Detection of AR-V7 in Liquid Biopsies of Castrate Resistant Prostate Cancer Patients: A Comparison of AR-V7 Analysis in Circulating Tumor Cells, Circulating Tumor RNA and Exosomes
Abstract
1. Introduction
2. Materials and Methods
2.1. Patients
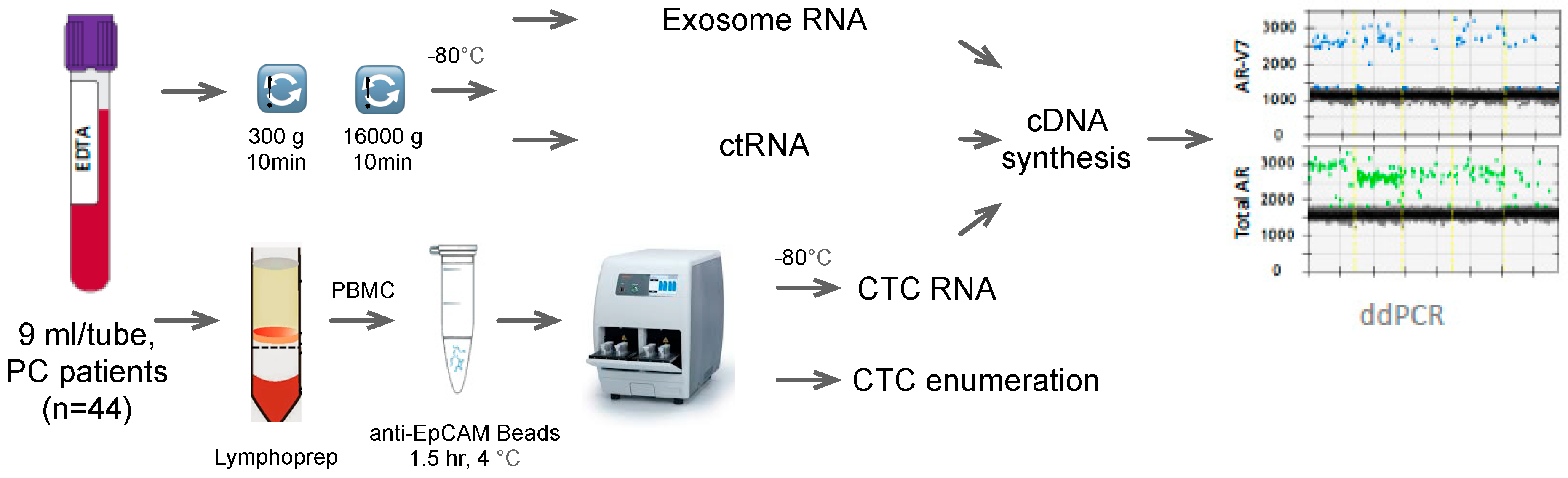
2.2. CTC Isolation
2.3. CTC Enumeration
2.4. Plasma Processing
2.5. RNA Extraction and cDNA Synthesis
2.6. Droplet Digital PCR (ddPCR)
2.7. Statistics
3. Results
3.1. Patients
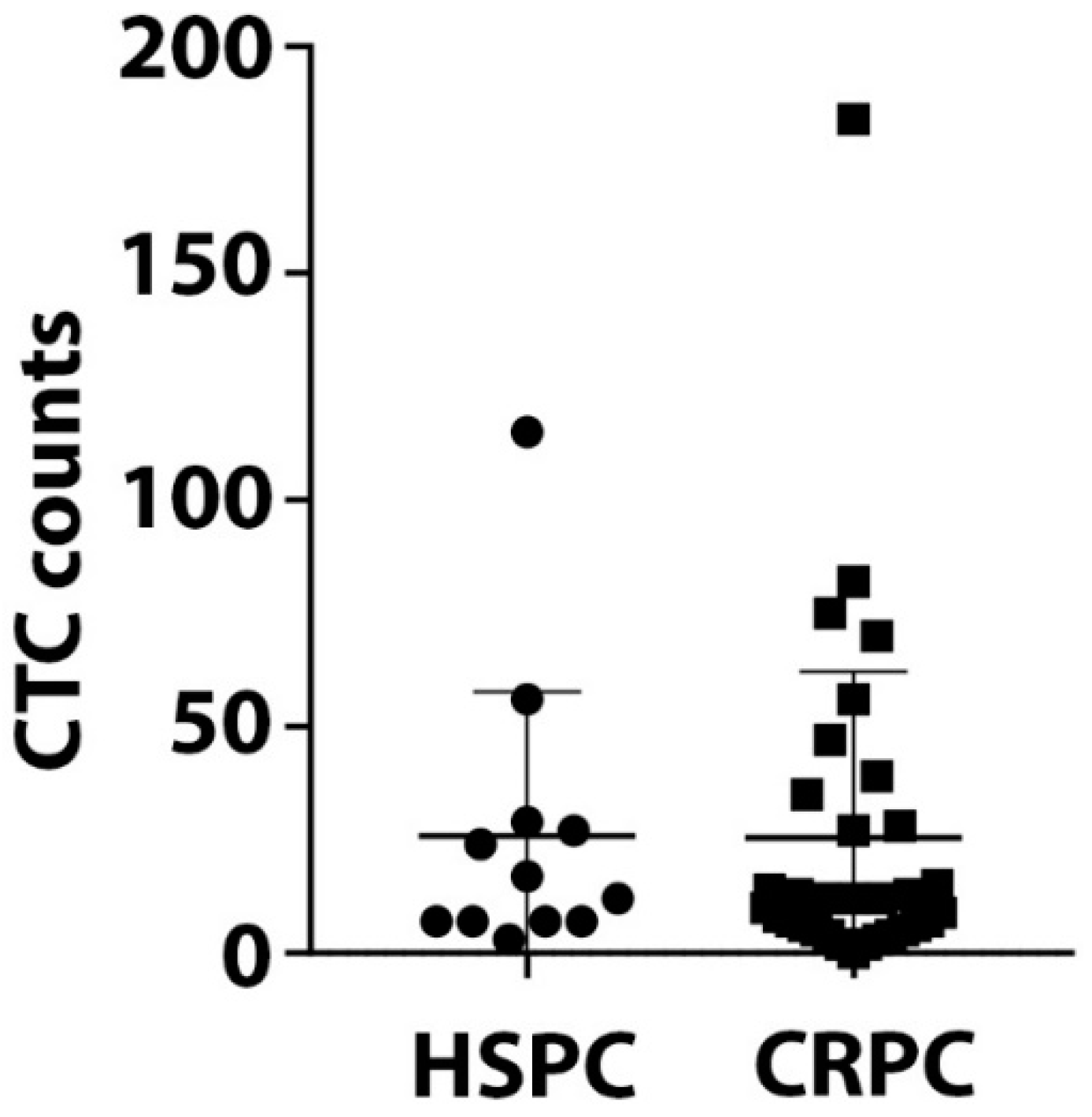
3.2. Prevalence of CTCs
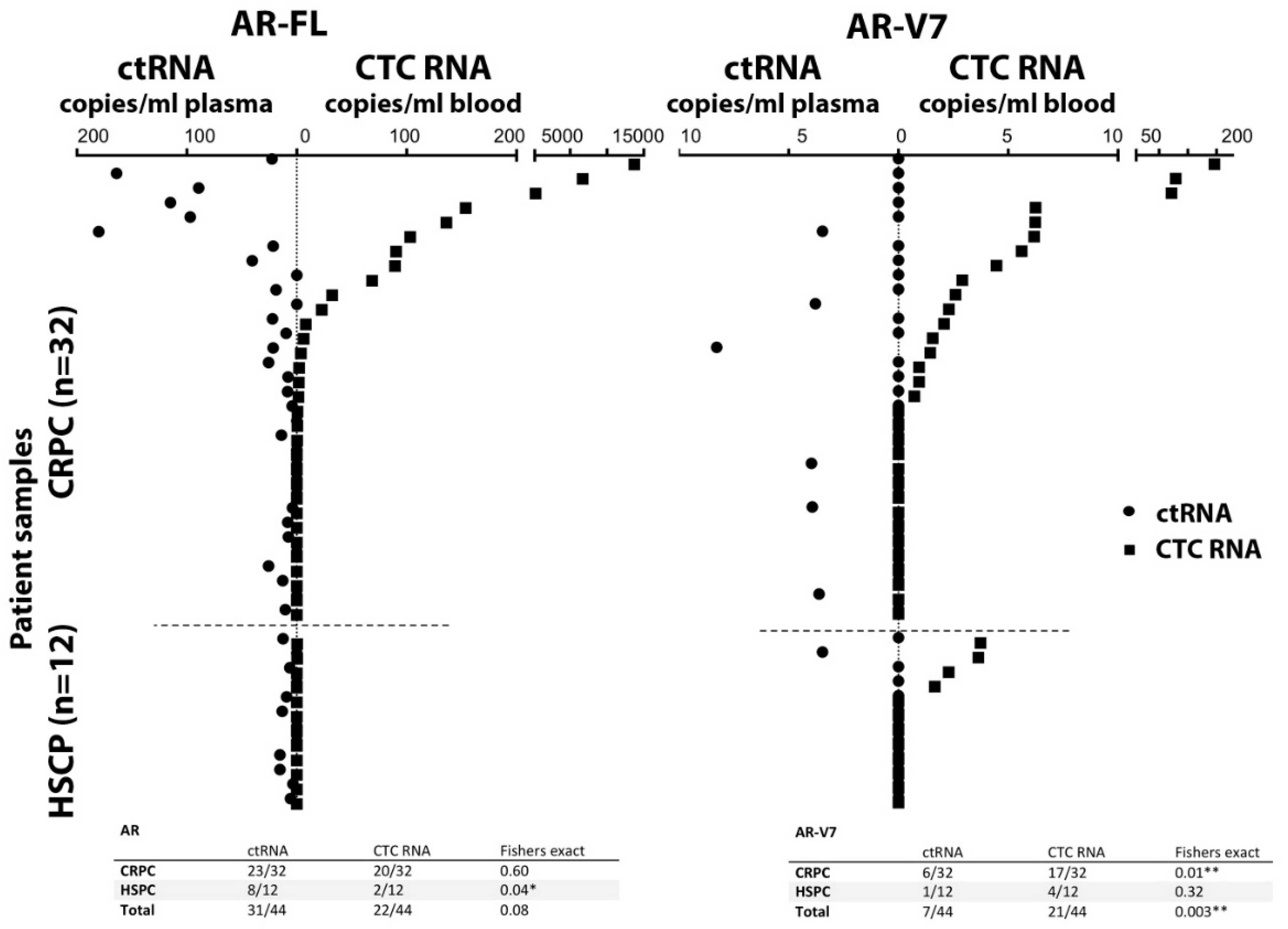
3.3. AR-V7 and AR-FL in CTC RNA and Plasma ctRNA
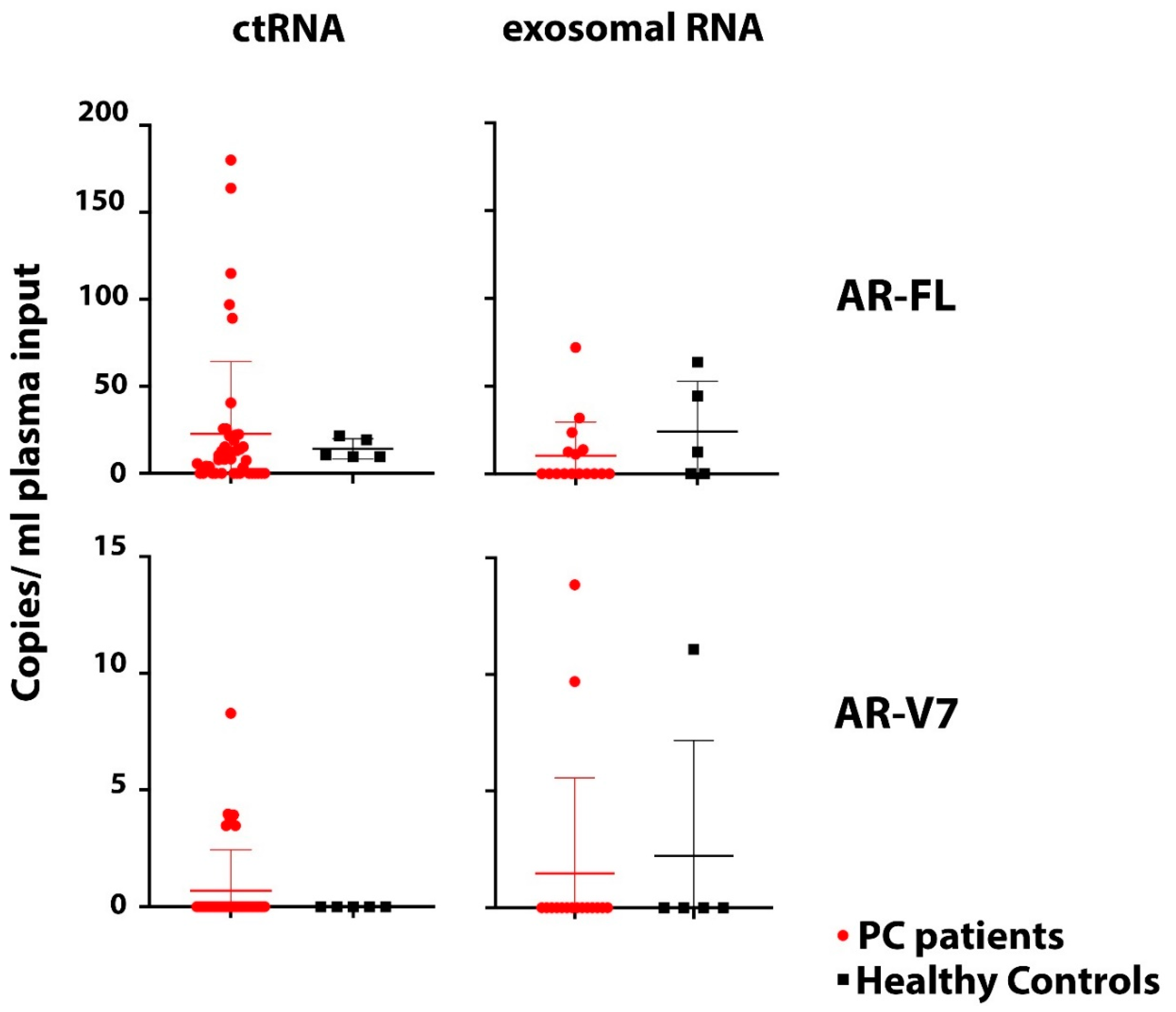
3.4. AR-V7 and AR-FL in Exosomal RNA, ctRNA and CTCs
3.5. Healthy Control Analysis and Implications for Assay Specificity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Knudsen, K.E.; Scher, H.I. Starving the addiction: New opportunities for durable suppression of AR signaling in prostate cancer. Clin. Cancer Res. 2009, 15, 4792–4798. [Google Scholar] [CrossRef] [PubMed]
- Wadosky, K.M.; Koochekpour, S. Molecular mechanisms underlying resistance to androgen deprivation therapy in prostate cancer. Oncotarget 2016, 7, 64447–64470. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Armstrong, A.J.; Dehm, S.M.; Luo, J. Androgen receptor variant-driven prostate cancer: clinical implications and therapeutic targeting. Prostate Cancer Prostatic Dis. 2016, 19, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Chan, S.C.; Brand, L.J.; Hwang, T.H.; Silverstein, K.A.; Dehm, S.M. Androgen receptor splice variants mediate enzalutamide resistance in