Ribosome and Translational Control in Stem Cells
Abstract
1. Introduction
2. Control of Translation in Stem Cells
2.1. Global Translation Increases During Stem Cell Differentiation
2.2. The Global Baseline of Translation Is Tightly Regulated in Stem Cells
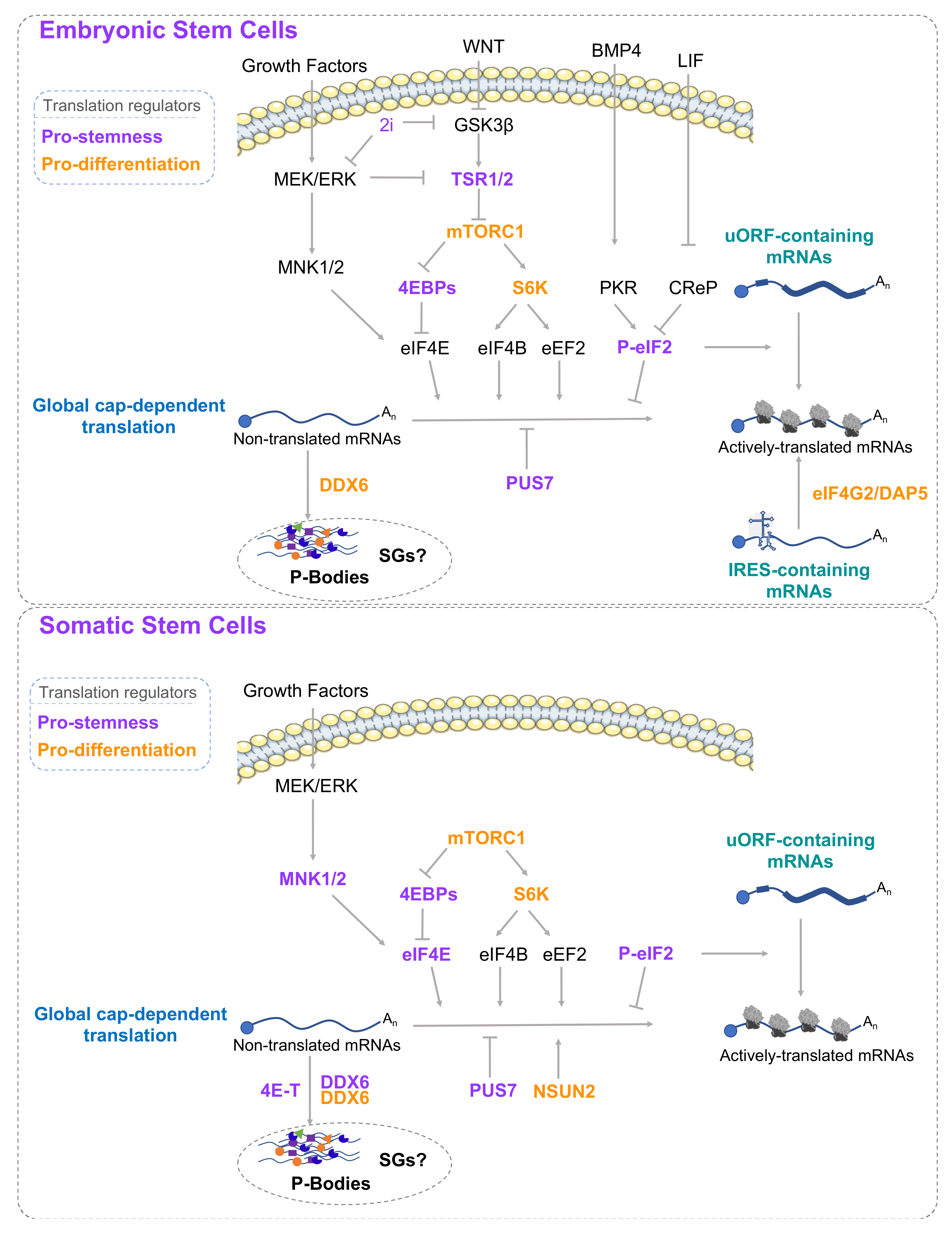
2.3. Mechanisms Regulating Translation in Stem Cells
2.3.1. 5’UTR Structures
2.3.2. The mTOR Pathway
2.3.3. The Mitogen-Activated Protein Kinase (MEK) Pathway
2.3.4. eIF2α Phosphorylation
2.3.5. RNA Granules
3. Ribosome Biogenesis in Stem Cells
3.1. Overview of Ribosome Biogenesis
3.2. The Expression and Modifications of rRNAs Are Highly Regulated in Stem Cells
3.3. Levels of Ribosome Biogenesis Factors Are Coordinated to Support Stem Cell Fate
3.3.1. Global Regulation of RBF Expression During Changes in Stem Cell Identity
3.3.2. Specific Ribosome Biogenesis Factors Are Preferentially Expressed in Stem Cells
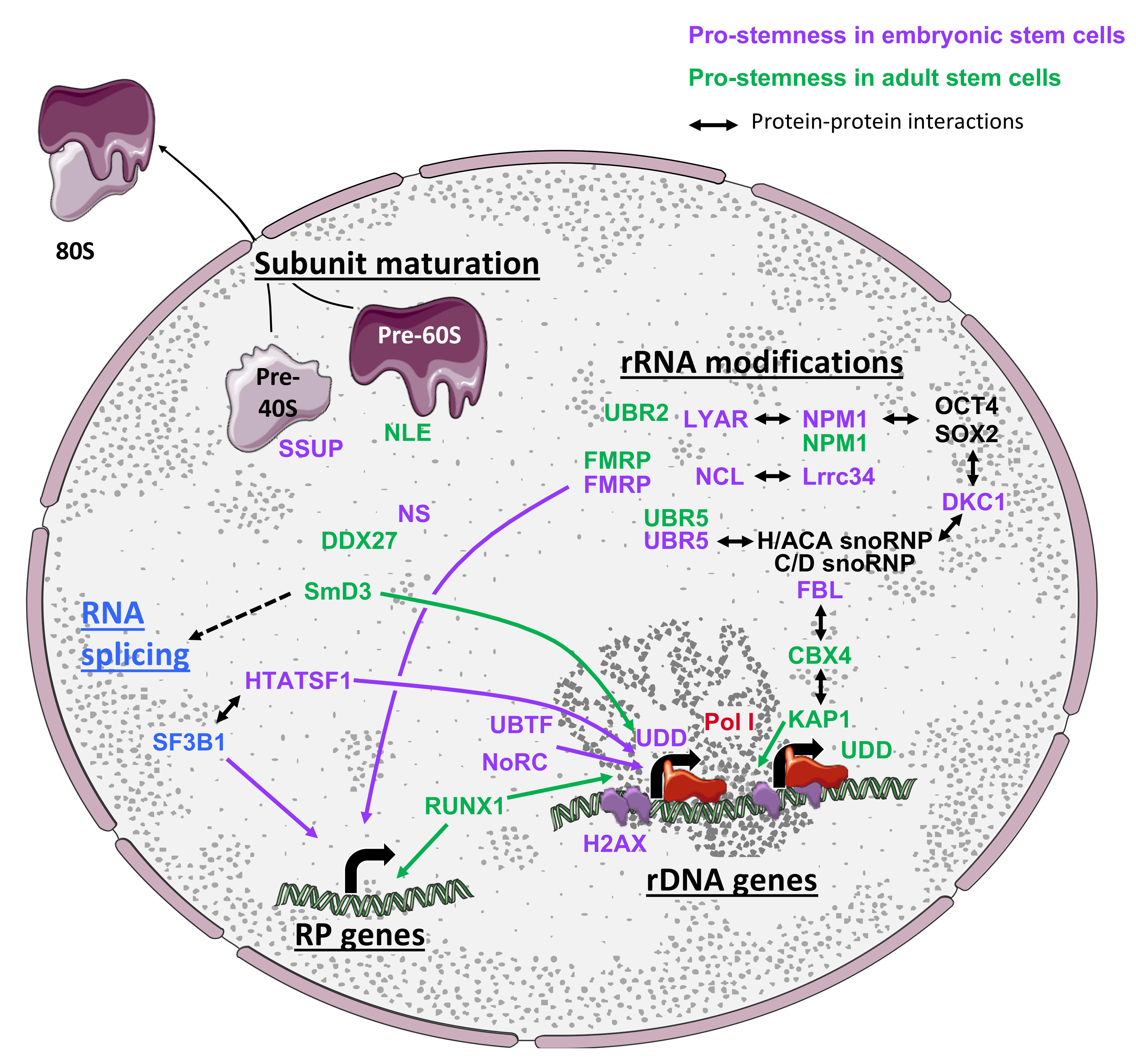
3.4. Ribosome Biogenesis Is Highly Regulated to Support Stem Cell Properties
3.4.1. Pre-Ribosomal Subunit Maturation Control in Stem Cells
3.4.2. Regulation of rRNA Modifications
3.4.3. Ribosome Biogenesis Surveillance by P53-Dependent Pathways in Stem Cells
3.5. Coordination Between Ribosome Biogenesis and Additional Steps of Gene Expression
4. The Ribosome, a New Regulator of Translation in Stem Cells
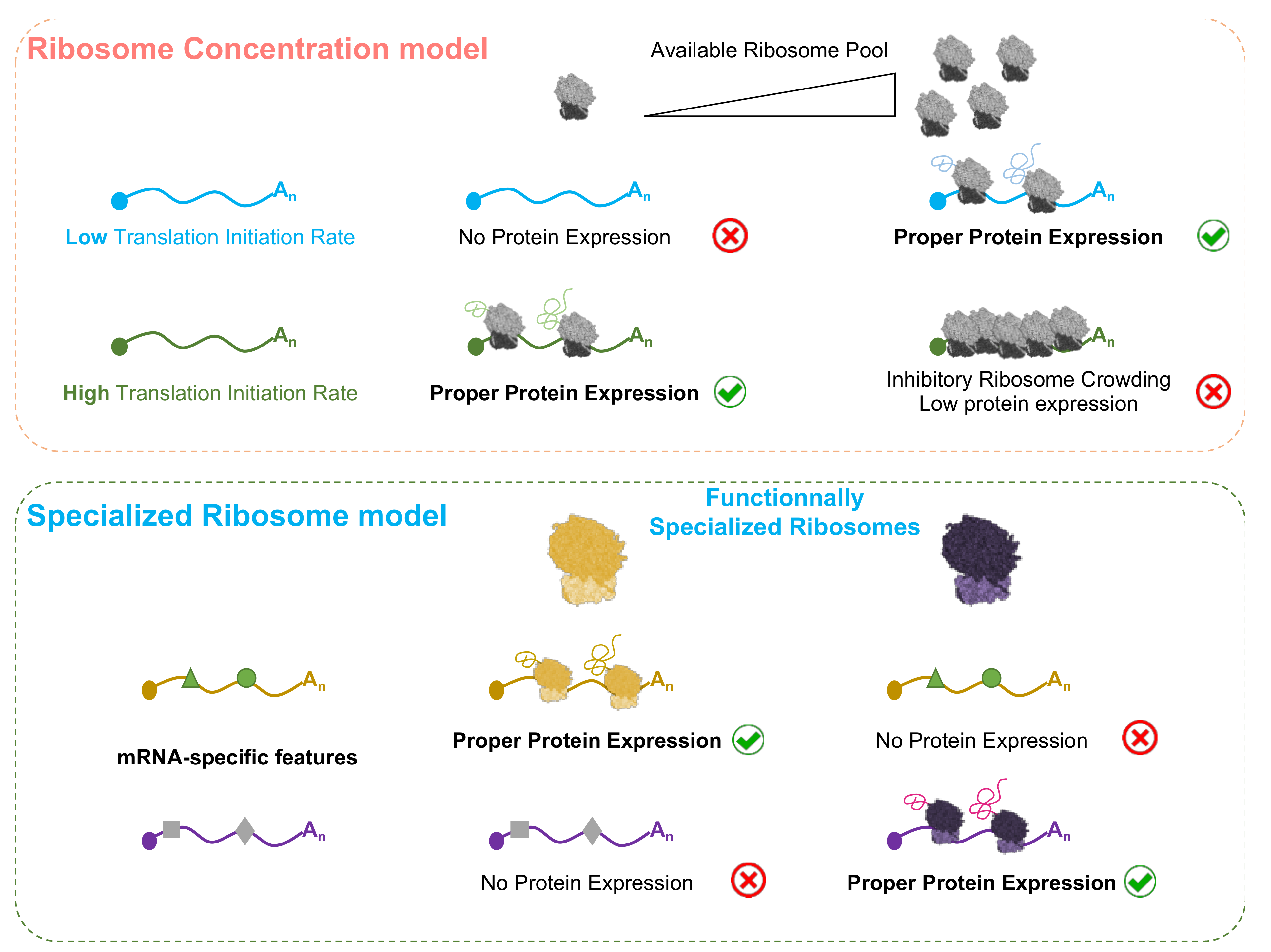
4.1. The Ribosome Concentration Regulates the Expression of Specific Transcripts
4.2. Specialized Ribosomes as Regulators of Gene Expression?
4.2.1. RP Tissue-Specific Expression and Functions
4.2.2. rRNA Modifications Contribute to Ribosome Heterogeneity
4.2.3. ESC Ribosomes Are Heterogeneous and Functionally Specialized
5. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- De Sousa Abreu, R.; Penalva, L.O.; Marcotte, E.M.; Vogel, C. Global signatures of protein and mRNA expression levels. Mol. Biosyst. 2009, 5, 1512–1526. [Google Scholar] [CrossRef] [PubMed]
- Maier, T.; Guell, M.; Serrano, L. Correlation of mRNA and protein in complex biological samples. FEBS Lett. 2009, 583, 3966–3973. [Google Scholar] [CrossRef] [PubMed]
- Schwanhausser, B.; Busse, D.; Li, N.; Dittmar, G.; Schuchhardt, J.; Wolf, J.; Chen, W.; Selbach, M. Global quantification of mammalian gene expression control. Nature 2011, 473, 337–342. [Google Scholar] [CrossRef] [PubMed]
- Munoz, J.; Low, T.Y.; Kok, Y.J.; Chin, A.; Frese, C.K.; Ding, V.; Choo, A.; Heck, A.J. The quantitative proteomes of human-induced pluripotent stem cells and embryonic stem cells. Mol. Syst. Biol. 2011, 7, 550. [Google Scholar] [CrossRef] [PubMed]
- Kristensen, A.R.; Gsponer, J.; Foster, L.J. Protein synthesis rate is the predominant regulator of protein expression during differentiation. Mol. Syst. Biol. 2013, 9, 689. [Google Scholar] [CrossRef]
- Lu, R.; Markowetz, F.; Unwin, R.D.; Leek, J.T.; Airoldi, E.M.; MacArthur, B.D.; Lachmann, A.; Rozov, R.; Ma’ayan, A.; Boyer, L.A.; et al. Systems-level dynamic analyses of fate change in murine embryonic stem cells. Nature 2009, 462, 358–362. [Google Scholar] [CrossRef]
- Ingolia, N.T.; Lareau, L.F.; Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 2011, 147, 789–802. [Google Scholar] [CrossRef]
- Liu, Y.; Beyer, A.; Aebersold, R. On the Dependency of Cellular Protein Levels on mRNA Abundance. Cell 2016, 165, 535–550. [Google Scholar] [CrossRef]
- Servier. Smart Servier Medical Art. Available online: https://smart.servier.com/ (accessed on 23 January 2020).
