Abstract
Environmentally-induced transgenerational epigenetic inheritance is an emerging field. The understanding of associated epigenetic mechanisms is currently in progress with open questions still remaining. In this review, we present an overview of the knowledge of environmentally-induced transgenerational inheritance and associated epigenetic mechanisms, mainly in animals. The second part focuses on the role of PIWI-interacting RNAs (piRNAs), a class of small RNAs involved in the maintenance of the germline genome, in epigenetic memory to put into perspective cases of environmentally-induced transgenerational inheritance involving piRNA production. Finally, the last part addresses how genomes are facing production of new piRNAs, and from a broader perspective, how this process might have consequences on evolution and on sporadic disease development.
1. Introduction
The impact of environmental stress on gene expression has been known since the 1960s. In the presence of lactose, glucose starvation in Escherichia coli induces derepression of the lactose operon, allowing cells to metabolize this sugar. These gene regulation changes are maintained until glucose is added to the nutritive medium without impacting subsequent generations [1].
In animals, plants, or fungi, some stress can induce epigenetic changes that can be maintained for at least more than one generation even if the stress is removed. Early studies analyzing human populations bring to light some physiological effects of parental food intake on their children’s health, for instance, in Sweden [2], during the Leningrad siege (1941–1944) [3] or the Dutch famine (1944–1945) [4,5]. A restricted number of studies identified epigenetic modifications associated with parental stress [6]. Identification of accurate molecular associated modifications remains difficult to pinpoint, because of (1) the obvious impossibility of establishing a classical design of experiment, such as reproducibility and control experiments during several human generations, (2) past cultural and ecological factor influences [7], and (3) multiple stresses (nutritional, psychological, or social) that go along with wars and famines. Therefore, to better understand the consequences of specific stress at the epigenetic level and their possible transmission, it is critical to develop animal models.
Epigenetic inheritance is characterized by a reversible phenotypic transmission to the next generation in the absence of DNA sequence modification. This definition can be applied in an extended sense, including social learning and communication or in a tight sense, like mother to daughter cell transmission, called ‘cellular epigenetic inheritance’ [8]. In the latter, it is also important to differentiate intergenerational, multigenerational, and transgenerational inheritance. If the stress condition impacts the F0 gestating female and the F1 progeny (embryos), it will be referred to as an intergenerational effect. If the F2 progeny (embryonic germ cells) are also affected, it will be referred to as a multigenerational effect. In contrast, transgenerational inheritance corresponds to an epigenetic transmission of stress passing through the germ cells, but without direct exposure. Hence, male and non-gestating female transgenerational phenotypes can be considered in F2, whereas for gestating female, only F3 has to be considered [9].
In the past twenty years, the development of next generation sequencing (NGS) approaches led to the accumulation of convincing evidence for environmentally-induced transgenerational epigenetic inheritance dependent on DNA methylation, specific histone modification, and non-coding RNA biology. However, the precise mechanisms remain unclear, leaving open questions to be tackled. PIWI-interacting RNAs (piRNAs), a class of small non-coding RNAs that tightly repress transposable element (TE) activity in animal gonads, have also been demonstrated to induce stable epigenetic memory. In this review, we present some examples of environmentally induced transgenerational epigenetic inheritance to underscore the role of piRNAs in epigenetic inheritance and more specifically, in transgenerational epigenetic inheritance in response to environmental stress. Finally, the implications of piRNAs in environmentally-induced transgenerational inheritance in evolution and potential implication in human health will be discussed.
2. Evidence and Mechanisms of Environmentally-Induced Transgenerational Epigenetic Inheritance
The universal aspect of transgenerational epigenetic inheritance is now well documented in several species from different taxa, including bacteria, protists, fungi, plants, and animals [8]. About one-third of these transgenerational effects are linked to environmental factors, such as nutritional stresses (starvation or supplementation), chemical stresses (arsenic), and physical stresses (temperature, pressure, light and dark exposure).
2.1. Multiple Molecular Supports for Environmentally-Induced Epigenetic Inheritance
Four main types of support have been identified including (1) DNA that can be methylated on CG, CH, and CHH motifs, where H corresponds to a nucleotide other than G, (2) histones that can be modified (trimethylation of histone 3 lysine 4 (H3K4me3) associated with transcriptionally active marks and di/trimethylation of histone 3 lysine 9 (H3K9me2/3) and trimethylation of histone 3 lysine 27 (H3K27me3) associated with transcriptionally repressive marks) or replaced in the late-stage spermatids of many animals by the protamines forming a highly compact genome [10]. Defect of histone replacement during gametogenesis, called ‘histone retention’, could be associated to some infertility in men [11], (3) small non-coding RNAs (microRNAs (miRNAs), short interfering RNAs (siRNAs), PIWI-interacting RNAs (piRNAs), and transfer RNA-derived small RNAs (tsRNAs) (also known as tRNAs Fragments (tRFs)), and (4) long non-coding RNAs (lncRNAs) [12,13,14].
Two additional supports for epigenetic inheritance are possible, although evidence of environmentally-induced epigenetic inheritance is still lacking. First, protein structures involving prions or the Hsp90 chaperone protein have both been linked to chromatin remodeling factors in Drosophila and could be good candidates for involvement in such mechanisms (for more details, see [15,16]). Second, over the years, more than 100 RNA modifications have been described, some linked to human diseases [17,18]. Among the enzymes involved in these modifications, the m5C Dnmt2 RNA methyltransferase has been shown to be required in mice for epigenetic transmission of Kit and Sox9 phenotypic variants [19].
Although evidence is still lacking concerning protein structure and RNA modifications, transgenerational environmentally-induced epigenetic inheritance associated with DNA methylation, histone modifications, or non-coding RNAs has been reported. Some examples are presented in Table 1, which highlights that one stress can induce multiple epigenetic modifications. For instance, in rat sperm, vinclozolin or DDT pesticide exposure modifies DNA methylation, drives histone retention, and decreases the global number of expressed small non-coding RNAs (miRNA, piRNAs, and tsRNAs) and long non-coding RNAs, affecting expression of genes implicated in metabolism, signaling, and transcription [20,21] and is correlated with the occurrence of several pathologies [22,23,24,25]. In Drosophila, the elevation of temperature during development causes functional piRNA emergence associated with H3K9me3 enrichment from a particular repeated locus mimicking a piRNA cluster for over 50 generations [26] (Table 1). Such cross-talk between several epigenetic modifications is not restricted to animals. In fission yeast, the RNA interference (RNAi) machinery is necessary to direct H3K9me3 mark deposition, inducing heterochromatin formation and silencing on a centromeric transgene. This new epigenetic state, sensitive to the H3K9 methyltransferase mutant, can then be maintained up to 32 generations [27]. Taken together, H3K9me3 marks and small non-coding RNA could act together to reinforce epigenetic stability through generations thanks to a feedback loop: on the one hand, H3K9me3 leads to heterochromatin formation and recruits the RNAi machinery, and on the other hand, the RNAi machinery targets H3K9 residues and induces their trimethylation [26,27].

Table 1.
A list of environmentally-induced transgenerational epigenetic inheritance.
2.2. How Long Does the Imprinting of Transgenerational Epigenetic Inheritance Last?
At each generation, the totipotency of primordial germ cells is ensured by the erasure of epigenetic modifications requiring numerous genetic factors, such as DNA demethylation, histone methyltransferases, and histone demethylases [55,56,57]. However, analyses of DNA methylation patterns show that some loci escape this resetting, suggesting that antagonist mechanisms provide maintenance of some epigenetic marks [58]. In fact, the DNA methyltransferases Dnmt3a and Dmnt3b, the PRC2 complex in mice and the PRC2 complex associated with H3K27me3 marks in worms have been shown to be involved in the maintenance of DNA methylation and histone modification pattern on discrete loci [59,60,61].
Some studies have addressed this question for small non-coding RNAs. In Caenorhabditis elegans, the RNAi machinery provides its own regulation by targeting RNAi genes and leads to a progressive decrease of small RNA at each generation, varying from one to four generations [62,63]. Antagonist molecular mechanisms govern the duration of the small RNA transgenerational response. As for other modifications, transgenerational small RNA inheritance relies on many genes coding for RNA-dependent RNA polymerase (RdRp), Argonaute proteins, or histone methyltransferases (Reviewed in [64,65,66]). The duration of the transgenerational response can be extended due to siRNA, which constitutes a second RNAi trigger that drives tertiary siRNA production [67]. Hence, the duration of epigenetic inheritance depends on the influence of these two opposite processes, namely resetting and stimulation of small RNA production [68,69]. Disruption of the endo-siRNA pathway leads to hypersensitivity to exogenous small RNAs, suggesting competition between these RNAi pathways, probably for molecular factors, such as DCR-1, which is the unique Dicer ortholog in C. elegans [70]. In contrast, piRNA producing loci transcription is actively maintained at each generation by maternal piRNA inheritance in Drosophila [71,72]. Disruption of this maternal deposition is followed by a complete erasure after only one generation [26,71].
The family of small non-coding RNAs is frequently considered in transgenerational epigenetic inheritance, probably because there are responsible for endogenous gene regulation and involved in important developmental programs (Reviewed in [73]). In the particular case of piRNAs, their function in epigenetic memory associated with their role in response to environmental factors has been recently demonstrated [26,46].
3. piRNAs in Environmentally-Induced Transgenerational Epigenetic Inheritance
3.1. Overview of piRNA Biogenesis and Functions
The piRNAs correspond to a class of small non-coding RNAs interacting with the PIWI proteins (Piwi, Aub, and Ago3 in Drosophila), belonging to the AGO subfamily proteins, mainly active in animal gonads to protect genome integrity against TEs. In Drosophila germ cells, piRNAs are produced from specific heterochromatic loci, composed of numerous TE fragments, called piRNA clusters [74,75]. Those clusters are enriched in the H3K9me3 mark and Rhino HP1-like protein [76] (Figure 1). Rhino interacts first with the co-factors Deadlock and Cutoff, then Deadlock recruits the transcriptional initiation machinery via the large subunit of transcription initiation factor II A (TFIIA-L) paralog called Moonshiner, as well as Bootlegger, UAP56, the THO complex, and nuclear export factors Nxt1 and Nxf3 [76,77,78,79]. The piRNA precursor transcripts emerging from piRNA clusters are then exported by the Crm1 (chromosomal maintenance 1) exportin to the cytoplasmic piRNA biogenesis site, where they undergo a single strand hydrolysis by the Zucchini RNase to finally give rise to piRNAs [78,79,80,81,82,83] (Figure 1). In contrast to si- and miRNA, piRNAs are charged directly as single guide RNA onto PIWI proteins to form piRNA-induced silencing complexes (piRISC) [74]. After nuclear translocation, piRISC complexes recognize nascent TE transcripts and perform transcriptional gene silencing (TGS) by local heterochromatinization of TEs. This silencing requires nuclear Piwi-piRNAs that direct Panoramix, Nxf2, its co-factor, Nxt1, and the histone methyltransferase Eggless to the TE loci [84,85,86,87,88,89,90,91] (Figure 1). In the cytoplasm, mRNAs of TEs are cleaved by piRISC complexes, leading to post-transcriptional gene silencing (PTGS) of mobile elements. Cleaved-transcripts are not degraded; they are charged by PIWI protein to form new piRISC complexes, corresponding to an amplification mechanism called the ‘Ping-Pong cycle’ [74] (Figure 1). piRNA-dependent silencing occurs at each gonadal developmental stage [92], and disruption of the piRNA pathway leads to TE mobilization [93], which induces DNA damage [94,95].
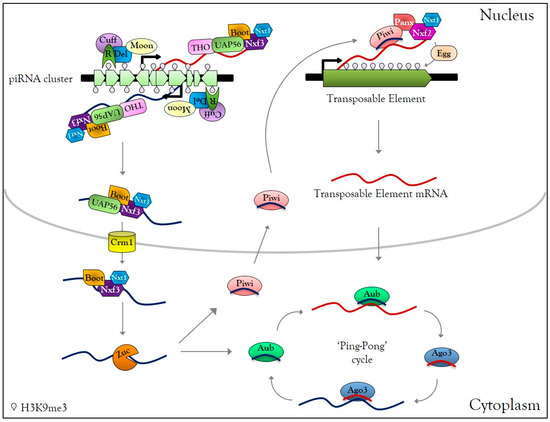
Figure 1.