castration-resistant prostate cancer cell lines. Cancer Res. 2013, 73, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Qi, Y.; Zhang, G.; Xu, D.; Zhan, Y.; Álvarez, X.; Guo, Z.; Fu, X.; Plymate, S.R.; Sartor, O.; et al. Androgen receptor splice variants activating the full-length receptor in mediating resistance to androgen-directed therapy. Oncotarget 2014, 5, 1646–1656. [Google Scholar] [CrossRef]
- Chan, S.C.; Li, Y.; Dehm, S.M. Androgen Receptor Splice Variants Activate Androgen Receptor Target Genes and Support Aberrant Prostate Cancer Cell Growth Independent of Canonical Androgen Receptor Nuclear Localization Signal*. J. Boil. Chem. 2012, 287, 19736–19749. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Zhan, Y.; Qi, Y.; Cao, B.; Bai, S.; Xu, W.; Gambhir, S.S.; Lee, P.; Sartor, O.; Flemington, E.K.; et al. Androgen receptor splice variants dimerize to transactivate target genes. Cancer Res. 2015, 75, 3663–3671. [Google Scholar] [CrossRef]
- Henzler, C.; Li, Y.; Yang, R.; McBride, T.; Ho, Y.; Sprenger, C.; Liu, G.; Coleman, I.; Lakely, B.; Li, R.; et al. Truncation and constitutive activation of the androgen receptor by diverse genomic rearrangements in prostate cancer. Nat. Commun. 2016, 7, 13668. [Google Scholar] [CrossRef]
- Conteduca, V.; Wetterskog, D.; Sharabiani, M.T.A.; Grande, E.; Fernandez-Perez, M.P.; Jayaram, A.; Salvi, S.; Castellano, D.; Romanel, A.; Lolli, C.; et al. Androgen receptor gene status in plasma DNA associates with worse outcome on enzalutamide or abiraterone for castration-resistant prostate cancer: a multi-institution correlative biomarker study. Ann. Oncol. 2017, 28, 1508–1516. [Google Scholar] [CrossRef]