- Sampath, P.; Pritchard, D.K.; Pabon, L.; Reinecke, H.; Schwartz, S.M.; Morris, D.R.; Murry, C.E. A hierarchical network controls protein translation during murine embryonic stem cell self-renewal and differentiation. Cell Stem Cell 2008, 2, 448–460. [Google Scholar] [CrossRef]
- Easley, C.A.t.; Ben-Yehudah, A.; Redinger, C.J.; Oliver, S.L.; Varum, S.T.; Eisinger, V.M.; Carlisle, D.L.; Donovan, P.J.; Schatten, G.P. mTOR-mediated activation of p70 S6K induces differentiation of pluripotent human embryonic stem cells. Cell Reprogram 2010, 12, 263–273. [Google Scholar] [CrossRef]
- Friend, K.; Brooks, H.A.; Propson, N.E.; Thomson, J.A.; Kimble, J. Embryonic Stem Cell Growth Factors Regulate eIF2alpha Phosphorylation. PLoS ONE 2015, 10, e0139076. [Google Scholar] [CrossRef] [PubMed]
- Guzzi, N.; Ciesla, M.; Ngoc, P.C.T.; Lang, S.; Arora, S.; Dimitriou, M.; Pimkova, K.; Sommarin, M.N.E.; Munita, R.; Lubas, M.; et al. Pseudouridylation of tRNA-Derived Fragments Steers Translational Control in Stem Cells. Cell 2018, 173, 1204–1216. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Shen, L.; Yu, J.; Wan, H.; Guo, A.; Chen, J.; Long, Y.; Zhao, J.; Pei, G. Rapamycin and other longevity-promoting compounds enhance the generation of mouse induced pluripotent stem cells. Aging Cell 2011, 10, 908–911. [Google Scholar] [CrossRef]
- He, J.; Kang, L.; Wu, T.; Zhang, J.; Wang, H.; Gao, H.; Zhang, Y.; Huang, B.; Liu, W.; Kou, Z.; et al. An elaborate regulation of Mammalian target of rapamycin activity is required for somatic cell reprogramming induced by defined transcription factors. Stem Cells Dev. 2012, 21, 2630–2641. [Google Scholar] [CrossRef] [PubMed]
- You, K.T.; Park, J.; Kim, V.N. Role of the small subunit processome in the maintenance of pluripotent stem cells. Genes Dev. 2015, 29, 2004–2009. [Google Scholar] [CrossRef] [PubMed]
- Freyria, A.M.; Ronziere, M.C.; Boutillon, M.M.; Herbage, D. Effect of retinoic acid on protein synthesis by foetal bovine chondrocytes in high-density culture: Down-regulation of the glucose-regulated protein, GRP-78, and type II collagen. Biochem. J. 1995, 305 ( Pt 2), 391–396. [Google Scholar] [CrossRef]
- Koziorowska, J.; Paczek, K.; Tautt, J. Influence of retinoic acid on protein synthesis and transport of L-methionine in cultured L cells. Exp. Cell Res. 1984, 150, 97–103. [Google Scholar] [CrossRef]
- Lotan, R.; Fischer, I.; Meromsky, L.; Moldave, K. Effects of retinoic acid on protein synthesis in cultured melanoma cells. J. Cell Physiol. 1982, 113, 47–55. [Google Scholar] [CrossRef]
- Novak, A.; Amit, M.; Ziv, T.; Segev, H.; Fishman, B.; Admon, A.; Itskovitz-Eldor, J. Proteomics profiling of human embryonic stem cells in the early differentiation stage. Stem Cell Rev. Rep. 2012, 8, 137–149. [Google Scholar] [CrossRef]
- Blanco, S.; Bandiera, R.; Popis, M.; Hussain, S.; Lombard, P.; Aleksic, J.; Sajini, A.; Tanna, H.; Cortes-Garrido, R.; Gkatza, N.; et al. Stem cell function and stress response are controlled by protein synthesis. Nature 2016, 534, 335–340. [Google Scholar] [CrossRef]
- Hartman, N.W.; Lin, T.V.; Zhang, L.; Paquelet, G.E.; Feliciano, D.M.; Bordey, A. mTORC1 targets the translational repressor 4E-BP2, but not S6 kinase 1/2, to regulate neural stem cell self-renewal in vivo. Cell Rep. 2013, 5, 433–444. [Google Scholar] [CrossRef] [PubMed]
- Llorens-Bobadilla, E.; Zhao, S.; Baser, A.; Saiz-Castro, G.; Zwadlo, K.; Martin-Villalba, A. Single-Cell Transcriptomics Reveals a Population of Dormant Neural Stem Cells that Become Activated upon Brain Injury. Cell Stem Cell 2015, 17, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, C.G.; Teixeira, F.K.; Czech, B.; Preall, J.B.; Zamparini, A.L.; Seifert, J.R.; Malone, C.D.; Hannon, G.J.; Lehmann, R. Regulation of Ribosome Biogenesis and Protein Synthesis Controls Germline Stem Cell Differentiation. Cell Stem Cell 2016, 18, 276–290. [Google Scholar] [CrossRef] [PubMed]
- Signer, R.A.; Magee, J.A.; Salic, A.; Morrison, S.J. Haematopoietic stem cells require a highly regulated protein synthesis rate. Nature 2014, 509, 49–54. [Google Scholar] [CrossRef]
- Signer, R.A.; Qi, L.; Zhao, Z.; Thompson, D.; Sigova, A.A.; Fan, Z.P.; DeMartino, G.N.; Young, R.A.; Sonenberg, N.; Morrison, S.J. The rate of protein synthesis in hematopoietic stem cells is limited partly by 4E-BPs. Genes Dev 2016, 30, 1698–1703. [Google Scholar] [CrossRef]
- Yang, G.; Smibert, C.A.; Kaplan, D.R.; Miller, F.D. An eIF4E1/4E-T complex determines the genesis of neurons from precursors by translationally repressing a proneurogenic transcription program. Neuron 2014, 84, 723–739. [Google Scholar] [CrossRef]
- Zismanov, V.; Chichkov, V.; Colangelo, V.; Jamet, S.; Wang, S.; Syme, A.; Koromilas, A.E.; Crist, C. Phosphorylation of eIF2alpha Is a Translational Control Mechanism Regulating Muscle Stem Cell Quiescence and Self-Renewal. Cell Stem Cell 2016, 18, 79–90. [Google Scholar] [CrossRef]
- Bulut-Karslioglu, A.; Macrae, T.A.; Oses-Prieto, J.A.; Covarrubias, S.; Percharde, M.; Ku, G.; Diaz, A.; McManus, M.T.; Burlingame, A.L.; Ramalho-Santos, M. The Transcriptionally Permissive Chromatin State of Embryonic Stem Cells Is Acutely Tuned to Translational Output. Cell Stem Cell 2018, 22, 369–383. [Google Scholar] [CrossRef]
- Blanco, S.; Dietmann, S.; Flores, J.V.; Hussain, S.; Kutter, C.; Humphreys, P.; Lukk, M.; Lombard, P.; Treps, L.; Popis, M.; et al. Aberrant methylation of tRNAs links cellular stress to neuro-developmental disorders. EMBO J. 2014, 33, 2020–2039. [Google Scholar] [CrossRef]
- Blanco, S.; Kurowski, A.; Nichols, J.; Watt, F.M.; Benitah, S.A.; Frye, M. The RNA-methyltransferase Misu (NSun2) poises epidermal stem cells to differentiate. PLoS Genet. 2011, 7, e1002403. [Google Scholar] [CrossRef]
- Tuorto, F.; Liebers, R.; Musch, T.; Schaefer, M.; Hofmann, S.; Kellner, S.; Frye, M.; Helm, M.; Stoecklin, G.; Lyko, F. RNA cytosine methylation by Dnmt2 and NSun2 promotes tRNA stability and protein synthesis. Nat. Struct. Mol. Biol. 2012, 19, 900–905. [Google Scholar] [CrossRef] [PubMed]
- Kong, S.Y.; Kim, W.; Lee, H.R.; Kim, H.J. The histone demethylase KDM5A is required for the repression of astrocytogenesis and regulated by the translational machinery in neural progenitor cells. FASEB J. 2018, 32, 1108–1119. [Google Scholar] [CrossRef] [PubMed]
