1. Introduction
Omega-3 and omega-6 polyunsaturated fatty acids (PUFAs), are critical for diverse biological functions including membrane stability, cell signaling and metabolism, particularly in the brain and retina [
1,
2]. Synthesis of PUFAs begins with the dietary intake of essential amino acids, linoleic acid, and alpha linoleic acid, which then goes through a series of elongation and desaturation reactions to form longer chain omega-3 and omega-6 fatty acids, including arachidonic acid, eicosapentaenoic acid (EPA), and docosahexaenoic acid (DHA). DHA, the major polyunsaturated fatty acid in the brain and retina, is a critical component of photoreceptor outer segments necessary for photoreceptor function [
1,
3,
4] while very long chain PUFAs (VLC-PUFAs; greater than 22 carbon length fatty acids [
5]) are believed to be indispensable in maintaining the curvature of the photoreceptor disk membrane. PUFAs have been implicated in eye diseases, as a decrease of dietary intake of food rich in omega-3 fatty acids, such as fish, have been linked to higher risk of age related eye diseases such as macular degeneration in multiple epidemiologic studies [
4,
6,
7].
ELOVL2 (Elongation Of Very Long Chain Fatty Acids-Like 2) encodes an enzyme involved in the elongation of long-chain omega-3 and omega-6 polyunsaturated fatty acids (LC-PUFAs) [
8]. In particular, ELOVL2 in mammals elongates docosapentaenoic acid (DPA—22:5n-3) to 24:5n-3, which in turn is the substrate for the formation of VLC-PUFAs as well as 22:6n-3, i.e., DHA [
9]. Interestingly, DNA methylation of the
Elovl2 regulatory element has been well established as an epigenetic biomarker of aging, as multiple studies have shown that the DNA methylation of the
ELOVL2 promoter across multiple human tissues highly correlates with chronological age [
10,
11].
We have previously studied the role of
Elovl2 in the mammalian retina. We have implicated
Elovl2 as a critical molecular regulator of aging, as loss of
Elovl2 activity in mice accelerates anatomical and functional surrogates of aging in the retina. Additionally, we observed sub-RPE deposits, which contained multiple proteins found in human drusen, a pathologic hallmark of age related macular degeneration [
12]. Pharmacological demethylation of
Elovl2 can increase
Elovl2 gene expression and prevent the progression of the age-related decline of the electroretinogram response, a functional surrogate of aging [
12].
Zebrafish (
Danio rerio) represents an excellent model system to study retinal development and biology, given its small size, transparent embryo, and the availability of robust visual behavior assays [
13]. The role of
elovl2 in zebrafish, particularly within the eye, is still poorly understood. Previous studies have demonstrated that zebrafish
elovl2 can elongate C18-C22 PUFAs in a heterologous yeast system, in contrast to human ELOVL2 which has been shown to only elongate C20 and C22 PUFAs [
8,
14,
15]. In addition, zebrafish
elovl5, which in humans is specific to the elongation of C18 PUFAs, can also elongate C20 and C22 PUFAs to a lesser extent. In a recent study of
elovl2 and
elovl5 knockout zebrafish, it was observed that
elovl2, but not
elovl5, is required for the conversion of C20 EPA to DPA, and thus synthesis of DHA [
16]. Finally, zebrafish
elovl4 is also able to elongate LC-PUFAs in striking contrast to the mammalian ortholog [
17].
Muller glia are macro glia found in the retina whose processes span the entire retina and contact most of the neurons in the retina. They play diverse functions including providing trophic support, removing metabolic waste, as well as regulating ion and water homeostasis, and regulating immune and inflammatory responses [
18]. Whether Muller glia can synthesize PUFAs, and what the role of Muller glia derived PUFAs is, is still poorly understood.
Here, we present the investigation of the function of
elovl2 in the zebrafish eye. We characterized the expression of
elovl2 in embryonic and adult wildtype zebrafish retina. We created zebrafish lacking
elovl2 function through the introduction of biallelic mutations in
elovl2 using CRISPR-Cas9 technology, termed “crispants” [
19]. Using lipidomics, retinal morphology, and visual behavior we showed the unexpected expression pattern and role of
elovl2 in zebrafish eye.
