Fibroblast Growth Factor Receptors (FGFRs): Structures and Small Molecule Inhibitors
Abstract
1. Introduction
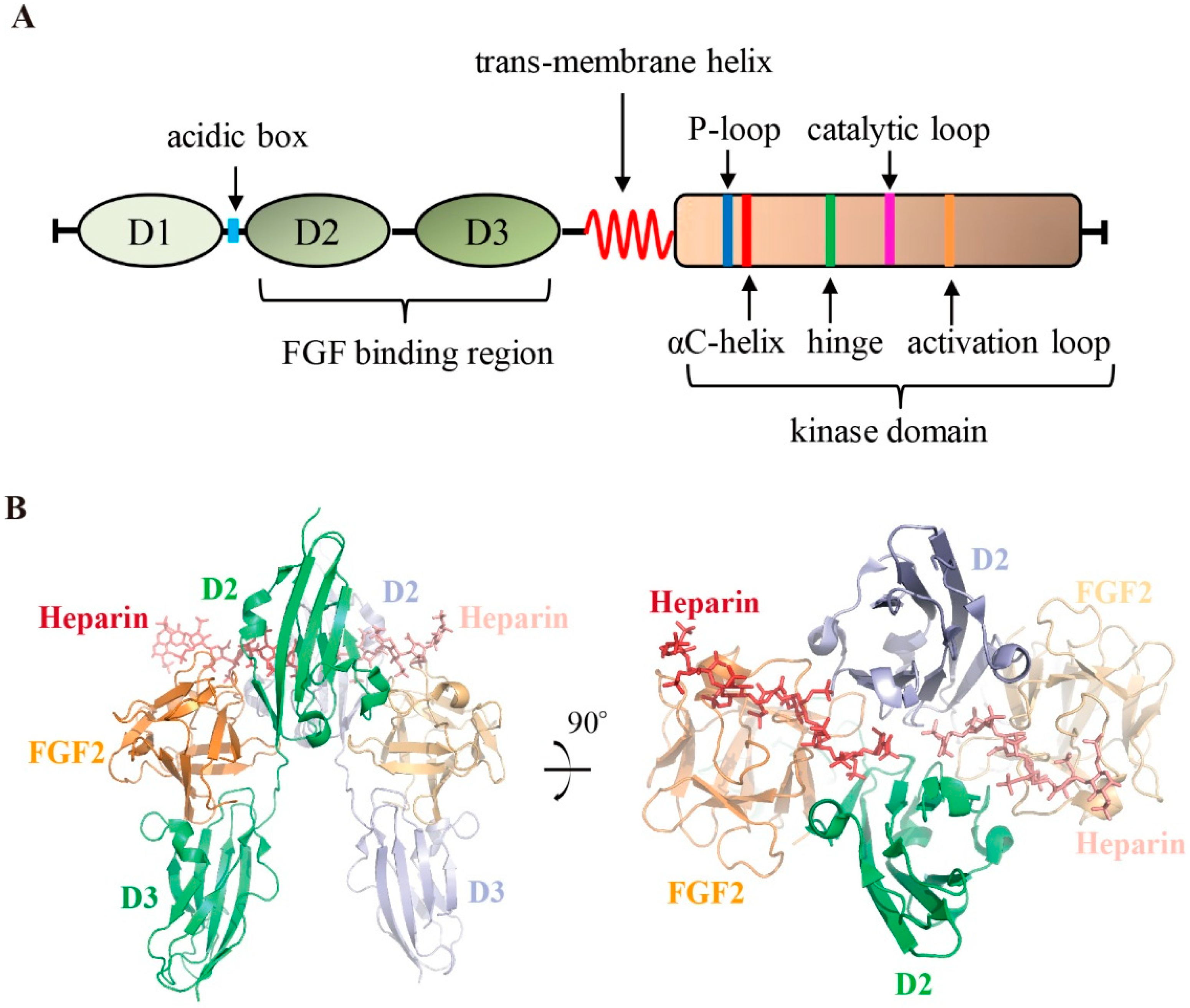
2. Organization of FGFR
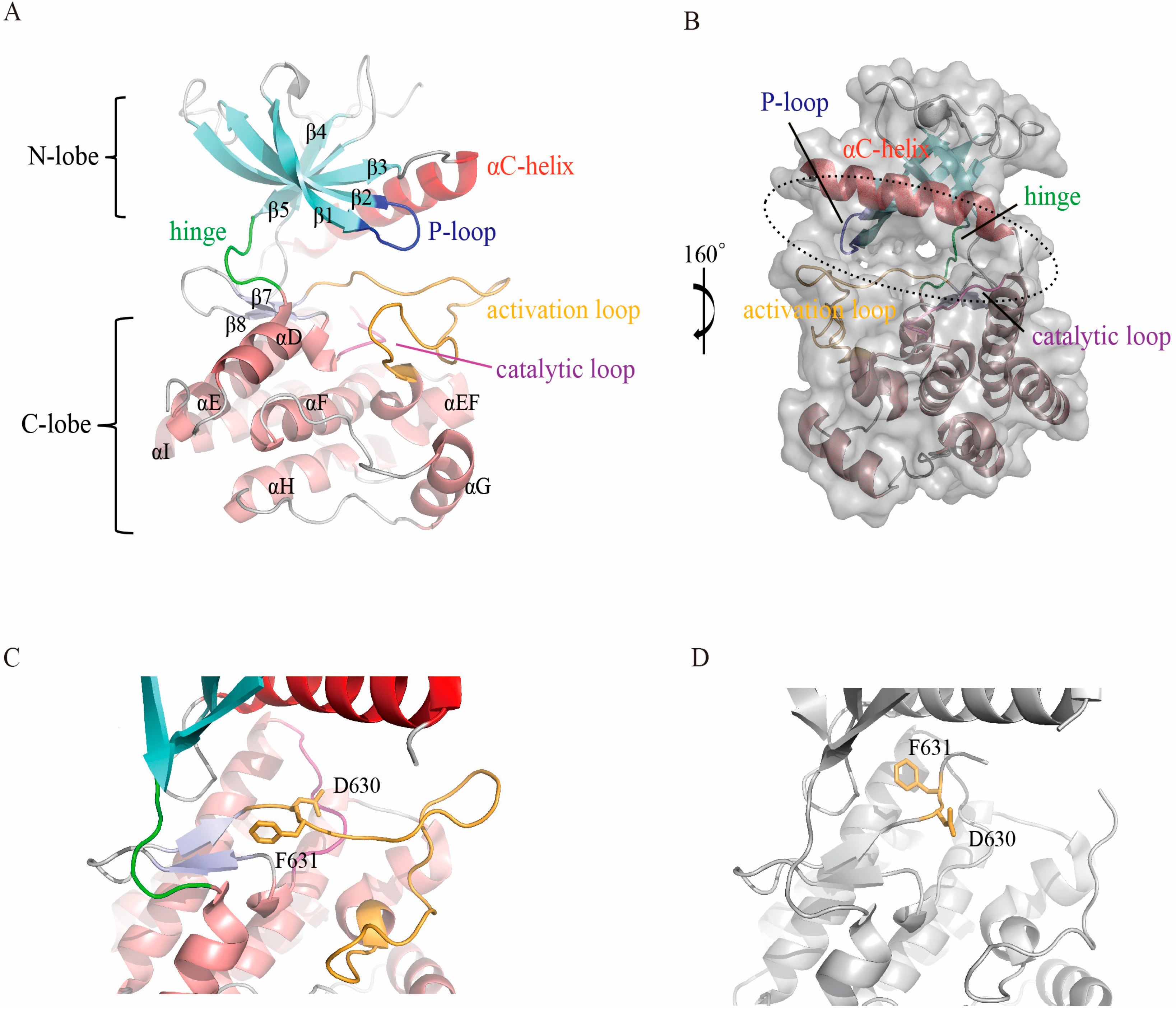
3. Structure of FGFR Kinase Domain
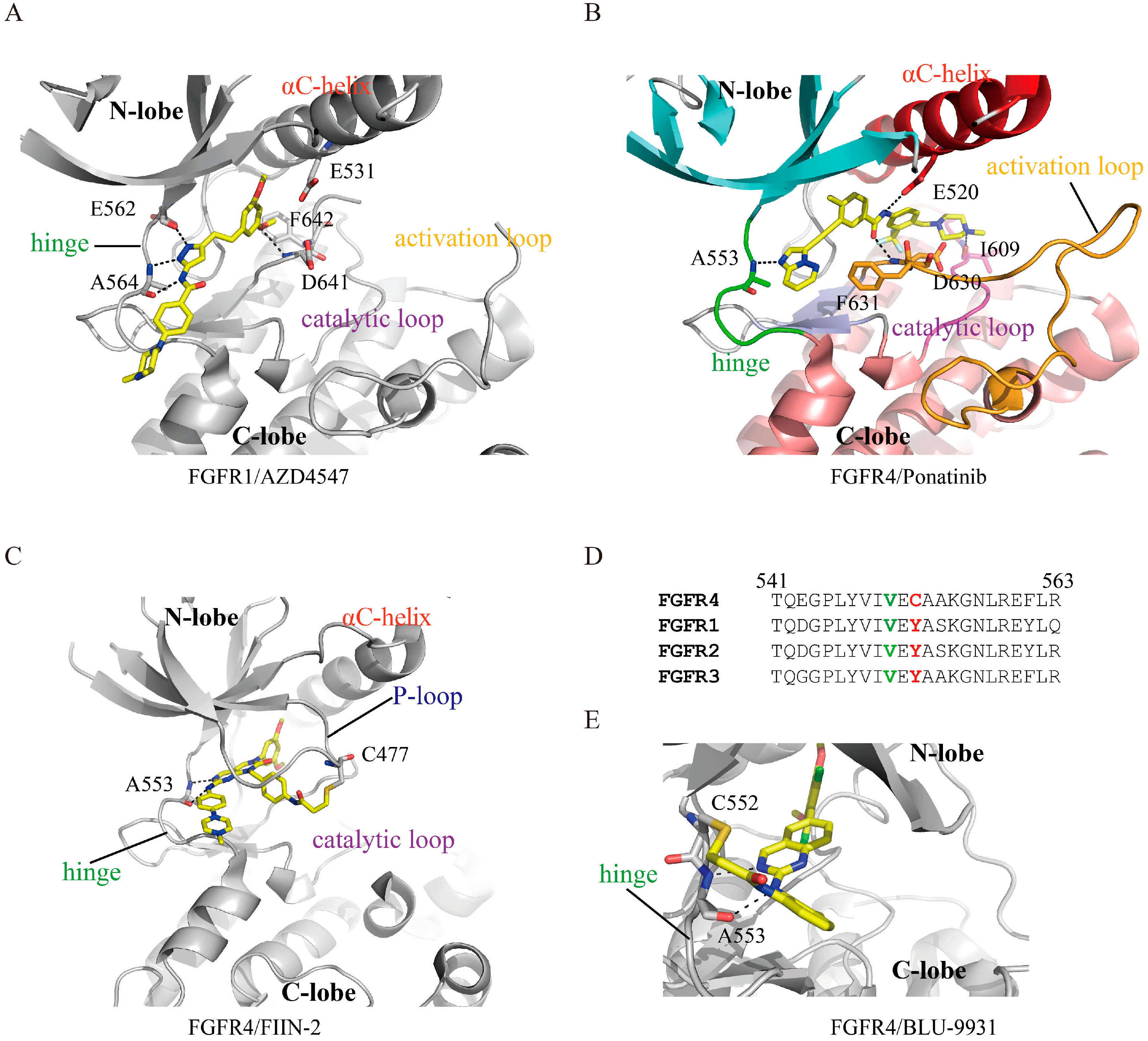
4. Characteristics of FGFR/Inhibitor Interaction
5. Current Status of Small Molecule FGFR Inhibitor Development
6. FGFR Gatekeeper Mutation and Drug Resistance
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Itoh, N.; Ornitz, D.M. Evolution of the Fgf and Fgfr gene families. Trends Genet. 2004, 20, 563–569. [Google Scholar] [CrossRef] [PubMed]
- Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 2000, 103, 211–225. [Google Scholar] [CrossRef]
- Weiner, H.L.; Zagzag, D. Growth factor receptor tyrosine kinases: Cell adhesion kinase family suggests a novel signaling mechanism in cancer. Cancer Investig. 2000, 18, 544–554. [Google Scholar] [CrossRef]
- Lemmon, M.A.; Schlessinger, J. Cell signaling by receptor tyrosine kinases. Cell 2010, 141, 1117–1134. [Google Scholar] [CrossRef] [PubMed]
- Schlessinger, J. Cell signaling by receptor tyrosine kinases: From basic concepts to clinical applications. Eur. J. Cancer Suppl. 2006, 4, 3–26. [Google Scholar] [CrossRef][Green Version]
- Schlessinger, J. Receptor Tyrosine Kinases: Legacy of the First Two Decades. Cold Spring Harb. Perspect. Biol. 2014. [Google Scholar] [CrossRef] [PubMed]
- Ornitz, D.M.; Itoh, N. The Fibroblast Growth Factor signaling pathway. Wiley Interdiscip. Rev. Dev. Biol. 2015, 4, 215–266. [Google Scholar] [CrossRef] [PubMed]
- Andrew, B.; Moosa, M. The FGF family: Biology, pathophysiology and therapy. Nat. Rev. Drug Discov. 2009, 8, 235–253. [Google Scholar]
- Karel, D.; Enrique, A. FGF signalling: Diverse roles during early vertebrate embryogenesis. Development 2010, 137, 3731–3742. [Google Scholar]
- Gowardhan, B.; Douglas, D.A.; Mathers, M.E.; McKie, A.B.; McCracken, S.R.C.; Robson, C.N.; Leung, H.Y. Evaluation of the fibroblast growth factor system as a potential target for therapy in human prostate cancer. Br. J. Cancer 2005, 92, 320–327. [Google Scholar] [CrossRef]
- Brooks, A.N.; Kilgour, E.; Smith, P.D. Molecular pathways: Fibroblast growth factor signaling: A new therapeutic opportunity in cancer. Clin. Cancer Res. 2012, 18, 1855–1862. [Google Scholar] [CrossRef] [PubMed]
- Turner, N.; Grose, R. Fibroblast growth factor signalling: From development to cancer. Nat. Rev. Cancer 2010, 10, 116–129. [Google Scholar] [CrossRef] [PubMed]
- Porta, D.G.; Weiss, A.; Fairhurst, R.A.; Wartmann, M.; Stamm, C.; Reimann, F.; Buhles, A.; Kinyamu-Akunda, J.; Sterker, D.; Murakami, M. Abstract 2098: NVP-FGF401, a first-in-class highly selective and potent FGFR4 inhibitor for the treatment of HCC. Cancer Res. 2017. [Google Scholar] [CrossRef]
- Gavine, P.R.; Mooney, L.; Kilgour, E.; Thomas, A.P.; Al-Kadhimi, K.; Beck, S.; Rooney, C.; Coleman, T.; Baker, D.; Mellor, M.J.; et al. AZD4547: An orally bioavailable, potent, and selective inhibitor of the fibroblast growth factor receptor tyrosine kinase family. Cancer Res. 2012, 72, 2045–2056. [Google Scholar] [CrossRef] [PubMed]
- Perera, T.P.S.; Jovcheva, E.; Mevellec, L.; Vialard, J.; De Lange, D.; Verhulst, T.; Paulussen, C.; Van De Ven, K.; King, P.; Freyne, E.; et al. Discovery and Pharmacological Characterization of JNJ-42756493 (Erdafitinib), a Functionally Selective Small-Molecule FGFR Family Inhibitor. Mol. Cancer Ther. 2017, 16, 1010–1020. [Google Scholar] [CrossRef]
- Farrell, B.; Breeze, A.L. Structure, activation and dysregulation of fibroblast growth factor receptor kinases: Perspectives for clinical targeting. Biochem. Soc. Trans. 2018, 46, 1753–1770. [Google Scholar] [CrossRef] [PubMed]
- Sanchez-Heras, E.; Howell, F.V.; Williams, G.; Doherty, P. The fibroblast growth factor receptor acid box is essential for interactions with N-cadherin and all of the major isoforms of neural cell adhesion molecule. J. Biol. Chem. 2006, 281, 35208–35216. [Google Scholar] [CrossRef]