Germline PIWI-interacting RNAs (piRNA) biogenesis. In Drosophila melanogaster germ cells, active piRNA clusters are enriched in H3K9me3 and HP1-like protein Rhino (R), which interact with Deadlock (Del), Cutoff (Cuff). Moonshiner (Moon) interacts with Del and recruits transcription initiation factors, allowing transcription of piRNA clusters in the sense and antisense direction. Bootlegger, UAP56, the THO complex, Nxt1, Nxf3 are recruited to the piRNA cluster transcription site. The transcripts are then exported to the cytoplasmic piRNAs biogenesis site via the CRM1 exportin. In the cytoplasm, piRNA precursors are cleaved by Zucchini (Zuc) into primary piRNA and charged by Piwi or Aubergine (Aub) proteins, forming Piwi-piRISC and Aub-piRISC complexes. Piwi-piRISC is translocated into the nucleus and recognizes nascent TE transcripts due to the interaction with Panoramix, the Nxf2, and Nxt1. The association of these factors to nascent TE transcripts causes recruitment of the histone methyltransferase Eggless that transcriptionally silences TEs by local heterochromatinization. In the cytoplasm, Aub-piRISC cleaves TE transcripts into secondary piRNAs, which are loaded onto Ago3, forming Ago3-piRISC complexes. Ago3-piRISC recognizes and cleaves piRNA precursors into new secondary piRNAs, loaded onto Aub. This mechanism, known as the ‘Ping-Pong’ cycle, allows the amplification of piRISC complexes and post-transcriptional regulation of TEs.
PIWI protein distributions reveal that the piRNA pathway is highly conserved in animals, but they exist neither in plants nor in prokaryotes [96]. Interestingly, recent studies show that piRNAs are also found in somatic tissues of mammalian, arthropods, and mollusks [97,98,99,100]. In Drosophila melanogaster, however, piRNA expression is restricted to the gonads, including the germline nurse cells and the somatic follicular cells [101,102]. These findings might suggest that the ancestral capacity to synthesize piRNAs in somatic tissues (outside of gonads) has been lost in D. melanogaster. However, this is still subject to debate [103,104,105].
3.2. Role of piRNAs Outside of TE Regulation
Genomic annotations of small RNA libraries isolated from various metazoans reveal that piRNAs are not restricted to TE regulation but can also target coding genes. In Drosophila, piRNAs derived from the 3′UTR of traffic-jam (tj) matching Fasciclin III (FasIII) can be identified in the ovarian somatic cells (OSC) line. This finding suggests that FasIII is regulated by piRNAs originated from the tj 3′UTR [102,106]. In Drosophila early embryos, nanos (nos) is regulated by piRNAs. Evidence suggests that maternal nos mRNAs are targeted by a complex composed of piRNAs, Aub, and CCR4-NOT deadenylation factors in the soma, whereas in the germ plasm, those transcripts are stabilized by the interaction with Aub [107,108]. In Aplysia, piRNAs seem to be implicated in CREB2 promoter methylation, a transcriptional repressor involved in long-term memory [109]. As the last example, sex determination in silkworm is dependent on the piRNA pathway that regulates Masculinizer, a locus required for the production of the doublesex female-specific isoform [110]. Taken together, these data highlight the idea that piRNAs are required in gene regulation of a broad range of cellular functions.
3.3. piRNAs in Epigenetic Memory
In Drosophila, the insertion of exogenous lacZ sequences into a subtelomeric piRNA cluster can functionally repress a euchromatic lacZ transgene in trans [111]. This epigenetic silencing mechanism, referred to as the trans-silencing effect (TSE), requires maternally but not paternally inherited piRNAs [71,112,113,114,115,116]. These results strongly suggest an active role for piRNAs in TSE and show that a piRNA cluster, without maternal piRNA inheritance, is not sufficient to promote this silencing process [112]. Involvement of piRNAs in epigenetic memory has been associated with epiallele emergence through a paramutation phenomenon in D. melanogaster and in C. elegans [71,117]. Paramutation is defined as a modification of one allele induced by another allele without DNA sequence modification and corresponds to an epigenetic conversion (for reviews see [118,119,120].
In Drosophila, using a locus made of repeated P-lacZ-white transgenes mimicking a piRNA cluster (called BX2 cluster), but inactive for piRNA synthesis, we have shown that homologous maternally-inherited piRNAs can stably convert this locus into an active piRNA cluster for over 200 generations [71] (Figure 2). This newly-activated cluster has the same properties as other Drosophila piRNA clusters. This piRNA-induced paramutation process requires molecular actors of the piRNA pathway [72] and performs TE-derived sequence control generation after generation. Finally, the reversibility of paramutation occurs when maternal piRNA inheritance is interrupted, underscoring the idea that this is an epigenetic process [26,71]. Taken together, based on sequence complementarity, these data show that maternally-inherited piRNAs are competent to initiate new piRNA production in the next generation and suggest that piRNAs are responsible for their own maintenance across generations. Two non-exclusive molecular mechanisms have been proposed to explain this case of epigenetic memory in which piRNA production requires piRNA inheritance [121]. In the first, inherited-piRNAs would induce heterochromatin formation on the piRNA cluster by leading to H3K9me3 deposition, inducing local Rhino enrichment [26]. The piRNA cluster-specific transcription machinery will then assemble at the locus to promote piRNA synthesis [77]. In the second, inherited piRNAs could trigger their production by Ping-Pong amplification from piRNA cluster precursors and TE transcripts [74,75].
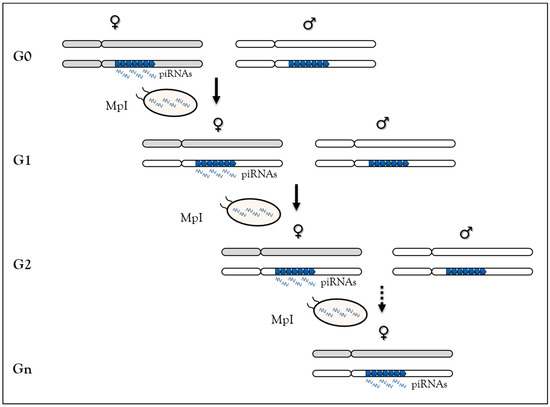
Figure 2.
The paramutation process in Drosophila melanogaster relies on maternal piRNA inheritance. At G0, females (grey chromosomes) carrying the BX2 clusters made of repeated P-lacZ-white transgenes (blue arrowheads) producing piRNAs are crossed to males (white chromosomes) carrying the same cluster not producing piRNA. At G1, maternal piRNA deposition in the embryo leads to activation of piRNA production from the paternal cluster allele. The newly activated cluster can activate the paternally-inherited inactive cluster in G2 due to maternal piRNA inheritance (MpI). Then, this process of activation leads to stable piRNAs production across generation.
In C. elegans, piRNAs correspond to 21 nucleotide (nt) long small RNAs with a uridine bias at the 5′-end (21U RNA) distinct from the endo-siRNAs that are 22 nt long small RNAs with a guanosine bias at the 5′-end (22G RNA) [122]. The biogenesis of nematode piRNAs depends on the Piwi protein, PRG-1 [66,123,124]. Like in Drosophila, transgenes are used to decipher the molecular mechanisms of germline repression. Analysis of the piRNA sensor indicates that piRNAs are required to trigger silencing through imperfect base-pairing [125,126,127]. This silencing is then transgenerationally propagated by the 22G RNA dependent on numerous factors, such as RdRp, NRDE-2, the Argonaute proteins WAGO-9/HRDE-1 and WAGO-4 and chromatin components, such as the histone methyltransferases SET-25, SET32, and HPL-2, a Heterochromatin 1 Protein (HP1) orthologue [45,128,129,130,131,132,133]. Indeed, specific H3K9 methyltransferases, SET-25 and SET-32, and endo-siRNAs have been shown to work together to promote silencing inheritance [134]. In addition, neuronal RDE-4 and germline HRDE-1 have been implicated in transgenerational regulation of endo-siRNAs and mRNAs controlling chemotaxis [135]. The authors proposed the existence of an intricate interaction between the neuronal system and the germline to control transgenerational behavior. Finally, precise dissection of the silencing mechanism determines that piRNAs lead to the production of secondary and tertiary siRNAs, necessary for full repression and leading to an epigenetic conversion form of paramutation [67].
Altogether, the piRNA silencing pathway in Drosophila involves piRNAs for the initiation and maintenance through generations [26,71,72], while in worms piRNAs are involved in triggering the silencing that is then maintained by siRNAs in the progeny, independent from piRNAs [124,136,137,138].
3.4. Heritable Stress-Induced piRNA Synthesis
In F0 mice, an early maternal separation causes downregulation of piRNA cluster 110 [139], and a Western-like diet leads to differential expression of 190 piRNAs distributed on 63 piRNA clusters [140]. In rat, a high-fat diet induces deregulation of 1092 piRNAs, and only three of them are still differentially expressed at the next generation [141]. In C. elegans, high temperature (25 °C) induces motif-dependent piRNA biogenesis downregulation [142]. This downregulation is associated with differential gene expression and fitness reduction, which persist at the next generation. However, in those studies, piRNA expression in subsequent generations, as well as their roles in a phenotypic response and the potential consequences on the epigenome, was not directly addressed. In the case of rat stressed with vinclozolin or DDT, differentially expressed piRNAs were identified in F3, but the mechanism and role of piRNAs were not questioned [20,21,22,23,25] (Table 1).
This year, two cases of environmental stresses inducing a genomic response dependent on piRNAs have been published in C. elegans and in D. melanogaster [26,46].
3.4.1. Behavior in C. elegans
Wild type male or female larvae of C. elegans exposed to pathogenic Pseudomonas aeruginosa learn to avoid it and switch their feeding preference to nonpathogenic bacteria. This learning to distinguish between pathogen and non-pathogen does not correspond to a general phenomenon but rather is specific for P. aeruginosa [46]. Surprisingly, this behavior of avoidance can be transgenerationally inherited to the naïve F1 progeny up to the fourth generation (F4). This peculiar inheritance is completed at the fifth generation, where progeny exhibits a naïve behavior. Differential transcriptome analyses were performed on treated vs. non-treated mothers fed with pathogenic P. aeruginosas and their respective progeny. Moore and colleagues showed that the TGF-β ligand DAF-7, previously identified as a neuronal responder of the pathogen [143], was expressed more in specific sensory neurons of treated mothers relative to non-treated animals as well as in the F1, F2, F3, and F4 progeny, thus mirroring transgenerational avoidance [46]. The authors further show that in the daf-7 loss of function mutant, while the mother can learn to avoid the pathogen, the F1 progeny did not receive the information of avoidance. The same pattern of inheritance was observed for mother defective for the Piwi encoding gene prg-1. This mutation is associated with a defect in piRNA expression in the mothers and in daf-7 induction in the ASI neurons in F1.
Altogether, the studies indicate that transgenerational inheritance involved in the avoidance of pathogenic P. aeruginosa depends on the TFG-β pathway linked to the small non-coding RNAs as well as piRNA molecular partners. Therefore, piRNAs might participate in the control of specific reversible behaviors, which might have consequences on adaptive survival, providing advantages to progeny. In addition, the loss of inheritance after five generations might be considered an advantage in worms that may colonize a new environment in their natural habitat. The precise molecular basis of this resetting occurring at the fifth generation was not addressed but might be reminiscent with previous studies [63].