2. Materials and Methods
2.1. Zebrafish Husbandry
The AB strain (Zebrafish International Research Center (ZIRC); Eugene, OR, USA) was used for all experiments. Zebrafish were maintained in the fish facility of the University of California, San Diego (UCSD) under a controlled 14/10 h light cycle between 27 and 29 degrees Celsius and fed with a standard brine shrimp diet twice daily with a recirculating aquarium system. Breeding and experimental procedures were approved by the Institute of Animal Care and Use Committee of the University of California, San Diego (S18067).
2.2. Zebrafish Crispant Creation
Potential gRNA sequences for zebrafish
elovl2 were searched for using the Chop-chop algorithm (
https://chopchop.cbu.uib.no). Two
elovl2 gRNA sequences were selected, GACAGCCTATTTGGAGAAAG in exon 2 and TTCCCAGGTAGATTGTTAGG in exon 3. The control gRNA used (TGAGTATTCGCATGCAACTA) does not target any known zebrafish nucleotide sequence.
GRNA oligonucleotides (Integrated DNA Technologies (IDT), Coraville Iowa) were synthesized by and were duplexed individually with tracrRNA (Integrated DNA Technologies (IDT) Coraville, IA, USA). The injection mix containing 250 ng/μL of gRNA duplex complex (both gRNAs), 500 ng/μL rCas9 protein (PNA Bio CP01-20 Thousand Oaks, CA, USA) and duplex buffer (IDT) was injected into one cell stage zebrafish embryos. The embryos were maintained in a 28 °C incubator. The level of DNA editing was determined through DNA Sanger sequencing (Genewiz La Jolla, CA, USA) and analysis using the ICE v2CRISPR Analysis Tool (Synthego (Redwood City, CA, USA), found at
https://www.synthego.com/products/bioinformatics/crispr-analysis) [
20] Sequencing primers for
elovl2 gRNA exon 2 were 5′ TTGAAGCTTGCAATCTGACTGT3′ and 5′ TGGAACGTTCTATTGAGTGTCG 3′. Sequencing primers for
elovl2 gRNA exon
3 were 5′ TTTGTTTGATGTCAGATACCCG3′ and 5′ ATGAGCACATGGACTGCTATTG3′.
2.3. Zebrafish In-Situ Hybridization
Whole mount RNA in situ hybridization was performed on fixed 1–3 dpf zebrafish embryos using previously described protocol [
21]. The stained embryos were imaged using a stereomicroscope (Zeiss STEMI 508 (Zeiss Oberkocken, Germany)). A 550 bp fragment of
elovl2 cDNA was PCR amplified and cloned into a TOPO vector (Invitrogen). (Primers 5′ F AGGCAGTCATTTAGGTGACACTATAGATGG and 3′ R CGTCGTGGACTAATACGACTCACTATAGAC). The plasmid was linearized and an in vitro transcription reaction with DIG labeling mix (Roche, cat. no. 1277073) was performed, as previously described [
21]. For staining of retinal sections, the RNAscope protocol was followed as per manufacturers protocol (ACD (Newark, CA, USA); elovl2 probe Cat. No. 550301). The optional step of target retrieval was excluded to allow better retention of the samples on the slides. The stained sections were flooded with ProlongTM Gold Antifade Mountant (Cat. No. P36930) and covered with coverslips (Fisher Scientific. Waltham, MA 12-545F, USA) before imaging.
2.4. Microscopic Imaging
Fluorescent imaging of the fluorescent in situ hybridization (RNAscope) as well as the retinal anatomy analysis were performed using the Keyence microscope (BZ-X710, Keyence, Itasca, IL, USA) using structured illumination and optical sectioning. Ten mm sections were imaged using the CFI PLAN APO 40x LAMBDA Lens (Nikon, Englewood, CO, USA) and analyzed using the BZX Analyzer Software v2.0 (Keyence, USA).
2.5. Zebrafish Visual Motor Response
Larvae at 6 dpf were placed in flat bottom clear 96 well plates (Corning CLS3370 Corning, NY, USA) and the assay was performed in DanioVision observation chamber (Noldus Information Technology, Wageningen, The Netherlands). The light program was designed based on a previously published protocol [
22]. The larvae were dark-adapted for three hours, followed by three cycles of light on/off program. In each light cycle, white light at 100% intensity was turned on for 5 min followed by a 30 min dark period. The activity of larvae was measured 30 s before and after light on/off using EthoVision XT software (Noldus Information Technology, Wageningen, The Netherlands). A 5 min five dark period was included in the light program to measure the activity of larvae 30 s before the first light was on. The activity data was exported from the software and analyzed using Microsoft Excel. Three rounds of experiments, each encompassing about 130 embryos were performed, with all data pooled for the analysis.
