Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development
Abstract
1. Introduction
2. Materials and Methods
2.1. Preparation of Gene Constructs for In Vivo Expression
2.2. Nuclear Transformations
2.3. Seedling Growth and Trangene Induction
2.4. RNA Extraction, Northern Bloting and RT-qPCR Analyses
2.5. Isolation of Mitochondria and Import Assessment
2.6. Microarray Assays
2.7. Microarray Data Analyses
3. Results
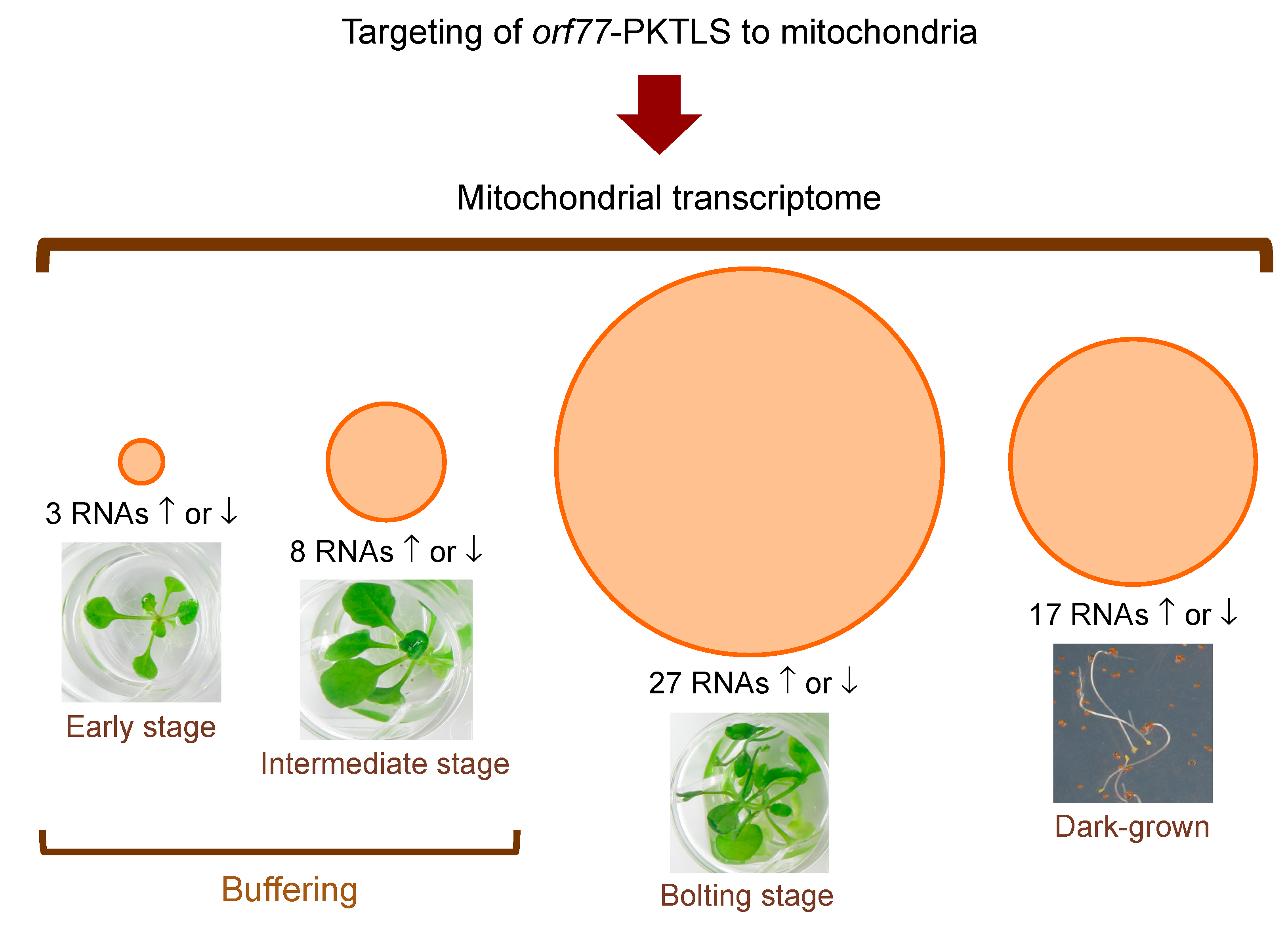
3.1. Trans-Ribozyme-Mediated Modulation of Mitochondrial RNA Steady State Levels Depends on the Plant Developmental Stage
3.2. CMS RNA-Triggered Modulation of Mitochondrial RNA Steady State Levels Depends on the Plant Developmental Stage
3.3. Ribozyme-Mediated Knockdown of a Mitochondrial mRNA Triggers a Nuclear Transcriptome Response
4. Discussion
5. Conclusions
6. Patents
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Shokolenko, I.N.; Alexeyev, M.F. Mitochondrial transcription in mammalian cells. Front. Biosci. 2017, 22, 835–853. [Google Scholar]
- Holec, S.; Lange, H.; Kühn, K.; Alioua, M.; Börner, T.; Gagliardi, D. Relaxed transcription in Arabidopsis mitochondria is counterbalanced by RNA stability control mediated by polyadenylation and polynucleotide phosphorylase. Mol. Cell. Biol. 2006, 26, 2869–2876. [Google Scholar] [CrossRef] [PubMed]
- Rackham, O.; Mercer, T.R.; Filipovska, A. The human mitochondrial transcriptome and the RNA-binding proteins that regulate its expression. Wiley Interdiscip. Rev. RNA 2012, 3, 675–695. [Google Scholar] [CrossRef] [PubMed]
- Small, I.D.; Rackham, O.; Filipovska, A. Organelle transcriptomes: Products of a deconstructed genome. Curr. Opin. Microbiol. 2013, 16, 652–658. [Google Scholar] [CrossRef] [PubMed]
- Pearce, S.F.; Rebelo-Guiomar, P.; D’Souza, A.R.; Powell, C.A.; Van Haute, L.; Minczuk, M. Regulation of mammalian mitochondrial gene expression: Recent advances. Trends Biochem. Sci. 2017, 42, 625–639. [Google Scholar] [CrossRef] [PubMed]
- Rebelo-Guiomar, P.; Powell, C.A.; Van Haute, L.; Minczuk, M. The mammalian mitochondrial epitranscriptome. Biochim. Biophys. Acta Gene Regul. Mech. 2019, 1862, 429–446. [Google Scholar] [CrossRef] [PubMed]
- Pohjoismaki, J.L.; Kruger, M.; Al-Furoukh, N.; Lagerstedt, A.; Karhunen, P.J.; Braun, T. Postnatal cardiomyocyte growth and mitochondrial reorganization cause multiple changes in the proteome of human cardiomyocytes. Mol. BioSyst. 2013, 9, 1210–1219. [Google Scholar] [CrossRef] [PubMed]
- Mposhi, A.; Van der Wijst, M.G.; Faber, K.N.; Rots, M.G. Regulation of mitochondrial gene expression, the epigenetic enigma. Front. Biosci. (Landmark Ed) 2017, 22, 1099–1113. [Google Scholar]
- Barshad, G.; Marom, S.; Cohen, T.; Mishmar, D. Mitochondrial DNA transcription and its regulation: An evolutionary perspective. Trends Genet. 2018, 34, 682–692. [Google Scholar] [CrossRef]
- Robles, P.; Quesada, V. Transcriptional and post-transcriptional regulation of organellar gene expression (OGE) and its roles in plant salt tolerance. Int. J. Mol. Sci. 2019, 20, 1056. [Google Scholar] [CrossRef]
- Woodson, J.D.; Chory, J. Coordination of gene expression between organellar and nuclear genomes. Nat. Rev. Genet. 2008, 9, 383–395. [Google Scholar] [CrossRef] [PubMed]
- Cogliati, S.; Lorenzi, I.; Rigoni, G.; Caicci, F.; Soriano, M.E. Regulation of mitochondrial electron transport chain assembly. J. Mol. Biol. 2018, 430, 4849–4873. [Google Scholar] [CrossRef] [PubMed]