- Wang, F.; Kan, M.; Xu, J.; Yan, G.; McKeehan, W.L. Ligand-specific structural domains in the fibroblast growth factor receptor. J. Biol. Chem. 1995, 270, 10222–10230. [Google Scholar] [CrossRef]
- Schlessinger, J.; Plotnikov, A.N.; Ibrahimi, O.A.; Eliseenkova, A.V.; Yeh, B.K.; Yayon, A.; Linhardt, R.J.; Mohammadi, M. Crystal structure of a ternary FGF-FGFR-heparin complex reveals a dual role for heparin in FGFR binding and dimerization. Mol. Cell 2000, 6, 743–750. [Google Scholar] [CrossRef]
- Pellegrini, L.; Burke, D.F.; von Delft, F.; Mulloy, B.; Blundell, T.L. Crystal structure of fibroblast growth factor receptor ectodomain bound to ligand and heparin. Nature 2000, 407, 1029–1034. [Google Scholar] [CrossRef]
- Saxena, K.; Schieborr, U.; Anderka, O.; Duchardt-Ferner, E.; Elshorst, B.; Gande, S.L.; Janzon, J.; Kudlinzki, D.; Sreeramulu, S.; Dreyer, M.K.; et al. Influence of heparin mimetics on assembly of the FGF.FGFR4 signaling complex. J. Biol. Chem. 2010, 285, 26628–26640. [Google Scholar] [CrossRef] [PubMed]
- Eriksson, A.E.; Cousens, L.S.; Weaver, L.H.; Matthews, B.W. Three-dimensional structure of human basic fibroblast growth factor. Proc. Natl. Acad. Sci. USA 1991, 88, 3441–3445. [Google Scholar] [CrossRef] [PubMed]
- Goetz, R.; Mohammadi, M. Exploring mechanisms of FGF signalling through the lens of structural biology. Nat. Rev. Mol. Cell Biol. 2013, 14, 166–180. [Google Scholar] [CrossRef] [PubMed]
- Razzaque, M.S. The FGF23-Klotho axis: Endocrine regulation of phosphate homeostasis. Nat. Rev. Endocrinol. 2009, 5, 611–619. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Liu, Y.; Goetz, R.; Fu, L.; Jayaraman, S.; Hu, M.C.; Moe, O.W.; Liang, G.; Li, X.; Mohammadi, M. alpha-Klotho is a non-enzymatic molecular scaffold for FGF23 hormone signalling. Nature 2018, 553, 461–466. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Choi, J.; Mohanty, J.; Sousa, L.P.; Tome, F.; Pardon, E.; Steyaert, J.; Lemmon, M.A.; Lax, I.; Schlessinger, J. Structures of beta-klotho reveal a ‘zip code’-like mechanism for endocrine FGF signalling. Nature 2018, 553, 501–505. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi, M.; Olsen, S.K.; Ibrahimi, O.A. Structural basis for fibroblast growth factor receptor activation. Cytokine Growth Factor Rev. 2005, 16, 107–137. [Google Scholar] [CrossRef]
- Kalinina, J.; Dutta, K.; Ilghari, D.; Beenken, A.; Goetz, R.; Eliseenkova, A.V.; Cowburn, D.; Mohammadi, M. The alternatively spliced acid box region plays a key role in FGF receptor autoinhibition. Structure 2012, 20, 77–88. [Google Scholar] [CrossRef]
- Olsen, S.K.; Ibrahimi, O.A.; Raucci, A.; Zhang, F.; Eliseenkova, A.V.; Yayon, A.; Basilico, C.; Linhardt, R.J.; Schlessinger, J.; Mohammadi, M. Insights into the molecular basis for fibroblast growth factor receptor autoinhibition and ligand-binding promiscuity. Proc. Natl. Acad. Sci. USA 2004, 101, 935–940. [Google Scholar] [CrossRef]
- Wang, F.; Kan, M.; Yan, G.; Xu, J.; McKeehan, W.L. Alternately spliced NH2-terminal immunoglobulin-like Loop I in the ectodomain of the fibroblast growth factor (FGF) receptor 1 lowers affinity for both heparin and FGF-1. J. Biol. Chem. 1995, 270, 10231–10235. [Google Scholar] [CrossRef]
- Mohammadi, M.; Schlessinger, J.; Hubbard, S.R. Structure of the FGF receptor tyrosine kinase domain reveals a novel autoinhibitory mechanism. Cell 1996, 86, 577–587. [Google Scholar] [CrossRef]
- Wu, D.; Guo, M.; Min, X.; Dai, S.; Li, M.; Tan, S.; Li, G.; Chen, X.; Ma, Y.; Li, J.; et al. LY2874455 potently inhibits FGFR gatekeeper mutants and overcomes mutation-based resistance. Chem. Commun. (Camb.) 2018, 54, 12089–12092. [Google Scholar] [CrossRef] [PubMed]