- Guedes, L.B.; Morais, C.L.; Almutairi, F.; Haffner, M.C.; Zheng, Q.; Isaacs, J.T.; Antonarakis, E.S.; Lu, C.; Tsai, H.; Luo, J.; et al. Analytic Validation of RNA In Situ Hybridization (RISH) for AR and AR-V7 Expression in Human Prostate Cancer. Clin. Cancer Res. 2016, 22, 4651–4663. [Google Scholar] [CrossRef]
- Welti, J.; Rodrigues, D.N.; Sharp, A.; Sun, S.; Lorente, D.; Riisnaes, R.; Figueiredo, I.; Zafeiriou, Z.; Rescigno, P.; De Bono, J.S.; et al. Analytical Validation and Clinical Qualification of a New Immunohistochemical Assay for Androgen Receptor Splice Variant-7 Protein Expression in Metastatic Castration-resistant Prostate Cancer. Eur. Urol. 2016, 70, 599–608. [Google Scholar] [CrossRef] [PubMed]
- Kallio, H.M.L.; Hieta, R.; Latonen, L.; Brofeldt, A.; Annala, M.; Kivinummi, K.; Tammela, T.L.; Nykter, M.; Isaacs, W.B.; Lilja, H.G.; et al. Constitutively active androgen receptor splice variants AR-V3, AR-V7 and AR-V9 are co-expressed in castration-resistant prostate cancer metastases. Br. J. Cancer 2018, 119, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Crowley, E.; Di Nicolantonio, F.; Loupakis, F.; Bardelli, A. Liquid biopsy: monitoring cancer-genetics in the blood. Nat. Rev. Clin. Oncol. 2013, 10, 472–484. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Lu, C.; Luber, B.; Wang, H.; Chen, Y.; Nakazawa, M.; Nadal, R.; Paller, C.J.; Denmeade, S.R.; Carducci, M.A.; et al. Androgen Receptor Splice Variant 7 and Efficacy of Taxane Chemotherapy in Patients With Metastatic Castration-Resistant Prostate Cancer. JAMA Oncol. 2015, 1, 582–591. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Luk, A.; Young, F.P.; Lynch, D.; Chua, W.; Balakrishnar, B.; De Souza, P.; Becker, T.M. Droplet Digital PCR Based Androgen Receptor Variant 7 (AR-V7) Detection from Prostate Cancer Patient Blood Biopsies. Int. J. Mol. Sci. 2016, 17, 1264. [Google Scholar] [CrossRef] [PubMed]
- Hodara, E.; Morrison, G.; Cunha, A.T.; Zainfeld, D.; Xu, T.; Xu, Y.; Dempsey, P.W.; Pagano, P.C.; Bischoff, F.; Khurana, A.; et al. Multiparametric liquid biopsy analysis in metastatic prostate cancer. JCI Insight 2019, 4, 4. [Google Scholar] [CrossRef]
- Todenhöfer, T.; Azad, A.; Stewart, C.; Gao, J.; Eigl, B.J.; Gleave, M.E.; Joshua, A.M.; Black, P.C.; Chi, K.N. AR-V7 Transcripts in Whole Blood RNA of Patients with Metastatic Castration Resistant Prostate Cancer Correlate with Response to Abiraterone Acetate. J. Urol. 2017, 197, 135–142. [Google Scholar] [CrossRef]
- Woo, H.K.; Park, J.; Ku, J.Y.; Lee, C.H.; Sunkara, V.; Ha, H.K.; Cho, Y.K. Urine-based liquid biopsy: non-invasive and sensitive AR-V7 detection in urinary EVs from patients with prostate cancer. Lab on a chip 2018, 19, 87–97. [Google Scholar] [CrossRef]
- Del Re, M.; Biasco, E.; Crucitta, S.; DeRosa, L.; Rofi, E.; Orlandini, C.; Miccoli, M.; Galli, L.; Falcone, A.; Jenster, G.W.; et al. The Detection of Androgen Receptor Splice Variant 7 in Plasma-derived Exosomal RNA Strongly Predicts Resistance to Hormonal Therapy in Metastatic Prostate Cancer Patients. Eur. Urol. 2017, 71, 680–687. [Google Scholar] [CrossRef]