- Weinhouse, C. Mitochondrial-epigenetic crosstalk in environmental toxicology. Toxicology 2017, 391, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Ng, S.; De Clercq, I.; Van Aken, O.; Law, S.R.; Ivanova, A.; Willems, P.; Giraud, E.; Van Breusegem, F.; Whelan, J. Anterograde and retrograde regulation of nuclear genes encoding mitochondrial proteins during growth, development, and stress. Mol. Plant 2014, 7, 1075–1093. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Butow, R.A. Mitochondrial retrograde signaling. Annu. Rev. Genet. 2006, 40, 159–185. [Google Scholar] [CrossRef]
- Dekkers, B.J.; Pearce, S.; van Bolderen-Veldkamp, R.P.; Marshall, A.; Widera, P.; Gilbert, J.; Drost, H.G.; Bassel, G.W.; Müller, K.; King, J.R.; et al. Transcriptional dynamics of two seed compartments with opposing roles in Arabidopsis seed germination. Plant Physiol. 2013, 163, 205–215. [Google Scholar] [CrossRef]
- Law, S.R.; Narsai, R.; Taylor, N.L.; Delannoy, E.; Carrie, C.; Giraud, E.; Millar, A.H.; Small, I.; Whelan, J. Nucleotide and RNA metabolism prime translational initiation in the earliest events of mitochondrial biogenesis during Arabidopsis germination. Plant Physiol. 2012, 158, 1610–1627. [Google Scholar] [CrossRef]
- Law, S.R.; Narsai, R.; Whelan, J. Mitochondrial biogenesis in plants during seed germination. Mitochondrion 2014, 19, 214–221. [Google Scholar] [CrossRef]
- Salinas, T.; Duchêne, A.M.; Maréchal-Drouard, L. Recent advances in tRNA mitochondrial import. Trends Biochem. Sci. 2008, 33, 320–329. [Google Scholar] [CrossRef] [PubMed]
- Val, R.; Wyszko, E.; Valentin, C.; Szymanski, M.; Cosset, A.; Alioua, M.; Dreher, T.W.; Barciszewski, J.; Dietrich, A. Organelle trafficking of chimeric ribozymes and genetic manipulation of mitochondria. Nucleic Acids Res. 2011, 39, 9262–9274. [Google Scholar] [CrossRef] [PubMed]
- Sultan, L.D.; Mileshina, D.; Grewe, F.; Rolle, K.; Abudraham, S.; Glodowicz, P.; Niazi, A.K.; Keren, I.; Shevtsov, S.; Klipcan, L.; et al. The reverse transcriptase/RNA maturase protein MatR is required for the splicing of various group II introns in Brassicaceae mitochondria. Plant Cell 2016, 28, 2805–2829. [Google Scholar] [CrossRef] [PubMed]
- Perrotta, A.T.; Been, M.D. A pseudoknot-like structure required for efficient self-cleavage of hepatitis delta virus RNA. Nature 1991, 350, 434–436. [Google Scholar] [CrossRef] [PubMed]
- Zuo, J.; Niu, Q.W.; Chua, N.H. Technical advance: An estrogen receptor-based transactivator XVE mediates highly inducible gene expression in transgenic plants. Plant J. 2000, 24, 265–273. [Google Scholar] [CrossRef] [PubMed]
- Mathews, D.H.; Sabina, J.; Zuker, M.; Turner, D.H. Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J. Mol. Biol. 1999, 288, 911–940. [Google Scholar] [CrossRef] [PubMed]
- van Engelen, F.A.; Molthoff, J.W.; Conner, A.J.; Nap, J.P.; Pereira, A.; Stiekema, W.J. pBINPLUS: An improved plant transformation vector based on pBIN19. Transgenic Res. 1995, 4, 288–290. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3–new capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef]
- Sweetlove, L.J.; Taylor, N.L.; Leaver, C.J. Isolation of intact, functional mitochondria from the model plant Arabidopsis thaliana. Methods Mol. Biol. 2007, 372, 125–136. [Google Scholar]
- Lurin, C.; Andrés, C.; Aubourg, S.; Bellaoui, M.; Bitton, F.; Bruyère, C.; Caboche, M.; Debast, C.; Gualberto, J.; Hoffmann, B.; et al. Genome-wide analysis of Arabidopsis pentatricopeptide repeat proteins reveals their essential role in organelle biogenesis. Plant Cell 2004, 16, 2089–2103. [Google Scholar] [CrossRef]
