Extracellular Matrix Features Discriminate Aggressive HER2-Positive Breast Cancer Patients Who Benefit from Trastuzumab Treatment
Abstract
:1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Immunohistochemistry and Fluorescence In Situ Hybridization
2.3. RNA Isolation and cDNA Microarray Techniques
2.4. Data Analysis
2.5. External Datasets and Signatures
2.6. In Vivo Study
2.7. Statistical Analysis
3. Results
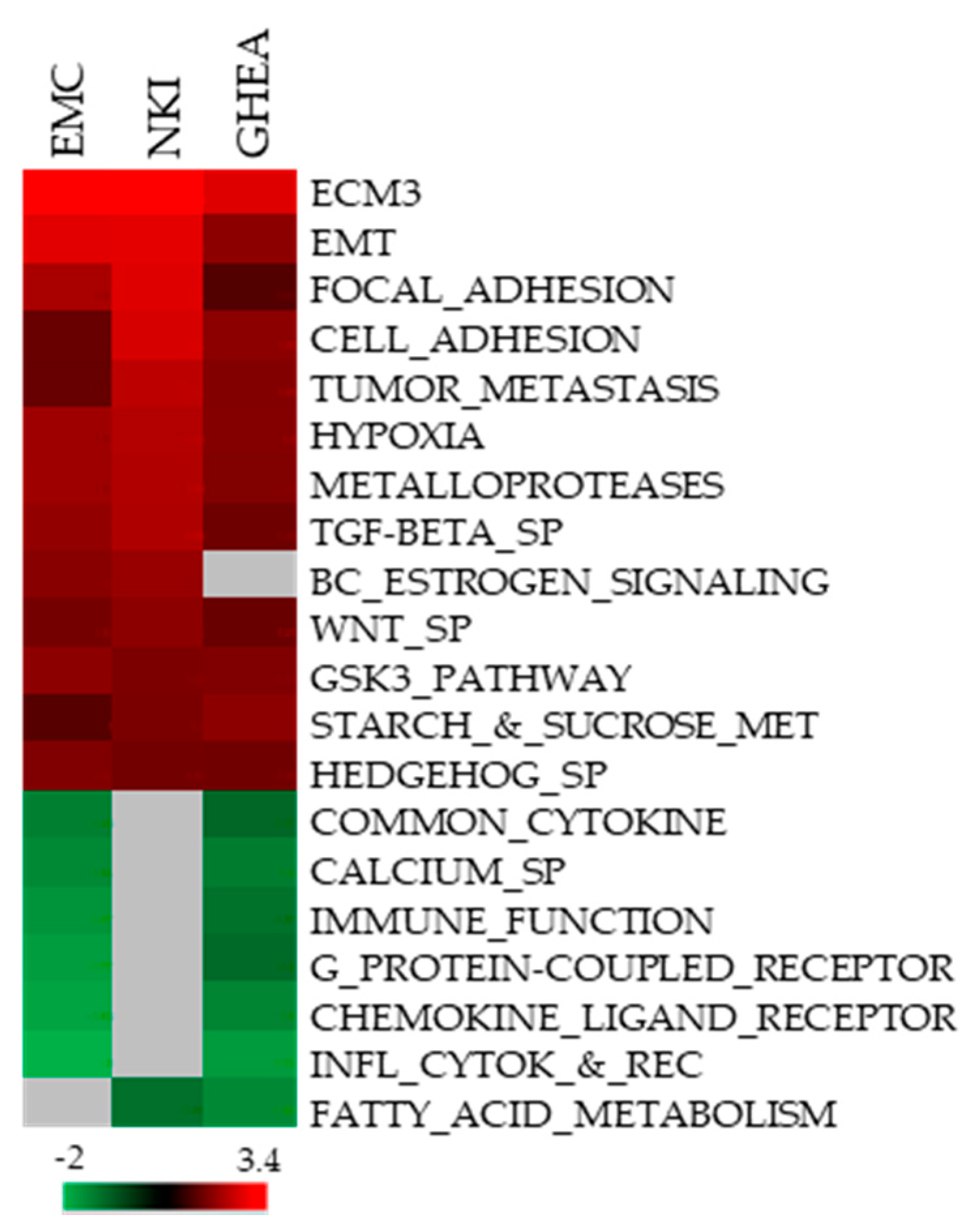
3.1. Identification of ECM3 HER2-Positive Breast Carcinomas
3.2. ECM3 in HER2-Positive BCs Not Treated with Trastuzumab
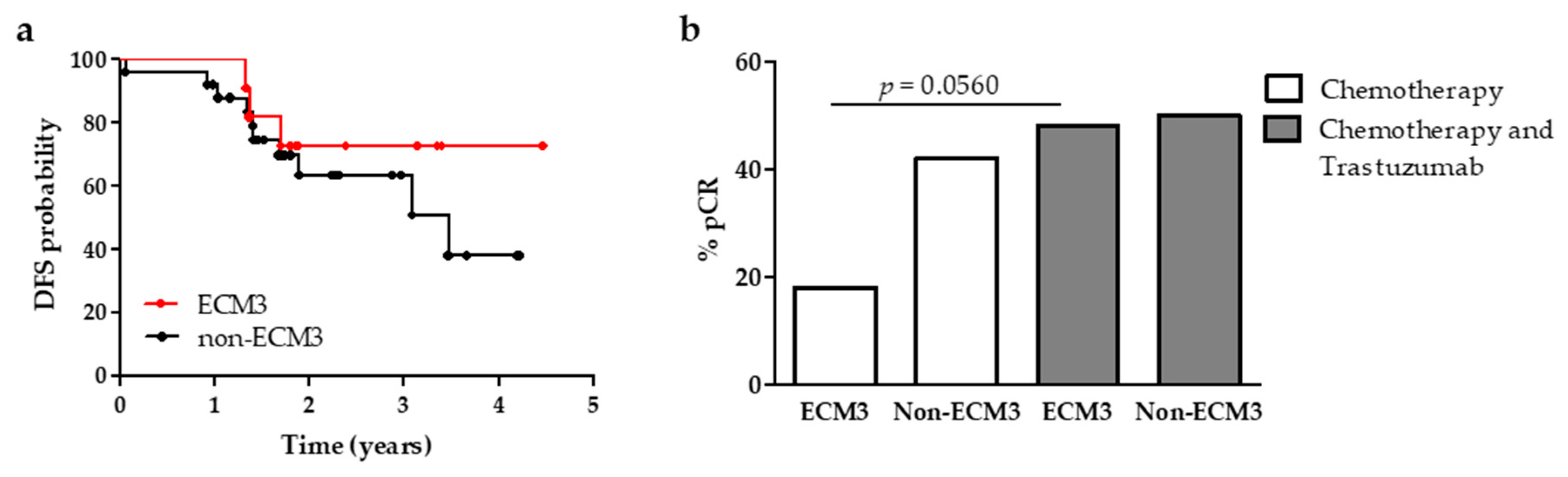
3.3. ECM3 in HER2-Positive BC Treated with Trastuzumab
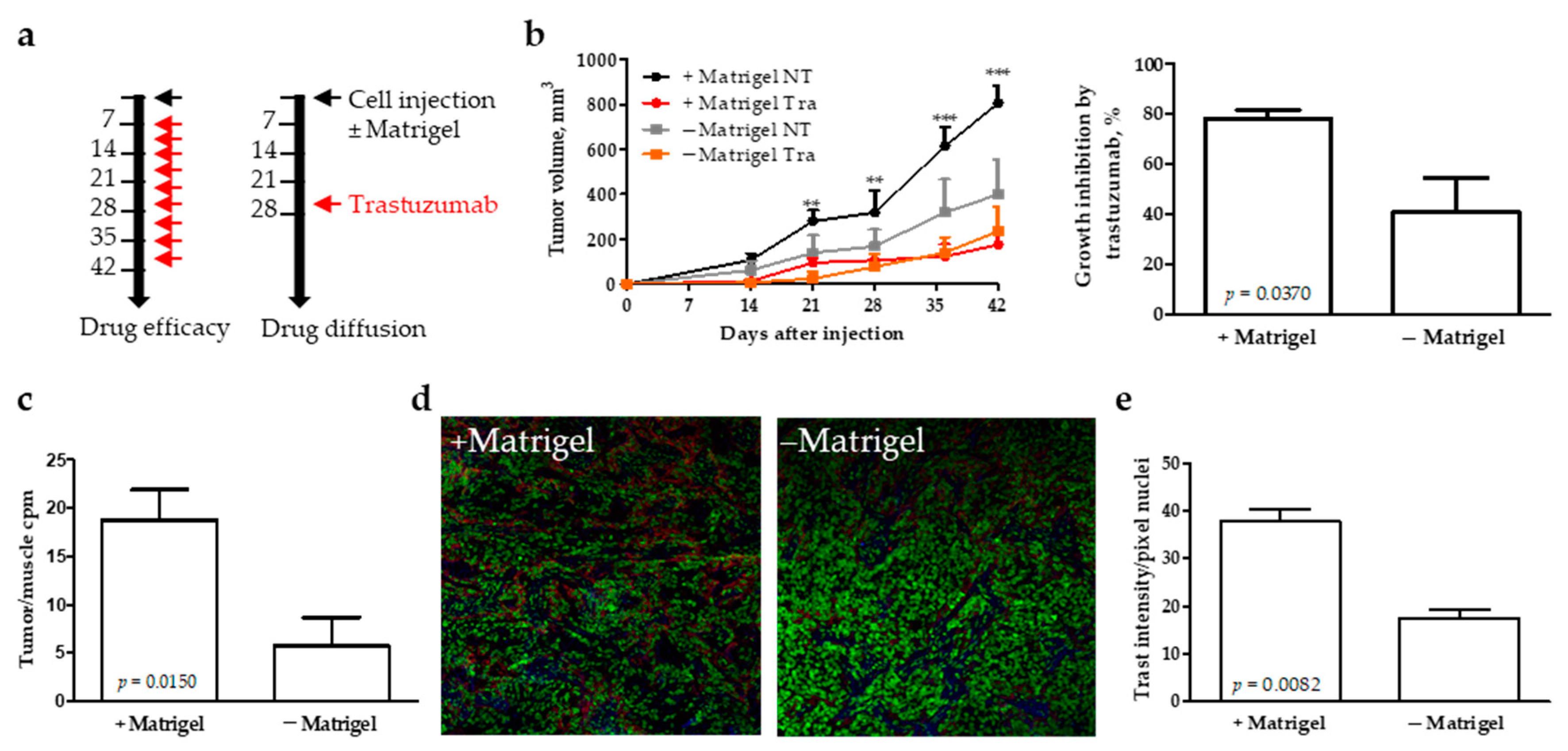
3.4. Extracellular Matrix and Response to Trastuzumab in a Xenograft Model
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Egeblad, M.; Nakasone, E.S.; Werb, Z. Tumors as Organs: Complex Tissues that Interface with the Entire Organism. Dev. Cell 2010, 18, 884–901. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Giussani, M.; Merlino, G.; Cappelletti, V.; Tagliabue, E.; Daidone, M.G. Tumor-extracellular matrix interactions: Identification of tools associated with breast cancer progression. Semin. Cancer Boil. 2015, 35, 3–10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bergamaschi, A.; Tagliabue, E.; Sorlie, T.; Naume, B.; Triulzi, T.; Orlandi, R.; Tammi, R.; Giercksky, H.; Nesland, J.M.; Ménard, S.; et al. Extracellular matrix signature identifies breast cancer subgroups with different clinical outcome. J. Pathol. 2008, 214, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Triulzi, T.; Casalini, P.; Sandri, M.; Ratti, M.; Carcangiu, M.L.; Colombo, M.P.; Balsari, A.; Ménard, S.; Orlandi, R.; Tagliabue, E. Neoplastic and Stromal Cells Contribute to an Extracellular Matrix Gene Expression Profile Defining a Breast Cancer Subtype Likely to Progress. PLoS ONE 2013, 8, e56761. [Google Scholar] [CrossRef] [PubMed]