- Wu, D.; Guo, M.; Philips, M.A.; Qu, L.; Jiang, L.; Li, J.; Chen, X.; Chen, Z.; Chen, L.; Chen, Y. Crystal Structure of the FGFR4/LY2874455 Complex Reveals Insights into the Pan-FGFR Selectivity of LY2874455. PLoS ONE 2016, 11, e0162491. [Google Scholar] [CrossRef] [PubMed]
- Tucker, J.A.; Klein, T.; Breed, J.; Breeze, A.L.; Overman, R.; Phillips, C.; Norman, R.A. Structural insights into FGFR kinase isoform selectivity: Diverse binding modes of AZD4547 and ponatinib in complex with FGFR1 and FGFR4. Structure 2014, 22, 1764–1774. [Google Scholar] [CrossRef] [PubMed]
- Ni, F.; Kung, A.; Duan, Y.; Shah, V.; Amador, C.D.; Guo, M.; Fan, X.; Chen, L.; Chen, Y.; McKenna, C.E.; et al. Remarkably Stereospecific Utilization of ATP alpha,beta-Halomethylene Analogues by Protein Kinases. J. Am. Chem. Soc. 2017, 139, 7701–7704. [Google Scholar] [CrossRef] [PubMed]
- Knighton, D.R.; Zheng, J.H.; Ten Eyck, L.F.; Ashford, V.A.; Xuong, N.H.; Taylor, S.S.; Sowadski, J.M. Crystal structure of the catalytic subunit of cyclic adenosine monophosphate-dependent protein kinase. Science 1991, 253, 407–414. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Ye, Q.; Jia, Z.; Cote, G.P. Characterization of the Catalytic and Nucleotide Binding Properties of the alpha-Kinase Domain of Dictyostelium Myosin-II Heavy Chain Kinase A. J. Biol. Chem. 2015, 290, 23935–23946. [Google Scholar] [CrossRef]
- Furdui, C.M.; Lew, E.D.; Schlessinger, J.; Anderson, K.S. Autophosphorylation of FGFR1 kinase is mediated by a sequential and precisely ordered reaction. Mol. Cell 2006, 21, 711–717. [Google Scholar] [CrossRef]
- Kornev, A.P.; Taylor, S.S.; Ten Eyck, L.F. A helix scaffold for the assembly of active protein kinases. Proc. Natl. Acad. Sci. USA 2008, 105, 14377–14382. [Google Scholar] [CrossRef]
- Duan, Y.; Chen, L.; Chen, Y.; Fan, X.G. c-Src binds to the cancer drug Ruxolitinib with an active conformation. PLoS ONE 2014, 9, e106225. [Google Scholar] [CrossRef]
- Hu, J.; Ahuja, L.G.; Meharena, H.S.; Kannan, N.; Kornev, A.P.; Taylor, S.S.; Shaw, A.S. Kinase regulation by hydrophobic spine assembly in cancer. Mol. Cell. Biol. 2015, 35, 264–276. [Google Scholar] [CrossRef] [PubMed]
- Hari, S.B.; Merritt, E.A.; Maly, D.J. Sequence determinants of a specific inactive protein kinase conformation. Chem. Biol. 2013, 20, 806–815. [Google Scholar] [CrossRef] [PubMed]
- Vijayan, R.S.; He, P.; Modi, V.; Duong-Ly, K.C.; Ma, H.; Peterson, J.R.; Dunbrack, R.L., Jr.; Levy, R.M. Conformational analysis of the DFG-out kinase motif and biochemical profiling of structurally validated type II inhibitors. J. Med. Chem. 2015, 58, 466–479. [Google Scholar] [CrossRef] [PubMed]
- Klein, T.; Vajpai, N.; Phillips, J.J.; Davies, G.; Holdgate, G.A.; Phillips, C.; Tucker, J.A.; Norman, R.A.; Scott, A.D.; Higazi, D.R.; et al. Structural and dynamic insights into the energetics of activation loop rearrangement in FGFR1 kinase. Nat. Commun. 2015. [Google Scholar] [CrossRef]
- Guimaraes, C.R.; Rai, B.K.; Munchhof, M.J.; Liu, S.; Wang, J.; Bhattacharya, S.K.; Buckbinder, L. Understanding the impact of the P-loop conformation on kinase selectivity. J. Chem. Inf. Model. 2011, 51, 1199–1204. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Ma, J.; Li, W.; Eliseenkova, A.V.; Xu, C.; Neubert, T.A.; Miller, W.T.; Mohammadi, M. A molecular brake in the kinase hinge region regulates the activity of receptor tyrosine kinases. Mol. Cell 2007, 27, 717–730. [Google Scholar] [CrossRef] [PubMed]
- Roskoski, R., Jr. Src protein-tyrosine kinase structure, mechanism, and small molecule inhibitors. Pharm. Res. 2015, 94, 9–25. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Li, W.Y.; Chen, D.; Henry, J.R.; Li, H.Y.; Chen, Z.; Zia-Ebrahimi, M.; Bloem, L.; Zhai, Y.; Huss, K.; et al. A novel, selective inhibitor of fibroblast growth factor receptors that shows a potent broad spectrum of antitumor activity in several tumor xenograft models. Mol. Cancer. Ther. 2011, 10, 2200–2210. [Google Scholar] [CrossRef]