- Yang, Y.H.; Dudoit, S.; Luu, P.; Lin, D.M.; Peng, V.; Ngai, J.; Speed, T.P. Normalization for cDNA microarray data: A robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 2002, 30, e15. [Google Scholar] [CrossRef]
- Matthew, E.R.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar]
- Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 2004, 3, 3. [Google Scholar] [CrossRef] [PubMed]
- Guedj, M.; Robin, S.; Celisse, A.; Nuel, G. Kerfdr: A semi-parametric kernel-based approach to local false discovery rate estimation. BMC Bioinform. 2009, 10, 84. [Google Scholar] [CrossRef] [PubMed]
- Storey, J.D.; Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 2003, 100, 9440–9445. [Google Scholar] [CrossRef] [PubMed]
- Chun, H.; Keleş, S. Sparse partial least squares regression for simultaneous dimension reduction and variable selection. J. R. Stat. Soc. Series B Stat. Methodol. 2010, 72, 3–25. [Google Scholar] [CrossRef] [PubMed]
- Gagnot, S.; Tamby, J.P.; Martin-Magniette, M.L.; Bitton, F.; Taconnat, L.; Balzergue, S.; Aubourg, S.; Renou, J.P.; Lecharny, A.; Brunaud, V. CATdb: A public access to Arabidopsis transcriptome data from the URGV-CATMA platform. Nucleic Acids Res. 2008, 36, D986–D990. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zaag, R.; Tamby, J.P.; Guichard, C.; Tariq, Z.; Rigaill, G.; Delannoy, E.; Renou, J.P.; Balzergue, S.; Mary-Huard, T.; Aubourg, S.; et al. GEM2Net: From gene expression modeling to–Omics networks, a new CATdb module to investigate Arabidopsis thaliana genes involved in stress response. Nucleic Acids Res. 2015, 43, D1010–D1017. [Google Scholar] [CrossRef][Green Version]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets--update. Nucleic Acids Res. 2013, 41, D991–D995. [Google Scholar] [CrossRef]
- Brazma, A.; Hingamp, P.; Quackenbush, J.; Sherlock, G.; Spellman, P.; Stoeckert, C.; Aach, J.; Ansorge, W.; Ball, C.A.; Causton, H.C.; et al. Minimum information about a microarray experiment (MIAME)–toward standards for microarray data. Nat. Genet. 2001, 29, 365–371. [Google Scholar] [CrossRef]
- Matsuda, D.; Dreher, T.W. The tRNA-like structure of Turnip yellow mosaic virus RNA is a 3’-translational enhancer. Virology 2004, 321, 36–46. [Google Scholar] [CrossRef]
- Delage, L.; Duchêne, A.M.; Zaepfel, M.; Maréchal-Drouard, L. The anticodon and the D-domain sequences are essential determinants for plant cytosolic tRNA(Val) import into mitochondria. Plant J. 2003, 34, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Dietrich, A.; Maréchal-Drouard, L.; Carneiro, V.; Cosset, A.; Small, I. A single base change prevents import of cytosolic tRNA(Ala) into mitochondria in transgenic plants. Plant J. 1996, 10, 913–918. [Google Scholar] [CrossRef]
- Matera, J.T.; Monroe, J.; Smelser, W.; Gabay-Laughnan, S.; Newton, K.J. Unique changes in mitochondrial genomes associated with reversions of S-type cytoplasmic male sterility in maizemar. PLoS ONE 2011, 6, e23405. [Google Scholar] [CrossRef] [PubMed]
- Heazlewood, J.L.; Verboom, R.E.; Tonti-Filippini, J.; Small, I.; Millar, A.H. SUBA: The Arabidopsis subcellular database. Nucleic Acids Res. 2007, 35, D213–D218. [Google Scholar] [CrossRef]
- Millar, A.H.; Whelan, J.; Soole, K.L.; Day, D.A. Organization and regulation of mitochondrial respiration in plants. Annu. Rev. Plant Biol. 2011, 62, 79–104. [Google Scholar] [CrossRef]
