HLA-E Polymorphism Determines Susceptibility to BK Virus Nephropathy after Living-Donor Kidney Transplant
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Population, BKPyV and PyVAN Screening
2.2. HLA-E Genotyping
2.3. Prediction of HLA-E Affinity for BKPyV Derived Peptides
2.4. Statistical Analysis
3. Results
3.1. HLA-E*01:01 May Be Able to Present Peptide Sequences Derived from BKPyV
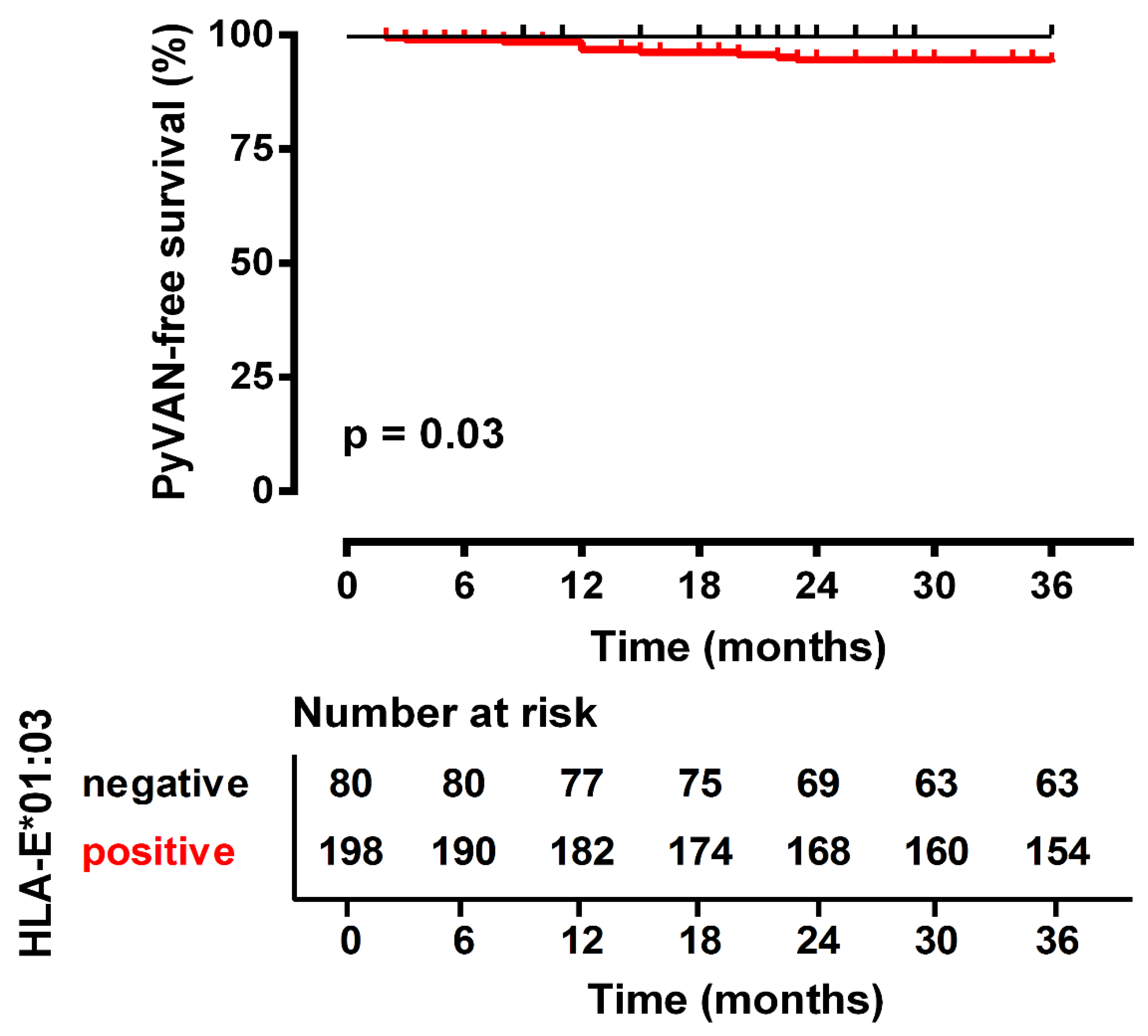
3.2. HLA-E*01:01 Homozygosity Exerts Protection against PyVAN
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Sullivan, L.C.; Clements, C.S.; Rossjohn, J.; Brooks, A.G. The major histocompatibility complex class Ib molecule HLA-E at the interface between innate and adaptive immunity. Tissue Antigens 2008, 72, 415–424. [Google Scholar] [CrossRef] [PubMed]
- Joosten, S.A.; Sullivan, L.C.; Ottenhoff, T.H. Characteristics of HLA-E Restricted T-Cell Responses and Their Role in Infectious Diseases. J. Immunol. Res. 2016, 2016, 2695396. [Google Scholar] [CrossRef] [PubMed]
- Romagnani, C.; Pietra, G.; Falco, M.; Mazzarino, P.; Moretta, L.; Mingari, M.C. HLA-E-restricted recognition of human cytomegalovirus by a subset of cytolytic T lymphocytes. Hum. Immunol. 2004, 65, 437–445. [Google Scholar] [CrossRef] [PubMed]
- Petrie, E.J.; Clements, C.S.; Lin, J.; Sullivan, L.C.; Johnson, D.; Huyton, T.; Heroux, A.; Hoare, H.L.; Beddoe, T.; Reid, H.H.; et al. CD94-NKG2A recognition of human leukocyte antigen (HLA)-E bound to an HLA class I leader sequence. J. Exp. Med. 2008, 205, 725–735. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lauterbach, N.; Voorter, C.E.; Tilanus, M.G. Molecular typing of HLA-E. Methods Mol. Biol. 2012, 882, 143–158. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.; Llano, M.; Carretero, M.; Ishitani, A.; Navarro, F.; Lopez-Botet, M.; Geraghty, D.E. HLA-E is a major ligand for the natural killer inhibitory receptor CD94/NKG2A. Proc. Natl. Acad. Sci. USA 1998, 95, 5199–5204. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Romagnani, C.; Pietra, G.; Falco, M.; Millo, E.; Mazzarino, P.; Biassoni, R.; Moretta, A.; Moretta, L.; Mingari, M.C. Identification of HLA-E-specific alloreactive T lymphocytes: A cell subset that undergoes preferential expansion in mixed lymphocyte culture and displays a broad cytolytic activity against allogeneic cells. Proc. Natl. Acad. Sci. USA 2002, 99, 11328–11333. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garcia, P.; Llano, M.; de Heredia, A.B.; Willberg, C.B.; Caparros, E.; Aparicio, P.; Braud, V.M.; Lopez-Botet, M. Human T cell receptor-mediated recognition of HLA-E. Eur. J. Immunol. 2002, 32, 936–944. [Google Scholar] [CrossRef]
- Braud, V.; Jones, E.Y.; McMichael, A. The human major histocompatibility complex class Ib molecule HLA-E binds signal sequence-derived peptides with primary anchor residues at positions 2 and 9. Eur. J. Immunol. 1997, 27, 1164–1169. [Google Scholar] [CrossRef]
- Felicio, L.P.; Porto, I.O.; Mendes-Junior, C.T.; Veiga-Castelli, L.C.; Santos, K.E.; Vianello-Brondani, R.P.; Sabbagh, A.; Moreau, P.; Donadi, E.A.; Castelli, E.C. Worldwide HLA-E nucleotide and haplotype variability reveals a conserved gene for coding and 3’ untranslated regions. Tissue Antigens 2014, 83, 82–93. [Google Scholar] [CrossRef]
