth, P.; Weller, M.; Campbell, D.S.; Deutsch, E.W.; Moritz, R.L.; Planz, O.; et al. A tissue-based draft map of the murine MHC class I immunopeptidome. Sci. Data 2018, 5. [Google Scholar] [CrossRef] [PubMed]
- Ignatowicz, L.; Kappler, J.; Marrack, P. The Repertoire of T Cells Shaped by a Single MHC/Peptide Ligand. Cell 1996, 84, 521–529. [Google Scholar] [CrossRef][Green Version]
- Jain, E.; Bairoch, A.; Duvaud, S.; Phan, I.; Redaschi, N.; Suzek, B.E.; Martin, M.J.; McGarvey, P.; Gasteiger, E. Infrastructure for the life sciences: Design and implementation of the UniProt website. BMC Bioinform. 2009, 10, 136. [Google Scholar] [CrossRef]
- UniProt Consortium. Ongoing and future developments at the Universal Protein Resource. Nucleic Acids Res. 2011, 39, D214–D219. [Google Scholar] [CrossRef]
- Textor, J.; Dannenberg, K.; Liśkiewicz, M. A Generic Finite Automata Based Approach to Implementing Lymphocyte Repertoire Models. In Proceedings of the 2014 Annual Conference on Genetic and Evolutionary Computation; ACM: New York, NY, USA, 2014; pp. 129–136. [Google Scholar] [CrossRef]
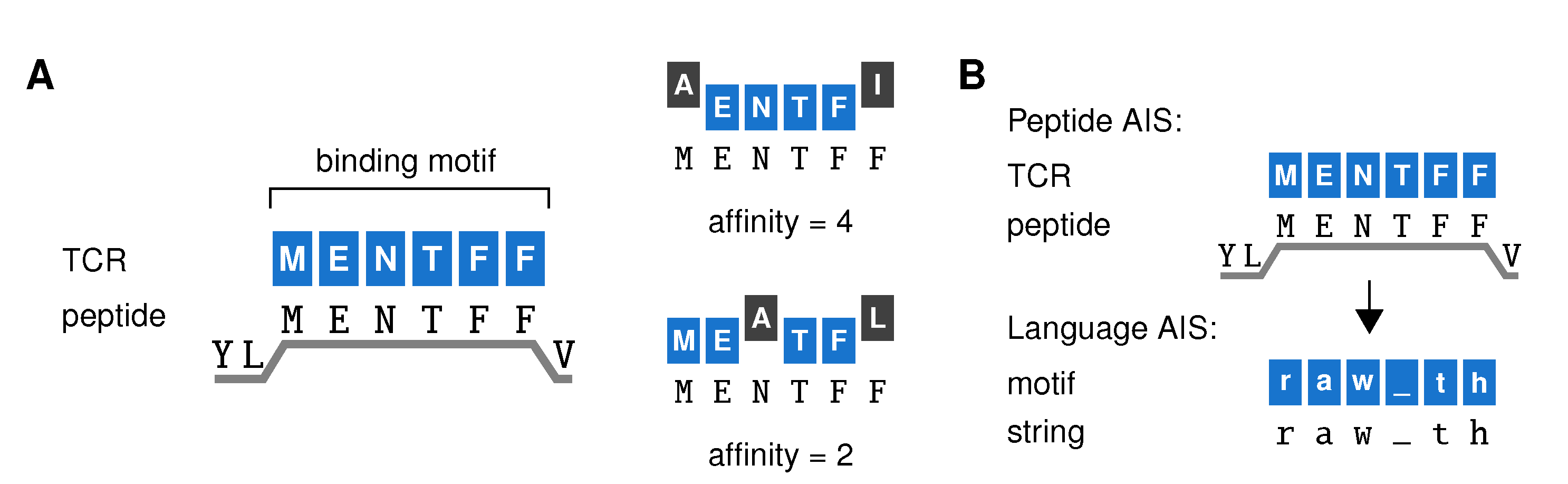

© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wortel, I.M.N.; Keşmir, C.; de Boer, R.J.; Mandl, J.N.; Textor, J. Is T Cell Negative Selection a Learning Algorithm? Cells 2020, 9, 690. https://doi.org/10.3390/cells9030690
Wortel IMN, Keşmir C, de Boer RJ, Mandl JN, Textor J. Is T Cell Negative Selection a Learning Algorithm? Cells. 2020; 9(3):690. https://doi.org/10.3390/cells9030690
Chicago/Turabian StyleWortel, Inge M. N., Can Keşmir, Rob J. de Boer, Judith N. Mandl, and Johannes Textor. 2020. "Is T Cell Negative Selection a Learning Algorithm?" Cells 9, no. 3: 690. https://doi.org/10.3390/cells9030690
APA StyleWortel, I. M. N., Keşmir, C., de Boer, R. J., Mandl, J. N., & Textor, J. (2020). Is T Cell Negative Selection a Learning Algorithm? Cells, 9(3), 690. https://doi.org/10.3390/cells9030690