3.4.2. Heat Response Induces de novo piRNAs
Recently, high temperature during development (29 °C instead of 25 °C) has been shown to be sufficient to induce de novo piRNA production and heterochromatin associated with H3K9me3 and Rhino enrichment from the BX2 cluster in Drosophila [26]. One generation raised at 29 °C is sufficient to activate piRNA production from the BX2 cluster, and once this production is started, it can be maintained over 50 generations even after return to normal temperature (25 °C) (Figure 3). These piRNAs can functionally silence homologous lacZ reporter transgenes, a phenomenon referred to previously as TSE. Hence, heat stress is able to induce piRNA production from a locus comprised of repeats. Once established, this production remains stable even after stress removal, i.e., when flies return to a normal temperature of 25 °C. Small RNA sequencing analyses were unable to identify other genomic loci used for de novo piRNA synthesis as if all loci competent to become piRNA cluster were already activated. Analyses performed at 29 °C to identify molecular factors required for the activation of de novo piRNA production suggest that there is an increase of transcription of the BX2 cluster flanking region along with the transcription of a homologous euchromatic sequence. When BX2 is maternally inherited, piRNA production is maintained, but when BX2 is paternally inherited, piRNA production is abolished. This result suggests that when the stress is removed, piRNA can be self-maintained over generations by the paramutation process, as long as the inheritance is maternal [26] (Table 1).
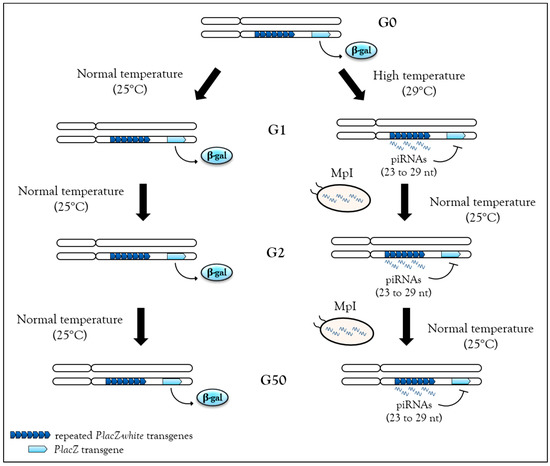
Figure 3.
Stable, heritable stress-induced de novo piRNAs. In flies, the BX2 cluster, a sequence made of repeated P-lacZ-white transgenes (dark blue arrowheads), is capable of producing piRNA after one generation at 29 °C, then these piRNAs can functionally repress a homologous P-lacZ sequence (light blue arrowheads) in trans. This silencing capacity is maintained at the second generation after return to normal temperature (25 °C) and then up to 50 generations. Hence, when stress is removed, piRNAs can be self-maintained by maternal piRNA inheritance (MpI) at each generation.
In Drosophila, environmental stresses can generate piRNA emergence, and this new piRNA production is maintained across generations, allowing stable repression of homologous sequences. How can environmental stress impact piRNA production? It remains unclear whether piRNA cluster chromatin containing H3K9me3 marks, piRISC complex formation and/or stability, or piRNA pathway genes are involved in the stress response, individually or simultaneously. To date, it is quite difficult to estimate the global effect of piRNA deregulation over many generations, in particular, if it might result in the alteration of inherited gene expression patterns. Future investigations will help our understanding of environmentally-induced piRNA transgenerational inheritance and allow the estimation of potential consequences.
4. Speculative Insights on Transgenerational piRNA Inheritance Consequences
4.1. Implications in Evolution
According to Lamarck’s theory, the environment and the experiences of the organism mediate changes that could be inherited in subsequent generations. These changes are then subject to natural selection and adaptation [144,145]. In the case of transgenerational inheritance, the environment can impact the epigenome by causing epimutations, through DNA methylation, histone modifications, and non-coding RNA misregulation. These epimutations lead to phenotypic variations and can give the organism a selective advantage. In C. elegans, it has been shown that stress conditions (starvation, supplemented diet, salt, or arsenic) enhance longevity and stress resistance in subsequent generations [146,147,148]. If they are stable enough to be maintained over time, epigenetic modifications and their gradual accumulation in response to various stresses, may have an important role in speciation [149]. For Brevik and colleagues, transgenerational epigenetic inheritance may also explain the rapid emergence of insect population resistance against pesticides [150]. Many studies have shown that pesticides, notably vinclozolin or DDT, induce transgenerational changes in DNA methylation, histone modification, and/or non-coding RNA deregulation in rodent models (Table 1).
Among the epigenetic mechanisms driving adaptation and evolution, piRNAs are playing a fundamental role in TE regulation across generations. For instance, during the 20th century, the D. melanogaster genome was invaded by the P element after a horizontal transfer [151]. It appears that in the absence of P element regulation, progeny display genetic defects, leading to gonad atrophy at high temperature, whereas no anomaly occurs when P element regulation is established [152,153]. It was later shown that this specific control of gonadal P element occurs through piRNAs [101]. Interestingly, the Drosophila simulans genome has been recently invaded by P elements [154], and the invasion dynamics are widely influenced by temperature. The transposition rate at elevated temperature is higher compared to the rate at low temperature [155]. At elevated temperature (29 °C), after 20 generations, the P element copy number in the genome reaches a plateau. These invasions are correlated with piRNA production against P element sequences, strongly suggesting that the regulation is established in response to invasion.
Analyses of P element distribution in the D. melanogaster genome showed that subtelomeric regions of the X chromosome are hotspots for P element insertions [156]. Those subtelomeric regions happened to be one of the active Drosophila piRNA clusters [74]. Once P element or P derived transgenes insert into subtelomeres, new piRNAs targeting those sequences will be synthesized, leading to repression of euchromatic copies [71,101,114,116,157]. Particular sequences in subtelomeric piRNA clusters, called TAS-L like (TLL), found in the X chromosome of D. melanogaster, are also present in autosomic subtelomeric piRNA clusters of other members of the melanogaster subgroup, including D. simulans. The finding that some P element insertions were found very close to TLL in D. simulans suggests that P element regulation could be conserved between D. melanogaster and D. simulans [155,157]. As with all piRNA clusters, the subtelomeric regions are made up of repeated domains that were once activated for piRNA production. However, the type of event that converted these loci remains unclear. Recently, we have shown that high temperature can convert a locus composed of tandemly repeated sequences into a stable piRNA cluster, highlighting the sensitivity of such regions to environment stresses for piRNA production [26]. Once activated, de novo piRNA production can be transmitted through generations by maternal inheritance [71]. Because these loci are hotspots of insertion for TEs, specific piRNAs are synthesized that functionally repress homologous sequences, leading to maintenance of genome integrity over time. Therefore, piRNA clusters constitute an epigenetic memory of TE invasion and are essential to provide genome defense.
TEs contain many regulatory sequences, including insulators, RNA polymerase signals, splice sites, and/or environment response elements [158]. Furthermore, TEs can modulate the expression of neighboring genes [159]. In addition, numerous stresses, including radiation, pollutants, temperature, or viral infection, can induce TE activation [160,161]. Thus, similar to P element activity under elevated temperature conditions, the emergence of piRNA production can occur in response to stress-induced TE activation. We can expect that organisms with TE regulatory mechanisms are likely to be selected as compared to organisms without this defense system. Indeed, TE mobilization induced by environmental stress can affect genome organization by creating mutations, chromosomal breaks, or genomic rearrangements [162,163]. In the long-term, these modifications can be beneficial for organisms by the creation of new molecular functions, but until this occurs, TEs must be regulated in the short-term to restrict genomic damage. In a way, if TEs are defined as molecular actors of evolution, piRNAs could act as molecular regulators of evolution.
Finally, piRNAs and molecular factors of the piRNA pathway have been described to play a role in reproductive isolation by maintaining interspecies sterility [164,165]. In D. melanogaster and D. simulans hybrids, D. simulans Rhino protein co-localizes with D. melanogaster Deadlock, but they cannot interact, because of the rapid evolution of their chromo domains. As a consequence, TE mobilization leads to hybrid sterility due to the absence of a functional piRNA pathway [164]. Additionally, the AT-chX locus, located close to the pericentromeric region of the X chromosome, is one of the D. melanogaster piRNA clusters absent from other Drosophila species. This locus contains sequences related to the vasa gene, which is required for germ cell development. Due to imperfect complementarity, piRNAs derived from the AT-chX locus are not able to repress vasa expression in D. melanogaster germ cells. Surprisingly, the AT-chX locus shares a high level of complementarity with the vasa gene from closely related species to D. melanogaster, leading to a repression of vasa in interspecies hybrid testes [165]. These studies suggest that piRNAs could play an important role in speciation.
4.2. Potential Implications for Disease Development
Environmental stresses, such as pesticide and pollutant exposure, have been shown to drive epigenetic inheritance of obesity and many associated diseases, including diabetes, infertility, immune disorders, and polycystic ovarian syndrome in rat. These transgenerational effects are associated with the emergence of differential DNA methylation regions, which contain genes involved in several processes, such as signaling, metabolism, transcription, development, and transport [23,24,25,32,35,36,38,166,167,168].
4.2.1. piRNA Pathway and Disease
To date, there is no evidence for piRNA-mediated transgenerational inheritance of disease risks. However, paramutation and small RNA inheritance could represent a part of ‘missing heritability’ in some rare diseases that genetic mutations cannot explain (discussed in [169,170]). Deregulation of piRNAs and PIWI proteins has been found in some diseases, including many types of cancer. Hence, piRNAs are considered as cancer biomarkers along with several circulating RNAs [171]. Transcriptome analyses highlight the deregulation of piRNAs in several malignant tumors [172,173,174,175]. For instance, piR-Hep1 is upregulated in a hepatocellular carcinoma cell line [173]. piR-Hep1 knockdown reduces cell motility and invasiveness, whereas piR-Hep1 overexpression strongly increases cell migration. Finally, the high expression of human PIWI proteins (PiwiL) has also been associated with various cancers. Interestingly, the expression level of PiwiL2 is correlated with tumors aggressiveness. Moreover, PiwiL2 knockdown decreases cellular migration, suggesting a role in invasion [176,177].
It also appears that PIWI proteins are involved in different cellular processes, such as stem-cell maintenance or cell differentiation, in the Drosophila germ line [178] and tumor cell proliferation through induction of c-Myc expression in human cells [179]. As a consequence, piRNA and PIWI protein deregulation could sustainably affect these essential processes and induce developmental defects. It is possible that piRNA deregulation could partly explain cases of family susceptibility to cancer. Indeed, if the piRNA pathway is impaired, TE regulation is affected, and genome integrity is compromised.
4.2.2. TEs and Disease
In different models (Drosophila, mouse, zebrafish, and human cultured cells), p53 mutation leads to TE derepression associated with a piRNA biogenesis defect, at least in Drosophila [180]. Accordingly, p53-derived cancers have been considered as ‘transposopathies’, diseases induced by TE mobilization resulting in genome instability [181]. However, even though TE insertions have been detected in several cases of cancers, it is still unclear whether they are the direct cause of cancer development or a consequence of another cellular defect causing cancer. For instance, in colon cancer, retrotransposon insertion into the APC tumor suppressor gene leads to its inactivation; in cancer cell lines, retrotransposon insertions are linked to long chimeric transcript emergence and thus, drive the expression of several genes [182,183]. Transcriptome-wide analyses of RNA-seq data for several cancer types and tumors samples identified TE-oncogene chimera transcripts, a phenomenon called a ‘TE onco-exaptation event’. Several TE onco-exaptation events lead to oncogene activation or overexpression. For instance, AluJb, which drives LIN28B expression, has been implicated in lung cancer. DNA methylation of the AluJb sequence leads to a decrease in LIN28B expression [184].
Other diseases are linked to TE activity, including hemophilia [185,186], psychiatric disorders [187], and Alzheimer’s disease [188]. In the particular case of Alzheimer’s disease, piRNA molecules seem to be more abundant in the brain and can be considered as a molecular marker of this disease [189]. Indeed, transcriptome analyses of samples isolated from human brain show that some piRNAs are upregulated in Alzheimer’s disease [190].
Altogether, control of TE expression, especially by piRNAs, might play a crucial role in genome stability. If impaired by stress, such as mutation or environmentally-induced piRNA biogenesis misregulation, aberrant gene expression can occur inducing cancer development or other diseases related to TEs in subsequent generations, leading to some of the unexplained family susceptibility.