- Topping, J.F.; Leaver, C.J. Mitochondrial gene expression during wheat leaf development. Planta 1990, 182, 399–407. [Google Scholar] [CrossRef]
- Howell, K.A.; Millar, A.H.; Whelan, J. Ordered assembly of mitochondria during rice germination begins with pro-mitochondrial structures rich in components of the protein import apparatus. Plant Mol. Biol. 2006, 60, 201–223. [Google Scholar] [CrossRef] [PubMed]
- Howell, K.A.; Cheng, K.; Murcha, M.W.; Jenkin, L.E.; Millar, A.H.; Whelan, J. Oxygen initiation of respiration and mitochondrial biogenesis in rice. J. Biol. Chem. 2007, 282, 15619–15631. [Google Scholar] [CrossRef] [PubMed]
- Ishizaki, K.; Larson, T.R.; Schauer, N.; Fernie, A.R.; Graham, I.A.; Leaver, C.J. The critical role of Arabidopsis electron-transfer flavoprotein:ubiquinone oxidoreductase during dark-induced starvation. Plant Cell 2005, 17, 2587–2600. [Google Scholar] [CrossRef]
- Sung, T.Y.; Tseng, C.C.; Hsieh, M.H. The SLO1 PPR protein is required for RNA editing at multiple sites with similar upstream sequences in Arabidopsis mitochondria. Plant J. 2010, 63, 499–511. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, W.Y.; Liao, J.C.; Chang, C.Y.; Harrison, T.; Boucher, C.; Hsieh, M.H. The SLOW GROWTH3 pentatricopeptide repeat protein is required for the splicing of mitochondrial NADH dehydrogenase subunit 7 intron 2 in Arabidopsis. Plant Physiol. 2015, 168, 490–501. [Google Scholar] [CrossRef] [PubMed]
- Koprivova, A.; Des Francs-Small, C.C.; Calder, G.; Mugford, S.T.; Tanz, S.; Lee, B.R.; Zechmann, B.; Small, I.; Kopriva, S. Identification of a pentatricopeptide repeat protein implicated in splicing of intron 1 of mitochondrial nad7 transcripts. J. Biol. Chem. 2010, 285, 32192–32199. [Google Scholar] [CrossRef] [PubMed]
- Nakagawa, N.; Sakurai, N. A mutation in At-nMat1a, which encodes a nuclear gene having high similarity to group II intron maturase, causes impaired splicing of mitochondrial NAD4 transcript and altered carbon metabolism in Arabidopsis thaliana. Plant Cell Physiol. 2006, 47, 772–783. [Google Scholar] [CrossRef] [PubMed]
- Woo, D.K.; Phang, T.L.; Trawick, J.D.; Poyton, R.O. Multiple pathways of mitochondrial-nuclear communication in yeast: Intergenomic signaling involves ABF1 and affects a different set of genes than retrograde regulation. Biochim. Biophys. Acta 2009, 1789, 135–145. [Google Scholar] [CrossRef] [PubMed]
- Börner, T. The discovery of plastid-to-nucleus retrograde signaling–A personal perspective. Protoplasma 2017, 254, 1845–1855. [Google Scholar] [CrossRef] [PubMed]
- Woodson, J.D.; Perez-Ruiz, J.M.; Schmit, R.J.; Ecker, J.R.; Chory, J. Sigma factor-mediated plastid retrograde signals control nuclear gene expression. Plant J. 2013, 73, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Holec, S.; Lange, H.; Canaday, J.; Gagliardi, D. Coping with cryptic and defective transcripts in plant mitochondria. Biochim. Biophys. Acta 2008, 1779, 566–573. [Google Scholar] [CrossRef]
- Nam, M.; Akie, T.E.; Sanosaka, M.; Craige, S.M.; Kant, S.; Keaney, J.F., Jr.; Cooper, M.P. Mitochondrial retrograde signaling connects respiratory capacity to thermogenic gene expression. Sci. Rep. 2017, 7, 2013. [Google Scholar] [CrossRef]
- da Cunha, F.M.; Torelli, N.Q.; Kowaltowski, A.J. Mitochondrial retrograde signaling: Triggers, pathways, and outcomes. Oxid. Med. Cell. Longev. 2015, 2015, 482582. [Google Scholar] [CrossRef]
- Jazwinski, S.M. The retrograde response: A conserved compensatory reaction to damage from within and from without. Prog. Mol. Biol. Transl. Sci. 2014, 127, 133–154. [Google Scholar]