- Sangaletti, S.; Tripodo, C.; Santangelo, A.; Castioni, N.; Portararo, P.; Gulino, A.; Botti, L.; Parenza, M.; Cappetti, B.; Orlandi, R.; et al. Mesenchymal transition of high-grade breast carcinomas depends on extracellular matrix control of myeloid suppressor cell activity. Cell Rep. 2016, 17, 233–248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- DeSantis, C.E.; Ma, J.; Gaudet, M.M.; Newman, L.A.; Miller, K.D.; Sauer, A.G.; Jemal, A.; Siegel, R.L. Breast cancer statistics, 2019. CA Cancer J. Clin. 2019, 69, 438–451. [Google Scholar] [CrossRef] [PubMed]
- Arteaga, C.L.; Sliwkowski, M.X.; Osborne, C.K.; Perez, E.A.; Puglisi, F.; Gianni, L. Treatment of HER2-positive breast cancer: Current status and future perspectives. Nat. Rev. Clin. Oncol. 2012, 9, 16–32. [Google Scholar] [CrossRef]
- Ross, J.S.; Fletcher, J.A. The HER-2/neu Oncogene in Breast Cancer: Prognostic Factor, Predictive Factor, and Target for Therapy. Stem Cells 1998, 16, 413–428. [Google Scholar] [CrossRef]
- O’Sullivan, C.C.; Bradbury, I.; Campbell, C.; Spielmann, M.; Perez, E.A.; Joensuu, H.; Costantino, J.P.; Delaloge, S.; Rastogi, P.; Zardavas, D.; et al. Efficacy of Adjuvant Trastuzumab for Patients With Human Epidermal Growth Factor Receptor 2-Positive Early Breast Cancer and Tumors Gen 2 cm: A Meta-Analysis of the Randomized Trastuzumab Trials. J. Clin. Oncol. 2015, 33, 2600–2608. [Google Scholar] [CrossRef]
- Triulzi, T.; Bianchi, G.V.; Tagliabue, E. Predictive biomarkers in the treatment of HER2-positive breast cancer: An ongoing challenge. Future Oncol. 2016, 12, 1413–1428. [Google Scholar] [CrossRef]
- Prat, A.; Carey, L.A.; Adamo, B.; Vidal, M.; Tabernero, J.; Cortés, J.; Parker, J.S.; Perou, C.M.; Baselga, J. Molecular features and survival outcomes of the intrinsic subtypes within HER2-positive breast cancer. J. Natl. Cancer Inst. 2014, 106, 152. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Cosimo, S.; Triulzi, T.; Pizzamiglio, S.; De Cecco, L.; De Azambuja, E.; Fumagalli, D.; Putzai, L.; Harbeck, N.; Izquierdo, M.; De La Peña, L.; et al. The 41-gene classifier TRAR predicts response of HER2 positive breast cancer patients in the NeoALTTO study. Eur. J. Cancer 2019, 118, 1–9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Di Modica, M.; Tagliabue, E.; Triulzi, T. Predicting the Efficacy of HER2-Targeted Therapies: A Look at the Host. Dis. Markers 2017, 2017, 1–14. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Velaei, K.; Samadi, N.; Barazvan, B.; Rad, J.S. Tumor microenvironment-mediated chemoresistance in breast cancer. Breast 2016, 30, 92–100. [Google Scholar] [CrossRef]
- Castiglioni, F.; Terenziani, M.; Carcangiu, M.L.; Miliano, R.; Aiello, P.; Bertola, L.; Triulzi, T.; Gasparini, P.; Camerini, T.; Sozzi, G.; et al. Radiation Effects on Development of HER2-Positive Breast Carcinomas. Clin. Cancer Res. 2007, 13, 46–51. [Google Scholar] [CrossRef] [Green Version]
- Dysvik, B.; Jonassen, I. J-Express: Exploring gene expression data using Java. Bioinformatics 2001, 17, 369–370. [Google Scholar] [CrossRef]
- E Canman, C.; Wolff, A.C.; Chen, C.Y.; Fornace, A.J.; Kastan, M.B. The p53-dependent G1 cell cycle checkpoint pathway and ataxia-telangiectasia. Cancer Res. 1994, 54, 5054–5058. [Google Scholar]
- Gentleman, R.C.; Carey, V.J.; Bates, D.M.; Bolstad, B.; Dettling, M.; Dudoit, S.; Ellis, B.; Gautier, L.; Ge, Y.; Gentry, J.; et al. Bioconductor: Open software development for computational biology and bioinformatics. Genome Boil. 2004, 5, R80. [Google Scholar] [CrossRef] [Green Version]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef] [Green Version]
- Van’t Veer, L.J.; Dai, H.; Van De Vijver, M.J.; He, Y.D.; Hart, A.A.; Mao, M.; Peterse, H.L.; Van Der Kooy, K.; Marton, M.J.; Witteveen, A.T.; et al. Gene expression profiling predicts clinical outcome of breast cance. Nature 2002, 415, 530–536. [Google Scholar] [CrossRef] [Green Version]
- Gluck, S.; Ross, J.S.; Royce, M.; McKenna, E.F., Jr.; Perou, C.M.; Avisar, E.; Wu, L. TP53 genomics predict higher clinical and pathologic tumor response in operable early-stage breast cancer treated with docetaxel-capecitabine +/− trastuzumab. Breast Cancer Res. Treat. 2012, 132, 781–791. [Google Scholar] [CrossRef] [PubMed]