5. Conclusions
There is increasing evidence of environmentally-induced epigenetic inheritance, and today we have a better understanding of their associated mechanisms. However, there are still many points in our understanding of the interaction between the environment and the epigenome that are lacking, in particular in metazoans. Three major mechanisms are usually reported as mediators of epigenetic inheritance, namely DNA methylation, histone modification, and non-coding RNAs.
One stress may cause multiple epigenetic changes. Among these changes, some of them are probably independent, whereas other modifications work together. Nature is a combination of numerous stressors including temperature, pressure variations, pollution, starvation, drought as well as competition for food, social stresses, and so on. It is unlikely that one stress can act only on one biological process, but even if this is the case, we can expect that the addition of all these stresses will impact several important pathways.
A balance between transgenerational epigenetic inheritance and epigenetic reprogramming occurs at each generation, and these processes are tightly regulated during development. Several factors are capable of tipping this epigenetic balance toward transgenerational inheritance, notably after stress exposure. piRNA molecules could be one of these factors. Molecular mechanisms of transgenerational piRNA inheritance have been partially studied, and some hypotheses to explain their maintenance have been proposed, which include other epigenetic mechanisms, such as chromatin modifications and several other potential factors discussed above. For the moment, piRNA function in environmentally-induced epigenetic inheritance has been little explored. TE activity is found in some diseases, and many environmental stresses are known to generate TE mobilization events. In this context, piRNA misregulation may have an important impact on genome integrity and structure. Changes in piRNA expression could also modify the regulation of several genes and influence the initiation and development of some diseases. Future work will surely improve our understanding of the piRNA transgenerational effect in response to environmental injury.
Author Contributions
Conceptualization, K.C. and L.T.; Writing—Original Draft Preparation, K.C.; Writing—Review and Editing, K.C., A.B., C.C., and L.T.; Visualization, K.C., A.B., and L.T.
Funding
This research was funded by University Pierre et Marie Curie, grant number EME1223 to L.T, Action Incitative 2018 and COST-Epitran, grant number CA16120 to CC, by financial supports of Sorbonne Université and the CNRS to L.T. and C.C., and by the Ph.D. fellowships from the Ministère de l’Enseignement Supérieur et de la Recherche and la Ligue Nationale Contre le Cancer to KC. We also thank the Réseau André Picard and la Société Française de Biologie du Développement for their supports on travel fellowship grants to K.C.
Acknowledgments
We thank FlyBase.org for providing valuable databases. We thank Lori Pile for critical reading of the manuscript, all the members of the Transgenerational Epigenetics & Small RNA Biology (TErBIO) Team for helpful discussions and the two reviewers for their help and comments during the revision process of this manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Jacob, F.; Monod, J. Genetic regulatory mechanisms in the synthesis of proteins. J. Mol. Biol. 1961, 3, 318–356. [Google Scholar] [CrossRef]
- Kaati, G.; Bygren, L.O.; Edvinsson, S. Cardiovascular and diabetes mortality determined by nutrition during parents’ and grandparents’ slow growth period. Eur. J. Hum. Genet. 2002, 10, 682–688. [Google Scholar] [CrossRef] [PubMed]
- Stanner, S.A.; Bulmer, K.; Andres, C.; Lantseva, O.E.; Borodina, V.; Poteen, V.V.; Yudkin, J.S. Does malnutrition in utero determine diabetes and coronary heart disease in adulthood? Results from the Leningrad siege study, a cross sectional study. BMJ 1997, 315, 1342–1348. [Google Scholar] [CrossRef] [PubMed]
- Painter, R.C.; Osmond, C.; Gluckman, P.; Hanson, M.; Phillips, D.I.; Roseboom, T.J. Transgenerational effects of prenatal exposure to the Dutch famine on neonatal adiposity and health in later life. BJOG 2008, 115, 1243–1249. [Google Scholar] [CrossRef] [PubMed]
- Veenendaal, M.V.; Painter, R.C.; de Rooij, S.R.; Bossuyt, P.M.; van der Post, J.A.; Gluckman, P.D.; Hanson, M.A.; Roseboom, T.J. Transgenerational effects of prenatal exposure to the 1944-45 Dutch famine. BJOG 2013, 120, 548–553. [Google Scholar] [CrossRef] [PubMed]
- Yehuda, R.; Daskalakis, N.P.; Bierer, L.M.; Bader, H.N.; Klengel, T.; Holsboer, F.; Binder, E.B. Holocaust exposure induced intergenerational effects on FKBP5 methylation. Biol. Psychiatry 2016, 80, 372–380. [Google Scholar] [CrossRef] [PubMed]
- Horsthemke, B. A critical view on transgenerational epigenetic inheritance in humans. Nat. Commun. 2018, 9, 2973. [Google Scholar] [CrossRef] [PubMed]
- Jablonka, E.; Raz, G. Transgenerational epigenetic inheritance: Prevalence, mechanisms, and implications for the study of heredity and evolution. Q. Rev. Biol. 2009, 84, 131–176. [Google Scholar] [CrossRef] [PubMed]
- Skinner, M.K. What is an epigenetic transgenerational phenotype? F3 or F2. Reprod. Toxicol. 2008, 25, 2–6. [Google Scholar] [CrossRef] [PubMed]
- Balhorn, R. The protamine family of sperm nuclear proteins. Genome Biol. 2007, 8, 227. [Google Scholar] [CrossRef] [PubMed]
- Hammoud, S.S.; Nix, D.A.; Hammoud, A.O.; Gibson, M.; Cairns, B.R.; Carrell, D.T. Genome-wide analysis identifies changes in histone retention and epigenetic modifications at developmental and imprinted gene loci in the sperm of infertile men. Hum. Reprod. 2011, 26, 2558–2569. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.P.; Brunet, A. Bridging the transgenerational gap with epigenetic memory. Trends Genet. 2013, 29, 176–186. [Google Scholar] [CrossRef] [PubMed]
- Heard, E.; Martienssen, R.A. Transgenerational epigenetic inheritance: Myths and mechanisms. Cell 2014, 157, 95–109. [Google Scholar] [CrossRef] [PubMed]
- Szyf, M. Nongenetic inheritance and transgenerational epigenetics. Trends Mol. Med. 2015, 21, 134–144. [Google Scholar] [CrossRef] [PubMed]
- Manjrekar, J. Epigenetic inheritance, prions and evolution. J. Genet. 2017, 96, 445–456. [Google Scholar] [CrossRef] [PubMed]
- Ruden, D.M.; Lu, X. Hsp90 affecting chromatin remodeling might explain transgenerational epigenetic inheritance in Drosophila. Curr. Genom. 2008, 9, 500–508. [Google Scholar] [CrossRef] [PubMed]
- Angelova, M.T.; Dimitrova, D.G.; Dinges, N.; Lence, T.; Worpenberg, L.; Carre, C.; Roignant, J.Y. The emerging field of epitranscriptomics in neurodevelopmental and neuronal disorders. Front. Bioeng. Biotechnol. 2018, 6, 46. [Google Scholar] [CrossRef] [PubMed]
- Dimitrova, D.G.; Teysset, L.; Carre, C. RNA 2′-O-methylation (Nm) modification in human diseases. Genes (Basel) 2019, 10, 117. [Google Scholar] [CrossRef] [PubMed]
- Kiani, J.; Grandjean, V.; Liebers, R.; Tuorto, F.; Ghanbarian, H.; Lyko, F.; Cuzin, F.; Rassoulzadegan, M. RNA-mediated epigenetic heredity requires the cytosine methyltransferase Dnmt2. PLoS Genet. 2013, 9, e1003498. [Google Scholar] [CrossRef]
- Ben Maamar, M.; Sadler-Riggleman, I.; Beck, D.; Skinner, M.K. Epigenetic transgenerational inheritance of altered sperm histone retention sites. Sci. Rep. 2018, 8, 5308. [Google Scholar] [CrossRef]
- Skinner, M.K.; Ben Maamar, M.; Sadler-Riggleman, I.; Beck, D.; Nilsson, E.; McBirney, M.; Klukovich, R.; Xie, Y.; Tang, C.; Yan, W. Alterations in sperm DNA methylation, non-coding RNA and histone retention associate with DDT-induced epigenetic transgenerational inheritance of disease. Epigenet. Chromatin 2018, 11, 8. [Google Scholar] [CrossRef] [PubMed]
- King, S.E.; McBirney, M.; Beck, D.; Sadler-Riggleman, I.; Nilsson, E.; Skinner, M.K. Sperm epimutation biomarkers of obesity and pathologies following DDT induced epigenetic transgenerational inheritance of disease. Environ. Epigenet. 2019, 5, dvz008. [Google Scholar] [CrossRef] [PubMed]
- Klukovich, R.; Nilsson, E.; Sadler-Riggleman, I.; Beck, D.; Xie, Y.; Yan, W.; Skinner, M.K. Environmental toxicant induced epigenetic transgenerational inheritance of prostate pathology and stromal-epithelial cell epigenome and transcriptome alterations: Ancestral origins of prostate disease. Sci. Rep. 2019, 9, 2209. [Google Scholar] [CrossRef] [PubMed]
- Kubsad, D.; Nilsson, E.E.; King, S.E.; Sadler-Riggleman, I.; Beck, D.; Skinner, M.K. Assessment of glyphosate induced epigenetic transgenerational inheritance of pathologies and sperm epimutations: Generational toxicology. Sci. Rep. 2019, 9, 6372. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, E.; Klukovich, R.; Sadler-Riggleman, I.; Beck, D.; Xie, Y.; Yan, W.; Skinner, M.K. Environmental toxicant induced epigenetic transgenerational inheritance of ovarian pathology and granulosa cell epigenome and transcriptome alterations: Ancestral origins of polycystic ovarian syndrome and primary ovarian insufiency. Epigenetics 2018, 13, 875–895. [Google Scholar] [CrossRef] [PubMed]
- Casier, K.; Delmarre, V.; Gueguen, N.; Hermant, C.; Viode, E.; Vaury, C.; Ronsseray, S.; Brasset, E.; Teysset, L.; Boivin, A. Environmentally-induced epigenetic conversion of a piRNA cluster. Elife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Yu, R.; Wang, X.; Moazed, D. Epigenetic inheritance mediated by coupling of RNAi and histone H3K9 methylation. Nature 2018, 558, 615–619. [Google Scholar] [CrossRef]
- Guerrero-Bosagna, C.; Settles, M.; Lucker, B.; Skinner, M.K. Epigenetic transgenerational actions of vinclozolin on promoter regions of the sperm epigenome. PLoS ONE 2010, 5, e13100. [Google Scholar] [CrossRef]