- Hinnebusch, A.G.; Ivanov, I.P.; Sonenberg, N. Translational control by 5′-untranslated regions of eukaryotic mRNAs. Science 2016, 352, 1413–1416. [Google Scholar] [CrossRef] [PubMed]
- Jackson, R.J.; Hellen, C.U.; Pestova, T.V. The mechanism of eukaryotic translation initiation and principles of its regulation. Nat. Rev. Mol. Cell Biol. 2010, 11, 113–127. [Google Scholar] [CrossRef] [PubMed]
- Sonenberg, N.; Dever, T.E. Eukaryotic translation initiation factors and regulators. Curr. Opin. Struct. Biol. 2003, 13, 56–63. [Google Scholar] [CrossRef]
- Calvo, S.E.; Pagliarini, D.J.; Mootha, V.K. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc. Natl. Acad. Sci. USA 2009, 106, 7507–7512. [Google Scholar] [CrossRef]
- Lu, P.D.; Harding, H.P.; Ron, D. Translation reinitiation at alternative open reading frames regulates gene expression in an integrated stress response. J. Cell Biol. 2004, 167, 27–33. [Google Scholar] [CrossRef]
- Mendell, J.T.; Sharifi, N.A.; Meyers, J.L.; Martinez-Murillo, F.; Dietz, H.C. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat. Genet. 2004, 36, 1073–1078. [Google Scholar] [CrossRef]
- Morris, D.R.; Geballe, A.P. Upstream open reading frames as regulators of mRNA translation. Mol. Cell. Biol. 2000, 20, 8635–8642. [Google Scholar] [CrossRef]
- Yoffe, Y.; David, M.; Kalaora, R.; Povodovski, L.; Friedlander, G.; Feldmesser, E.; Ainbinder, E.; Saada, A.; Bialik, S.; Kimchi, A. Cap-independent translation by DAP5 controls cell fate decisions in human embryonic stem cells. Genes Dev. 2016, 30, 1991–2004. [Google Scholar] [CrossRef]
- Henis-Korenblit, S.; Shani, G.; Sines, T.; Marash, L.; Shohat, G.; Kimchi, A. The caspase-cleaved DAP5 protein supports internal ribosome entry site-mediated translation of death proteins. Proc. Natl. Acad. Sci. USA 2002, 99, 5400–5405. [Google Scholar] [CrossRef] [PubMed]
- Lewis, S.M.; Cerquozzi, S.; Graber, T.E.; Ungureanu, N.H.; Andrews, M.; Holcik, M. The eIF4G homolog DAP5/p97 supports the translation of select mRNAs during endoplasmic reticulum stress. Nucleic Acids Res. 2008, 36, 168–178. [Google Scholar] [CrossRef] [PubMed]
- Liberman, N.; Gandin, V.; Svitkin, Y.V.; David, M.; Virgili, G.; Jaramillo, M.; Holcik, M.; Nagar, B.; Kimchi, A.; Sonenberg, N. DAP5 associates with eIF2beta and eIF4AI to promote Internal Ribosome Entry Site driven translation. Nucleic Acids Res. 2015, 43, 3764–3775. [Google Scholar] [CrossRef] [PubMed]
- Marash, L.; Liberman, N.; Henis-Korenblit, S.; Sivan, G.; Reem, E.; Elroy-Stein, O.; Kimchi, A. DAP5 promotes cap-independent translation of Bcl-2 and CDK1 to facilitate cell survival during mitosis. Mol. Cell. 2008, 30, 447–459. [Google Scholar] [CrossRef] [PubMed]
- Gingras, A.C.; Raught, B.; Sonenberg, N. mTOR signaling to translation. Curr. Top. Microbiol. Immunol. 2004, 279, 169–197. [Google Scholar] [CrossRef] [PubMed]
- Laplante, M.; Sabatini, D.M. mTOR signaling in growth control and disease. Cell 2012, 149, 274–293. [Google Scholar] [CrossRef] [PubMed]
- Bulut-Karslioglu, A.; Biechele, S.; Jin, H.; Macrae, T.A.; Hejna, M.; Gertsenstein, M.; Song, J.S.; Ramalho-Santos, M. Inhibition of mTOR induces a paused pluripotent state. Nature 2016, 540, 119–123. [Google Scholar] [CrossRef]
- Tahmasebi, S.; Alain, T.; Rajasekhar, V.K.; Zhang, J.P.; Prager-Khoutorsky, M.; Khoutorsky, A.; Dogan, Y.; Gkogkas, C.G.; Petroulakis, E.; Sylvestre, A.; et al. Multifaceted regulation of somatic cell reprogramming by mRNA translational control. Cell Stem Cell 2014, 14, 606–616. [Google Scholar] [CrossRef]
- Ohtsuka, S.; Nakai-Futatsugi, Y.; Niwa, H. LIF signal in mouse embryonic stem cells. JAKSTAT 2015, 4, e1086520. [Google Scholar] [CrossRef]
- Wray, J.; Kalkan, T.; Smith, A.G. The ground state of pluripotency. Biochem. Soc. Trans. 2010, 38, 1027–1032. [Google Scholar] [CrossRef]
- Guzikowski, A.R.; Chen, Y.S.; Zid, B.M. Stress-induced mRNP granules: Form and function of processing bodies and stress granules. Wiley Interdiscip. Rev. RNA 2019, 10, e1524. [Google Scholar] [CrossRef] [PubMed]
- Sfakianos, A.P.; Whitmarsh, A.J.; Ashe, M.P. Ribonucleoprotein bodies are phased in. Biochem. Soc. Trans. 2016, 44, 1411–1416. [Google Scholar] [CrossRef] [PubMed]
- Kedersha, N.L.; Gupta, M.; Li, W.; Miller, I.; Anderson, P. RNA-binding proteins TIA-1 and TIAR link the phosphorylation of eIF-2 alpha to the assembly of mammalian stress granules. J. Cell. Biol. 1999, 147, 1431–1442. [Google Scholar] [CrossRef] [PubMed]
- Di Stefano, B.; Luo, E.C.; Haggerty, C.; Aigner, S.; Charlton, J.; Brumbaugh, J.; Ji, F.; Rabano Jimenez, I.; Clowers, K.J.; Huebner, A.J.; et al. The RNA Helicase DDX6 Controls Cellular Plasticity by Modulating P-Body Homeostasis. Cell Stem Cell 2019. [Google Scholar] [CrossRef] [PubMed]
- Bastide, A.; David, A. The ribosome, (slow) beating heart of cancer (stem) cell. Oncogenesis 2018, 7, 34. [Google Scholar] [CrossRef] [PubMed]
- De la Cruz, J.; Gomez-Herreros, F.; Rodriguez-Galan, O.; Begley, V.; de la Cruz Munoz-Centeno, M.; Chavez, S. Feedback regulation of ribosome assembly. Curr. Genet. 2018, 64, 393–404. [Google Scholar] [CrossRef] [PubMed]
- Klinge, S.; Woolford, J.L., Jr. Ribosome assembly coming into focus. Nat. Rev. Mol. Cell Biol. 2019, 20, 116–131. [Google Scholar] [CrossRef]
- Pelletier, J.; Thomas, G.; Volarevic, S. Ribosome biogenesis in cancer: New players and therapeutic avenues. Nat. Rev. Cancer 2018, 18, 51–63. [Google Scholar] [CrossRef]
- Zhang, Q.; Shalaby, N.A.; Buszczak, M. Changes in rRNA transcription influence proliferation and cell fate within a stem cell lineage. Science 2014, 343, 298–301. [Google Scholar] [CrossRef]
- Woolnough, J.L.; Atwood, B.L.; Liu, Z.; Zhao, R.; Giles, K.E. The Regulation of rRNA Gene Transcription during Directed Differentiation of Human Embryonic Stem Cells. PLoS ONE 2016, 11, e0157276. [Google Scholar] [CrossRef]
- Jarzebowski, L.; Le Bouteiller, M.; Coqueran, S.; Raveux, A.; Vandormael-Pournin, S.; David, A.; Cumano, A.; Cohen-Tannoudji, M. Mouse adult hematopoietic stem cells actively synthesize ribosomal RNA. RNA 2018, 24, 1803–1812. [Google Scholar] [CrossRef] [PubMed]