- Prat, A.; Bianchini, G.; Thomas, M.; Belousov, A.; Cheang, M.C.U.; Koehler, A.; Gomez, P.; Semiglazov, V.; Eiermann, W.; Tjulandin, S.; et al. Research-Based PAM50 Subtype Predictor Identifies Higher Responses and Improved Survival Outcomes in HER2-Positive Breast Cancer in the NOAH Study. Clin. Cancer Res. 2014, 20, 511–521. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Triulzi, T.; De Cecco, L.; Sandri, M.; Prat, A.; Giussani, M.; Paolini, B.; Carcangiu, M.L.; Canevari, S.; Bottini, A.; Balsari, A.; et al. Whole-transcriptome analysis links trastuzumab sensitivity of breast tumors to both HER2 dependence and immune cell nfiltration. Oncotarget 2015, 6, 28173–28182. [Google Scholar] [CrossRef] [PubMed]
- Staaf, J.; Ringnér, M.; Vallon-Christersson, J.; Jonsson, G.; Bendahl, P.-O.; Holm, K.; Arason, A.; Gunnarsson, H.; Hegardt, C.; Agnarsson, B.A.; et al. Identification of Subtypes in Human Epidermal Growth Factor Receptor 2–Positive Breast Cancer Reveals a Gene Signature Prognostic of Outcome. J. Clin. Oncol. 2010, 28, 1813–1820. [Google Scholar] [CrossRef] [PubMed]
- Shabalin, A.A.; Weighman, V.J.; Perou, C.M.; Nobel, A.B. Finding Large Average Submatricies in High Dimensional Data. Ann. Appl. Stat. 2009, 3, 985–1012. [Google Scholar] [CrossRef]
- Brock, G.; Pihur, V.; Datta, S. clValid: An R Package for Cluster Validation. J. Stat. Softw. 2008, 25, 25. [Google Scholar] [CrossRef] [Green Version]
- Rody, A.; Holtrich, U.; Pusztai, L.; Liedtke, C.; Gaetje, R.; Ruckhaeberle, E.; Solbach, C.; Hanker, L.; Ahr, A.; Metzler, D.; et al. T-cell metagene predicts a favorable prognosis in estrogen receptor-negative and HER2-positive breast cancers. Breast Cancer Res. 2009, 11, R15. [Google Scholar] [CrossRef] [Green Version]
- Bianchini, G.; Pusztai, L.; Pienkowski, T.; Im, Y.-H.; Bianchi, G.V.; Tseng, L.-M.; Liu, M.-C.; Lluch, A.; Galeota, E.; Magazzù, D.; et al. Immune modulation of pathologic complete response after neoadjuvant HER2-directed therapies in the NeoSphere trial. Ann. Oncol. 2015, 26, 2429–2436. [Google Scholar] [CrossRef]
- Guo, S.; Deng, C.-X. Effect of Stromal Cells in Tumor Microenvironment on Metastasis Initiation. Int. J. Boil. Sci. 2018, 14, 2083–2093. [Google Scholar] [CrossRef]