- Antonarakis, E.S.; Lu, C.; Wang, H.; Luber, B.; Nakazawa, M.; Roeser, J.C.; Chen, Y.; Mohammad, T.A.; Chen, Y.; Fedor, H.L.; et al. AR-V7 and Resistance to Enzalutamide and Abiraterone in Prostate Cancer. N. Eng. J. Med. 2014, 371, 1028–1038. [Google Scholar] [CrossRef]
- Bernemann, C.; Schnoeller, T.J.; Luedeke, M.; Steinestel, K.; Boegemann, M.; Schrader, A.J.; Steinestel, J. Expression of AR-V7 in Circulating Tumour Cells Does Not Preclude Response to Next Generation Androgen Deprivation Therapy in Patients with Castration Resistant Prostate Cancer. Eur. Urol. 2017, 71, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Antonarakis, E.S.; Lu, C.; Luber, B.; Wang, H.; Chen, Y.; Zhu, Y.; Silberstein, J.L.; Taylor, M.N.; Maughan, B.L.; Denmeade, S.R.; et al. Clinical Significance of Androgen Receptor Splice Variant-7 mRNA Detection in Circulating Tumor Cells of Men With Metastatic Castration-Resistant Prostate Cancer Treated With First- and Second-Line Abiraterone and Enzalutamide. J. Clin. Oncol. 2017, 35, 2149–2156. [Google Scholar] [CrossRef] [PubMed]
- Onstenk, W.; Sieuwerts, A.M.; Kraan, J.; Van, M.; Nieuweboer, A.J.; Mathijssen, R.H.; Hamberg, P.; Meulenbeld, H.J.; De Laere, B.; Dirix, L.Y.; et al. Efficacy of Cabazitaxel in Castration-resistant Prostate Cancer Is Independent of the Presence of AR-V7 in Circulating Tumor Cells. Eur. Urol. 2015, 68, 939–945. [Google Scholar] [CrossRef] [PubMed]
- Mantalaris, A.; Panoskaltsis, N.; Sakai, Y.; Bourne, P.; Chang, C.; Messing, E.M.; Wu, J.H.D. Localization of androgen receptor expression in human bone marrow. J. Pathol. 2001, 193, 361–366. [Google Scholar] [CrossRef]