- Fichelson, P.; Huynh, J.R. [Asymetric growth in Drosophila stem cells is related to ribosomal biogenesis]. Med. Sci. 2009, 25, 780–781. [Google Scholar] [CrossRef]
- Eleuteri, B.; Aranda, S.; Ernfors, P. NoRC Recruitment by H2A.X Deposition at rRNA Gene Promoter Limits Embryonic Stem Cell Proliferation. Cell Rep. 2018, 23, 1853–1866. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.; Hu, B.; Song, M.; Ding, Z.; Dang, Y.; Liu, Z.; Zhang, W.; Ji, Q.; Ren, R.; Ding, J.; et al. Maintenance of Nucleolar Homeostasis by CBX4 Alleviates Senescence and Osteoarthritis. Cell Rep. 2019, 26, 3643–3656. [Google Scholar] [CrossRef]
- Cai, X.; Gao, L.; Teng, L.; Ge, J.; Oo, Z.M.; Kumar, A.R.; Gilliland, D.G.; Mason, P.J.; Tan, K.; Speck, N.A. Runx1 Deficiency Decreases Ribosome Biogenesis and Confers Stress Resistance to Hematopoietic Stem and Progenitor Cells. Cell Stem Cell 2015, 17, 165–177. [Google Scholar] [CrossRef]
- Kwon, S.C.; Yi, H.; Eichelbaum, K.; Fohr, S.; Fischer, B.; You, K.T.; Castello, A.; Krijgsveld, J.; Hentze, M.W.; Kim, V.N. The RNA-binding protein repertoire of embryonic stem cells. Nat. Struct. Mol. Biol. 2013, 20, 1122–1130. [Google Scholar] [CrossRef]
- Saha, S.; Murmu, K.C.; Biswas, M.; Chakraborty, S.; Basu, J.; Madhulika, S.; Kolapalli, S.P.; Chauhan, S.; Sengupta, A.; Prasad, P. Transcriptomic Analysis Identifies RNA Binding Proteins as Putative Regulators of Myelopoiesis and Leukemia. Front. Oncol. 2019, 9, 692. [Google Scholar] [CrossRef]
- Athanasiadis, E.I.; Botthof, J.G.; Andres, H.; Ferreira, L.; Lio, P.; Cvejic, A. Single-cell RNA-sequencing uncovers transcriptional states and fate decisions in haematopoiesis. Nat. Commun. 2017, 8, 2045. [Google Scholar] [CrossRef]
- Ximerakis, M.; Lipnick, S.L.; Innes, B.T.; Simmons, S.K.; Adiconis, X.; Dionne, D.; Mayweather, B.A.; Nguyen, L.; Niziolek, Z.; Ozek, C.; et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 2019, 22, 1696–1708. [Google Scholar] [CrossRef]
- Recher, G.; Jouralet, J.; Brombin, A.; Heuze, A.; Mugniery, E.; Hermel, J.M.; Desnoulez, S.; Savy, T.; Herbomel, P.; Bourrat, F.; et al. Zebrafish midbrain slow-amplifying progenitors exhibit high levels of transcripts for nucleotide and ribosome biogenesis. Development 2013, 140, 4860–4869. [Google Scholar] [CrossRef]
- Li, P.; Wu, M.; Lin, Q.; Wang, S.; Chen, T.; Jiang, H. Key genes and integrated modules in hematopoietic differentiation of human embryonic stem cells: A comprehensive bioinformatic analysis. Stem Cell Res. Ther. 2018, 9, 301. [Google Scholar] [CrossRef] [PubMed]
- Koyuncu, S.; Saez, I.; Lee, H.J.; Gutierrez-Garcia, R.; Pokrzywa, W.; Fatima, A.; Hoppe, T.; Vilchez, D. The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients. Nat. Commun. 2018, 9, 2886. [Google Scholar] [CrossRef] [PubMed]
- Watanabe-Susaki, K.; Takada, H.; Enomoto, K.; Miwata, K.; Ishimine, H.; Intoh, A.; Ohtaka, M.; Nakanishi, M.; Sugino, H.; Asashima, M.; et al. Biosynthesis of ribosomal RNA in nucleoli regulates pluripotency and differentiation ability of pluripotent stem cells. Stem Cells 2014, 32, 3099–3111. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.; Shi, G.; Zhou, C.; Lu, R.; Li, H.; Sun, L.; Jin, Y. Nucleolin maintains embryonic stem cell self-renewal by suppression of p53 protein-dependent pathway. J. Biol. Chem 2011, 286, 43370–43382. [Google Scholar] [CrossRef] [PubMed]
- Fong, Y.W.; Ho, J.J.; Inouye, C.; Tjian, R. The dyskerin ribonucleoprotein complex as an OCT4/SOX2 coactivator in embryonic stem cells. Elife 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, B.; Yang, A.; Lu, R.; Wang, W.; Zhou, Y.; Shi, G.; Kwon, S.W.; Zhao, Y.; Jin, Y. Ly-1 antibody reactive clone is an important nucleolar protein for control of self-renewal and differentiation in embryonic stem cells. Stem Cells 2009, 27, 1244–1254. [Google Scholar] [CrossRef]
- Le Bouteiller, M.; Souilhol, C.; Beck-Cormier, S.; Stedman, A.; Burlen-Defranoux, O.; Vandormael-Pournin, S.; Bernex, F.; Cumano, A.; Cohen-Tannoudji, M. Notchless-dependent ribosome synthesis is required for the maintenance of adult hematopoietic stem cells. J. Exp. Med. 2013, 210, 2351–2369. [Google Scholar] [CrossRef]
- Stedman, A.; Beck-Cormier, S.; Le Bouteiller, M.; Raveux, A.; Vandormael-Pournin, S.; Coqueran, S.; Lejour, V.; Jarzebowski, L.; Toledo, F.; Robine, S.; et al. Ribosome biogenesis dysfunction leads to p53-mediated apoptosis and goblet cell differentiation of mouse intestinal stem/progenitor cells. Cell Death Differ. 2015, 22, 1865–1876. [Google Scholar] [CrossRef]
- Gayraud-Morel, B.; Le Bouteiller, M.; Commere, P.H.; Cohen-Tannoudji, M.; Tajbakhsh, S. Notchless defines a stage-specific requirement for ribosome biogenesis during lineage progression in adult skeletal myogenesis. Development 2018, 145. [Google Scholar] [CrossRef]
- Romanova, L.; Grand, A.; Zhang, L.; Rayner, S.; Katoku-Kikyo, N.; Kellner, S.; Kikyo, N. Critical role of nucleostemin in pre-rRNA processing. J. Biol. Chem. 2009, 284, 4968–4977. [Google Scholar] [CrossRef]
- Romanova, L.; Kellner, S.; Katoku-Kikyo, N.; Kikyo, N. Novel role of nucleostemin in the maintenance of nucleolar architecture and integrity of small nucleolar ribonucleoproteins and the telomerase complex. J. Biol. Chem. 2009, 284, 26685–26694. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Zhou, X.; Wu, R.; Mosley, A.; Zeng, S.X.; Xing, Z.; Lu, H. The role of IMP dehydrogenase 2 in Inauhzin-induced ribosomal stress. Elife 2014, 3. [Google Scholar] [CrossRef] [PubMed]
- Baddoo, M.; Hill, K.; Wilkinson, R.; Gaupp, D.; Hughes, C.; Kopen, G.C.; Phinney, D.G. Characterization of mesenchymal stem cells isolated from murine bone marrow by negative selection. J. Cell. Biochem. 2003, 89, 1235–1249. [Google Scholar] [CrossRef]
- Ohmura, M.; Naka, K.; Hoshii, T.; Muraguchi, T.; Shugo, H.; Tamase, A.; Uema, N.; Ooshio, T.; Arai, F.; Takubo, K.; et al. Identification of stem cells during prepubertal spermatogenesis via monitoring of nucleostemin promoter activity. Stem Cells 2008, 26, 3237–3246. [Google Scholar] [CrossRef] [PubMed]
- Qu, J.; Bishop, J.M. Nucleostemin maintains self-renewal of embryonic stem cells and promotes reprogramming of somatic cells to pluripotency. J. Cell Biol. 2012, 197, 731–745. [Google Scholar] [CrossRef] [PubMed]