- Berkowitz, O.; De Clercq, I.; Van Breusegem, F.; Whelan, J. Interaction between hormonal and mitochondrial signalling during growth, development and in plant defence responses. Plant Cell Environ. 2016, 39, 1127–1139. [Google Scholar] [CrossRef] [PubMed]
- Kühn, K.; Yin, G.; Duncan, O.; Law, S.R.; Kubiszewski-Jakubiak, S.; Kaur, P.; Meyer, E.; Wang, Y.; Colas des Francs-Small, C.; Giraud, E.; et al. Decreasing electron flux through the cytochrome and/or alternative respiratory pathways triggers common and distinct cellular responses dependent on growth conditions. Plant Physiol. 2015, 167, 228–250. [Google Scholar] [CrossRef]
- Zhang, F.; Zhao, J.; Xu, S.; Fang, W.; Chen, F.; Teng, N. MicroRNA and putative target discoveries in Chrysanthemum polyploidy breeding. Int. J. Genomics 2017, 2017, 6790478. [Google Scholar] [CrossRef] [PubMed]
- Ma, C.; Yang, J.; Cheng, Q.; Mao, A.; Zhang, J.; Wang, S.; Weng, Y.; Wen, C. Comparative analysis of miRNA and mRNA abundance in determinate cucumber by high-throughput sequencing. PLoS ONE 2018, 13, e0190691. [Google Scholar]
- Scott, M.S.; Ono, M. From snoRNA to miRNA: Dual function regulatory non-coding RNAs. Biochimie 2011, 93, 1987–1992. [Google Scholar] [CrossRef] [PubMed]
- Stepanov, G.A.; Filippova, J.A.; Komissarov, A.B.; Kuligina, E.V.; Richter, V.A.; Semenov, D.V. Regulatory role of small nucleolar RNAs in human diseases. Biomed. Res. Int. 2015, 2015, 206849. [Google Scholar] [CrossRef] [PubMed]
- Stortenbeker, N.; Bemer, M. The SAUR gene family: The plant’s toolbox for adaptation of growth and development. J. Exp. Bot. 2019, 70, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Li, Y.X.; Li, C.; Shi, Y.; Song, Y.; Zhang, D.; Li, Y.; Wang, T. Genome-wide analysis of the pentatricopeptide repeat gene family in different maize genomes and its important role in kernel development. BMC Plant Biol. 2018, 18, 366. [Google Scholar] [CrossRef] [PubMed]
- Gorchs Rovira, A.; Smith, A.G. PPR proteins–Orchestrators of organelle RNA metabolism. Physiol. Plant 2019. [Google Scholar] [CrossRef]
- Xing, H.; Fu, X.; Yang, C.; Tang, X.; Guo, L.; Li, C.; Xu, C.; Luo, K. Genome-wide investigation of pentatricopeptide repeat gene family in poplar and their expression analysis in response to biotic and abiotic stresses. Sci. Rep. 2018, 8, 2817. [Google Scholar] [CrossRef]
| Conditions and Developmental Stage | Day 1 | Day 2 | Day 3 | Day 4 |
|---|---|---|---|---|
| Light Early stage | ↓nad5a* | |||
| ↓atp4** | ||||
| ↑rps7* | ||||
| ↑AOX1d* | ↓AOX1d*** | ↓AOX1d*** | ||
| Light Intermediate stage | ↑cob** | ↑rps7* | ||
| ↑nad5a** | ↓ccmFN2*** | ↑rpl2** | ||
| ↑nad5b** | ↑atp4* | ↑rpl5* | ||
| ↓ccmFN2** | ↑rpl2* | |||
| ↑rpl5* | ||||
| ↑AOX1d*** | ↑AOX1d*** | |||
| Light Bolting stage | ↓nad2b** | ↓atp4* | ↑nad1b* | |
| ↑nad5a** | ↓atp9**** | ↓nad2b* | ||
| ↓nad5b** | ↓rps4** | ↓nad4** | ||
| ↓nad9** | ↓rps7**** | ↑nad5a*** | ||
| ↓cox1** ↓ccmB** | ↓rpl2**** | ↓nad5b* | ||
| ↓mttB* | ↓rpl5** | ↓nad6* | ||
| ↓rpl5* | ↓rrn18** | ↓nad9** | ||
| ↓cob*** | ||||
| ↓cox1* | ↓cox1**** | |||
| ↓cox2** | ↓atp9* | |||
| ↓cox3**** | ↓ccmFC*** | |||
| ↑atp9* | ↑mttB* | |||
| ↓ccmB*** | ↑rpl5** | |||
| ↓ccmFC* | ↓rrn26* | |||
| ↓ccmFN1*** | ||||
| ↓ccmFN2** | ||||
| ↓mttB** | ||||
| ↓rrn18*** | ||||
| ↓rrn26** | ||||
| ↓AOX1a* | ||||
| ↓AOX1d** | ↓AOX1d* | ↓AOX1d* | ||
| Dark | ↑nad2b** | ↑nad4* | ↓nad1b*** | ↓ccmFC*** |