- Onstenk, W.; De Klaver, W.; De Wit, R.; Lolkema, M.; Foekens, J.; Sleijfer, S. The use of circulating tumor cells in guiding treatment decisions for patients with metastatic castration-resistant prostate cancer. Cancer Treat. Rev. 2016, 46, 42–50. [Google Scholar] [CrossRef] [PubMed]
- Luk, A.W.S.; Ma, Y.; Ding, P.N.; Young, F.P.; Chua, W.; Balakrishnar, B.; Dransfield, D.T.; De Souza, P.; Becker, T.M. CTC-mRNA (AR-V7) Analysis from Blood Samples—Impact of Blood Collection Tube and Storage Time. Int. J. Mol. Sci. 2017, 18, 1047. [Google Scholar] [CrossRef] [PubMed]
- Hu, R.; Dunn, T.A.; Wei, S.; Isharwal, S.; Veltri, R.W.; Humphreys, E.; Han, M.; Partin, A.W.; Vessella, R.L.; Isaacs, W.B.; et al. Ligand-independent Androgen Receptor Variants Derived from Splicing of Cryptic Exons Signify Hormone Refractory Prostate Cancer. Cancer Res. 2009, 69, 16–22. [Google Scholar] [CrossRef]
- Qu, Y.; Dai, B.; Ye, D.; Kong, Y.; Chang, K.; Jia, Z.; Yang, X.; Zhang, H.; Zhu, Y.; Shi, G. Constitutively Active AR-V7 Plays an Essential Role in the Development and Progression of Castration-Resistant Prostate Cancer. Sci. Rep. 2015, 5, 7654. [Google Scholar] [CrossRef]
- El-Heliebi, A.; Hille, C.; Laxman, N.; Svedlund, J.; Haudum, C.; Ercan, E.; Kroneis, T.; Chen, S.; Smolle, M.; Rossmann, C.; et al. In Situ Detection and Quantification of AR-V7, AR-FL, PSA, and KRAS Point Mutations in Circulating Tumor Cells. Clin. Chem. 2018, 64, 536–546. [Google Scholar] [CrossRef]
- Scher, H.I.; Lu, D.; Schreiber, N.A.; Louw, J.; Graf, R.P.; Vargas, H.A.; Johnson, A.; Jendrisak, A.; Bambury, R.; Danila, D.; et al. Association of AR-V7 on Circulating Tumor Cells as a Treatment-Specific Biomarker With Outcomes and Survival in Castration-Resistant Prostate Cancer. JAMA Oncol. 2016, 2, 1441–1449. [Google Scholar] [CrossRef]