- Nakanishi, Y.; Akiyama, N.; Tsukaguchi, T.; Fujii, T.; Sakata, K.; Sase, H.; Isobe, T.; Morikami, K.; Shindoh, H.; Mio, T.; et al. The fibroblast growth factor receptor genetic status as a potential predictor of the sensitivity to CH5183284/Debio 1347, a novel selective FGFR inhibitor. Mol. Cancer. Ther. 2014, 13, 2547–2558. [Google Scholar] [CrossRef]
- Guagnano, V.; Furet, P.; Spanka, C.; Bordas, V.; Le Douget, M.; Stamm, C.; Brueggen, J.; Jensen, M.R.; Schnell, C.; Schmid, H.; et al. Discovery of 3-(2,6-dichloro-3,5-dimethoxy-phenyl)-1-{6-[4-(4-ethyl-piperazin-1-yl)-phenylamin o]-pyrimidin-4-yl}-1-methyl-urea (NVP-BGJ398), a potent and selective inhibitor of the fibroblast growth factor receptor family of receptor tyrosine kinase. J. Med. Chem. 2011, 54, 7066–7083. [Google Scholar] [CrossRef]
- Karkera, J.D.; Cardona, G.M.; Bell, K.; Gaffney, D.; Portale, J.C.; Santiago-Walker, A.; Moy, C.H.; King, P.; Sharp, M.; Bahleda, R.; et al. Oncogenic Characterization and Pharmacologic Sensitivity of Activating Fibroblast Growth Factor Receptor (FGFR) Genetic Alterations to the Selective FGFR Inhibitor Erdafitinib. Mol. Cancer. Ther. 2017, 16, 1717–1726. [Google Scholar] [CrossRef] [PubMed]
- Collin, M.P.; Lobell, M.; Hubsch, W.; Brohm, D.; Schirok, H.; Jautelat, R.; Lustig, K.; Bomer, U.; Vohringer, V.; Heroult, M.; et al. Discovery of Rogaratinib (BAY 1163877): A pan-FGFR Inhibitor. Chem. Med. Chem. 2018, 13, 437–445. [Google Scholar] [CrossRef] [PubMed]
- Brameld, K.A.; Owens, T.D.; Verner, E.; Venetsanakos, E.; Bradshaw, J.M.; Phan, V.T.; Tam, D.; Leung, K.; Shu, J.; LaStant, J.; et al. Discovery of the Irreversible Covalent FGFR Inhibitor 8-(3-(4-Acryloylpiperazin-1-yl)propyl)-6-(2,6-dichloro-3,5-dimethoxyphenyl)-2-(me thylamino)pyrido[2,3-d]pyrimidin-7(8H)-one (PRN1371) for the Treatment of Solid Tumors. J. Med. Chem. 2017, 60, 6516–6527. [Google Scholar] [CrossRef] [PubMed]
- Kalyukina, M.; Yosaatmadja, Y.; Middleditch, M.J.; Patterson, A.V.; Smaill, J.B.; Squire, C.J. TAS-120 Cancer Target Binding: Defining Reactivity and Revealing the First Fibroblast Growth Factor Receptor 1 (FGFR1) Irreversible Structure. ChemMedChem 2019, 14, 494–500. [Google Scholar] [CrossRef] [PubMed]
- Kim, R.; Sharma, S.; Meyer, T.; Sarker, D.; Macarulla, T.; Sung, M.; Choo, S.P.; Shi, H.; Schmidt-Kittler, O.; Clifford, C.; et al. First-in-human study of BLU-554, a potent, highly-selective FGFR4 inhibitor designed for hepatocellular carcinoma (HCC) with FGFR4 pathway activation. Eur. J. Cancer 2016, 69, S41. [Google Scholar] [CrossRef]
- Joshi, J.J.; Coffey, H.; Corcoran, E.; Tsai, J.; Huang, C.L.; Ichikawa, K.; Prajapati, S.; Hao, M.H.; Bailey, S.; Wu, J.; et al. H3B-6527 Is a Potent and Selective Inhibitor of FGFR4 in FGF19-Driven Hepatocellular Carcinoma. Cancer Res. 2017, 77, 6999–7013. [Google Scholar] [CrossRef]
- Zhou, Z.; Chen, X.; Fu, Y.; Zhang, Y.; Dai, S.; Li, J.; Chen, L.; Xu, G.; Chen, Z.; Chen, Y. Characterization of FGF401 as a reversible covalent inhibitor of fibroblast growth factor receptor 4. Chem. Commun. (Camb.) 2019. [Google Scholar] [CrossRef]
- Roskoski, R., Jr. ERK1/2 MAP kinases: Structure, function, and regulation. Pharm. Res. 2012, 66, 105–143. [Google Scholar]
- Dar, A.C.; Shokat, K.M. The evolution of protein kinase inhibitors from antagonists to agonists of cellular signaling. Annu. Rev. Biochem. 2011, 80, 769–795. [Google Scholar] [CrossRef]
- Norman, R.A.; Schott, A.K.; Andrews, D.M.; Breed, J.; Foote, K.M.; Garner, A.P.; Ogg, D.; Orme, J.P.; Pink, J.H.; Roberts, K.; et al. Protein-ligand crystal structures can guide the design of selective inhibitors of the FGFR tyrosine kinase. J. Med. Chem. 2012, 55, 5003–5012. [Google Scholar] [CrossRef]
- Liu, Y.; Gray, N.S. Rational design of inhibitors that bind to inactive kinase conformations. Nat. Chem. Biol. 2006, 2, 358–364. [Google Scholar] [CrossRef] [PubMed]