| ↓nad5a** | ↑nad5b* | ↓nad2b** | ↓matR* | |
| ↑nad5b**** | ↑cox1* | ↑nad5b** | ↓mttB* | |
| ↑nad9* | ↓ccmFN2** | ↓ccmFN2** | ↓rrn18**** | |
| ↓cox2* | ↓mttB** | ↑rrn26** | ||
| ↓matR* | ↓rrn26** | |||
| ↑rps7** | ||||
| ↓rpl5* | ||||
| ↓rrn26* | ||||
| ↑AOX1d** |
| Modulation | Annotation / Function / Organellar Localization | Gene |
|---|---|---|
| Upregulated (opposite to nad9) | Basic helix-loop-helix (bHLH) DNA-binding superfamily protein; transcription factor; response to ethylene | AT1G05710 |
| ABI3-interacting protein 3, AIP3, PFD4, PREFOLDIN 4; protein chaperone; ABI3 is an auxin-inducible transcription factor | AT1G08780 | |
| ATMYB60, Myb domain protein 60, MYB60; transcription factor; response to abscisic acid, jasmonic acid, salicylic acid | AT1G08810 | |
| Agamous-like 87, AGL87, MADS-box family protein; transcription factor | AT1G22590 | |
| GL2, GLABRA 2, HD-ZIP IV family of homeobox-leucine zipper protein with lipid-binding START domain; transcription factor | AT1G79840 | |
| DREB subfamily A-6 of ERF/AP2 transcription factor family; one AP2 domain; ethylene-activated | AT2G22200 | |
| AP2/B3-like transcriptional factor family protein; transcription factor | AT2G33720 | |
| FMA (FAMA), basic helix-loop-helix (bHLH) DNA-binding superfamily protein; transcription factor/ transcriptional activator | AT3G24140 | |
| EDF3, ethylene response DNA-binding factor 3; transcription factor; AP2 domain; ethylene responding | AT3G25730 | |
| IAA30, indole-3-acetic acid inducible protein 30; transcription factor; response to auxin | AT3G62100 | |
| B-BOX domain protein 23, BBX23; transcription factor | AT4G10240 | |
| PUCHI, ethylene response factor (ERF) subfamily B-1 of ERF/AP2 transcription factor family; one AP2 domain; ethylene response | AT5G18560 | |
| ARGOS, Auxin-regulated gene involved in organ size; response to ethylene, auxin; membrane, cytoplasm, mitochondrion | AT3G59900 | |
| EMB3103, Embryo-defective 3103, PDM2, Pigment-Defective Mutant2; pentatricopeptide repeat (PPR) superfamily protein; endonuclease; chloroplast/mitochondrion | AT1G10910 | |
| RNH1C, RNase H family protein, RNase H domain-containing protein; chloroplast | AT1G24090 | |
| RING-finger, DEAD-like helicase, PHD and SNF2 domain-containing protein | AT2G40770 | |
| AT-SR34B, Serine/arginine-rich protein splicing factor 34B, SR34B | AT4G02430 | |
| Mitochondrial nuclease 1, MNU1; putative endonuclease or glycosyl hydrolase; mitochondrial RNA 5'-end processing; chloroplast/mitochondrion | AT5G64710 | |
| EMB2394, Embryo-defective 2394; structural constituent of chloroplast ribosome; response to cytokinin | AT1G05190 | |
| rRNA, cytosolic small ribosomal subunit | AT2G01010 | |
| 5.8S rRNA, cytosolic large ribosomal subunit | AT2G01020 | |
| 40S ribosomal protein S23 (RPS23A), ribosomal protein S12/S23 family protein; cytosolic small ribosomal subunit | AT3G09680 | |
| Ribosomal protein L15, RPL15; large subunit of the chloroplast ribosome; response to cytokinin | AT3G25920 | |
| EMB3126, Embryo-defective 3126, plastid ribosomal protein L1, PRPL1; ribosomal protein L1p/L10e family; chloroplast large ribosomal subunit | AT3G63490 | |
| 60S ribosomal protein L31, RPL31B; cytosolic large ribosomal subunit | AT4G26230 | |
| U3 ribonucleoprotein, Utp family protein; rRNA processing | AT5G08600 | |
| MA3 domain-containing translation regulatory factor 1, MRF1; colocalizes with cytosolic large ribosomal subunit; isomerase activity | AT5G63190 | |