2.6. Zebrafish Retinal Anatomy Analysis
The embryos were fixed in 4% PFA for 72 h and transferred into 100% ethanol solution. The samples were submitted to HistoWiz (
https://home.histowiz.com) for sectioning and hematoxylin and eosin (H&E) staining for histological analysis. For uniformity, only sections close to nerve fibers were selected for the measurements. Measurements were taken at three different regions in each selected section: near optic nerve (middle), anterior to the optic nerve (anterior) and posterior to optic nerve (posterior). Measurements were recorded using Fiji ImageJ software version 1.52p (National Institutes of Health, Maryland, MA, USA).
2.7. Lipid Analysis
For lipidomics sample preparation, two and seven days and post fertilization (dpf) larvae were transferred in an Eppendorf tube and excess water was removed. The tubes were then placed in a dish containing dry ice and ethanol to flash freeze the larvae. Twelve embryos were placed in each tube, with three replicates per timepoint. Lipids were extracted using a modified version of the Bligh-Dyer method [
23]. Briefly, zebrafish embryos were homogenized in 1 mL PBS and shaken in a glass vial (VWR International, Radnor, PA, USA) with 1 mL methanol and 2 mL chloroform containing internal standards (
13C16-palmitic acid, d7-Cholesterol) for 30 s. The resulting mixture was vortexed for 15 s and centrifuged at 2400×
g for 6 min to induce phase separation. The organic (bottom) layer was retrieved using a Pasteur pipette, dried under a gentle stream of nitrogen, and reconstituted in 2:1 chloroform:methanol for LC/MS analysis.
Extracted lipids were resuspended in 200 μL of EtOH, incubated with 0.1 M KOH at room temperature for 24 h for saponification. The reaction was stopped by addition of 0.2 M HCl. Lipids were extracted as described above with d31-palmitic acid as internal standard.
Untargeted lipidomic analysis was performed on a Vanquish HPLC online with a Q-Exactive quadrupole-orbitrap mass spectrometer equipped with an electrospray ion source (Thermo Fisher Scientific, Milan, Italy). A Bio-Bond C4 column (Dikma, 5 μm, 4.6 mm × 50 mm) was used. Solvent A consisted of 95:5 water:methanol, Solvent B was 60:35:5 isopropanol:methanol:water. Solvents A and B contained 5 mM ammonium formate with 0.1% formic acid. The gradient was held at 0% B between 0 and 5 min, raised to 20% B at 5.1 min, increased linearly from 20% to 100% B between 5.1 and 55 min, held at 100% B between 55 min and 63 min, returned to 0% B at 63.1 min, and held at 0% B until 70 min. Flow rate was 0.1 mL/min from 0 to 5 min, 0.4 mL/min between 5.1 min and 55 min, and 0.5 mL/min between 55 min and 70 min. Data was acquired in negative ionization mode. Spray voltage was −2.5 kV. Sheath, auxiliary, and sweep gases were 53, 14, and 3 arbitrary units (a.u.), respectively. Capillary temperature was 275 °C. Data was collected in full MS/dd-MS2 (top 5). Full MS was acquired from 100 to 1500 m/z with a resolution of 70,000, AGC target of 1 × 106 and a maximum injection time of 100 ms. MS2 was acquired with resolution of 17,500, a fixed first mass of 50 m/z, AGC target of 1 × 105 and a maximum injection time of 200 ms. Stepped normalized collision energies were 20, 30, and 40%.
Targeted lipidomic analysis was performed on a Dionex Ultimate 3000 LC system (Thermo Fisher Scientific, Milan, Italy) coupled to a TSQ Quantiva mass spectrometer (Thermo Fisher Scientific, Milan, Italy). A XBridge C8 column (Waters, 5 μm, 4.6 mm × 50 mm) was used. The solvents and gradient were as described above. MS analyses were performed using electrospray ionization in negative mode, with spray voltages of −2.5 kV, ion transfer tube temperature of 325 °C, and vaporizer temperature of 200 °C. Pseudo multiple reaction monitoring (MRM) was performed to detect fatty acids.