- Skinner, M.K.; Nilsson, E.; Sadler-Riggleman, I.; Beck, D.; Ben Maamar, M.; McCarrey, J.R. Transgenerational sperm DNA methylation epimutation developmental origins following ancestral vinclozolin exposure. Epigenetics 2019, 14, 721–739. [Google Scholar] [CrossRef]
- Schuster, A.; Skinner, M.K.; Yan, W. Ancestral vinclozolin exposure alters the epigenetic transgenerational inheritance of sperm small noncoding RNAs. Environ. Epigenet 2016, 2, dvw001. [Google Scholar]
- Brieno-Enriquez, M.A.; Garcia-Lopez, J.; Cardenas, D.B.; Guibert, S.; Cleroux, E.; Ded, L.; Hourcade Jde, D.; Peknicova, J.; Weber, M.; Del Mazo, J. Exposure to endocrine disruptor induces transgenerational epigenetic deregulation of microRNAs in primordial germ cells. PLoS ONE 2015, 10, e0124296. [Google Scholar] [CrossRef] [PubMed]
- Skinner, M.K.; Manikkam, M.; Tracey, R.; Guerrero-Bosagna, C.; Haque, M.; Nilsson, E.E. Ancestral dichlorodiphenyltrichloroethane (DDT) exposure promotes epigenetic transgenerational inheritance of obesity. BMC Med. 2013, 11, 228. [Google Scholar] [CrossRef] [PubMed]
- Ben Maamar, M.; Nilsson, E.; Sadler-Riggleman, I.; Beck, D.; McCarrey, J.R.; Skinner, M.K. Developmental origins of transgenerational sperm DNA methylation epimutations following ancestral DDT exposure. Dev. Biol. 2019, 445, 280–293. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, E.; Larsen, G.; Manikkam, M.; Guerrero-Bosagna, C.; Savenkova, M.I.; Skinner, M.K. Environmentally induced epigenetic transgenerational inheritance of ovarian disease. PLoS ONE 2012, 7, e36129. [Google Scholar] [CrossRef] [PubMed]
- Manikkam, M.; Guerrero-Bosagna, C.; Tracey, R.; Haque, M.M.; Skinner, M.K. Transgenerational actions of environmental compounds on reproductive disease and identification of epigenetic biomarkers of ancestral exposures. PLoS ONE 2012, 7, e31901. [Google Scholar] [CrossRef]
- Manikkam, M.; Tracey, R.; Guerrero-Bosagna, C.; Skinner, M.K. Plastics derived endocrine disruptors (BPA, DEHP and DBP) induce epigenetic transgenerational inheritance of obesity, reproductive disease and sperm epimutations. PLoS ONE 2013, 8, e55387. [Google Scholar] [CrossRef] [PubMed]
- Gely-Pernot, A.; Hao, C.; Legoff, L.; Multigner, L.; D’Cruz, S.C.; Kervarrec, C.; Jegou, B.; Tevosian, S.; Smagulova, F. Gestational exposure to chlordecone promotes transgenerational changes in the murine reproductive system of males. Sci. Rep. 2018, 8, 10274. [Google Scholar] [CrossRef]
- Tracey, R.; Manikkam, M.; Guerrero-Bosagna, C.; Skinner, M.K. Hydrocarbons (jet fuel JP-8) induce epigenetic transgenerational inheritance of obesity, reproductive disease and sperm epimutations. Reprod. Toxicol. 2013, 36, 104–116. [Google Scholar] [CrossRef]
- Camacho, J.; Truong, L.; Kurt, Z.; Chen, Y.W.; Morselli, M.; Gutierrez, G.; Pellegrini, M.; Yang, X.; Allard, P. The memory of environmental chemical exposure in C. elegans is dependent on the jumonji demethylases jmjd-2 and jmjd-3/utx-1. Cell Rep. 2018, 23, 2392–2404. [Google Scholar] [CrossRef]
- Ou, X.; Zhang, Y.; Xu, C.; Lin, X.; Zang, Q.; Zhuang, T.; Jiang, L.; von Wettstein, D.; Liu, B. Transgenerational inheritance of modified DNA methylation patterns and enhanced tolerance induced by heavy metal stress in rice (Oryza sativa L.). PLoS ONE 2012, 7, e41143. [Google Scholar] [CrossRef]
- Franklin, T.B.; Russig, H.; Weiss, I.C.; Graff, J.; Linder, N.; Michalon, A.; Vizi, S.; Mansuy, I.M. Epigenetic transmission of the impact of early stress across generations. Biol. Psychiatry 2010, 68, 408–415. [Google Scholar] [CrossRef] [PubMed]
- Dias, B.G.; Ressler, K.J. Parental olfactory experience influences behavior and neural structure in subsequent generations. Nat. Neurosci. 2014, 17, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Hoile, S.P.; Lillycrop, K.A.; Thomas, N.A.; Hanson, M.A.; Burdge, G.C. Dietary protein restriction during F0 pregnancy in rats induces transgenerational changes in the hepatic transcriptome in female offspring. PLoS ONE 2011, 6, e21668. [Google Scholar] [CrossRef] [PubMed]
- Rechavi, O.; Houri-Ze’evi, L.; Anava, S.; Goh, W.S.S.; Kerk, S.Y.; Hannon, G.J.; Hobert, O. Starvation-induced transgenerational inheritance of small RNAs in C. elegans. Cell 2014, 158, 277–287. [Google Scholar] [CrossRef] [PubMed]
- Ashe, A.; Sapetschnig, A.; Weick, E.-M.; Mitchell, J.; Bagijn, M.P.; Cording, A.C.; Doebley, A.-L.; Goldstein, L.D.; Lehrbach, N.J.; Le Pen, J.; et al. piRNAs can trigger a multigenerational epigenetic memory in the germline of C. elegans. Cell 2012, 150, 88–99. [Google Scholar] [CrossRef] [PubMed]
- Moore, R.S.; Kaletsky, R.; Murphy, C.T. Piwi/PRG-1 argonaute and TGF-β mediate transgenerational learned pathogenic avoidance. Cell 2019, 177, 1827–1841.e12. [Google Scholar] [CrossRef]
- Govorko, D.; Bekdash, R.A.; Zhang, C.; Sarkar, D.K. Male germline transmits fetal alcohol adverse effect on hypothalamic proopiomelanocortin gene across generations. Biol. Psychiatry 2012, 72, 378–388. [Google Scholar] [CrossRef]
- Kou, H.P.; Li, Y.; Song, X.X.; Ou, X.F.; Xing, S.C.; Ma, J.; Von Wettstein, D.; Liu, B. Heritable alteration in DNA methylation induced by nitrogen-deficiency stress accompanies enhanced tolerance by progenies to the stress in rice (Oryza sativa L.). J. Plant. Physiol. 2011, 168, 1685–1693. [Google Scholar] [CrossRef]
- Zheng, X.; Chen, L.; Li, M.; Lou, Q.; Xia, H.; Wang, P.; Li, T.; Liu, H.; Luo, L. Transgenerational variations in DNA methylation induced by drought stress in two rice varieties with distinguished difference to drought resistance. PLoS ONE 2013, 8, e80253. [Google Scholar] [CrossRef]
- Jeremias, G.; Barbosa, J.; Marques, S.M.; De Schamphelaere, K.A.C.; Van Nieuwerburgh, F.; Deforce, D.; Goncalves, F.J.M.; Pereira, J.L.; Asselman, J. Transgenerational inheritance of DNA hypomethylation in daphnia magna in response to salinity stress. Environ. Sci. Technol. 2018, 52, 10114–10123. [Google Scholar] [CrossRef]
- Seong, K.-H.; Li, D.; Shimizu, H.; Nakamura, R.; Ishii, S. Inheritance of stress-induced, ATF-2-dependent epigenetic change. Cell 2011, 145, 1049–1061. [Google Scholar] [CrossRef] [PubMed]
- Klosin, A.; Casas, E.; Hidalgo-Carcedo, C.; Vavouri, T.; Lehner, B. Transgenerational transmission of environmental information in C. elegans. Science 2017, 356, 320–323. [Google Scholar] [CrossRef] [PubMed]
- Ni, J.Z.; Kalinava, N.; Chen, E.; Huang, A.; Trinh, T.; Gu, S.G. A transgenerational role of the germline nuclear RNAi pathway in repressing heat stress-induced transcriptional activation in C. elegans. Epigenet. Chromatin 2016, 9, 3. [Google Scholar] [CrossRef] [PubMed]
- Schott, D.; Yanai, I.; Hunter, C.P. Natural RNA interference directs a heritable response to the environment. Sci. Rep. 2014, 4, 7387. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Jacobsen, S.E.; Reik, W. Epigenetic reprogramming in plant and animal development. Science 2010, 330, 622–627. [Google Scholar] [CrossRef]
- Hackett, J.A.; Zylicz, J.J.; Surani, M.A. Parallel mechanisms of epigenetic reprogramming in the germline. Trends Genet. 2012, 28, 164–174. [Google Scholar] [CrossRef]
- Shen, H.; Xu, W.; Lan, F. Histone lysine demethylases in mammalian embryonic development. Exp. Mol. Med. 2017, 49, e325. [Google Scholar] [CrossRef]
- Seisenberger, S.; Andrews, S.; Krueger, F.; Arand, J.; Walter, J.; Santos, F.; Popp, C.; Thienpont, B.; Dean, W.; Reik, W. The dynamics of genome-wide DNA methylation reprogramming in mouse primordial germ cells. Mol. Cell 2012, 48, 849–862. [Google Scholar] [CrossRef]
- Chen, T.; Ueda, Y.; Dodge, J.E.; Wang, Z.; Li, E. Establishment and maintenance of genomic methylation patterns in mouse embryonic stem cells by Dnmt3a and Dnmt3b. Mol. Cell Biol. 2003, 23, 5594–5605. [Google Scholar] [CrossRef]
- Gaydos, L.J.; Wang, W.; Strome, S. H3K27me and PRC2 transmit a memory of repression across generations and during development. Science 2014, 345, 1515–1518. [Google Scholar] [CrossRef]
- Maenohara, S.; Unoki, M.; Toh, H.; Ohishi, H.; Sharif, J.; Koseki, H.; Sasaki, H. Role of UHRF1 in de novo DNA methylation in oocytes and maintenance methylation in preimplantation embryos. PLoS Genet. 2017, 13, e1007042. [Google Scholar] [CrossRef] [PubMed]
- Houri-Ze’evi, L.; Korem, Y.; Sheftel, H.; Faigenbloom, L.; Toker, I.A.; Dagan, Y.; Awad, L.; Degani, L.; Alon, U.; Rechavi, O. A tunable mechanism determines the duration of the transgenerational small RNA inheritance in C. elegans. Cell 2016, 165, 88–99. [Google Scholar] [CrossRef] [PubMed]
- Alcazar, R.M.; Lin, R.; Fire, A.Z. Transmission dynamics of heritable silencing induced by double-stranded RNA in Caenorhabditis elegans. Genetics 2008, 180, 1275–1288. [Google Scholar] [CrossRef] [PubMed]
- Rechavi, O.; Lev, I. Principles of transgenerational small RNA inheritance in Caenorhabditis elegans. Curr. Biol. 2017, 27, R720–R730. [Google Scholar] [CrossRef] [PubMed]
- Minkina, O.; Hunter, C.P. Intergenerational transmission of gene regulatory information in Caenorhabditis elegans. Trends Genet. 2018, 34, 54–64. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Guang, S.; Feng, X. Distinct nuclear and cytoplasmic machineries cooperatively promote the inheritance of RNAi in Caenorhabditis elegans: The inheritance of RNAi. Biol. Cell 2018, 110, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Sapetschnig, A.; Sarkies, P.; Lehrbach, N.J.; Miska, E.A. Tertiary siRNAs mediate paramutation in C. elegans. PLoS Genet. 2015, 11, e1005078. [Google Scholar] [CrossRef] [PubMed]