- Mohammadi, M.; Froum, S.; Hamby, J.M.; Schroeder, M.C.; Panek, R.L.; Lu, G.H.; Eliseenkova, A.V.; Green, D.; Schlessinger, J.; Hubbard, S.R. Crystal structure of an angiogenesis inhibitor bound to the FGF receptor tyrosine kinase domain. EMBO J. 1998, 17, 5896–5904. [Google Scholar] [CrossRef] [PubMed]
- Davis, M.I.; Hunt, J.P.; Herrgard, S.; Ciceri, P.; Wodicka, L.M.; Pallares, G.; Hocker, M.; Treiber, D.K.; Zarrinkar, P.P. Comprehensive analysis of kinase inhibitor selectivity. Nat. Biotechnol. 2011, 29, 1046–1051. [Google Scholar] [CrossRef]
- Baillie, T.A. Targeted Covalent Inhibitors for Drug Design. Angew. Chem. Int. Ed. 2016, 55, 13408–13421. [Google Scholar] [CrossRef] [PubMed]
- Awoonor-Williams, E.; Walsh, A.G.; Rowley, C.N. Modeling covalent-modifier drugs. Biochim. Biophys. Acta 2017, 1865, 1664–1675. [Google Scholar] [CrossRef]
- Liu, Q.; Sabnis, Y.; Zhao, Z.; Zhang, T.; Buhrlage, S.J.; Jones, L.H.; Gray, N.S. Developing irreversible inhibitors of the protein kinase cysteinome. Chem. Biol. 2013, 20, 146–159. [Google Scholar] [CrossRef] [PubMed]
- Serafimova, I.M.; Pufall, M.A.; Krishnan, S.; Duda, K.; Cohen, M.S.; Maglathlin, R.L.; McFarland, J.M.; Miller, R.M.; Frodin, M.; Taunton, J. Reversible targeting of noncatalytic cysteines with chemically tuned electrophiles. Nat. Chem. Biol. 2012, 8, 471–476. [Google Scholar] [CrossRef]
- Zhou, W.; Ercan, D.; Chen, L.; Yun, C.H.; Li, D.; Capelletti, M.; Cortot, A.B.; Chirieac, L.; Iacob, R.E.; Padera, R.; et al. Novel mutant-selective EGFR kinase inhibitors against EGFR T790M. Nature 2009, 462, 1070–1074. [Google Scholar] [CrossRef]
- Zhang, J.; Yang, P.L.; Gray, N.S. Targeting cancer with small molecule kinase inhibitors. Nat. Rev. Cancer 2009, 9, 28–39. [Google Scholar] [CrossRef]
- Powis, G.; Bonjouklian, R.; Berggren, M.M.; Gallegos, A.; Abraham, R.; Ashendel, C.; Zalkow, L.; Matter, W.F.; Dodge, J.; Grindey, G.; et al. Wortmannin, a potent and selective inhibitor of phosphatidylinositol-3-kinase. Cancer Res. 1994, 54, 2419–2423. [Google Scholar]
- Fox, T.; Fitzgibbon, M.J.; Fleming, M.A.; Hsiao, H.M.; Brummel, C.L.; Su, M.S. Kinetic mechanism and ATP-binding site reactivity of p38gamma MAP kinase. FEBS Lett. 1999, 461, 323–328. [Google Scholar] [CrossRef]
- Shannon, D.A.; Weerapana, E. Covalent protein modification: The current landscape of residue-specific electrophiles. Curr. Opin. Chem. Biol. 2015, 24, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Wang, J.; Tanizaki, J.; Huang, Z.; Aref, A.R.; Rusan, M.; Zhu, S.J.; Zhang, Y.; Ercan, D.; Liao, R.G.; et al. Development of covalent inhibitors that can overcome resistance to first-generation FGFR kinase inhibitors. Proc. Natl. Acad. Sci. USA 2014, 111, E4869–E4877. [Google Scholar] [CrossRef] [PubMed]
- Paul, S.M.; Mytelka, D.S.; Dunwiddie, C.T.; Persinger, C.C.; Munos, B.H.; Lindborg, S.R.; Schacht, A.L. How to improve R&D productivity: The pharmaceutical industry’s grand challenge. Nat. Rev. Drug Discov. 2010, 9, 203–214. [Google Scholar] [CrossRef] [PubMed]
- Fedorov, O.; Muller, S.; Knapp, S. The (un)targeted cancer kinome. Nat. Chem. Biol. 2010, 6, 166–169. [Google Scholar] [CrossRef] [PubMed]
- O’Hare, T.; Shakespeare, W.C.; Zhu, X.; Eide, C.A.; Rivera, V.M.; Wang, F.; Adrian, L.T.; Zhou, T.; Huang, W.S.; Xu, Q.; et al. AP24534, a pan-BCR-ABL inhibitor for chronic myeloid leukemia, potently inhibits the T315I mutant and overcomes mutation-based resistance. Cancer Cell 2009, 16, 401–412. [Google Scholar] [CrossRef] [PubMed]
- Trudel, S.; Li, Z.H.; Wei, E.; Wiesmann, M.; Chang, H.; Chen, C.; Reece, D.; Heise, C.; Stewart, A.K. CHIR-258, a novel, multitargeted tyrosine kinase inhibitor for the potential treatment of t(4;14) multiple myeloma. Blood 2005, 105, 2941–2948. [Google Scholar] [CrossRef]