- Ulbrecht, M.; Couturier, A.; Martinozzi, S.; Pla, M.; Srivastava, R.; Peterson, P.A.; Weiss, E.H. Cell surface expression of HLA-E: Interaction with human beta2-microglobulin and allelic differences. Eur. J. Immunol. 1999, 29, 537–547. [Google Scholar] [CrossRef]
- Strong, R.K.; Holmes, M.A.; Li, P.; Braun, L.; Lee, N.; Geraghty, D.E. HLA-E allelic variants. Correlating differential expression, peptide affinities, crystal structures, and thermal stabilities. J. Biol. Chem. 2003, 278, 5082–5090. [Google Scholar] [CrossRef] [PubMed]
- Pyo, C.W.; Williams, L.M.; Moore, Y.; Hyodo, H.; Li, S.S.; Zhao, L.P.; Sageshima, N.; Ishitani, A.; Geraghty, D.E. HLA-E, HLA-F, and HLA-G polymorphism: Genomic sequence defines haplotype structure and variation spanning the nonclassical class I genes. Immunogenetics 2006, 58, 241–251. [Google Scholar] [CrossRef] [PubMed]
- Lajoie, J.; Hargrove, J.; Zijenah, L.S.; Humphrey, J.H.; Ward, B.J.; Roger, M. Genetic variants in nonclassical major histocompatibility complex class I human leukocyte antigen (HLA)-E and HLA-G molecules are associated with susceptibility to heterosexual acquisition of HIV-1. J. Infect. Dis. 2006, 193, 298–301. [Google Scholar] [CrossRef] [PubMed]
- Zidi, I.; Laaribi, A.B.; Bortolotti, D.; Belhadj, M.; Mehri, A.; Yahia, H.B.; Babay, W.; Chaouch, H.; Zidi, N.; Letaief, A.; et al. HLA-E polymorphism and soluble HLA-E plasma levels in chronic hepatitis B patients. Hla 2016, 87, 153–159. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Guberina, H.; da Silva Nardi, F.; Michita, R.T.; Dolff, S.; Bienholz, A.; Heinemann, F.M.; Wilde, B.; Trilling, M.; Horn, P.A.; Kribben, A.; et al. Susceptibility of HLA-E*01:03 Allele Carriers to Develop Cytomegalovirus Replication After Living-Donor Kidney Transplantation. J. Infect. Dis. 2018, 217, 1918–1922. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, H.H.; Randhawa, P. AST Infectious Diseases Community of Practice. BK polyomavirus in solid organ transplantation-Guidelines from the American Society of Transplantation Infectious Diseases Community of Practice. Clin. Transplant. 2019. [Google Scholar] [CrossRef]
- Hirsch, H.H.; Randhawa, P. AST Infectious Diseases Community of Practice. BK polyomavirus in solid organ transplantation. Am. J. Transplant. 2013, 13 (Suppl. 4), 179–188. [Google Scholar] [CrossRef] [PubMed]
- Babel, N.; Volk, H.D.; Reinke, P. BK polyomavirus infection and nephropathy: The virus-immune system interplay. Nat. Rev. Nephrol. 2011, 7, 399–406. [Google Scholar] [CrossRef]
- Bressollette-Bodin, C.; Coste-Burel, M.; Hourmant, M.; Sebille, V.; Andre-Garnier, E.; Imbert-Marcille, B.M. A prospective longitudinal study of BK virus infection in 104 renal transplant recipients. Am. J. Transplant. 2005, 5, 1926–1933. [Google Scholar] [CrossRef]