- Houri-Ze’evi, L.; Rechavi, O. Plastic germline reprogramming of heritable small RNAs enables maintenance or erasure of epigenetic memories. RNA Biol. 2016, 13, 1212–1217. [Google Scholar] [CrossRef]
- Houri-Zeevi, L.; Rechavi, O. A matter of time: Small RNAs regulate the duration of epigenetic inheritance. Trends Genet. 2017, 33, 46–57. [Google Scholar] [CrossRef]
- Zhuang, J.J.; Hunter, C.P. The influence of competition among C. elegans small RNA pathways on development. Genes (Basel) 2012, 3, 671–685. [Google Scholar] [CrossRef]
- de Vanssay, A.; Bougé, A.-L.; Boivin, A.; Hermant, C.; Teysset, L.; Delmarre, V.; Antoniewski, C.; Ronsseray, S. Paramutation in Drosophila linked to emergence of a piRNA-producing locus. Nature 2012, 490, 112–115. [Google Scholar] [CrossRef] [PubMed]
- Hermant, C.; Boivin, A.; Teysset, L.; Delmarre, V.; Asif-Laidin, A.; van den Beek, M.; Antoniewski, C.; Ronsseray, S. Paramutation in Drosophila requires both nuclear and cytoplasmic actors of the piRNA pathway and induces cis-spreading of piRNA production. Genetics 2015, 201, 1381–1396. [Google Scholar] [CrossRef] [PubMed]
- Ghildiyal, M.; Zamore, P.D. Small silencing RNAs: An expanding universe. Nat. Rev. Genet. 2009, 10, 94–108. [Google Scholar] [CrossRef] [PubMed]
- Brennecke, J.; Aravin, A.A.; Stark, A.; Dus, M.; Kellis, M.; Sachidanandam, R.; Hannon, G.J. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 2007, 128, 1089–1103. [Google Scholar] [CrossRef] [PubMed]
- Gunawardane, L.S.; Saito, K.; Nishida, K.M.; Miyoshi, K.; Kawamura, Y.; Nagami, T.; Siomi, H.; Siomi, M.C. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 2007, 315, 1587–1590. [Google Scholar] [CrossRef]
- Mohn, F.; Sienski, G.; Handler, D.; Brennecke, J. The rhino-deadlock-cutoff complex licenses noncanonical transcription of dual-strand piRNA clusters in Drosophila. Cell 2014, 157, 1364–1379. [Google Scholar] [CrossRef]
- Andersen, P.R.; Tirian, L.; Vunjak, M.; Brennecke, J. A heterochromatin-dependent transcription machinery drives piRNA expression. Nature 2017, 549, 54–59. [Google Scholar] [CrossRef]
- ElMaghraby, M.F.; Andersen, P.R.; Pühringer, F.; Hohmann, U.; Meixner, K.; Lendl, T.; Tirian, L.; Brennecke, J. A heterochromatin-specific RNA export pathway facilitates piRNA production. Cell 2019, 178, 964–979.e20. [Google Scholar] [CrossRef]
- Kneuss, E.; Munafò, M.; Eastwood, E.L.; Deumer, U.-S.; Preall, J.B.; Hannon, G.J.; Czech, B. Specialization of the Drosophila nuclear export family protein Nxf3 for piRNA precursor export. Genes Dev. 2019, 33, 1208–1220. [Google Scholar] [CrossRef]
- Ipsaro, J.J.; Haase, A.D.; Knott, S.R.; Joshua-Tor, L.; Hannon, G.J. The structural biochemistry of Zucchini implicates it as a nuclease in piRNA biogenesis. Nature 2012, 491, 279–283. [Google Scholar] [CrossRef]
- Nishimasu, H.; Ishizu, H.; Saito, K.; Fukuhara, S.; Kamatani, M.K.; Bonnefond, L.; Matsumoto, N.; Nishizawa, T.; Nakanaga, K.; Aoki, J.; et al. Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature 2012, 491, 284–287. [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Wang, J.; Xu, J.; Zhang, Z.; Koppetsch, B.S.; Schultz, N.; Vreven, T.; Meignin, C.; Davis, I.; Zamore, P.D.; et al. UAP56 couples piRNA clusters to the perinuclear transposon silencing machinery. Cell 2012, 151, 871–884. [Google Scholar] [CrossRef] [PubMed]
- Hur, J.K.; Luo, Y.; Moon, S.; Ninova, M.; Marinov, G.K.; Chung, Y.D.; Aravin, A.A. Splicing-independent loading of TREX on nascent RNA is required for efficient expression of dual-strand piRNA clusters in Drosophila. Genes Dev. 2016, 30, 840–855. [Google Scholar] [CrossRef] [PubMed]
- Batki, J.; Schnabl, J.; Wang, J.; Handler, D.; Andreev, V.I.; Stieger, C.E.; Novatchkova, M.; Lampersberger, L.; Kauneckaite, K.; Xie, W.; et al. The nascent RNA binding complex SFiNX licenses piRNA-guided heterochromatin formation. Nat. Struct. Mol. Biol. 2019, 26, 720–731. [Google Scholar] [CrossRef] [PubMed]
- Brower-Toland, B.; Findley, S.D.; Jiang, L.; Liu, L.; Yin, H.; Dus, M.; Zhou, P.; Elgin, S.C.; Lin, H. Drosophila PIWI associates with chromatin and interacts directly with HP1a. Genes Dev. 2007, 21, 2300–2311. [Google Scholar] [CrossRef] [PubMed]
- Fabry, M.H.; Ciabrelli, F.; Munafò, M.; Eastwood, E.L.; Kneuss, E.; Falciatori, I.; Falconio, F.A.; Hannon, G.J.; Czech, B. piRNA-guided co-transcriptional silencing coopts nuclear export factors. eLife 2019, 8, e47999. [Google Scholar] [CrossRef]
- Klenov, M.S.; Sokolova, O.A.; Yakushev, E.Y.; Stolyarenko, A.D.; Mikhaleva, E.A.; Lavrov, S.A.; Gvozdev, V.A. Separation of stem cell maintenance and transposon silencing functions of Piwi protein. Proc. Natl. Acad. Sci. USA 2011, 108, 18760–18765. [Google Scholar] [CrossRef]
- Klenov, M.S.; Lavrov, S.A.; Korbut, A.P.; Stolyarenko, A.D.; Yakushev, E.Y.; Reuter, M.; Pillai, R.S.; Gvozdev, V.A. Impact of nuclear Piwi elimination on chromatin state in Drosophila melanogaster ovaries. Nucleic Acids Res. 2014, 42, 6208–6218. [Google Scholar] [CrossRef]
- Le Thomas, A.; Rogers, A.K.; Webster, A.; Marinov, G.K.; Liao, S.E.; Perkins, E.M.; Hur, J.K.; Aravin, A.A.; Tóth, K.F. Piwi induces piRNA-guided transcriptional silencing and establishment of a repressive chromatin state. Genes Dev. 2013, 27, 390–399. [Google Scholar] [CrossRef]
- Rangan, P.; Malone, C.D.; Navarro, C.; Newbold, S.P.; Hayes, P.S.; Sachidanandam, R.; Hannon, G.J.; Lehmann, R. piRNA production requires heterochromatin formation in Drosophila. Curr. Biol. 2011, 21, 1373–1379. [Google Scholar] [CrossRef]
- Murano, K.; Iwasaki, Y.W.; Ishizu, H.; Mashiko, A.; Shibuya, A.; Kondo, S.; Adachi, S.; Suzuki, S.; Saito, K.; Natsume, T.; et al. Nuclear RNA export factor variant initiates piRNA-guided co-transcriptional silencing. EMBO J. 2019, 38. [Google Scholar] [CrossRef] [PubMed]
- Marie, P.P.; Ronsseray, S.; Boivin, A. From embryo to adult: piRNA-mediated silencing throughout germline development in Drosophila. G3 (Bethesda) 2017, 7, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Reiss, D.; Josse, T.; Anxolabehere, D.; Ronsseray, S. aubergine mutations in Drosophila melanogaster impair P cytotype determination by telomeric P elements inserted in heterochromatin. Mol. Genet. Genom. 2004, 272, 336–343. [Google Scholar] [CrossRef] [PubMed]
- Klattenhoff, C.; Bratu, D.P.; McGinnis-Schultz, N.; Koppetsch, B.S.; Cook, H.A.; Theurkauf, W.E. Drosophila rasiRNA pathway mutations disrupt embryonic axis specification through activation of an ATR/Chk2 DNA damage response. Dev. Cell 2007, 12, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Molla-Herman, A.; Valles, A.M.; Ganem-Elbaz, C.; Antoniewski, C.; Huynh, J.-R. tRNA processing defects induce replication stress and Chk2-dependent disruption of piRNA transcription. EMBO J. 2015, 34, 3009–3027. [Google Scholar] [CrossRef] [PubMed]
- Grimson, A.; Srivastava, M.; Fahey, B.; Woodcroft, B.J.; Chiang, H.R.; King, N.; Degnan, B.M.; Rokhsar, D.S.; Bartel, D.P. Early origins and evolution of microRNAs and Piwi-interacting RNAs in animals. Nature 2008, 455, 1193–1197. [Google Scholar] [CrossRef] [PubMed]
- Zuo, L.; Wang, Z.; Tan, Y.; Chen, X.; Luo, X. piRNAs and their functions in the brain. Int. J. Hum. Genet. 2016, 16, 53–60. [Google Scholar] [CrossRef]
- Lewis, S.H.; Quarles, K.A.; Yang, Y.; Tanguy, M.; Frezal, L.; Smith, S.A.; Sharma, P.P.; Cordaux, R.; Gilbert, C.; Giraud, I.; et al. Pan-arthropod analysis reveals somatic piRNAs as an ancestral defence against transposable elements. Nat. Ecol. Evol. 2018, 2, 174–181. [Google Scholar] [CrossRef] [PubMed]
- Jehn, J.; Gebert, D.; Pipilescu, F.; Stern, S.; Kiefer, J.S.T.; Hewel, C.; Rosenkranz, D. PIWI genes and piRNAs are ubiquitously expressed in mollusks and show patterns of lineage-specific adaptation. Commun. Biol. 2018, 1, 137. [Google Scholar] [CrossRef]
- Perera, B.P.U.; Tsai, Z.T.-Y.; Colwell, M.L.; Jones, T.R.; Goodrich, J.M.; Wang, K.; Sartor, M.A.; Faulk, C.; Dolinoy, D.C. Somatic expression of piRNA and associated machinery in the mouse identifies short, tissue-specific piRNA. Epigenetics 2019, 14, 504–521. [Google Scholar] [CrossRef]
- Brennecke, J.; Malone, C.D.; Aravin, A.A.; Sachidanandam, R.; Stark, A.; Hannon, G.J. An epigenetic role for maternally inherited piRNAs in transposon silencing. Science 2008, 322, 1387–1392. [Google Scholar] [CrossRef] [PubMed]
- Saito, K.; Inagaki, S.; Mituyama, T.; Kawamura, Y.; Ono, Y.; Sakota, E.; Kotani, H.; Asai, K.; Siomi, H.; Siomi, M.C. A regulatory circuit for piwi by the large Maf gene traffic jam in Drosophila. Nature 2009, 461, 1296–1299. [Google Scholar] [CrossRef] [PubMed]
- Jones, B.C.; Wood, J.G.; Chang, C.; Tam, A.D.; Franklin, M.J.; Siegel, E.R.; Helfand, S.L. A somatic piRNA pathway in the Drosophila fat body ensures metabolic homeostasis and normal lifespan. Nat. Commun. 2016, 7, 13856. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Samimi, H.; Gamez, M.; Zare, H.; Frost, B. Pathogenic tau-induced piRNA depletion promotes neuronal death through transposable element dysregulation in neurodegenerative tauopathies. Nat. Neurosci. 2018, 21, 1038–1048. [Google Scholar] [CrossRef] [PubMed]
- van den Beek, M.; da Silva, B.; Pouch, J.; Ali Chaouche, M.E.A.; Carre, C.; Antoniewski, C. Dual-layer transposon repression in heads of Drosophila melanogaster. RNA 2018, 24, 1749–1760. [Google Scholar] [CrossRef]
- Robine, N.; Lau, N.C.; Balla, S.; Jin, Z.; Okamura, K.; Kuramochi-Miyagawa, S.; Blower, M.D.; Lai, E.C. A broadly conserved pathway generates 3′UTR-directed primary piRNAs. Curr. Biol. 2009, 19, 2066–2076. [Google Scholar] [CrossRef]
- Rouget, C.; Papin, C.; Boureux, A.; Meunier, A.-C.; Franco, B.; Robine, N.; Lai, E.C.; Pelisson, A.; Simonelig, M. Maternal mRNA deadenylation and decay by the piRNA pathway in the early Drosophila embryo. Nature 2010, 467, 1128–1132. [Google Scholar] [CrossRef]