Lipid identification was performed with LipidSearch (Thermo Fisher Scientific, Milan, Italy). Mass accuracy, chromatography and peak integration of all LipidSearch-identified lipids were verified with Skyline [
24]. Peak integration of targeted fatty acids was also performed with Skyline. Peak areas were used in data reporting, data was normalized using internal standards. The relative abundance of lipid classes were calculated by the percent relative area method with proper normalization using internal standard and considering the sum of all relative areas of the identified lipids.
Statistical analyses were conducted using Prism 7 (GraphPad Prism, La Jolla, CA, USA). All values are expressed as means ± SD (standard deviations). One-way ANOVA was performed to determine significant differences between different groups. Significant calls were made based on P values < 0.05 and the Fold Change (FC) >1.5.
4. Discussion
We have recently implicated ELOVL2 as a molecular regulator of aging in the mouse retina [
12]. In this work, we investigated the expression and function of
elovl2 in the embryonic zebrafish retina. We found that
elovl2 is strongly expressed in the zebrafish retina during early embryogenesis as well as in the adult retina, with expression primarily confined to Muller glia. Analysis of
elovl2 crispants results in changes in the lipid levels in the zebrafish embryo, in particular in lipids containing polyunsaturated fatty acids longer than 22 carbons. Analysis of
elovl2 crispants demonstrates no gross changes in retinal morphology, but significant changes in the VMR-OFF response further underlying importance of the enzyme in a healthy vision.
The
elovl2 enzyme is strongly conserved between zebrafish and other species. Interestingly, while work has shown that zebrafish
elovl2, elongates C20 and C22 carbons, it also has some elongation activity for C18 fatty acids, which is different from mammalian
Elovl2 [
11,
12]. This is likely due to some overlap of functions between
elovl2 and
elovl5 in zebrafish (
Figure 2A), however recent work suggests that
elovl2 is the primary elongase required for the synthesis of DHA [
16]. Interestingly, the zebrafish
elovl4 enzyme is also able to elongate C20 and C22 fatty acids in addition to the activity towards VLC-PUFAs.