| snoRNA | AT1G03743; AT1G19373; AT1G19376; AT1G75166; AT2G35387; AT3G27865; AT3G47342; AT3G47347; AT3G58193; AT3G58196; AT4G39366; AT5G44286 |
| Modulation | Annotation/Function/Organellar Localization | Gene |
|---|---|---|
| Downregulated (as nad9) | MicroRNA414, mir414, short open reading frame 16, SORF16; miRNA; identified as a translated small open reading frame by ribosome profiling | AT1G67195 |
| B-BOX domain protein 25, BBX25, Salt tolerance homologue, STH; transcription factor; zinc ion binding | AT2G31380 | |
| ATIBH1, IBH1, ILI1 binding BHLH 1 ILI1 binding bHLH 1; transcription factor; brassinosteroid signaling; gibberellic acid signaling | AT2G43060 | |
| Arabidopsis thaliana response regulator 2, ARR5, ATRR2, IBC6, induced by cytokinin 6, response regulator 5, RR5; transcription repressor; cytokinin signaling | AT3G48100 | |
| Basic helix-loop-helix (bHLH) DNA-binding superfamily protein; transcription factor | AT4G01460 | |
| KELP; homodimers or heterodimers with the kiwi protein; transcriptional co-activator | AT4G10920 | |
| WRKY DNA-binding protein 24, ATWRKY24, WRKY24; WRKY transcription factor group II-c | AT5G41570 | |
| BOA, Brother of lux ARRHYTHMO; transcription factor; circadian clock; mRNA cell-to-cell mobile | AT5G59570 | |
| SAUR53, small auxin-upregulated RNA 53, SAUR-like auxin-responsive protein family; mitochondrion | AT1G19840 | |
| SAUR65, small auxin-upregulated RNA 65, SAUR-like auxin-responsive protein family; membrane; mitochondrion | AT1G29460 | |
| ATCLE19, CLAVATA3/ESR-related 19, CLE19, embryo surrounding region 19, ESR19; receptor binding; signal transduction; mitochondrion | AT3G24225 | |
| VQ motif-containing protein 29, VQ29; response to hypoxia | AT4G37710 | |
| RNA-binding (RRM/RBD/RNP motifs) family protein | AT1G33470 | |
| ATRNS1, Ribonuclease 1, RNS1; endoribonuclease | AT2G02990 | |
| Eukaryotic translation initiation factor 2 (eIF-2) family protein; cytosol | AT1G76820 | |
| EMB3113, Embryo-defective 3113, ribosomal protein S5, RPS5, SCA1, SCABRA 1; structural component of the 70S chloroplast ribosome; mitochondrial small ribosomal subunit | AT2G33800 | |
| U3 containing 90S pre-ribosomal complex subunit | AT2G43110 | |
| ATRAB8D, ATRABE1B, RAB GTPase homolog E1B, RABE1B; translation elongation factor; membrane; chloroplast | AT4G20360 | |
| snoRNA | AT2G35382; AT2G43137; AT2G43138; AT2G43139; AT2G43141; AT4G02550; AT4G02555; AT4G13245 |
| Modulation | Annotation/Function/Organellar Localization | Gene |
|---|---|---|
| Upregulated (opposite to nad9) | Small nuclear RNA U6acat, mRNA splicing | AT5G40395 |
| Downregulated (as nad9) | ANAC028, NAC domain-containing protein 28, NAC028; transcription factor | AT1G65910 |
| Transcription elongation factor Spt5; KOW domain | AT2G34210 | |
| MYR2, homeodomain-like superfamily protein; transcription factor | AT3G04030 | |
| HVA22-like protein F, HVA22F; membrane protein; response to abscisic acid | AT2G42820 | |
| ALY2 RNA-binding (RRM/RBD/RNP motifs) family protein; mRNA transport | AT5G02530 |
| Modulation | Annotation/Function/Organellar Localization | Gene |
|---|---|---|
| Upregulated (opposite to nad9) | auxin-responsive family protein (SAUR72) | AT3G12830 |
| auxin-responsive protein, putative (SAUR9) | AT4G36110 | |
| auxin-responsive family protein (SAUR41) | AT1G16510 | |
| auxin-responsive protein-related (SAUR77) | AT1G17345 | |
| auxin-responsive family protein (SAUR53); mitochondrion | AT1G19840 | |
| auxin-responsive family protein (SAUR71) | AT1G56150 | |