- Barckmann, B.; Pierson, S.; Dufourt, J.; Papin, C.; Armenise, C.; Port, F.; Grentzinger, T.; Chambeyron, S.; Baronian, G.; Desvignes, J.-P.; et al. Aubergine iCLIP reveals piRNA-dependent decay of mRNAs involved in germ cell development in the early embryo. Cell Rep. 2015, 12, 1205–1216. [Google Scholar] [CrossRef]
- Rajasethupathy, P.; Antonov, I.; Sheridan, R.; Frey, S.; Sander, C.; Tuschl, T.; Kandel, E.R. A role for neuronal piRNAs in the epigenetic control of memory-related synaptic plasticity. Cell 2012, 149, 693–707. [Google Scholar] [CrossRef]
- Kiuchi, T.; Koga, H.; Kawamoto, M.; Shoji, K.; Sakai, H.; Arai, Y.; Ishihara, G.; Kawaoka, S.; Sugano, S.; Shimada, T.; et al. A single female-specific piRNA is the primary determiner of sex in the silkworm. Nature 2014, 509, 633–636. [Google Scholar] [CrossRef]
- Roche, S.E.; Rio, D.C. Trans-silencing by P elements inserted in subtelomeric heterochromatin involves the Drosophila polycomb group gene, enhancer of zeste. Genetics 1998, 149, 1839–1855. [Google Scholar] [PubMed]
- Josse, T.; Teysset, L.; Todeschini, A.-L.; Sidor, C.M.; Anxolabéhère, D.; Ronsseray, S. Telomeric trans-silencing: An epigenetic repression combining RNA silencing and heterochromatin formation. PLoS Genet. 2007, 3, 1633–1643. [Google Scholar] [CrossRef] [PubMed]
- Josse, T.; Maurel-Zaffran, C.; de Vanssay, A.; Teysset, L.; Todeschini, A.-L.; Delmarre, V.; Chaminade, N.; Anxolabehere, D.; Ronsseray, S. Telomeric trans-silencing in Drosophila melanogaster: Tissue specificity, development and functional interactions between non-homologous telomeres. PLoS ONE 2008, 3, e3249. [Google Scholar] [CrossRef] [PubMed]
- Todeschini, A.-L.; Teysset, L.; Delmarre, V.; Ronsseray, S. The epigenetic trans-silencing effect in Drosophila involves maternally-transmitted small RNAs whose production depends on the piRNA pathway and HP1. PLoS ONE 2010, 5, e11032. [Google Scholar] [CrossRef] [PubMed]
- Poyhonen, M.; de Vanssay, A.; Delmarre, V.; Hermant, C.; Todeschini, A.L.; Teysset, L.; Ronsseray, S. Homology-dependent silencing by an exogenous sequence in the Drosophila germline. G3 (Bethesda) 2012, 2, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Muerdter, F.; Olovnikov, I.; Molaro, A.; Rozhkov, N.V.; Czech, B.; Gordon, A.; Hannon, G.J.; Aravin, A.A. Production of artificial piRNAs in flies and mice. RNA 2012, 18, 42–52. [Google Scholar] [CrossRef]
- Shirayama, M.; Seth, M.; Lee, H.C.; Gu, W.; Ishidate, T.; Conte, D., Jr.; Mello, C.C. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 2012, 150, 65–77. [Google Scholar] [CrossRef] [PubMed]
- Chandler, V.L. Paramutation: From maize to mice. Cell 2007, 128, 641–645. [Google Scholar] [CrossRef]
- Hollick, J.B. Paramutation: A trans-homolog interaction affecting heritable gene regulation. Curr. Opin. Plant. Biol. 2012, 15, 536–543. [Google Scholar] [CrossRef]
- Ronsseray, S. Paramutation phenomena in non-vertebrate animals. Semin. Cell Dev. Biol. 2015, 44, 39–46. [Google Scholar] [CrossRef]
- Stuwe, E.; Toth, K.F.; Aravin, A.A. Small but sturdy: Small RNAs in cellular memory and epigenetics. Genes Dev. 2014, 28, 423–431. [Google Scholar] [CrossRef] [PubMed]
- Almeida, M.V.; Andrade-Navarro, M.A.; Ketting, R.F. Function and evolution of nematode RNAi pathways. Noncoding RNA 2019, 5, 8. [Google Scholar] [CrossRef] [PubMed]
- Batista, P.J.; Ruby, J.G.; Claycomb, J.M.; Chiang, R.; Fahlgren, N.; Kasschau, K.D.; Chaves, D.A.; Gu, W.; Vasale, J.J.; Duan, S.; et al. PRG-1 and 21U-RNAs interact to form the piRNA complex required for fertility in C. elegans. Mol. Cell 2008, 31, 67–78. [Google Scholar] [CrossRef] [PubMed]
- Das, P.P.; Bagijn, M.P.; Goldstein, L.D.; Woolford, J.R.; Lehrbach, N.J.; Sapetschnig, A.; Buhecha, H.R.; Gilchrist, M.J.; Howe, K.L.; Stark, R.; et al. Piwi and piRNAs act upstream of an endogenous siRNA pathway to suppress Tc3 transposon mobility in the Caenorhabditis elegans germline. Mol. Cell 2008, 31, 79–90. [Google Scholar] [CrossRef] [PubMed]
- Antoniewski, C.; Carré, C. New rules for regulation of genes by piRNAs in C. elegans. Non-coding RNA Investig. 2018, 2, 33. [Google Scholar] [CrossRef]
- Shen, E.-Z.; Chen, H.; Ozturk, A.R.; Tu, S.; Shirayama, M.; Tang, W.; Ding, Y.-H.; Dai, S.-Y.; Weng, Z.; Mello, C.C. Identification of piRNA binding sites reveals the argonaute regulatory landscape of the C. elegans germline. Cell 2018, 172, 937–951. [Google Scholar] [CrossRef]
- Zhang, D.; Tu, S.; Stubna, M.; Wu, W.-S.; Huang, W.-C.; Weng, Z.; Lee, H.-C. The piRNA targeting rules and the resistance to piRNA silencing in endogenous genes. Science 2018, 359, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Buckley, B.A.; Burkhart, K.B.; Gu, S.G.; Spracklin, G.; Kershner, A.; Fritz, H.; Kimble, J.; Fire, A.; Kennedy, S. A nuclear Argonaute promotes multigenerational epigenetic inheritance and germline immortality. Nature 2012, 489, 447–451. [Google Scholar] [CrossRef]
- Guang, S.; Bochner, A.F.; Burkhart, K.B.; Burton, N.; Pavelec, D.M.; Kennedy, S. Small regulatory RNAs inhibit RNA polymerase II during the elongation phase of transcription. Nature 2010, 465, 1097–1101. [Google Scholar] [CrossRef]
- Kalinava, N.; Ni, J.Z.; Gajic, Z.; Kim, M.; Ushakov, H.; Gu, S.G. C. elegans heterochromatin factor SET-32 plays an essential role in transgenerational establishment of nuclear RNAi-mediated epigenetic silencing. Cell Rep. 2018, 25, 2273–2284. [Google Scholar] [CrossRef]
- Pak, J.; Fire, A. Distinct populations of primary and secondary effectors during RNAi in C. elegans. Science 2007, 315, 241–244. [Google Scholar] [CrossRef] [PubMed]
- Woodhouse, R.M.; Buchmann, G.; Hoe, M.; Harney, D.J.; Low, J.K.K.; Larance, M.; Boag, P.R.; Ashe, A. Chromatin modifiers SET-25 and SET-32 are required for establishment but not long-term maintenance of transgenerational epigenetic inheritance. Cell Rep. 2018, 25, 2259–2272. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Feng, X.; Chen, X.; Weng, C.; Yan, Q.; Xu, T.; Hong, M.; Guang, S. A cytoplasmic argonaute protein promotes the inheritance of RNAi. Cell Rep. 2018, 23, 2482–2494. [Google Scholar] [CrossRef] [PubMed]
- Lev, I.; Gingold, H.; Rechavi, O. H3K9me3 is required for inheritance of small RNAs that target a unique subset of newly evolved genes. eLife 2019, 8. [Google Scholar] [CrossRef] [PubMed]
- Posner, R.; Toker, I.A.; Antonova, O.; Star, E.; Anava, S.; Azmon, E.; Hendricks, M.; Bracha, S.; Gingold, H.; Rechavi, O. Neuronal small RNAs control behavior transgenerationally. Cell 2019, 177, 1814–1826. [Google Scholar] [CrossRef] [PubMed]
- Bagijn, M.P.; Goldstein, L.D.; Sapetschnig, A.; Weick, E.-M.; Bouasker, S.; Lehrbach, N.J.; Simard, M.J.; Miska, E.A. Function, targets, and evolution of Caenorhabditis elegans piRNAs. Science 2012, 337, 574–578. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.-C.; Gu, W.; Shirayama, M.; Youngman, E.; Conte, D.; Mello, C.C. C. elegans piRNAs mediate the genome-wide surveillance of germline transcripts. Cell 2012, 150, 78–87. [Google Scholar] [CrossRef]
- Gent, J.I.; Lamm, A.T.; Pavelec, D.M.; Maniar, J.M.; Parameswaran, P.; Tao, L.; Kennedy, S.; Fire, A.Z. Distinct Phases of siRNA synthesis in an endogenous RNAi pathway in C. elegans soma. Mol. Cell 2010, 37, 679–689. [Google Scholar] [CrossRef]
- Gapp, K.; Jawaid, A.; Sarkies, P.; Bohacek, J.; Pelczar, P.; Prados, J.; Farinelli, L.; Miska, E.; Mansuy, I.M. Implication of sperm RNAs in transgenerational inheritance of the effects of early trauma in mice. Nat. Neurosci. 2014, 17, 667–669. [Google Scholar] [CrossRef]
- Grandjean, V.; Fourre, S.; De Abreu, D.A.F.; Derieppe, M.-A.; Remy, J.-J.; Rassoulzadegan, M. RNA-mediated paternal heredity of diet-induced obesity and metabolic disorders. Sci. Rep. 2015, 5, 18193. [Google Scholar] [CrossRef]
- de Castro Barbosa, T.; Ingerslev, L.R.; Alm, P.S.; Versteyhe, S.; Massart, J.; Rasmussen, M.; Donkin, I.; Sjogren, R.; Mudry, J.M.; Vetterli, L.; et al. High-fat diet reprograms the epigenome of rat spermatozoa and transgenerationally affects metabolism of the offspring. Mol. Metab. 2016, 5, 184–197. [Google Scholar] [CrossRef] [PubMed]
- Belicard, T.; Jareosettasin, P.; Sarkies, P. The piRNA pathway responds to environmental signals to establish intergenerational adaptation to stress. BMC Biol. 2018, 16, 103. [Google Scholar] [CrossRef] [PubMed]
- Meisel, J.D.; Panda, O.; Mahanti, P.; Schroeder, F.C.; Kim, D.H. Chemosensation of bacterial secondary metabolites modulates neuroendocrine signaling and behavior of C. elegans. Cell 2014, 159, 267–280. [Google Scholar] [CrossRef] [PubMed]
- Richards, E.J. Inherited epigenetic variation--revisiting soft inheritance. Nat. Rev. Genet. 2006, 7, 395–401. [Google Scholar] [CrossRef] [PubMed]
- Skinner, M.K. Environmental epigenetics and a unified theory of the molecular aspects of evolution: A neo-lamarckian concept that facilitates neo-darwinian evolution. Genome Biol. Evol. 2015, 7, 1296–1302. [Google Scholar] [CrossRef] [PubMed]
- Tauffenberger, A.; Parker, J.A. Heritable transmission of stress resistance by high dietary glucose in Caenorhabditis elegans. PLoS Genet. 2014, 10, e1004346. [Google Scholar] [CrossRef] [PubMed]
- Jobson, M.A.; Jordan, J.M.; Sandrof, M.A.; Hibshman, J.D.; Lennox, A.L.; Baugh, L.R. Transgenerational effects of early life starvation on growth, reproduction, and stress resistance in Caenorhabditis elegans. Genetics 2015, 201, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Kishimoto, S.; Uno, M.; Okabe, E.; Nono, M.; Nishida, E. Environmental stresses induce transgenerationally inheritable survival advantages via germline-to-soma communication in Caenorhabditis elegans. Nat. Commun. 2017, 8, 14031. [Google Scholar] [CrossRef] [PubMed]