- Bello, E.; Colella, G.; Scarlato, V.; Oliva, P.; Berndt, A.; Valbusa, G.; Serra, S.C.; D’Incalci, M.; Cavalletti, E.; Giavazzi, R.; et al. E-3810 is a potent dual inhibitor of VEGFR and FGFR that exerts antitumor activity in multiple preclinical models. Cancer Res. 2011, 71, 1396–1405. [Google Scholar] [CrossRef]
- Hilberg, F.; Roth, G.J.; Krssak, M.; Kautschitsch, S.; Sommergruber, W.; Tontsch-Grunt, U.; Garin-Chesa, P.; Bader, G.; Zoephel, A.; Quant, J.; et al. BIBF 1120: Triple angiokinase inhibitor with sustained receptor blockade and good antitumor efficacy. Cancer Res. 2008, 68, 4774–4782. [Google Scholar] [CrossRef]
- Nogova, L.; Sequist, L.V.; Perez Garcia, J.M.; Andre, F.; Delord, J.P.; Hidalgo, M.; Schellens, J.H.; Cassier, P.A.; Camidge, D.R.; Schuler, M.; et al. Evaluation of BGJ398, a Fibroblast Growth Factor Receptor 1–3 Kinase Inhibitor, in Patients with Advanced Solid Tumors Harboring Genetic Alterations in Fibroblast Growth Factor Receptors: Results of a Global Phase I, Dose-Escalation and Dose-Expansion Study. J. Clin. Oncol. 2017, 35, 157–165. [Google Scholar] [CrossRef]
- Degirolamo, C.; Sabba, C.; Moschetta, A. Therapeutic potential of the endocrine fibroblast growth factors FGF19, FGF21 and FGF23. Nat. Rev. Drug Discov. 2016, 15, 51–69. [Google Scholar] [CrossRef]
- Chell, V.; Balmanno, K.; Little, A.S.; Wilson, M.; Andrews, S.; Blockley, L.; Hampson, M.; Gavine, P.R.; Cook, S.J. Tumour cell responses to new fibroblast growth factor receptor tyrosine kinase inhibitors and identification of a gatekeeper mutation in FGFR3 as a mechanism of acquired resistance. Oncogene 2013, 32, 3059–3070. [Google Scholar] [CrossRef] [PubMed]
- Zhou, W.; Hur, W.; McDermott, U.; Dutt, A.; Xian, W.; Ficarro, S.B.; Zhang, J.; Sharma, S.V.; Brugge, J.; Meyerson, M.; et al. A structure-guided approach to creating covalent FGFR inhibitors. Chem. Biol. 2010, 17, 285–295. [Google Scholar] [CrossRef] [PubMed]
- Hagel, M.; Miduturu, C.; Sheets, M.; Rubin, N.; Weng, W.; Stransky, N.; Bifulco, N.; Kim, J.L.; Hodous, B.; Brooijmans, N.; et al. First Selective Small Molecule Inhibitor of FGFR4 for the Treatment of Hepatocellular Carcinomas with an Activated FGFR4 Signaling Pathway. Cancer Discov. 2015, 5, 424–437. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Tan, L.; Wang, H.; Liu, Y.; Blais, S.; Deng, J.; Neubert, T.A.; Gray, N.S.; Li, X.; Mohammadi, M. DFG-out mode of inhibition by an irreversible type-1 inhibitor capable of overcoming gate-keeper mutations in FGF receptors. ACS Chem. Biol. 2015, 10, 299–309. [Google Scholar] [CrossRef] [PubMed]
- Fairhurst, R.A.; Knoepfel, T.; Leblanc, C.; Buschmann, N.; Gaul, C.; Blank, J.; Galuba, I.; Trappe, J.; Zou, C.; Voshol, J.; et al. Approaches to selective fibroblast growth factor receptor 4 inhibition through targeting the ATP-pocket middle-hinge region. MedChemComm 2017, 8, 1604–1613. [Google Scholar] [CrossRef]
- Ho, H.K.; Yeo, A.H.; Kang, T.S.; Chua, B.T. Current strategies for inhibiting FGFR activities in clinical applications: Opportunities, challenges and toxicological considerations. Drug Discov. Today 2014, 19, 51–62. [Google Scholar] [CrossRef]
- Katoh, M. FGFR inhibitors: Effects on cancer cells, tumor microenvironment and whole-body homeostasis (Review). Int. J. Mol. Med. 2016, 38, 3–15. [Google Scholar] [CrossRef]
- Lu, X.; Chen, H.; Patterson, A.V.; Smaill, J.B.; Ding, K. Fibroblast Growth Factor Receptor 4 (FGFR4) Selective Inhibitors as Hepatocellular Carcinoma Therapy: Advances and Prospects. J. Med. Chem. 2019, 62, 2905–2915. [Google Scholar] [CrossRef]
- Hierro, C.; Rodon, J.; Tabernero, J. Fibroblast Growth Factor (FGF) Receptor/FGF Inhibitors: Novel Targets and Strategies for Optimization of Response of Solid Tumors. Semin. Oncol. 2015, 42, 801–819. [Google Scholar] [CrossRef]
- Bradshaw, J.M.; McFarland, J.M.; Paavilainen, V.O.; Bisconte, A.; Tam, D.; Phan, V.T.; Romanov, S.; Finkle, D.; Shu, J.; Patel, V.; et al. Prolonged and tunable residence time using reversible covalent kinase inhibitors. Nat. Chem. Biol. 2015, 11, 525–531. [Google Scholar] [CrossRef] [PubMed]