- Tsai, R.Y.; McKay, R.D. A nucleolar mechanism controlling cell proliferation in stem cells and cancer cells. Genes Dev. 2002, 16, 2991–3003. [Google Scholar] [CrossRef]
- Yamashita, M.; Nitta, E.; Nagamatsu, G.; Ikushima, Y.M.; Hosokawa, K.; Arai, F.; Suda, T. Nucleostemin is indispensable for the maintenance and genetic stability of hematopoietic stem cells. Biochem. Biophys. Res. Commun. 2013, 441, 196–201. [Google Scholar] [CrossRef]
- Tsai, R.Y. Turning a new page on nucleostemin and self-renewal. J. Cell Sci. 2014, 127, 3885–3891. [Google Scholar] [CrossRef]
- Yu, J.; Luan, X.; Yan, Y.; Qiao, C.; Liu, Y.; Zhao, D.; Xie, B.; Zheng, Q.; Wang, M.; Chen, W.; et al. Small ribonucleoprotein particle protein SmD3 governs the homeostasis of germline stem cells and the crosstalk between the spliceosome and ribosome signals in Drosophila. FASEB J. 2019, 33, 8125–8137. [Google Scholar] [CrossRef]
- Bennett, A.H.; O’Donohue, M.F.; Gundry, S.R.; Chan, A.T.; Widrick, J.; Draper, I.; Chakraborty, A.; Zhou, Y.; Zon, L.I.; Gleizes, P.E.; et al. RNA helicase, DDX27 regulates skeletal muscle growth and regeneration by modulation of translational processes. PLoS Genet. 2018, 14, e1007226. [Google Scholar] [CrossRef]
- Cai, P.; Mao, X.; Zhao, J.; Luo, L. Ribosome biogenesis protein Urb2 regulates hematopoietic stem cells development via P53 pathway in zebrafish. Biochem. Biophys. Res. Commun. 2018, 497, 776–782. [Google Scholar] [CrossRef]
- Saez, I.; Gerbracht, J.V.; Koyuncu, S.; Lee, H.J.; Horn, M.; Kroef, V.; Denzel, M.S.; Dieterich, C.; Gehring, N.H.; Vilchez, D. The E3 ubiquitin ligase UBR5 interacts with the H/ACA ribonucleoprotein complex and regulates ribosomal RNA biogenesis in embryonic stem cells. FEBS Lett. 2019. [Google Scholar] [CrossRef]
- Luhrig, S.; Siamishi, I.; Tesmer-Wolf, M.; Zechner, U.; Engel, W.; Nolte, J. Lrrc34, a novel nucleolar protein, interacts with npm1 and ncl and has an impact on pluripotent stem cells. Stem Cells Dev. 2014, 23, 2862–2874. [Google Scholar] [CrossRef] [PubMed]
- Nachmani, D.; Bothmer, A.H.; Grisendi, S.; Mele, A.; Bothmer, D.; Lee, J.D.; Monteleone, E.; Cheng, K.; Zhang, Y.; Bester, A.C.; et al. Germline NPM1 mutations lead to altered rRNA 2′-O-methylation and cause dyskeratosis congenita. Nat. Genet. 2019, 51, 1518–1529. [Google Scholar] [CrossRef] [PubMed]
- Johansson, H.; Simonsson, S. Core transcription factors, Oct4, Sox2 and Nanog, individually form complexes with nucleophosmin (Npm1) to control embryonic stem (ES) cell fate determination. Aging (Albany NY) 2010, 2, 815–822. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Shan, G.; Guo, W.; Smrt, R.D.; Johnson, E.B.; Li, X.; Pfeiffer, R.L.; Szulwach, K.E.; Duan, R.; Barkho, B.Z.; et al. Fragile x mental retardation protein regulates proliferation and differentiation of adult neural stem/progenitor cells. PLoS Genet. 2010, 6, e1000898. [Google Scholar] [CrossRef]
- Guo, W.; Allan, A.M.; Zong, R.; Zhang, L.; Johnson, E.B.; Schaller, E.G.; Murthy, A.C.; Goggin, S.L.; Eisch, A.J.; Oostra, B.A.; et al. Ablation of Fmrp in adult neural stem cells disrupts hippocampus-dependent learning. Nat. Med. 2011, 17, 559–565. [Google Scholar] [CrossRef]
- Telias, M.; Segal, M.; Ben-Yosef, D. Neural differentiation of Fragile X human Embryonic Stem Cells reveals abnormal patterns of development despite successful neurogenesis. Dev. Biol. 2013, 374, 32–45. [Google Scholar] [CrossRef]
- Liu, B.; Li, Y.; Stackpole, E.E.; Novak, A.; Gao, Y.; Zhao, Y.; Zhao, X.; Richter, J.D. Regulatory discrimination of mRNAs by FMRP controls mouse adult neural stem cell differentiation. Proc. Natl. Acad. Sci. USA 2018, 115, E11397–E11405. [Google Scholar] [CrossRef]
- Chen, E.; Sharma, M.R.; Shi, X.; Agrawal, R.K.; Joseph, S. Fragile X mental retardation protein regulates translation by binding directly to the ribosome. Mol. Cell. 2014, 54, 407–417. [Google Scholar] [CrossRef]
- D’Souza, M.N.; Gowda, N.K.C.; Tiwari, V.; Babu, R.O.; Anand, P.; Dastidar, S.G.; Singh, R.; James, O.G.; Selvaraj, B.; Pal, R.; et al. FMRP Interacts with C/D Box snoRNA in the Nucleus and Regulates Ribosomal RNA Methylation. iScience 2019, 12, 368. [Google Scholar] [CrossRef] [PubMed]
- Aubert, M.; O’Donohue, M.F.; Lebaron, S.; Gleizes, P.E. Pre-Ribosomal RNA Processing in Human Cells: From Mechanisms to Congenital Diseases. Biomolecules 2018, 8, 123. [Google Scholar] [CrossRef] [PubMed]
- James, A.; Wang, Y.; Raje, H.; Rosby, R.; DiMario, P. Nucleolar stress with and without p53. Nucleus 2014, 5, 402–426. [Google Scholar] [CrossRef] [PubMed]
- Yang, K.; Yang, J.; Yi, J. Nucleolar Stress: Hallmarks, sensing mechanism and diseases. Cell Stress 2018, 2, 125–140. [Google Scholar] [CrossRef] [PubMed]
- Armistead, J.; Triggs-Raine, B. Diverse diseases from a ubiquitous process: The ribosomopathy paradox. FEBS Lett. 2014, 588, 1491–1500. [Google Scholar] [CrossRef]
- Danilova, N.; Gazda, H.T. Ribosomopathies: How a common root can cause a tree of pathologies. Dis Model Mech. 2015, 8, 1013–1026. [Google Scholar] [CrossRef]
- Mills, E.W.; Green, R. Ribosomopathies: There’s strength in numbers. Science 2017, 358. [Google Scholar] [CrossRef]
- Sulima, S.O.; Kampen, K.R.; De Keersmaecker, K. Cancer Biogenesis in Ribosomopathies. Cells 2019, 8, 229. [Google Scholar] [CrossRef]
- Tahmasebi, S.; Khoutorsky, A.; Mathews, M.B.; Sonenberg, N. Translation deregulation in human disease. Nat. Rev. Mol. Cell Biol. 2018, 19, 791–807. [Google Scholar] [CrossRef]
- Morgado-Palacin, L.; Llanos, S.; Serrano, M. Ribosomal stress induces L11- and p53-dependent apoptosis in mouse pluripotent stem cells. Cell Cycle 2012, 11, 503–510. [Google Scholar] [CrossRef]
- Dutt, S.; Narla, A.; Lin, K.; Mullally, A.; Abayasekara, N.; Megerdichian, C.; Wilson, F.H.; Currie, T.; Khanna-Gupta, A.; Berliner, N.; et al. Haploinsufficiency for ribosomal protein genes causes selective activation of p53 in human erythroid progenitor cells. Blood 2011, 117, 2567–2576. [Google Scholar] [CrossRef] [PubMed]
