A Hybrid Prediction Method for Plant lncRNA-Protein Interaction
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset
2.2. Interaction between LncRNA and Protein
2.3. Sequence Feature Encoding
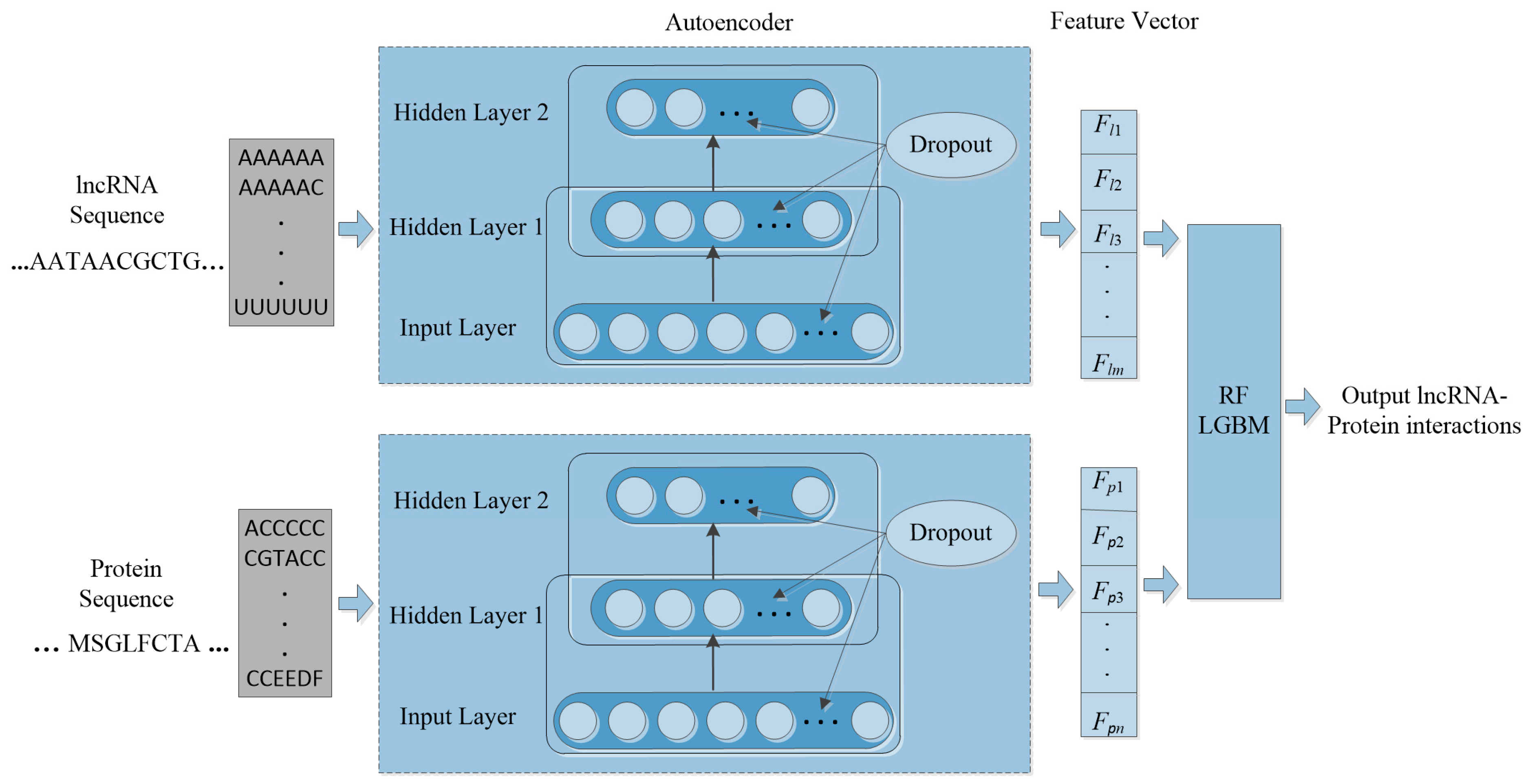
2.4. The PLRPIM Pipeline
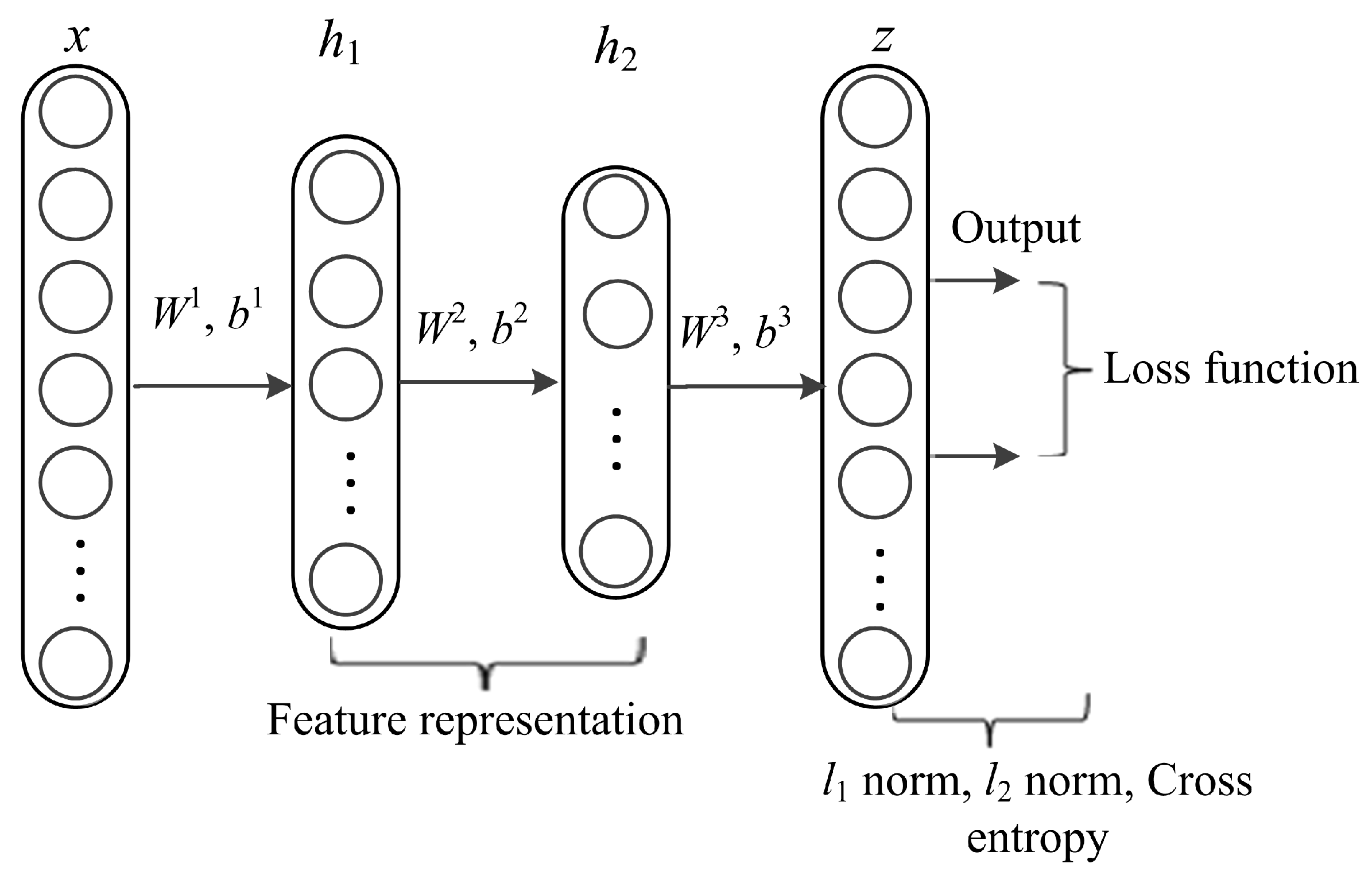
2.4.1. Feature Extraction
2.4.2. Hybrid Learning Method
2.4.3. Training of PLRPIM Model with Mixed Norm Constraint
2.4.4. Shallow Classification Models
2.4.5. Experimental Setup
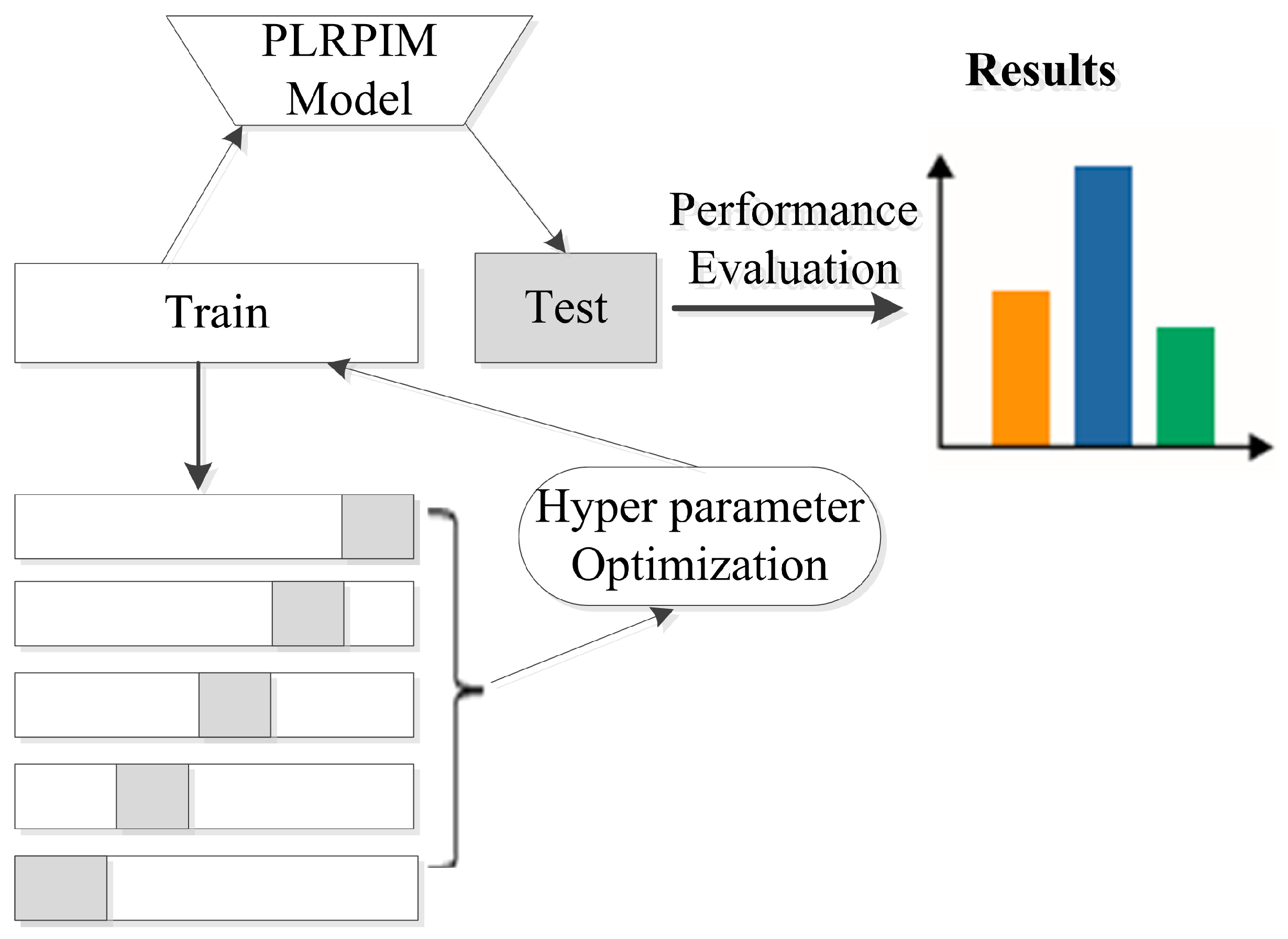
2.4.6. PLRPIM Method
| Algorithm 1: Pseudo code of PLRPIM | |
| Input: Y—adjacency matrix of interacting lncRNA i and protein j; (Y(li, pj)), SlncRNA—set of lncRNA sequence of interest, Sprotein—set of protein sequence of interest, number of stacked AEs = T, number of iterations (epoch) = R | |