- Takebe, N.; Miele, L.; Harris, P.J.; Jeong, W.; Bando, H.; Kahn, M.; Yang, S.X.; Ivy, S.P. Targeting Notch, Hedgehog, and Wnt pathways in cancer stem cells: Clinical update. Nat. Rev. Clin. Oncol. 2015, 12, 445–464. [Google Scholar] [CrossRef]
- Kirkland, S.C. Type I collagen inhibits differentiation and promotes a stem cell-like phenotype in human colorectal carcinoma cells. Br. J. Cancer 2009, 101, 320–326. [Google Scholar] [CrossRef] [Green Version]
- Gupta, P.B.; Onder, T.T.; Jiang, G.; Tao, K.; Kuperwasser, C.; Weinberg, R.A.; Lander, E.S. Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell 2009, 138, 645–659. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dittmer, J.; Leyh, B. The impact of tumor stroma on drug response in breast cancer. Semin. Cancer Boil. 2015, 31, 3–15. [Google Scholar] [CrossRef]
- Tannock, I.F.; Lee, C.M.; Tunggal, J.K.; Cowan, D.S.M.; Egorin, M.J. Limited penetration of anticancer drugs through tumor tissue: A potential cause of resistance of solid tumors to chemotherapy. Clin. Cancer Res. 2002, 8, 878–884. [Google Scholar]
- Jansen, M.P.; Foekens, J.A.; Van Staveren, I.L.; Dirkzwager-Kiel, M.M.; Ritstier, K.; Look, M.P.; Gelder, M.E.M.-V.; Sieuwerts, A.M.; Portengen, H.; Dorssers, L.C.; et al. Molecular Classification of Tamoxifen-Resistant Breast Carcinomas by Gene Expression Profiling. J. Clin. Oncol. 2005, 23, 732–740. [Google Scholar] [CrossRef]
- Farmer, P.; Bonnefoi, H.; Anderle, P.; Cameron, D.; Wirapati, P.; Becette, V.; André, S.; Piccart, M.; Campone, M.; Brain, E.; et al. A stroma-related gene signature predicts resistance to neoadjuvant chemotherapy in breast cancer. Nat. Med. 2009, 15, 68–74. [Google Scholar] [CrossRef]
- Azim, H.A., Jr.; Singhal, S.; Ignatiadis, M.; Desmedt, C.; Fumagalli, D.; Veys, I.; Larsimont, D.; Piccart, M.; Michiels, S.; Sotiriou, C. Association between SPARC mRNA expression, prognosis and response to neoadjuvant chemotherapy in early breast cancer: A pooled in-silico analysis. PLoS ONE 2013, 8, e62451. [Google Scholar] [CrossRef]