- Korth, J.; Widera, M.; Dolff, S.; Guberina, H.; Bienholz, A.; Brinkhoff, A.; Anastasiou, O.E.; Kribben, A.; Dittmer, U.; Verheyen, J.; et al. Impact of low-level BK polyomavirus viremia on intermediate-term renal allograft function. Transpl. Infect. Dis. 2018, 20, e12817. [Google Scholar] [CrossRef]
- Berger, S.P.; Sommerer, C.; Witzke, O.; Tedesco, H.; Chadban, S.; Mulgaonkar, S.; Qazi, Y.; de Fijter, J.W.; Oppenheimer, F.; Cruzado, J.M.; et al. Two-year outcomes in de novo renal transplant recipients receiving everolimus-facilitated calcineurin inhibitor reduction regimen from TRANSFORM study. Am. J. Transplant. 2019. [Google Scholar] [CrossRef]
- Budde, K.; Lehner, F.; Sommerer, C.; Reinke, P.; Arns, W.; Eisenberger, U.; Wuthrich, R.P.; Muhlfeld, A.; Heller, K.; Porstner, M.; et al. Five-year outcomes in kidney transplant patients converted from cyclosporine to everolimus: The randomized ZEUS study. Am. J. Transplant. 2015, 15, 119–128. [Google Scholar] [CrossRef] [PubMed]
- Jouve, T.; Rostaing, L.; Malvezzi, P. Place of mTOR inhibitors in management of BKV infection after kidney transplantation. J. Nephropathol. 2016, 5, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Korth, J.; Anastasiou, O.E.; Verheyen, J.; Dickow, J.; Sertznig, H.; Frericks, N.; Bleekmann, B.; Kribben, A.; Brinkhoff, A.; Wilde, B.; et al. Impact of immune suppressive agents on the BK-Polyomavirus non coding control region. Antivir. Res. 2018, 159, 68–76. [Google Scholar] [CrossRef]
- Acott, P.; Babel, N. BK virus replication following kidney transplant: Does the choice of immunosuppressive regimen influence outcomes? Ann. Transplant. 2012, 17, 86–99. [Google Scholar] [CrossRef]
- Kopp, J.B. Banff Classification of Polyomavirus Nephropathy: A New Tool for Research and Clinical Practice. J. Am. Soc. Nephrol. 2018, 29, 354–355. [Google Scholar] [CrossRef]
- Kotton, C.N.; Kumar, D.; Caliendo, A.M.; Huprikar, S.; Chou, S.; Danziger-Isakov, L.; Humar, A. The Third International Consensus Guidelines on the Management of Cytomegalovirus in Solid-organ Transplantation. Transplantation 2018, 102, 900–931. [Google Scholar] [CrossRef] [Green Version]
- Sagedal, S.; Hartmann, A.; Nordal, K.P.; Osnes, K.; Leivestad, T.; Foss, A.; Degre, M.; Fauchald, P.; Rollag, H. Impact of early cytomegalovirus infection and disease on long-term recipient and kidney graft survival. Kidney Int. 2004, 66, 329–337. [Google Scholar] [CrossRef] [Green Version]
- Sagedal, S.; Nordal, K.P.; Hartmann, A.; Sund, S.; Scott, H.; Degre, M.; Foss, A.; Leivestad, T.; Osnes, K.; Fauchald, P.; et al. The impact of cytomegalovirus infection and disease on rejection episodes in renal allograft recipients. Am. J. Transplant. 2002, 2, 850–856. [Google Scholar] [CrossRef]
- Petersen, P.; Schneeberger, H.; Schleibner, S.; Illner, W.D.; Hofmann, G.O.; Land, W. Positive donor and negative recipient cytomegalovirus status is a detrimental factor for long-term renal allograft survival. Transpl. Int. 1994, 7 (Suppl. 1), S336–S338. [Google Scholar] [CrossRef] [PubMed]