| auxin-responsive family protein (SAUR52); mitochondrion | AT1G75590 | |
| auxin-responsive protein-related (SAUR36); mitochondrion/nucleus (nucleus confirmed by GFP targeting experiments) | AT2G45210 | |
| auxin-responsive family protein (SAUR59); mitochondrion | AT3G60690 | |
| auxin-responsive protein, putative (SAUR25); mitochondrion | AT4G13790 | |
| auxin-responsive family protein (SAUR1); chloroplast | AT4G34770 | |
| auxin-responsive protein, putative (SAUR23) | AT5G18060 | |
| MIF1 (MINI ZINC FINGER 1); transcription factor; response to abscisic acid, auxin, brassinosteroid, cytokinin, gibberellin; nucleus (cytosol reported from GFP data) | AT1G74660 | |
| PLS (POLARIS), cytokinin and auxin responses; mitochondrion | AT4G39403 | |
| Tetratricopeptide repeat (TPR)-like superfamily protein; mitochondrion | AT1G28690 | |
| S-RBP11, SMALL RNA-BINDING PROTEIN 11; salt stress response; chloroplast | AT5G06210 | |
| Downregulated (as nad9) | ARGAH2, Arginine amidohydrolase 2, response to jasmonate; chloroplast/mitochondrion (confirmed by MS data) | AT4G08870 |
| Tetratricopeptide repeat (TPR)-like superfamily protein; chloroplast/mitochondrion (chloroplast confirmed by MS data) | AT2G37230 | |
| pentatricopeptide (PPR) repeat-containing protein | AT3G62470 | |
| pentatricopeptide (PPR) repeat-containing protein | AT4G01030 | |
| EMB1417 (embryo-defective 1417), PPR protein; RNA binding; endonuclease activity | AT4G21190 | |
| pentatricopeptide (PPR) repeat-containing protein | AT4G21880 | |
| pentatricopeptide (PPR) repeat-containing protein | AT5G65560 | |
| EMB1586, Embryo-defective 1586, increased size exclusion limit 1, ISE1; DEAD-box RNA helicase; chloroplast/mitochondrion (mitochondrion confirmed by GFP data) | AT1G12770 | |
| AGS1, AHG2-1 suppressor 1, bacterial-type poly(A) polymerase; mRNA polyadenylation; chloroplast/mitochondrion | AT2G17580 | |
| ATTRM2A, TRM2A, tRNA methyltransferase 2A, RNA methyltransferase family protein; cytoplasm | AT3G21300 | |
| ATP-dependent RNA helicase | AT5G39840 | |
| ARFB1A, ATARFB1A (ADP-ribosylation factor B1A); GTP binding; protein transport; Golgi apparatus | AT2G15310 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niazi, A.K.; Delannoy, E.; Iqbal, R.K.; Mileshina, D.; Val, R.; Gabryelska, M.; Wyszko, E.; Soubigou-Taconnat, L.; Szymanski, M.; Barciszewski, J.; et al. Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development. Cells 2019, 8, 583. https://doi.org/10.3390/cells8060583
Niazi AK, Delannoy E, Iqbal RK, Mileshina D, Val R, Gabryelska M, Wyszko E, Soubigou-Taconnat L, Szymanski M, Barciszewski J, et al. Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development. Cells. 2019; 8(6):583. https://doi.org/10.3390/cells8060583
Chicago/Turabian StyleNiazi, Adnan Khan, Etienne Delannoy, Rana Khalid Iqbal, Daria Mileshina, Romain Val, Marta Gabryelska, Eliza Wyszko, Ludivine Soubigou-Taconnat, Maciej Szymanski, Jan Barciszewski, and et al. 2019. "Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development" Cells 8, no. 6: 583. https://doi.org/10.3390/cells8060583
APA StyleNiazi, A. K., Delannoy, E., Iqbal, R. K., Mileshina, D., Val, R., Gabryelska, M., Wyszko, E., Soubigou-Taconnat, L., Szymanski, M., Barciszewski, J., Weber-Lotfi, F., Gualberto, J. M., & Dietrich, A. (2019). Mitochondrial Transcriptome Control and Intercompartment Cross-Talk During Plant Development. Cells, 8(6), 583. https://doi.org/10.3390/cells8060583