| Output: lncRNA-protein interaction matrix Mij | |
| 1. | B = {1, 0}—Binary domain; |
| 2. | Interact: SlncRNA × Sprotein B; |
| 3. | Interact: (S(li), S(pi))- check whether lncRNA li and protein pi interact; if true, return 1; otherwise, return 0; |
| 4. | Concatenate: SlncRNA × SproteinY; |
| 5. | Initialize training examples labels (Y(li, pj)) = 0; |
| 6. | Fort = 1 to T do; |
| 7. | Forr = 1 to R do; |
| 8. | Minimize objective function using formula (10); |
| 9. | Compute number of features from similarity matrices and ; |
| 10. | Compute hidden vector with learned AE; |
| 11. | Update ; |
| 12. | End for; |
| 13. | Compute feature vectors {Fl1, Fl2,…, Flm} and {Fp1, Fp2,…, Fpn}; |
| 14. | Predict class labels of the test dataset based on ensemble voting process using formula (11); |
| 15. | Update training examples labels (Y(li, pj)) = 1; |
| 16. | End for; |
| 17. | Return:Mij. |
2.5. Evaluation of PLRPIM
3. Results
3.1. Comparison between Classification Models
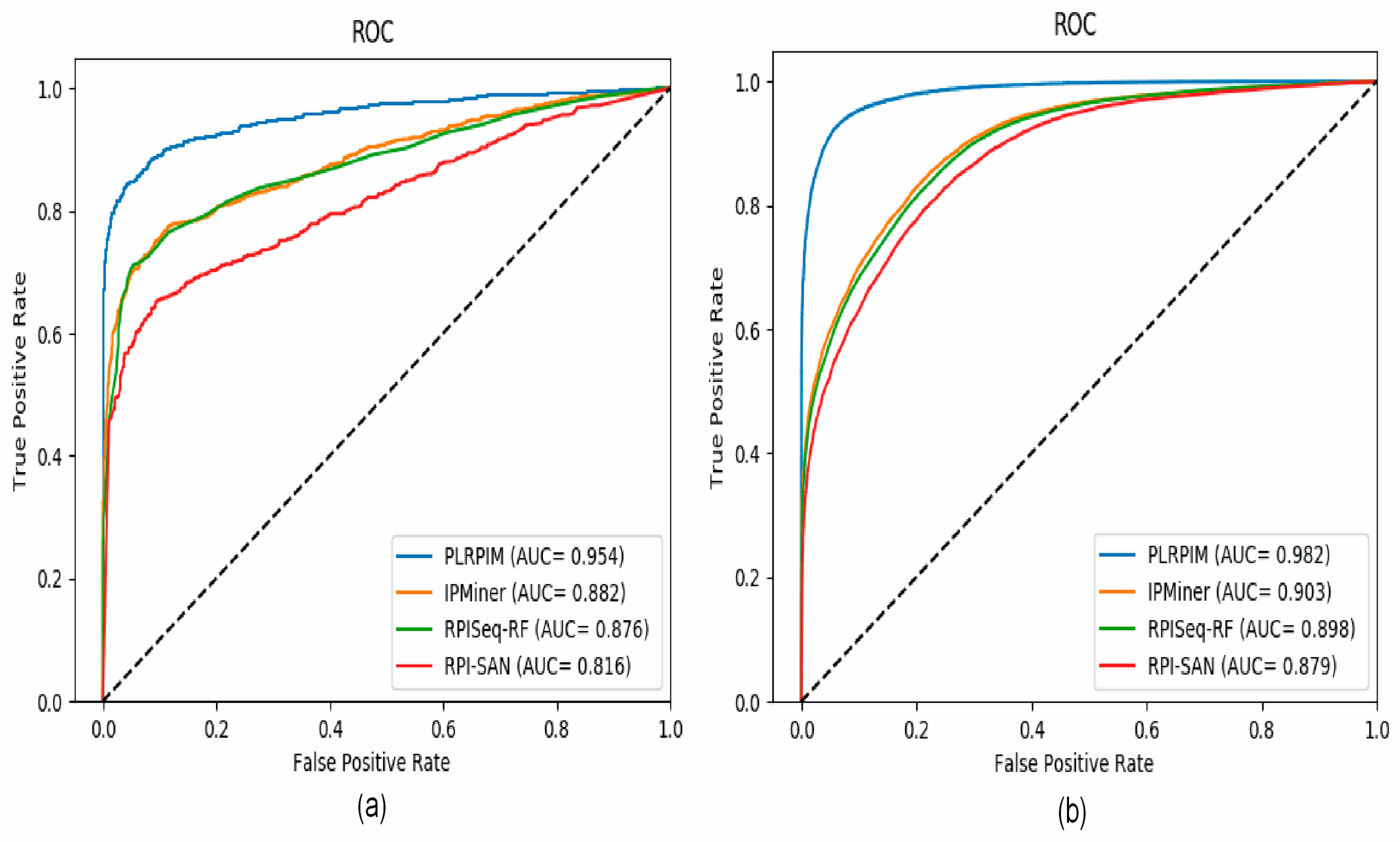
3.2. Comparison of PLRPIM with Other Existing Tools
3.3. Functional Analysis of lncRNAs
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- D’Aniello, S.; Spagnuolo, A.; Ceccarelli, M.; Cerulo, L.; Ventola, G.M.M.; Noviello, T.M.R.; D’Aniello, S. Identification of long non-coding transcripts with feature selection: a comparative study. BMC Bioinform. 2017, 18, 187. [Google Scholar]