- Ljungman, P.; Boeckh, M.; Hirsch, H.H.; Josephson, F.; Lundgren, J.; Nichols, G.; Pikis, A.; Razonable, R.R.; Miller, V.; Griffiths, P.D.; et al. Definitions of Cytomegalovirus Infection and Disease in Transplant Patients for Use in Clinical Trials. Clin. Infect. Dis. 2017, 64, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Grimsley, C.; Kawasaki, A.; Gassner, C.; Sageshima, N.; Nose, Y.; Hatake, K.; Geraghty, D.E.; Ishitani, A. Definitive high resolution typing of HLA-E allelic polymorphisms: Identifying potential errors in existing allele data. Tissue Antigens 2002, 60, 206–212. [Google Scholar] [CrossRef] [PubMed]
- Guberina, H.; Rebmann, V.; Wagner, B.; da Silva Nardi, F.; Dziallas, P.; Dolff, S.; Bienholz, A.; Wohlschlaeger, J.; Bankfalvi, A.; Heinemann, F.M.; et al. Association of high HLA-E expression during acute cellular rejection and numbers of HLA class I leader peptide mismatches with reduced renal allograft survival. Immunobiology 2017, 222, 536–543. [Google Scholar] [CrossRef] [PubMed]
- Kraemer, T.; Celik, A.A.; Huyton, T.; Kunze-Schumacher, H.; Blasczyk, R.; Bade-Doding, C. HLA-E: Presentation of a Broader Peptide Repertoire Impacts the Cellular Immune Response-Implications on HSCT Outcome. Stem Cells Int. 2015, 2015, 346714. [Google Scholar] [CrossRef] [PubMed]
- Andreatta, M.; Nielsen, M. Gapped sequence alignment using artificial neural networks: Application to the MHC class I system. Bioinformatics 2016, 32, 511–517. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, M.; Lundegaard, C.; Worning, P.; Lauemoller, S.L.; Lamberth, K.; Buus, S.; Brunak, S.; Lund, O. Reliable prediction of T-cell epitopes using neural networks with novel sequence representations. Protein Sci. 2003, 12, 1007–1017. [Google Scholar] [CrossRef]
- Heatley, S.L.; Pietra, G.; Lin, J.; Widjaja, J.M.; Harpur, C.M.; Lester, S.; Rossjohn, J.; Szer, J.; Schwarer, A.; Bradstock, K.; et al. Polymorphism in human cytomegalovirus UL40 impacts on recognition of human leukocyte antigen-E (HLA-E) by natural killer cells. J. Biol. Chem. 2013, 288, 8679–8690. [Google Scholar] [CrossRef]
- Michaelsson, J.; Teixeira de Matos, C.; Achour, A.; Lanier, L.L.; Karre, K.; Soderstrom, K. A signal peptide derived from hsp60 binds HLA-E and interferes with CD94/NKG2A recognition. J. Exp. Med. 2002, 196, 1403–1414. [Google Scholar] [CrossRef]
- Nattermann, J.; Nischalke, H.D.; Hofmeister, V.; Kupfer, B.; Ahlenstiel, G.; Feldmann, G.; Rockstroh, J.; Weiss, E.H.; Sauerbruch, T.; Spengler, U. HIV-1 infection leads to increased HLA-E expression resulting in impaired function of natural killer cells. Antivir. Ther. 2005, 10, 95–107. [Google Scholar]
- Walters, L.C.; Harlos, K.; Brackenridge, S.; Rozbesky, D.; Barrett, J.R.; Jain, V.; Walter, T.S.; O’Callaghan, C.A.; Borrow, P.; Toebes, M.; et al. Pathogen-derived HLA-E bound epitopes reveal broad primary anchor pocket tolerability and conformationally malleable peptide binding. Nat. Commun. 2018, 9, 3137. [Google Scholar] [CrossRef] [PubMed]
- Schulte, D.; Vogel, M.; Langhans, B.; Kramer, B.; Korner, C.; Nischalke, H.D.; Steinberg, V.; Michalk, M.; Berg, T.; Rockstroh, J.K.; et al. The HLA-E(R)/HLA-E(R) genotype affects the natural course of hepatitis C virus (HCV) infection and is associated with HLA-E-restricted recognition of an HCV-derived peptide by interferon-gamma-secreting human CD8(+) T cells. J. Infect. Dis. 2009, 200, 1397–1401. [Google Scholar] [CrossRef] [PubMed]