- Sui, H.; Zhu, L.; Deng, W.; Li, Q. Epithelial-Mesenchymal Transition and Drug Resistance: Role, Molecular Mechanisms, and Therapeutic Strategies. Oncol. Res. Treat. 2014, 37, 9. [Google Scholar] [CrossRef]
- Khawar, I.A.; Kim, J.H.; Kuh, H.-J. Improving drug delivery to solid tumors: Priming the tumor microenvironment. J. Control. Release 2015, 201, 78–89. [Google Scholar] [CrossRef]
- Mohammad, N.; Singh, S.V.; Malvi, P.; Chaube, B.; Athavale, D.; Vanuopadath, M.; Nair, S.S.; Nair, B.; Bhat, M.K. Strategy to enhance efficacy of doxorubicin in solid tumor cells by methyl-β-cyclodextrin: Involvement of p53 and Fas receptor ligand complex. Sci. Rep. 2015, 5, 11853. [Google Scholar] [CrossRef] [Green Version]
- García-Mendoza, M.G.; Inman, D.R.; Ponik, S.M.; Jeffery, J.J.; Sheerar, D.S.; Van Doorn, R.R.; Keely, P.J. Neutrophils drive accelerated tumor progression in the collagen-dense mammary tumor microenvironment. Breast Cancer Res. 2016, 18, 49. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Magnifico, A.; Albano, L.; Campaner, S.; Delia, D.; Castiglioni, F.; Gasparini, P.; Sozzi, G.; Fontanella, E.; Menard, S.; Tagliabue, E. Tumor-Initiating Cells of HER2-Positive Carcinoma Cell Lines Express the Highest Oncoprotein Levels and Are Sensitive to Trastuzumab. Clin. Cancer Res. 2009, 15, 2010–2021. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Minchinton, A.I.; Tannock, I.F. Drug penetration in solid tumours. Nat. Rev. Cancer 2006, 6, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Baker, J.H.; Lindquist, K.E.; Huxham, L.A.; Kyle, A.H.; Sy, J.T.; Minchinton, A.I. Direct Visualization of Heterogeneous Extravascular Distribution of Trastuzumab in Human Epidermal Growth Factor Receptor Type 2 Overexpressing Xenografts. Clin. Cancer Res. 2008, 14, 2171–2179. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Triulzi, T.; Ratti, M.; Tortoreto, M.; Ghirelli, C.; Aiello, P.; Regondi, V.; Modica, M.D.; Cominetti, D.; Carcangiu, M.L.; Moliterni, A.; et al. Maspin influences response to doxorubicin by changing the tumor microenvironment organization. Int. J. Cancer 2014, 134, 2789–2797. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shah, C.; Miller, T.W.; Wyatt, S.K.; McKinley, E.T.; Olivares, M.G.; Sanchez, V.; Nolting, D.D.; Buck, J.R.; Zhao, P.; Ansari, M.S.; et al. Imaging biomarkers predict response to anti-HER2 (ErbB2) therapy in preclinical models of breast cancer. Clin. Cancer Res. 2009, 15, 4712–4721. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohsin, S.K.; Weiss, H.L.; Gutierrez, M.C.; Chamness, G.C.; Schiff, R.; DiGiovanna, M.P.; Wang, C.-X.; Hilsenbeck, S.G.; Osborne, C.K.; Allred, D.C.; et al. Neoadjuvant Trastuzumab Induces Apoptosis in Primary Breast Cancers. J. Clin. Oncol. 2005, 23, 2460–2468. [Google Scholar] [CrossRef]
- Sorace, A.G.; Quarles, C.C.; Whisenant, J.G.; Hanker, A.B.; McIntyre, J.O.; Sanchez, V.M.; Yankeelov, T.E. Trastuzumab improves tumor perfusion and vascular delivery of cytotoxic therapy in a murine model of HER2+ breast cancer: Preliminary results. Breast Cancer Res. Treat. 2016, 155, 273–284. [Google Scholar] [CrossRef] [Green Version]
- He, L.; Du, Z.; Xiong, X.; Ma, H.; Zhu, Z.; Gao, H.; Cao, J.; Li, T.; Li, H.; Yang, K.; et al. Targeting Androgen Receptor in Treating HER2 Positive Breast Cancer. Sci. Rep. 2017, 7, 14584. [Google Scholar] [CrossRef] [Green Version]
- I Koukourakis, M.; Giatromanolaki, A.; Bottini, A.; Cappelletti, M.R.; Zanotti, L.; Allevi, G.; Strina, C.; Ardine, M.; Milani, M.; Brugnoli, G.; et al. Prospective neoadjuvant analysis of PET imaging and mechanisms of resistance to Trastuzumab shows role of HIF1 and autophagy. Br. J. Cancer 2014, 110, 2209–2216. [Google Scholar] [CrossRef] [Green Version]


| EMC 29% | NKI 25% | FIRB 34% | GHEA 31% | NOAH 37% | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Variable | ECM3 | Non-ECM3 | ECM3 | Non-ECM3 | ECM3 | Non-ECM3 | ECM3 | Non-ECM3 | ECM3 | Non-ECM3 |
| (n = 14) | (n = 35) | (n = 13) | (n = 39) | (n = 15) | (n = 29) | (n = 11) | (n = 25) | (n = 42) | (n = 72) | |
| Estrogen receptor | ||||||||||
| Positive | 9 (64%) | 24 (69%) | 8 (62%) | 25 (64%) | 5 (33%) | 16 (55%) | 6 (55%) | 12 (48%) | 11 (26%) | 16 (22%) |
| Tumor size | ||||||||||