| Patient | n = 44 |
|---|---|
| Mean age (years) | 77 (55–94) |
| Gleason score (%) | |
| ≥7 | 34 (77%) |
| <7 | 4 (9%) |
| N.A. | 6 (14%) |
| Metastatic status (%) | 41 (93%) |
| Bone metastases | 35 (79%) |
| Lymph metastases | 22 (50%) |
| Visceral metastases | 13 (29%) |
| Definitive therapy (%) | |
| Radical Prostatectomy | 12 (27%) |
| Radiotherapy | 12 (27%) |
| Both | 2 (4%) |
| None | 22 (50%) |
| Past systemic therapy (%) | |
| None | 2 (4%) |
| ADT | 42 (95%) |
| Chemotherapy | 17 (38%) |
| Novel antiandrogens | |
| Enzalutamide | 14 (31%) |
| Abiraterone | 9 (20%) |
| Both | 4 (9%) |
| Name | Sequence |
|---|---|
| GAPDH-fwrd | CGGGAAGCTTGTCATCAATGG |
| GAPDH-rev | CTCCACGACGTACTCAGCG |
| GAPDH-probe | FAM-5′-TCTTCCAGGAGCGAGATCCCT-3-BQ1 |
| Patient | Status | AR-FL CTCs | AR-V7 CTCs | AR-FL ctRNA | AR-V7 ctRNA | AR-FL Exosomal | AR-V7 Exosomal |
|---|---|---|---|---|---|---|---|
| 1 | CRPC | − | − | − | − | + | − |
| 2 | CRPC | + | − | + | − | − | − |
| 3 | CRPC | + | − | + | − | − | − |
| 4 | CRPC | + | + | + | − | − | − |
| 5 | CRPC | − | + | + | − | + | + |
| 6 | CRPC | − | + | − | − | + | − |
| 7 | CRPC | + | + | + | − | + | + |
| 8 | CRPC | + | + | − | − | − | − |
| 9 | CRPC | + | − | + | − | − | − |
| 10 | HSPC | + | − | − | − | + | − |
| 11 | HSPC | + | − | + | − | − | − |
| 12 | HSPC | − | − | + | − | − | − |
| 13 | HSPC | − | + | + | − | − | − |
| 14 | HSPC | − | + | − | + | − | − |
| 15 | HSPC | − | − | + | − | + | − |