- Fishman, J.A. Infection in Organ Transplantation. Am. J. Transplant. 2017, 17, 856–879. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Reischig, T.; Kacer, M.; Hes, O.; Machova, J.; Nemcova, J.; Lysak, D.; Jindra, P.; Pivovarcikova, K.; Kormunda, S.; Bouda, M. Cytomegalovirus prevention strategies and the risk of BK polyomavirus viremia and nephropathy. Am. J. Transplant. 2019. [Google Scholar] [CrossRef] [PubMed]
- Celik, A.A.; Kraemer, T.; Huyton, T.; Blasczyk, R.; Bade-Doding, C. The diversity of the HLA-E-restricted peptide repertoire explains the immunological impact of the Arg107Gly mismatch. Immunogenetics 2016, 68, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Lampen, M.H.; Hassan, C.; Sluijter, M.; Geluk, A.; Dijkman, K.; Tjon, J.M.; de Ru, A.H.; van der Burg, S.H.; van Veelen, P.A.; van Hall, T. Alternative peptide repertoire of HLA-E reveals a binding motif that is strikingly similar to HLA-A2. Mol. Immunol. 2013, 53, 126–131. [Google Scholar] [CrossRef]
- Hansen, S.G.; Wu, H.L.; Burwitz, B.J.; Hughes, C.M.; Hammond, K.B.; Ventura, A.B.; Reed, J.S.; Gilbride, R.M.; Ainslie, E.; Morrow, D.W.; et al. Broadly targeted CD8(+) T cell responses restricted by major histocompatibility complex E. Science 2016, 351, 714–720. [Google Scholar] [CrossRef]
- Llano, M.; Lee, N.; Navarro, F.; Garcia, P.; Albar, J.P.; Geraghty, D.E.; Lopez-Botet, M. HLA-E-bound peptides influence recognition by inhibitory and triggering CD94/NKG2 receptors: Preferential response to an HLA-G-derived nonamer. Eur. J. Immunol. 1998, 28, 2854–2863. [Google Scholar] [CrossRef]
- Stevens, J.; Joly, E.; Trowsdale, J.; Butcher, G.W. Peptide binding characteristics of the non-classical class Ib MHC molecule HLA-E assessed by a recombinant random peptide approach. BMC Immunol. 2001, 2, 5. [Google Scholar] [CrossRef]
- Miller, J.D.; Weber, D.A.; Ibegbu, C.; Pohl, J.; Altman, J.D.; Jensen, P.E. Analysis of HLA-E peptide-binding specificity and contact residues in bound peptide required for recognition by CD94/NKG2. J. Immunol. 2003, 171, 1369–1375. [Google Scholar] [CrossRef]
- Vales-Gomez, M.; Reyburn, H.T.; Erskine, R.A.; Lopez-Botet, M.; Strominger, J.L. Kinetics and peptide dependency of the binding of the inhibitory NK receptor CD94/NKG2-A and the activating receptor CD94/NKG2-C to HLA-E. EMBO J. 1999, 18, 4250–4260. [Google Scholar] [CrossRef]
- Kaiser, B.K.; Barahmand-Pour, F.; Paulsene, W.; Medley, S.; Geraghty, D.E.; Strong, R.K. Interactions between NKG2x immunoreceptors and HLA-E ligands display overlapping affinities and thermodynamics. J. Immunol. 2005, 174, 2878–2884. [Google Scholar] [CrossRef]
- Guma, M.; Angulo, A.; Vilches, C.; Gomez-Lozano, N.; Malats, N.; Lopez-Botet, M. Imprint of human cytomegalovirus infection on the NK cell receptor repertoire. Blood 2004, 104, 3664–3671. [Google Scholar] [CrossRef] [Green Version]