- Fortier, S.; MacRae, T.; Bilodeau, M.; Sargeant, T.; Sauvageau, G. Haploinsufficiency screen highlights two distinct groups of ribosomal protein genes essential for embryonic stem cell fate. Proc. Natl. Acad. Sci. USA 2015, 112, 2127–2132. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Bellini, M.; Ceman, S. Fragile X mental retardation protein FMRP binds mRNAs in the nucleus. Mol. Cell Biol. 2009, 29, 214–228. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, V.; Kishore, A.H.; Febitha, K.K.; Kundu, T.K. Human histone chaperone nucleophosmin enhances acetylation-dependent chromatin transcription. Mol. Cell Biol. 2005, 25, 7534–7545. [Google Scholar] [CrossRef] [PubMed]
- Tasaki, T.; Mulder, L.C.; Iwamatsu, A.; Lee, M.J.; Davydov, I.V.; Varshavsky, A.; Muesing, M.; Kwon, Y.T. A family of mammalian E3 ubiquitin ligases that contain the UBR box motif and recognize N-degrons. Mol. Cell Biol. 2005, 25, 7120–7136. [Google Scholar] [CrossRef] [PubMed]
- Corsini, N.S.; Peer, A.M.; Moeseneder, P.; Roiuk, M.; Burkard, T.R.; Theussl, H.C.; Moll, I.; Knoblich, J.A. Coordinated Control of mRNA and rRNA Processing Controls Embryonic Stem Cell Pluripotency and Differentiation. Cell Stem Cell 2018, 22, 543–558. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, T.L.; Stoneley, M.; Spriggs, K.A.; Bushell, M. TOPs and their regulation. Biochem. Soc. Trans. 2006, 34, 12–16. [Google Scholar] [CrossRef]
- Yoshihama, M.; Uechi, T.; Asakawa, S.; Kawasaki, K.; Kato, S.; Higa, S.; Maeda, N.; Minoshima, S.; Tanaka, T.; Shimizu, N.; et al. The human ribosomal protein genes: Sequencing and comparative analysis of 73 genes. Genome Res. 2002, 12, 379–390. [Google Scholar] [CrossRef]
- Meyuhas, O.a.H.E. Translational Control of Gene Expression, 2nd ed.; Nahum Sonenberg, J.W.B.H., Michael, M., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, USA, 2000. [Google Scholar]
- Meyuhas, O.; Kahan, T. The race to decipher the top secrets of TOP mRNAs. Biochim. Biophys. Acta 2015, 1849, 801–811. [Google Scholar] [CrossRef]
- Stolovich, M.; Tang, H.; Hornstein, E.; Levy, G.; Cohen, R.; Bae, S.S.; Birnbaum, M.J.; Meyuhas, O. Transduction of growth or mitogenic signals into translational activation of TOP mRNAs is fully reliant on the phosphatidylinositol 3-kinase-mediated pathway but requires neither S6K1 nor rpS6 phosphorylation. Mol. Cell Biol. 2002, 22, 8101–8113. [Google Scholar] [CrossRef]
- Kampen, K.R.; Sulima, S.O.; Vereecke, S.; De Keersmaecker, K. Hallmarks of ribosomopathies. Nucleic Acids Res. 2019. [Google Scholar] [CrossRef] [PubMed]
- Yelick, P.C.; Trainor, P.A. Ribosomopathies: Global process, tissue specific defects. Rare Dis. 2015, 3, e1025185. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, L.S.; Gazda, H.T.; Eng, J.C.; Eichhorn, S.W.; Thiru, P.; Ghazvinian, R.; George, T.I.; Gotlib, J.R.; Beggs, A.H.; Sieff, C.A.; et al. Altered translation of GATA1 in Diamond-Blackfan anemia. Nat. Med. 2014, 20, 748–753. [Google Scholar] [CrossRef] [PubMed]
- Missra, A.; Ernest, B.; Lohoff, T.; Jia, Q.; Satterlee, J.; Ke, K.; von Arnim, A.G. The Circadian Clock Modulates Global Daily Cycles of mRNA Ribosome Loading. Plant. Cell 2015, 27, 2582–2599. [Google Scholar] [CrossRef] [PubMed]
- Jouffe, C.; Cretenet, G.; Symul, L.; Martin, E.; Atger, F.; Naef, F.; Gachon, F. The circadian clock coordinates ribosome biogenesis. PLoS Biol. 2013, 11, e1001455. [Google Scholar] [CrossRef]
- Sawicka, K.; Hale, C.R.; Park, C.Y.; Fak, J.J.; Gresack, J.E.; Van Driesche, S.J.; Kang, J.J.; Darnell, J.C.; Darnell, R.B. FMRP has a cell-type-specific role in CA1 pyramidal neurons to regulate autism-related transcripts and circadian memory. Elife 2019, 8. [Google Scholar] [CrossRef]
- Wang, J.; Mauvoisin, D.; Martin, E.; Atger, F.; Galindo, A.N.; Dayon, L.; Sizzano, F.; Palini, A.; Kussmann, M.; Waridel, P.; et al. Nuclear Proteomics Uncovers Diurnal Regulatory Landscapes in Mouse Liver. Cell Metab. 2017, 25, 102–117. [Google Scholar] [CrossRef]
- Janich, P.; Arpat, A.B.; Castelo-Szekely, V.; Lopes, M.; Gatfield, D. Ribosome profiling reveals the rhythmic liver translatome and circadian clock regulation by upstream open reading frames. Genome Res. 2015, 25, 1848–1859. [Google Scholar] [CrossRef]
- Dierickx, P.; Van Laake, L.W.; Geijsen, N. Circadian clocks: From stem cells to tissue homeostasis and regeneration. EMBO Rep. 2018, 19, 18–28. [Google Scholar] [CrossRef]
- Janich, P.; Toufighi, K.; Solanas, G.; Luis, N.M.; Minkwitz, S.; Serrano, L.; Lehner, B.; Benitah, S.A. Human epidermal stem cell function is regulated by circadian oscillations. Cell Stem Cell 2013, 13, 745–753. [Google Scholar] [CrossRef]
- Weger, M.; Diotel, N.; Dorsemans, A.C.; Dickmeis, T.; Weger, B.D. Stem cells and the circadian clock. Dev. Biol. 2017, 431, 111–123. [Google Scholar] [CrossRef] [PubMed]
- Guydosh, N.R.; Green, R. Dom34 rescues ribosomes in 3′ untranslated regions. Cell 2014, 156, 950–962. [Google Scholar] [CrossRef] [PubMed]
- Tsuboi, T.; Kuroha, K.; Kudo, K.; Makino, S.; Inoue, E.; Kashima, I.; Inada, T. Dom34:hbs1 plays a general role in quality-control systems by dissociation of a stalled ribosome at the 3′ end of aberrant mRNA. Mol Cell 2012, 46, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Yang, F.; Xuan, Y.; Xi, R.; Zhao, R. Pelota-interacting G protein Hbs1 is required for spermatogenesis in Drosophila. Sci. Rep. 2019, 9, 3226. [Google Scholar] [CrossRef] [PubMed]
- Raju, P.; Nyamsuren, G.; Elkenani, M.; Kata, A.; Tsagaan, E.; Engel, W.; Adham, I.M. Pelota mediates gonocyte maturation and maintenance of spermatogonial stem cells in mouse testes. Reproduction 2015, 149, 213–221. [Google Scholar] [CrossRef]
- Xi, R.; Doan, C.; Liu, D.; Xie, T. Pelota controls self-renewal of germline stem cells by repressing a Bam-independent differentiation pathway. Development 2005, 132, 5365–5374. [Google Scholar] [CrossRef]
- Nyamsuren, G.; Kata, A.; Xu, X.; Raju, P.; Dressel, R.; Engel, W.; Pantakani, D.V.; Adham, I.M. Pelota regulates the development of extraembryonic endoderm through activation of bone morphogenetic protein (BMP) signaling. Stem Cell Res 2014, 13, 61–74. [Google Scholar] [CrossRef]
- Bortoluzzi, S.; d’Alessi, F.; Romualdi, C.; Danieli, G.A. Differential expression of genes coding for ribosomal proteins in different human tissues. Bioinformatics 2001, 17, 1152–1157. [Google Scholar] [CrossRef]