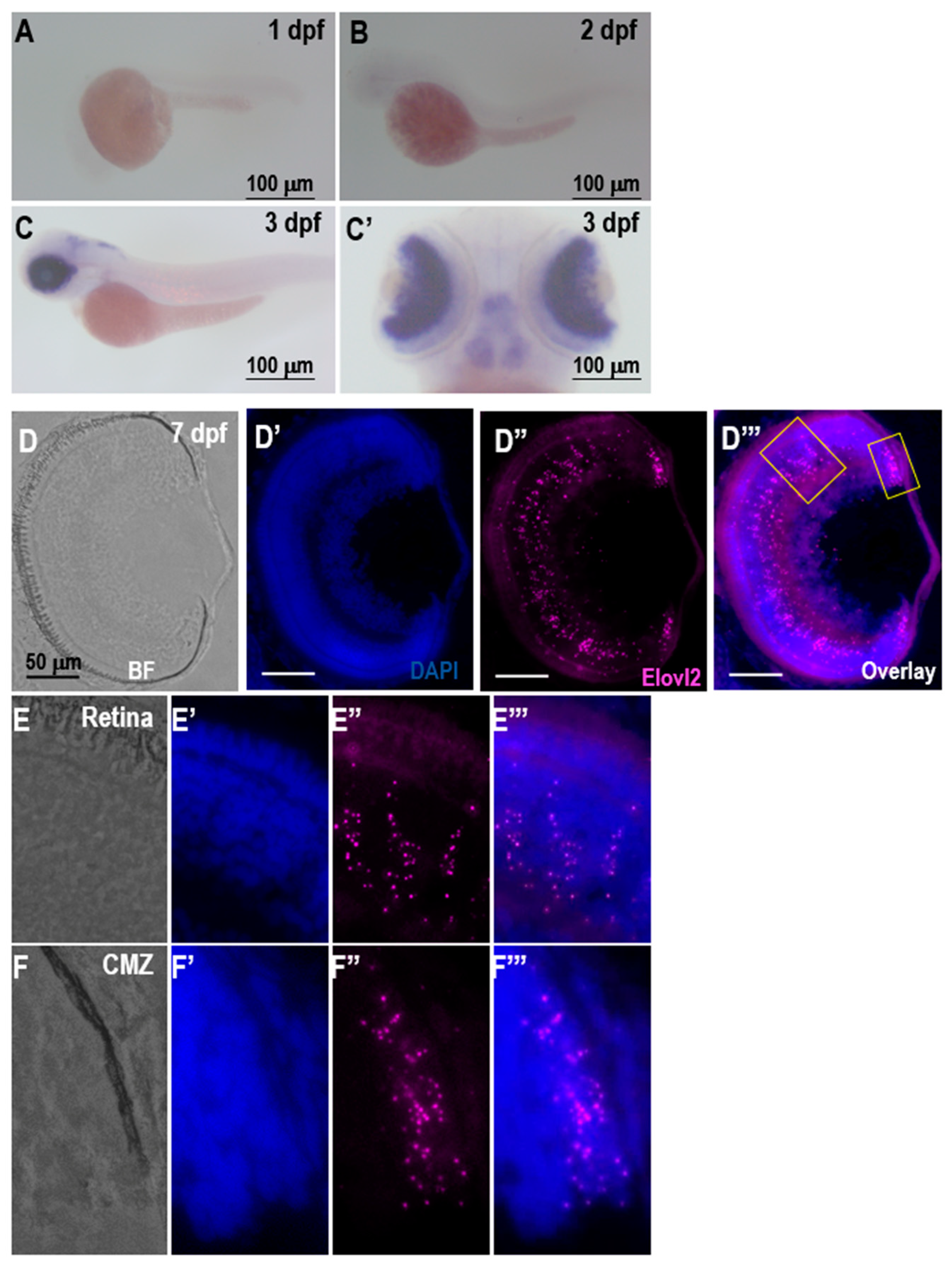
When investigating the expression of the
elovl2, we observed a strong retinal expression at 3 dpf. On histological sections at 7 days, clear RNAscope signal can be observed throughout the retina. In situ hybridization of retina sections in adult zebrafish showed a similar pattern of the expression, with no observable signal on negative control slides (
Supplementary Figure S8). A lower expression of
elovl2 in old zebrafish compared to young adult zebrafish was observed, suggesting that there may also be an age dependent decrease in
elovl2 expression that has been seen in human and mouse tissues [
10,
12]. Several groups analyzed expression patterns of
elovl2, 4, and
5 in zebrafish retina [
15,
17,
32]. Our data is in general agreement with published data, although one of the groups detected the first expression of Elovl2 earlier than at 3 dpf [
32]. Intriguingly, none of the studies convincingly presented the expression of
elovl5 in the eye while
elovl4 seems to be expressed mostly in retinal pigmented epithelium cells in striking contrast to the mammalian gene. Taken together, the data suggests that the
elovl2 is the sole fatty acid elongase that is able to elongate C22 to C24 in the inner nuclear layer.
We find expression of
elovl2 primarily in the Muller glia, which is in contrast to the mouse data where it is primarily expressed in photoreceptors in the retinal pigment epithelium [
12]. The role of the Muller glia synthesized PUFAs is still poorly understood. Interestingly, Muller glia synthesis of 9,20-dihydroxydocosapentaenoic acid (DHDP), a diol of docosahexenoic acid (DHA), has been observed inhibit endothelial Notch signaling and regulate retinal angiogenesis [
33]. In the mammalian retina,
Elovl2 is expressed in photoreceptors, especially in cones. Since the VLC-PUFAs are the key fatty acids in photoreceptor disks membranes, the striking difference in the expression pattern of mammalian and zebrafish
elovl2 gene suggests a potential dependence of photoreceptor cells on Muller glia. This interesting interaction would involve the Muller glia in zebrafish eye to provide VLC-PUFAs for the photoreceptor disks during the daily disk regeneration. Further studies are needed to elucidate the role of Muller glia derived PUFAs in retinal structure and function.