- Guma, M.; Budt, M.; Saez, A.; Brckalo, T.; Hengel, H.; Angulo, A.; Lopez-Botet, M. Expansion of CD94/NKG2C+ NK cells in response to human cytomegalovirus-infected fibroblasts. Blood 2006, 107, 3624–3631. [Google Scholar] [CrossRef] [Green Version]
- Hammer, Q.; Ruckert, T.; Borst, E.M.; Dunst, J.; Haubner, A.; Durek, P.; Heinrich, F.; Gasparoni, G.; Babic, M.; Tomic, A.; et al. Peptide-specific recognition of human cytomegalovirus strains controls adaptive natural killer cells. Nat. Immunol. 2018, 19, 453–463. [Google Scholar] [CrossRef]
- Rolle, A.; Jager, D.; Momburg, F. HLA-E Peptide Repertoire and Dimorphism-Centerpieces in the Adaptive NK Cell Puzzle? Front. Immunol. 2018, 9, 2410. [Google Scholar] [CrossRef]
- Rolle, A.; Meyer, M.; Calderazzo, S.; Jager, D.; Momburg, F. Distinct HLA-E Peptide Complexes Modify Antibody-Driven Effector Functions of Adaptive NK Cells. Cell Rep. 2018, 24, 1967–1976. [Google Scholar] [CrossRef]
- Boudreau, J.E.; Hsu, K.C. Natural Killer Cell Education and the Response to Infection and Cancer Therapy: Stay Tuned. Trends Immunol. 2018, 39, 222–239. [Google Scholar] [CrossRef]
- Horowitz, A.; Djaoud, Z.; Nemat-Gorgani, N.; Blokhuis, J.; Hilton, H.G.; Beziat, V.; Malmberg, K.J.; Norman, P.J.; Guethlein, L.A.; Parham, P. Class I HLA haplotypes form two schools that educate NK cells in different ways. Sci. Immunol. 2016, 1. [Google Scholar] [CrossRef]
- Nattermann, J.; Feldmann, G.; Ahlenstiel, G.; Langhans, B.; Sauerbruch, T.; Spengler, U. Surface expression and cytolytic function of natural killer cell receptors is altered in chronic hepatitis C. Gut 2006, 55, 869–877. [Google Scholar] [CrossRef] [Green Version]
- Moser, J.M.; Gibbs, J.; Jensen, P.E.; Lukacher, A.E. CD94-NKG2A receptors regulate antiviral CD8(+) T cell responses. Nat. Immunol. 2002, 3, 189–195. [Google Scholar] [CrossRef]
- Moser, J.M.; Byers, A.M.; Lukacher, A.E. NK cell receptors in antiviral immunity. Curr. Opin. Immunol. 2002, 14, 509–516. [Google Scholar] [CrossRef]
- Byers, A.M.; Kemball, C.C.; Andrews, N.P.; Lukacher, A.E. Regulation of antiviral CD8+ T cells by inhibitory natural killer cell receptors. Microbes Infect. 2003, 5, 169–177. [Google Scholar] [CrossRef]
- Byers, A.M.; Andrews, N.P.; Lukacher, A.E. CD94/NKG2A expression is associated with proliferative potential of CD8 T cells during persistent polyoma virus infection. J. Immunol. 2006, 176, 6121–6129. [Google Scholar] [CrossRef]
- Wagner, B.; da Silva Nardi, F.; Schramm, S.; Kraemer, T.; Celik, A.A.; Durig, J.; Horn, P.A.; Duhrsen, U.; Nuckel, H.; Rebmann, V. HLA-E allelic genotype correlates with HLA-E plasma levels and predicts early progression in chronic lymphocytic leukemia. Cancer 2017, 123, 814–823. [Google Scholar] [CrossRef]
| A | B | C | ||
|---|---|---|---|---|
| Total | HLA-E*01:03 Carrier# | HLA-E*01:03 Non-Carrier | p-Value B vs. C | |
| Donor | N = 278 | N = 211 | N = 67 | |
| Sex (men/women) | 115/163 | 89/122 | 26/41 | 0.67 a |
| Age (y) ±SD | 51.41 ± 10.03 | 51.99 ± 10.06 | 43.30 ± 9.83 | 0.10 b |
| Recipient | N = 278 | N = 198 | N = 80 | |
| Sex (men/women) | 162/116 | 116/82 | 46/34 | 0.87 a |
| Age (y) ±SD | 41.06 ± 15.43 | 40.46 ± 15.42 | 42.53 ± 15.47 | 0.32 b |