- Guimaraes, J.C.; Zavolan, M. Patterns of ribosomal protein expression specify normal and malignant human cells. Genome Biol. 2016, 17, 236. [Google Scholar] [CrossRef] [PubMed]
- Barakat, A.; Szick-Miranda, K.; Chang, I.F.; Guyot, R.; Blanc, G.; Cooke, R.; Delseny, M.; Bailey-Serres, J. The organization of cytoplasmic ribosomal protein genes in the Arabidopsis genome. Plant Physiol. 2001, 127, 398–415. [Google Scholar] [CrossRef]
- Komili, S.; Farny, N.G.; Roth, F.P.; Silver, P.A. Functional specificity among ribosomal proteins regulates gene expression. Cell 2007, 131, 557–571. [Google Scholar] [CrossRef] [PubMed]
- Balasubramanian, S.; Zheng, D.; Liu, Y.J.; Fang, G.; Frankish, A.; Carriero, N.; Robilotto, R.; Cayting, P.; Gerstein, M. Comparative analysis of processed ribosomal protein pseudogenes in four mammalian genomes. Genome Biol. 2009, 10, R2. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, Y.; Honda, H.; Iida, T.; Morinaga, T.; Hino, S.; Okajima, T.; Matsuda, T.; Nadano, D. Proteomic analysis of rodent ribosomes revealed heterogeneity including ribosomal proteins L10-like, L22-like 1, and L39-like. J. Proteome Res. 2010, 9, 1351–1366. [Google Scholar] [CrossRef] [PubMed]
- Jiang, L.; Li, T.; Zhang, X.; Zhang, B.; Yu, C.; Li, Y.; Fan, S.; Jiang, X.; Khan, T.; Hao, Q.; et al. RPL10L Is Required for Male Meiotic Division by Compensating for RPL10 during Meiotic Sex Chromosome Inactivation in Mice. Curr. Biol. 2017, 27, 1498–1505. [Google Scholar] [CrossRef] [PubMed]
- Lopes, A.M.; Miguel, R.N.; Sargent, C.A.; Ellis, P.J.; Amorim, A.; Affara, N.A. The human RPS4 paralogue on Yq11.223 encodes a structurally conserved ribosomal protein and is preferentially expressed during spermatogenesis. BMC Mol. Biol. 2010, 11, 33. [Google Scholar] [CrossRef] [PubMed]
- Sugihara, Y.; Sadohara, E.; Yonezawa, K.; Kugo, M.; Oshima, K.; Matsuda, T.; Nadano, D. Identification and expression of an autosomal paralogue of ribosomal protein S4, X-linked, in mice: Potential involvement of testis-specific ribosomal proteins in translation and spermatogenesis. Gene 2013, 521, 91–99. [Google Scholar] [CrossRef] [PubMed]
- Wong, Q.W.; Li, J.; Ng, S.R.; Lim, S.G.; Yang, H.; Vardy, L.A. RPL39L is an example of a recently evolved ribosomal protein paralog that shows highly specific tissue expression patterns and is upregulated in ESCs and HCC tumors. RNA Biol. 2014, 11, 33–41. [Google Scholar] [CrossRef]
- Kondrashov, N.; Pusic, A.; Stumpf, C.R.; Shimizu, K.; Hsieh, A.C.; Ishijima, J.; Shiroishi, T.; Barna, M. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 2011, 145, 383–397. [Google Scholar] [CrossRef]
- Zhang, Y.; Duc, A.C.; Rao, S.; Sun, X.L.; Bilbee, A.N.; Rhodes, M.; Li, Q.; Kappes, D.J.; Rhodes, J.; Wiest, D.L. Control of hematopoietic stem cell emergence by antagonistic functions of ribosomal protein paralogs. Dev. Cell 2013, 24, 411–425. [Google Scholar] [CrossRef]
- Zhang, Y.; O’Leary, M.N.; Peri, S.; Wang, M.; Zha, J.; Melov, S.; Kappes, D.J.; Feng, Q.; Rhodes, J.; Amieux, P.S.; et al. Ribosomal Proteins Rpl22 and Rpl22l1 Control Morphogenesis by Regulating Pre-mRNA Splicing. Cell Rep. 2017, 18, 545–556. [Google Scholar] [CrossRef]
- Incarnato, D.; Anselmi, F.; Morandi, E.; Neri, F.; Maldotti, M.; Rapelli, S.; Parlato, C.; Basile, G.; Oliviero, S. High-throughput single-base resolution mapping of RNA 2-O-methylated residues. Nucleic Acids Res. 2017, 45, 1433–1441. [Google Scholar] [CrossRef] [PubMed]
- Marchand, V.; Blanloeil-Oillo, F.; Helm, M.; Motorin, Y. Illumina-based RiboMethSeq approach for mapping of 2′-O-Me residues in RNA. Nucleic Acids Res. 2016, 44, e135. [Google Scholar] [CrossRef] [PubMed]
- Sloan, K.E.; Warda, A.S.; Sharma, S.; Entian, K.D.; Lafontaine, D.L.J.; Bohnsack, M.T. Tuning the ribosome: The influence of rRNA modification on eukaryotic ribosome biogenesis and function. RNA Biol. 2017, 14, 1138–1152. [Google Scholar] [CrossRef] [PubMed]
- Erales, J.; Marchand, V.; Panthu, B.; Gillot, S.; Belin, S.; Ghayad, S.E.; Garcia, M.; Laforets, F.; Marcel, V.; Baudin-Baillieu, A.; et al. Evidence for rRNA 2′-O-methylation plasticity: Control of intrinsic translational capabilities of human ribosomes. Proc. Natl. Acad. Sci. USA 2017, 114, 12934–12939. [Google Scholar] [CrossRef]
- Marcel, V.; Catez, F.; Diaz, J.J. Ribosome heterogeneity in tumorigenesis: The rRNA point of view. Mol. Cell Oncol. 2015, 2, e983755. [Google Scholar] [CrossRef]
- Hebras, J.; Krogh, N.; Marty, V.; Nielsen, H.; Cavaille, J. Developmental changes of rRNA ribose methylations in the mouse. RNA Biol. 2020, 17, 150–164. [Google Scholar] [CrossRef]
- Castle, J.C.; Armour, C.D.; Lower, M.; Haynor, D.; Biery, M.; Bouzek, H.; Chen, R.; Jackson, S.; Johnson, J.M.; Rohl, C.A.; et al. Digital genome-wide ncRNA expression, including SnoRNAs, across 11 human tissues using polyA-neutral amplification. PLoS ONE 2010, 5, e11779. [Google Scholar] [CrossRef]
- Ferretti, M.B.; Ghalei, H.; Ward, E.A.; Potts, E.L.; Karbstein, K. Rps26 directs mRNA-specific translation by recognition of Kozak sequence elements. Nat. Struct. Mol. Biol. 2017, 24, 700–707. [Google Scholar] [CrossRef]
- Slavov, N.; Semrau, S.; Airoldi, E.; Budnik, B.; van Oudenaarden, A. Differential Stoichiometry among Core Ribosomal Proteins. Cell Rep. 2015, 13, 865–873. [Google Scholar] [CrossRef]
- Shi, Z.; Fujii, K.; Kovary, K.M.; Genuth, N.R.; Rost, H.L.; Teruel, M.N.; Barna, M. Heterogeneous Ribosomes Preferentially Translate Distinct Subpools of mRNAs Genome-wide. Mol. Cell 2017, 67, 71–83. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gabut, M.; Bourdelais, F.; Durand, S. Ribosome and Translational Control in Stem Cells. Cells 2020, 9, 497. https://doi.org/10.3390/cells9020497
Gabut M, Bourdelais F, Durand S. Ribosome and Translational Control in Stem Cells. Cells. 2020; 9(2):497. https://doi.org/10.3390/cells9020497
Chicago/Turabian StyleGabut, Mathieu, Fleur Bourdelais, and Sébastien Durand. 2020. "Ribosome and Translational Control in Stem Cells" Cells 9, no. 2: 497. https://doi.org/10.3390/cells9020497
APA StyleGabut, M., Bourdelais, F., & Durand, S. (2020). Ribosome and Translational Control in Stem Cells. Cells, 9(2), 497. https://doi.org/10.3390/cells9020497