We then assessed the function of
elovl2 in zebrafish embryos by creating
elovl2 crispants. This was done by injecting 2 gRNAs specific to
elovl2 as well as Cas9 protein into single cell embryos. High rates of gene editing were observed at the sites of both gRNAs (
Supplementary Figure S3), causing high biallelic changes in the
elovl2 gene. One caveat of this approach is that off target effects of CRISPR editing can occur.
Elovl2 crispants had significant changes in levels of various fatty acids, consistent with loss of Elovl2 function compared to control crispants. Lipidomics performed on whole zebrafish embryo showed a significant decrease of LC-PUFAs in lipids isolated from
elovl2 crispants. These FAs, including 20:4, 20:5, 22:4, 22:5, 22:6, 24:5 and 24:6, are potential products of Elovl2 enzymatic activity. Our results are in general agreement with the data presented by Liu et al. [
16], in particular regarding the key role of Elovl2 in DHA synthesis. However, we did not observe the accumulation of 20:5n-3 in our extracts. This might be due to the stage difference (2 dpf vs. 3 dpf) or different detection methods; more sensitive GC-MS used in Liu et al., vs. the LC-MS used in our studies. Our approach however allowed us to detect VLC-PUFAs and their lower abundance in 7 dpf retina. In addition, the availability of specific fatty acids affected the overall composition of lipids in the zebrafish embryos. In particular, levels of phospholipids were significantly changed while availability of free fatty acids substantially increased. Importantly, the samples analyzed at day 2 were those of whole zebrafish embryos. Therefore, these changes reflect a diversity of tissue types and not specifically changes in the zebrafish retina. At day 7, we analyzed dissected eyes. Nevertheless, this data shows that lack of specific fatty acids can affect the lipid biosynthesis, and therefore the overall compositions of membranes.
To analyze the impact of the lack of
elovl2 enzymatic activity on the retinal morphology thickness of the retinal layers in the 7 dpf control and
elovl2 crispants, sections were examined and quantified. No gross disruptions of the retinal architecture were observed, however, the thickening of several retinal layers in
elovl2 crispants when compared to controls was detected. Other studies have reported changes in the thickness of retinal layers after the inactivation of Muller glia cells [
28,
34]. MacDonald et al. showed that inhibition of Muller glia formation during zebrafish development results into wider RGL but no effect was observed in other retinal layers. This suggest that other factors play role in maintenance of retinal layers in the absence of Muller glia. While our observation further supports the indispensable role of Muller glia in maintenance of RGL in zebrafish retina, the factors involved in the maintenance of other retinal layers in the absence of Muller glia cells remain unstudied. Another study targeting the deletion of Dicer1 miRNA specifically in Muller glia cells present in INL of mice retina reported an increase in the thickness of INL. Intriguingly, the group also observed reduction in thickness of ONL. Considering the global change in the membrane lipids composition in
elovl2 crispants (
Supplementary Figure S5) and therefore different physical properties of the membranes, our studies further supports the observation of the key role of the Muller glia in the maintenance of retinal architecture.
Finally, we examined the changes in visual behavior in zebrafish
elovl2 crispants. We performed the VMR assay, a well characterized visual reflex that develops by day 5. We assessed the visual motor response at day 6 and did not find any changes in the VMR-ON response. However, we did see a significantly weaker, albeit still present, VMR-OFF response after
elovl2 knockdown. It is still unclear what parts of the visual circuit mediate this VMR-OFF response. Interesting,
slc7a14 mutant zebrafish, which affects rod photoreceptors more than cones, shows greater defects in the VMR-OFF response than the VMR-ON response [
35]. This is in contrast to
Pde6c mutation, which results in cone degeneration, and leads to defects in the VMR-ON response rather than VMR-OFF response 22. This suggests a possibility that VMR-ON and VMR-OFF responses may be primarily influenced by different classes of photoreceptors [
35]. Further studies are needed to examine the effect of
elovl2 on visual function in the zebrafish retina.
In conclusion, this is the first study to investigate the role of elovl2 in the zebrafish retina. Loss of elovl2 activity results in broad changes across the production of lipids compatible with its function as an elongase of LC-PUFAs as well as in impact on visual function. Elovl2 expression in the Muller glia may suggest additional and unique functions of zebrafish elovl2 compared to mammals. Taken together, our work further advances our understanding of the role of elovl2 and LC-PUFAs in retina structure and function.