| >T1 | NA | NA | 7 (54%) | 20 (51%) | 13 (93%) | 23 (79%) | 5 (45%) | 12 (48%) | NA | NA |
| Histological grade | ||||||||||
| III | NA | NA | 6 (46%) | 24 (62%) | 9 (69%) | 19 (68%) | 8 (73%) | 20 (80%) | 21 (50%) | 40 (56%) |
| Lymph node status | ||||||||||
| Positive | 0 | 0 | 8 (62%) | 18 (46%) | 10 (71%) | 19 (66%) | 10 (91%) | 21 (84%) | NA | NA |
| EMC | NKI | FIRB | ||||
|---|---|---|---|---|---|---|
| Variable | HR (95%CI) | p-Value | HR (95%CI) | p-Value | HR (95%CI) | p-Value |
| ECM3 | 5.50 (2.07–14.62) | 0.0006 | 2.57 (1.06–6.19) | 0.0361 | 2.29 (0.76–6.87) | 0.1383 |
| ER pos | 0.58 (0.22–1.54) | 0.2730 | 0.46 (0.20–1.09) | 0.0781 | 1.09 (0.36–3.29) | 0.8800 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rybinska, I.; Sandri, M.; Bianchi, F.; Orlandi, R.; De Cecco, L.; Gasparini, P.; Campiglio, M.; Paolini, B.; Sfondrini, L.; Tagliabue, E.; et al. Extracellular Matrix Features Discriminate Aggressive HER2-Positive Breast Cancer Patients Who Benefit from Trastuzumab Treatment. Cells 2020, 9, 434. https://doi.org/10.3390/cells9020434
Rybinska I, Sandri M, Bianchi F, Orlandi R, De Cecco L, Gasparini P, Campiglio M, Paolini B, Sfondrini L, Tagliabue E, et al. Extracellular Matrix Features Discriminate Aggressive HER2-Positive Breast Cancer Patients Who Benefit from Trastuzumab Treatment. Cells. 2020; 9(2):434. https://doi.org/10.3390/cells9020434
Chicago/Turabian StyleRybinska, Ilona, Marco Sandri, Francesca Bianchi, Rosaria Orlandi, Loris De Cecco, Patrizia Gasparini, Manuela Campiglio, Biagio Paolini, Lucia Sfondrini, Elda Tagliabue, and et al. 2020. "Extracellular Matrix Features Discriminate Aggressive HER2-Positive Breast Cancer Patients Who Benefit from Trastuzumab Treatment" Cells 9, no. 2: 434. https://doi.org/10.3390/cells9020434
APA StyleRybinska, I., Sandri, M., Bianchi, F., Orlandi, R., De Cecco, L., Gasparini, P., Campiglio, M., Paolini, B., Sfondrini, L., Tagliabue, E., & Triulzi, T. (2020). Extracellular Matrix Features Discriminate Aggressive HER2-Positive Breast Cancer Patients Who Benefit from Trastuzumab Treatment. Cells, 9(2), 434. https://doi.org/10.3390/cells9020434