- Boffelli, D.; Martin, D.I.K. Epigenetic inheritance: A contributor to species differentiation? DNA Cell Biol. 2012, 31, S11. [Google Scholar] [CrossRef] [PubMed]
- Brevik, K.; Lindstrom, L.; McKay, S.D.; Chen, Y.H. Transgenerational effects of insecticides-implications for rapid pest evolution in agroecosystems. Curr. Opin. Insect Sci. 2018, 26, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Daniels, S.B.; Peterson, K.R.; Strausbaugh, L.D.; Kidwell, M.G.; Chovnick, A. Evidence for horizontal transmission of the P transposable element between Drosophila species. Genetics 1990, 124, 339–355. [Google Scholar] [PubMed]
- Engels, W.R. Germ line aberrations associated with a case of hybrid dysgenesis in Drosophila melanogaster males. Genet. Res. 1979, 33, 137–146. [Google Scholar] [CrossRef] [PubMed]
- Ronsseray, S.; Anxolabehere, D.; Periquet, G. Hybrid dysgenesis in Drosophila melanogaster: Influence of temperature on cytotype determination in the P-M system. Mol. Gen. Genet. 1984, 196, 17–23. [Google Scholar] [CrossRef] [PubMed]
- Kofler, R.; Hill, T.; Nolte, V.; Betancourt, A.J.; Schlötterer, C. The recent invasion of natural Drosophila simulans populations by the P-element. Proc. Natl. Acad. Sci. USA 2015, 112, 6659–6663. [Google Scholar] [CrossRef] [PubMed]
- Kofler, R.; Senti, K.-A.; Nolte, V.; Tobler, R.; Schlötterer, C. Molecular dissection of a natural transposable element invasion. Genome Res. 2018, 28, 824–835. [Google Scholar] [CrossRef] [PubMed]
- Ronsseray, S.; Lehmann, M.; Anxolabéhère, D. Copy number and distribution of P and I mobile elements in Drosophila melanogaster populations. Chromosoma 1989, 98, 207–214. [Google Scholar] [CrossRef] [PubMed]
- Asif-Laidin, A.; Delmarre, V.; Laurentie, J.; Miller, W.J.; Ronsseray, S.; Teysset, L. Short and long-term evolutionary dynamics of subtelomeric piRNA clusters in Drosophila. DNA Res. 2017, 24, 459–472. [Google Scholar] [CrossRef]
- Rebollo, R.; Romanish, M.T.; Mager, D.L. Transposable elements: An abundant and natural source of regulatory sequences for host genes. Annu. Rev. Genet. 2012, 46, 21–42. [Google Scholar] [CrossRef]
- van de Lagemaat, L.N.; Landry, J.R.; Mager, D.L.; Medstrand, P. Transposable elements in mammals promote regulatory variation and diversification of genes with specialized functions. Trends Genet. 2003, 19, 530–536. [Google Scholar] [CrossRef]
- Garcia Guerreiro, M.P. What makes transposable elements move in the Drosophila genome? Heredity 2012, 108, 461–468. [Google Scholar] [CrossRef]
- Miousse, I.R.; Chalbot, M.-C.G.; Lumen, A.; Ferguson, A.; Kavouras, I.G.; Koturbash, I. Response of transposable elements to environmental stressors. Mutat Res. Rev. Mutat. Res. 2015, 765, 19–39. [Google Scholar] [CrossRef] [PubMed]
- Feschotte, C.; Pritham, E.J. DNA transposons and the evolution of eukaryotic genomes. Annu. Rev. Genet. 2007, 41, 331–368. [Google Scholar] [CrossRef] [PubMed]
- Jangam, D.; Feschotte, C.; Betrán, E. Transposable element domestication as an adaptation to evolutionary conflicts. Trends Genet. 2017, 33, 817–831. [Google Scholar] [CrossRef] [PubMed]
- Parhad, S.S.; Tu, S.; Weng, Z.; Theurkauf, W.E. Adaptive evolution leads to cross-species incompatibility in the piRNA transposon silencing machinery. Dev. Cell 2017, 43, 60–70. [Google Scholar] [CrossRef] [PubMed]
- Kotov, A.A.; Adashev, V.E.; Godneeva, B.K.; Ninova, M.; Shatskikh, A.S.; Bazylev, S.S.; Aravin, A.A.; Olenina, L.V. piRNA silencing contributes to interspecies hybrid sterility and reproductive isolation in Drosophila melanogaster. Nucleic Acids Res. 2019, 47, 4255–4271. [Google Scholar] [CrossRef] [PubMed]
- Skinner, M.K. Environmental stress and epigenetic transgenerational inheritance. BMC Med. 2014, 12, 153. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, E.E.; Sadler-Riggleman, I.; Skinner, M.K. Environmentally induced epigenetic transgenerational inheritance of disease. Environ. Epigenet. 2018, 4, dvy016. [Google Scholar] [CrossRef]
- Perera, B.P.U.; Faulk, C.; Svoboda, L.K.; Goodrich, J.M.; Dolinoy, D.C. The role of environmental exposures and the epigenome in health and disease. Environ. Mol. Mutagen. 2019. [Google Scholar] [CrossRef]
- Manolio, T.A.; Collins, F.S.; Cox, N.J.; Goldstein, D.B.; Hindorff, L.A.; Hunter, D.J.; McCarthy, M.I.; Ramos, E.M.; Cardon, L.R.; Chakravarti, A.; et al. Finding the missing heritability of complex diseases. Nature 2009, 461, 747–753. [Google Scholar] [CrossRef]
- Rassoulzadegan, M.; Cuzin, F. From paramutation to human disease: RNA-mediated heredity. Semin. Cell Dev. Biol. 2015, 44, 47–50. [Google Scholar] [CrossRef]
- Sole, C.; Arnaiz, E.; Manterola, L.; Otaegui, D.; Lawrie, C.H. The circulating transcriptome as a source of cancer liquid biopsy biomarkers. Semin. Cancer Biol. 2019, 58, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Huang, G.; Hu, H.; Xue, X.; Shen, S.; Gao, E.; Guo, G.; Shen, X.; Zhang, X. Altered expression of piRNAs and their relation with clinicopathologic features of breast cancer. Clin. Transl. Oncol. 2013, 15, 563–568. [Google Scholar] [CrossRef] [PubMed]
- Law, P.T.Y.; Qin, H.; Ching, A.K.K.; Lai, K.P.; Co, N.N.; He, M.; Lung, R.W.M.; Chan, A.W.H.; Chan, T.-F.; Wong, N. Deep sequencing of small RNA transcriptome reveals novel non-coding RNAs in hepatocellular carcinoma. J. Hepatol. 2013, 58, 1165–1173. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wu, X.; Gao, H.; Jin, J.M.; Li, A.X.; Kim, Y.S.; Pal, S.K.; Nelson, R.A.; Lau, C.M.; Guo, C.; et al. Piwi-interacting RNAs (piRNAs) are dysregulated in renal cell carcinoma and associated with tumor metastasis and cancer-specific survival. Mol. Med. 2015, 21, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Mai, D.; Zhang, B.; Jiang, X.; Zhang, J.; Bai, R.; Ye, Y.; Li, M.; Pan, L.; Su, J.; et al. PIWI-interacting RNA-36712 restrains breast cancer progression and chemoresistance by interaction with SEPW1 pseudogene SEPW1P RNA. Mol. Cancer 2019, 18, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Sun, X.; Yan, D.; Huang, J.; Luo, Q.; Tang, H.; Peng, Z. Piwil2 modulates the proliferation and metastasis of colon cancer via regulation of matrix metallopeptidase 9 transcriptional activity. Exp. Biol. Med. 2012, 237, 1231–1240. [Google Scholar] [CrossRef] [PubMed]
- Mei, Y.; Clark, D.; Mao, L. Novel dimensions of piRNAs in cancer. Cancer Lett. 2013, 336, 46–52. [Google Scholar] [CrossRef] [PubMed]
- Rojas-Rios, P.; Chartier, A.; Pierson, S.; Simonelig, M. Aubergine and piRNAs promote germline stem cell self-renewal by repressing the proto-oncogene Cbl. EMBO J. 2017, 36, 3194–3211. [Google Scholar] [CrossRef]
- Yao, Y.; Li, C.; Zhou, X.; Zhang, Y.; Lu, Y.; Chen, J.; Zheng, X.; Tao, D.; Liu, Y.; Ma, Y. PIWIL2 induces c-Myc expression by interacting with NME2 and regulates c-Myc-mediated tumor cell proliferation. Oncotarget 2014, 5, 8466–8477. [Google Scholar] [CrossRef]
- Wylie, A.; Jones, A.E.; D’Brot, A.; Lu, W.-J.; Kurtz, P.; Moran, J.V.; Rakheja, D.; Chen, K.S.; Hammer, R.E.; Comerford, S.A.; et al. p53 genes function to restrain mobile elements. Genes Dev. 2016, 30, 64–77. [Google Scholar] [CrossRef]
- Wylie, A.; Jones, A.E.; Abrams, J.M. p53 in the game of transposons. Bioessays 2016, 38, 1111–1116. [Google Scholar] [CrossRef] [PubMed]
- Miki, Y.; Nishisho, I.; Horii, A.; Miyoshi, Y.; Utsunomiya, J.; Kinzler, K.W.; Vogelstein, B.; Nakamura, Y. Disruption of the APC gene by a retrotransposal insertion of L1 sequence in a colon cancer. Cancer Res. 1992, 52, 643–645. [Google Scholar] [PubMed]
- Cruickshanks, H.A.; Tufarelli, C. Isolation of cancer-specific chimeric transcripts induced by hypomethylation of the LINE-1 antisense promoter. Genomics 2009, 94, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.S.; Shah, N.M.; Du, A.Y.; Dailey, Z.Z.; Pehrsson, E.C.; Godoy, P.M.; Zhang, D.; Li, D.; Xing, X.; Kim, S.; et al. Transposable elements drive widespread expression of oncogenes in human cancers. Nat. Genet. 2019, 51, 611–617. [Google Scholar] [CrossRef] [PubMed]
- Kazazian, H.H., Jr.; Wong, C.; Youssoufian, H.; Scott, A.F.; Phillips, D.G.; Antonarakis, S.E. Haemophilia A resulting from de novo insertion of L1 sequences represents a novel mechanism for mutation in man. Nature 1988, 332, 164–166. [Google Scholar] [CrossRef] [PubMed]
- Ganguly, A.; Dunbar, T.; Chen, P.; Godmilow, L.; Ganguly, T. Exon skipping caused by an intronic insertion of a young Alu Yb9 element leads to severe hemophilia A. Hum. Genet. 2003, 113, 348–352. [Google Scholar] [CrossRef] [PubMed]
- Guffanti, G.; Gaudi, S.; Fallon, J.H.; Sobell, J.; Potkin, S.G.; Pato, C.; Macciardi, F. Transposable elements and psychiatric disorders. Am. J. Med. Genet. B Neuropsychiatr. Genet. 2014, 165B, 201–216. [Google Scholar] [CrossRef]
- Guo, C.; Jeong, H.-H.; Hsieh, Y.-C.; Klein, H.-U.; Bennett, D.A.; De Jager, P.L.; Liu, Z.; Shulman, J.M. Tau activates transposable elements in Alzheimer’s disease. Cell Rep. 2018, 23, 2874–2880. [Google Scholar] [CrossRef]
- Qiu, W.; Guo, X.; Lin, X.; Yang, Q.; Zhang, W.; Zhang, Y.; Zuo, L.; Zhu, Y.; Li, C.-S.R.; Ma, C.; et al. Transcriptome-wide piRNA profiling in human brains of Alzheimer’s disease. Neurobiol. Aging 2017, 57, 170–177. [Google Scholar] [CrossRef]
- Roy, J.; Sarkar, A.; Parida, S.; Ghosh, Z.; Mallick, B. Small RNA sequencing revealed dysregulated piRNAs in Alzheimer’s disease and their probable role in pathogenesis. Mol. Biosyst. 2017, 13, 565–576. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).