- Knoepfel, T.; Furet, P.; Mah, R.; Buschmann, N.; Leblanc, C.; Ripoche, S.; Graus-Porta, D.; Wartmann, M.; Galuba, I.; Fairhurst, R.A. 2-Formylpyridyl Ureas as Highly Selective Reversible-Covalent Inhibitors of Fibroblast Growth Factor Receptor 4. ACS Med. Chem. Lett. 2018, 9, 215–220. [Google Scholar] [CrossRef] [PubMed]
- Shabani, M.; Hojjat-Farsangi, M. Targeting Receptor Tyrosine Kinases Using Monoclonal Antibodies: The Most Specific Tools for Targeted-Based Cancer Therapy. Curr. Drug Targets 2016, 17, 1687–1703. [Google Scholar] [CrossRef] [PubMed]
- Pierce, K.L.; Deshpande, A.M.; Stohr, B.A.; Gemo, A.T.; Patil, N.S.; Brennan, T.J.; Bellovin, D.I.; Palencia, S.; Giese, T.; Huang, C.; et al. FPA144, a humanized monoclonal antibody for both FGFR2-amplified and nonamplified, FGFR2b-overexpressing gastric cancer patients. J. Clin. Oncol. 2014. [Google Scholar] [CrossRef]
- Sommer, A.; Kopitz, C.; Schatz, C.A.; Nising, C.F.; Mahlert, C.; Lerchen, H.G.; Stelte-Ludwig, B.; Hammer, S.; Greven, S.; Schuhmacher, J.; et al. Preclinical Efficacy of the Auristatin-Based Antibody-Drug Conjugate BAY 1187982 for the Treatment of FGFR2-Positive Solid Tumors. Cancer Res. 2016, 76, 6331–6339. [Google Scholar] [CrossRef] [PubMed]
- Schatz, C.A.; Kopitz, C.; Wittemer-Rump, S.; Sommer, A.; Lindbom, L.; Osada, M.; Yamanouchi, H.; Huynh, H.; Krahn, T.; Asadullah, K. Abstract 4766: Pharmacodynamic and stratification biomarker for the anti-FGFR2 antibody (BAY1179470) and the FGFR2-ADC. Cancer Res. 2014. [Google Scholar] [CrossRef]
- Trudel, S.; Bergsagel, P.L.; Singhal, S.; Niesvizky, R.; Comenzo, R.L.; Bensinger, W.I.; Lebovic, D.; Choi, Y.; Lu, D.; French, D.; et al. A Phase I Study of the Safety and Pharmacokinetics of Escalating Doses of MFGR1877S, a Fibroblast Growth Factor Receptor 3 (FGFR3) Antibody, in Patients with Relapsed or Refractory t(4;14)-Positive Multiple Myeloma. Blood 2012, 120, 4029. [Google Scholar]
- Blackwell, C.; Sherk, C.; Fricko, M.; Ganji, G.; Barnette, M.; Hoang, B.; Tunstead, J.; Skedzielewski, T.; Alsaid, H.; Jucker, B.M.; et al. Inhibition of FGF/FGFR autocrine signaling in mesothelioma with the FGF ligand trap, FP-1039/GSK3052230. Oncotarget 2016, 7, 39861–39871. [Google Scholar] [CrossRef]
- Bono, F.; De Smet, F.; Herbert, C.; De Bock, K.; Georgiadou, M.; Fons, P.; Tjwa, M.; Alcouffe, C.; Ny, A.; Bianciotto, M.; et al. Inhibition of tumor angiogenesis and growth by a small-molecule multi-FGF receptor blocker with allosteric properties. Cancer Cell 2013, 23, 477–488. [Google Scholar] [CrossRef]
- Herbert, C.; Schieborr, U.; Saxena, K.; Juraszek, J.; De Smet, F.; Alcouffe, C.; Bianciotto, M.; Saladino, G.; Sibrac, D.; Kudlinzki, D.; et al. Molecular mechanism of SSR128129E, an extracellularly acting, small-molecule, allosteric inhibitor of FGF receptor signaling. Cancer Cell 2013, 23, 489–501. [Google Scholar] [CrossRef]
- Babina, I.S.; Turner, N.C. Advances and challenges in targeting FGFR signalling in cancer. Nat. Rev. Cancer 2017, 17, 318–332. [Google Scholar] [CrossRef] [PubMed]
- Cheetham, G.M.; Charlton, P.A.; Golec, J.M.; Pollard, J.R. Structural basis for potent inhibition of the Aurora kinases and a T315I multi-drug resistant mutant form of Abl kinase by VX-680. Cancer Lett. 2007, 251, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Weisberg, E.; Choi, H.G.; Ray, A.; Barrett, R.; Zhang, J.; Sim, T.; Zhou, W.; Seeliger, M.; Cameron, M.; Azam, M.; et al. Discovery of a small-molecule type II inhibitor of wild-type and gatekeeper mutants of BCR-ABL, PDGFRalpha, Kit, and Src kinases: Novel type II inhibitor of gatekeeper mutants. Blood 2010, 115, 4206–4216. [Google Scholar] [CrossRef] [PubMed]
- Ryan, M.R.; Sohl, C.D.; Luo, B.; Anderson, K.S. The FGFR1 V561M Gatekeeper Mutation Drives AZD4547 Resistance through STAT3 Activation and EMT. Mol. Cancer Res. 2019, 17, 532–543. [Google Scholar] [CrossRef] [PubMed]