| KTx Related Variables | ||||
| HLA A, B mismatches, mean ± SD | 1.97 ± 1.14 | 1.83 ± 1.1 | 2.31 ± 1.17 | 0.002 **b |
| HLA-DR mismatch, mean ± SD | 1.12 ± 0.62 | 1.09 ± 0.60 | 1.19 ± 0.66 | 0.22 b |
| Panel of Antibodies (%) | ||||
| 0% | 244 | 172 | 72 | 0.55 a |
| 1–10% | 14 | 9 | 5 | 0.55 a |
| 10–50% | 14 | 12 | 2 | 0.36 a |
| >50% | 6 | 5 | 1 | 0.68 a |
| Cold ischemia time, median, in minutes (range) | 134.99 ± 48.90 | 136.07 ± 48.63 | 131.32 ± 48.86 | 0.57 b |
| Warm ischemia time, median, in minutes (range) | 19.97 ± 6.87 | 19.38 ± 5.84 | 21.46 ± 8.83 | 0.057 b |
| Immunosuppressive Therapy | ||||
| ATG-based induction therapy, yes/no | 13/265 | 9/189 | 4/76 | 1 a |
| CNI administration, yes/no | 278/0 | 198/0 | 80/0 | nd |
| MMF co-administration, yes/no | 260/18 | 189/9 | 75/5 | 0.55 a |
| Steroid co-administration, yes/no | 278/0 | 198/0 | 80/0 | nd |
| PyVAN and CMV | ||||
| PyVAN, yes/no | 11/267 | 11/187 | 0/80 | 0.031 *a |
| CMV positive recipient | 159/119 | 116/82 | 43/37 | 0.46 a |
| CMV positive donor | 177/101 | 128/70 | 49/31 | 0.59 a |
| CMV Infection, yes/no | 38/240 | 32/166 | 6/74 | 0.057 a |
| PyVAN and CMV infection, yes/no | 3/275 | 3/195 | 0/80 | 0.27 a |
| PyVAN or CMV infection vs. no PyVAN and no CMV infection | 46/232 | 40/158 | 6/74 | 0.01 a |
| Large-T and Small-t Antigen | |||
|---|---|---|---|
| Position | Peptide | 1 − log50k(aff) | Affinity (nM) |
| 416 | NVPKRRYWL | 0.202 | 5619.93 |
| 568 | RILQSGMTL | 0.173 | 7666.40 |
| 454 | PMERLTFEL | 0.165 | 8432.78 |
| 557 | SLQNSEFLL | 0.160 | 8874.14 |
| 570 | LQSGMTLLL | 0.150 | 9850.58 |
| BK Polyomavirus Major Capsid Protein VP1 | |||
|---|---|---|---|
| Position | Peptide | 1 − log50k(aff) | Affinity (nM) |
| 18 | KEPVQVPKL | 0.186 | 6716.19 |
| 237 | TNTATTVLL | 0.142 | 10,773.00 |
| 297 | NPYPISFLL | 0.139 | 11,067.54 |
| BK Polyomavirus Major Capsid Protein VP2 | |||
|---|---|---|---|
| Position | Peptide | 1 − log50k(aff) | Affinity(nM) |
| 292 | WMLPLLLGL | 0.152 | 9685.81 |
| BK polyomavirus agnoprotein | |||
|---|---|---|---|
| Position | Peptide | 1 − log50k(aff) | Affinity (nM) |
| 46 | SXXPESVMF | 0.139 | 11,081.08 |
| 52 | VMFCEPKNL | 0.139 | 11,162.18 |
| Derived from | Peptide | 1 − log50k(aff) | Affinity (nM) |
|---|---|---|---|
| Human leukocyte antigen-Cw*3 | VMAPRTLIL | 0.589 | 85.69 |
| CMV protein pUL40 | VMAPRTLIL | 0.589 | 85.69 |
| Human leukocyte antigen-G | VMAPRTLFL | 0.564 | 112.12 |
| Mycobacterium tuberculosis enoyl-[acyl-carrier-protein] reductase [NADH] | RLPAKAPLL | 0.547 | 134.77 |
| CMV phosphorylated matrix protein pp65 (UL83) | VLPHETRLL | 0.330 | 1405.23 |
| CMV immediate-early protein 1 (pUL123) | VMLAKRPLI | 0.303 | 1887.52 |
| Human heat shock protein 60 (hsp60) | QMRPVSRVL | 0.276 | 2530.17 |
| CMV protein pUL18 | SEPQCNPLL | 0.273 | 2614.63 |
| BK polyomavirus large-T and small-t antigen | NVPKRRYWL | 0.202 | 5619.93 |
| BK polyomavirus major capsid protein VP1 | KEPVQVPKL | 0.186 | 6716.19 |
| Human immunodeficiency virus 1 Gag protein | RMYSPVSIL | 0.173 | 7728.11 |