| 16 | HSPC | − | − | + | − | − | − |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nimir, M.; Ma, Y.; Jeffreys, S.A.; Opperman, T.; Young, F.; Khan, T.; Ding, P.; Chua, W.; Balakrishnar, B.; Cooper, A.; et al. Detection of AR-V7 in Liquid Biopsies of Castrate Resistant Prostate Cancer Patients: A Comparison of AR-V7 Analysis in Circulating Tumor Cells, Circulating Tumor RNA and Exosomes. Cells 2019, 8, 688. https://doi.org/10.3390/cells8070688
Nimir M, Ma Y, Jeffreys SA, Opperman T, Young F, Khan T, Ding P, Chua W, Balakrishnar B, Cooper A, et al. Detection of AR-V7 in Liquid Biopsies of Castrate Resistant Prostate Cancer Patients: A Comparison of AR-V7 Analysis in Circulating Tumor Cells, Circulating Tumor RNA and Exosomes. Cells. 2019; 8(7):688. https://doi.org/10.3390/cells8070688
Chicago/Turabian StyleNimir, Mohammed, Yafeng Ma, Sarah A. Jeffreys, Thomas Opperman, Francis Young, Tanzila Khan, Pei Ding, Wei Chua, Bavanthi Balakrishnar, Adam Cooper, and et al. 2019. "Detection of AR-V7 in Liquid Biopsies of Castrate Resistant Prostate Cancer Patients: A Comparison of AR-V7 Analysis in Circulating Tumor Cells, Circulating Tumor RNA and Exosomes" Cells 8, no. 7: 688. https://doi.org/10.3390/cells8070688
APA StyleNimir, M., Ma, Y., Jeffreys, S. A., Opperman, T., Young, F., Khan, T., Ding, P., Chua, W., Balakrishnar, B., Cooper, A., De Souza, P., & Becker, T. M. (2019). Detection of AR-V7 in Liquid Biopsies of Castrate Resistant Prostate Cancer Patients: A Comparison of AR-V7 Analysis in Circulating Tumor Cells, Circulating Tumor RNA and Exosomes. Cells, 8(7), 688. https://doi.org/10.3390/cells8070688