- Byron, S.A.; Chen, H.; Wortmann, A.; Loch, D.; Gartside, M.G.; Dehkhoda, F.; Blais, S.P.; Neubert, T.A.; Mohammadi, M.; Pollock, P.M. The N550K/H mutations in FGFR2 confer differential resistance to PD173074, dovitinib, and ponatinib ATP-competitive inhibitors. Neoplasia 2013, 15, 975–988. [Google Scholar] [CrossRef] [PubMed]
- Yoza, K.; Himeno, R.; Amano, S.; Kobashigawa, Y.; Amemiya, S.; Fukuda, N.; Kumeta, H.; Morioka, H.; Inagaki, F. Biophysical characterization of drug-resistant mutants of fibroblast growth factor receptor 1. Genes Cells 2016, 21, 1049–1058. [Google Scholar] [CrossRef]
- Goyal, L.; Saha, S.K.; Liu, L.Y.; Siravegna, G.; Leshchiner, I.; Ahronian, L.G.; Lennerz, J.K.; Vu, P.; Deshpande, V.; Kambadakone, A.; et al. Polyclonal Secondary FGFR2 Mutations Drive Acquired Resistance to FGFR Inhibition in Patients with FGFR2 Fusion-Positive Cholangiocarcinoma. Cancer Discov. 2017, 7, 252–263. [Google Scholar] [CrossRef]
- Ang, D.; Ballard, M.; Beadling, C.; Warrick, A.; Schilling, A.; O’Gara, R.; Pukay, M.; Neff, T.L.; West, R.B.; Corless, C.L.; et al. Novel mutations in neuroendocrine carcinoma of the breast: Possible therapeutic targets. Appl. Immunohistochem. Mol. Morphol. 2015, 23, 97–103. [Google Scholar] [CrossRef]
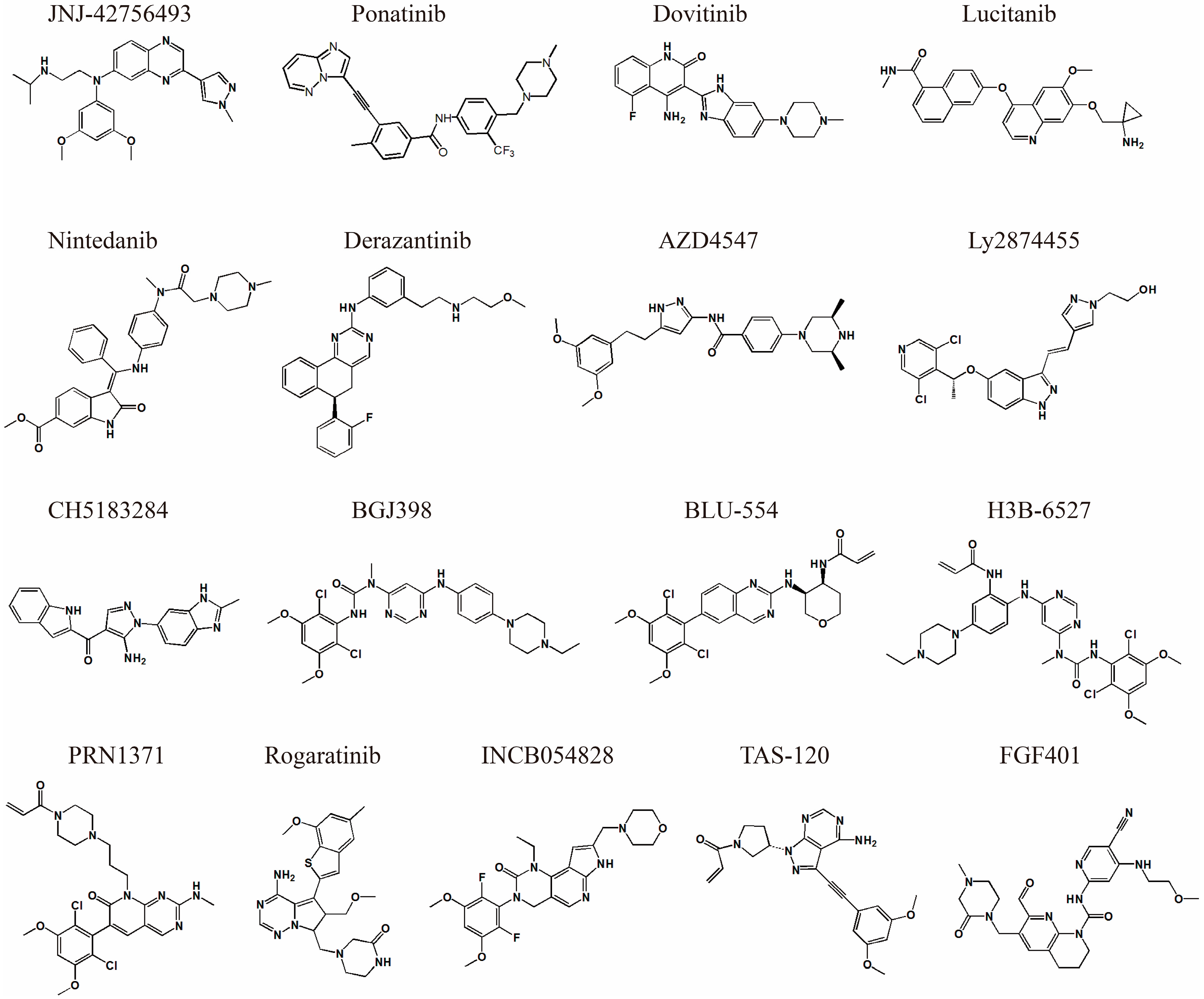
| Inhibitor Name | Binding Features | IC50 (nM) | PDB ID | Clinical Trial Phase/Number | Reference |
|---|---|---|---|---|---|
| JNJ-42756493 (Erdafitinib) | Pan-FGFR Reversible Type I | FGFR1: 1.2 FGFR2: 2.5 FGFR3: 3.0 FGFR4: 5.7 | n/a | FDA approved | [15] |
| AZD4547 | Pan-FGFR Reversible Type I | FGFR1: 0.2 FGFR2: 2.5 FGFR3: 1.8 FGFR4: 165 | 4V05 | Phase I/II NCT02824133 | [14,34] |
| Ly2874455 | Pan-FGFR Reversible Type I | FGFR1: 2.8 FGFR2: 2.6 FGFR3: 6.4 FGFR4: 6 | 5JKG | Phase I NCT01212107 | [33,48] |
| CH5183284 | Pan-FGFR Reversible Type I | FGFR1: 9.3 FGFR2: 7.6 FGFR3: 22 FGFR4: 290 | 5N7V | Phase II/III NCT03344536 | [49] |
| NVP-BGJ398 | Pan-FGFR Reversible Type I | FGFR1: 0.9 FGFR2: 1.4 FGFR3: 1 FGFR4: 60 | 3TT0 | Phase II NCT02706691 | [50] |
| INCB054828 | Pan-FGFR Reversible Type I | FGFR1: 0.4 FGFR2: 0.5 FGFR3: 1.2 FGFR4: 30 | n/a | Phase II NCT03011372 | [51] |
| Rogaratinib | Pan-FGFR Reversible Type I | FGFR1: 12–15 FGFR2: <1 FGFR3: 19 FGFR4: 33 | n/a | Phase II/III NCT03410693 | [52] |
| PRN1371 | Pan-FGFR Irreversible Type I | FGFR1: 0.6 FGFR2: 1.3 FGFR3: 4.1 FGFR4: 19.3 | n/a | Phase I NCT02608125 | [53] |
| TAS-120 | Pan-FGFR Irreversible Type I | FGFR1: 3.9 FGFR2: 1.3 FGFR3: 1.6 FGFR4: 8.3 | 6M2Q | Phase I/II NCT02052778 | [54] |
| BLU-554 | FGFR4 selective Irreversible, Type I | FGFR1: 624 FGFR2: 1202 FGFR3: 2203 FGFR4: 5 | n/a | Phase I NCT02508467 | [55] |
| H3B-6527 | FGFR4 selective Irreversible, Type I | FGFR1: 320 FGFR2: 1290 FGFR3: 1060 FGFR4: <1.2 | 5VND | Phase I NCT02834780 | [56] |
| FGF401 | FGFR4 selective Reversible Covalent, Type I | FGFR1-3: >10,000 FGFR4: 1.1 | 6JPJ | Phase I/II NCT02325739 | [13,57] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dai, S.; Zhou, Z.; Chen, Z.; Xu, G.; Chen, Y. Fibroblast Growth Factor Receptors (FGFRs): Structures and Small Molecule Inhibitors. Cells 2019, 8, 614. https://doi.org/10.3390/cells8060614
Dai S, Zhou Z, Chen Z, Xu G, Chen Y. Fibroblast Growth Factor Receptors (FGFRs): Structures and Small Molecule Inhibitors. Cells. 2019; 8(6):614. https://doi.org/10.3390/cells8060614
Chicago/Turabian StyleDai, Shuyan, Zhan Zhou, Zhuchu Chen, Guangyu Xu, and Yongheng Chen. 2019. "Fibroblast Growth Factor Receptors (FGFRs): Structures and Small Molecule Inhibitors" Cells 8, no. 6: 614. https://doi.org/10.3390/cells8060614
APA StyleDai, S., Zhou, Z., Chen, Z., Xu, G., & Chen, Y. (2019). Fibroblast Growth Factor Receptors (FGFRs): Structures and Small Molecule Inhibitors. Cells, 8(6), 614. https://doi.org/10.3390/cells8060614