| BK polyomavirus major capsid protein VP2 | WMLPLLLGL | 0.152 | 9685.81 |
| BK polyomavirus agnoprotein | SXXPESVMF | 0.139 | 11081.08 |
| (A) Recipient HLA-E genotype | PyVAN N = 11 | No PyVAN N = 267 | p-Value | OR | 95% CI |
| 01:03/01:03 | 3 (27.3%) | 61 (22.8%) | 0.73 | 1.26 | 0.33–4.92 |
| 01:01/01:03 | 8 (72.7%) | 126 (47.2%) | 0.10 | 2.98 | 0.77–11.50 |
| 01:01/01:01 | 0 (0%) | 80 (30%) | 0.025 * | 0.09 | 0.005–1.59 |
| Allele Frequencies | |||||
| 01:03 | 14 | 248 | 0.11 | 2.02 | 0.83–4.89 |
| 01:01 | 8 | 286 | |||
| (B) Donor HLA-E genotype | PyVAN N = 11 | No PyVAN N = 267 | p-Value | OR | CI (95%) |
| 01:03/01:03 | 1 (9.1%) | 62 (23.2%) | 0.27 | 0.33 | 0.04–2.63 |
| 01:01/01:03 | 9 (81.8%) | 139 (52.1%) | 0.052 | 4.14 | 0.89–19.55 |
| 01:01/01:01 | 1 (9.1%) | 66 (24.7%) | 0.24 | 0.3 | 0.04–2.42 |
| Allele Frequencies | |||||
| 01:03 | 11 | 263 | 0.77 | 1.1 | 0.47–2.7 |
| 01:01 | 10 | 271 | |||
| (A) Recipient HLA-E genotype | PyVAN or CMV Infection N = 46 | No PyVAN and no CMV Infection N = 267 | p-Value | OR | 95% CI |
| 01:03/01:03 | 12 (26.1%) | 52 (22.4%) | 0.57 | 1.22 | 0.59–2.53 |
| 01:01/01:03 | 28 (60.9%) | 106 (45.7%) | 0.07 | 1.85 | 0.96–3.53 |
| 01:01/01:01 | 6 (13.0%) | 74 (31.9%) | 0.012 * | 0.32 | 0.13–0.79 |
| Allele Frequencies | |||||
| 01:03 | 52 | 210 | 0.052 | 1.57 | 1.00–2.47 |
| 01:01 | 40 | 254 | |||
| (B) Donor HLA-E genotype | PyVAN or CMV Infection N = 46 | No PyVAN and no CMV Infection N = 267 | p-Value | OR | CI (95%) |
| 01:03/01:03 | 12 (9.1%) | 51 (22.0%) | 0.44 | 1.33 | 0.64–2.79 |
| 01:01/01:03 | 26 (81.8%) | 122 (52.6%) | 0.75 | 1.17 | 0.62–2.22 |
| 01:01/01:01 | 8 (9.1%) | 59 (25.4%) | 0.34 | 0.62 | 0.27–1.39 |
| Allele Frequencies | |||||
| 01:03 | 50 | 224 | 0.31 | 1.3 | 0.81–1.99 |
| 01:01 | 42 | 240 | |||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rohn, H.; Michita, R.T.; Schramm, S.; Dolff, S.; Gäckler, A.; Korth, J.; Heinemann, F.M.; Wilde, B.; Trilling, M.; Horn, P.A.; et al. HLA-E Polymorphism Determines Susceptibility to BK Virus Nephropathy after Living-Donor Kidney Transplant. Cells 2019, 8, 847. https://doi.org/10.3390/cells8080847
Rohn H, Michita RT, Schramm S, Dolff S, Gäckler A, Korth J, Heinemann FM, Wilde B, Trilling M, Horn PA, et al. HLA-E Polymorphism Determines Susceptibility to BK Virus Nephropathy after Living-Donor Kidney Transplant. Cells. 2019; 8(8):847. https://doi.org/10.3390/cells8080847
Chicago/Turabian StyleRohn, Hana, Rafael Tomoya Michita, Sabine Schramm, Sebastian Dolff, Anja Gäckler, Johannes Korth, Falko M. Heinemann, Benjamin Wilde, Mirko Trilling, Peter A. Horn, and et al. 2019. "HLA-E Polymorphism Determines Susceptibility to BK Virus Nephropathy after Living-Donor Kidney Transplant" Cells 8, no. 8: 847. https://doi.org/10.3390/cells8080847
APA StyleRohn, H., Michita, R. T., Schramm, S., Dolff, S., Gäckler, A., Korth, J., Heinemann, F. M., Wilde, B., Trilling, M., Horn, P. A., Kribben, A., Witzke, O., & Rebmann, V. (2019). HLA-E Polymorphism Determines Susceptibility to BK Virus Nephropathy after Living-Donor Kidney Transplant. Cells, 8(8), 847. https://doi.org/10.3390/cells8080847