- Lu, X.; Chen, X.; Mu, M.; Wang, J.; Wang, X.; Wang, D.; Yin, Z.; Fan, W.; Wang, S.; Guo, L.; Ye, W. Genome-Wide Analysis of Long Noncoding RNAs and Their Responses to Drought Stress in Cotton (Gossypium hirsutum L.). PLoS ONE 2016, 11, e0156723. [Google Scholar] [CrossRef]
- Nejat, N.; Mantri, N. Emerging roles of long non-coding RNAs in plant response to biotic and abiotic stresses. Crit. Rev. Biotechnol. 2018, 38, 93–105. [Google Scholar] [CrossRef]
- Kim, D.H.; Xi, Y.; Sung, S. Modular function of long noncoding RNA, COLDAIR, in the vernalization response. PLOS Genet. 2017, 13, e1006939. [Google Scholar] [CrossRef]
- Bhatia, G.; Goyal, N.; Sharma, S.; Upadhyay, S.K.; Singh, K. Present Scenario of Long Non-Coding RNAs in Plants. Non-Coding RNA 2017, 3, 16. [Google Scholar] [CrossRef] [PubMed]
- Kashi, K.; Henderson, L.; Bonetti, A.; Carninci, P. Discovery and functional analysis of lncRNAs: Methodologies to investigate an uncharacterized transcriptome. Biochim. et Biophys. Acta (BBA)-Gene Regul. Mech. 2016, 1859, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Wu, W.; Han, Q.; Wang, Y.; Li, C.; Zhang, P.; Xu, H. New Insights into the Interplay between Non-Coding RNAs and RNA-Binding Protein HnRNPK in Regulating Cellular Functions. Cells 2019, 8, 62. [Google Scholar] [CrossRef]
- Camborde, L.; Jauneau, A.; Brière, C.; Deslandes, L.; Dumas, B.; Gaulin, E. Detection of nucleic acid-protein interactions in plant leaves using fluorescence lifetime imaging microscopy. Nat. Protoc. 2017, 12, 1933–1950. [Google Scholar] [CrossRef]
- Bierhoff, H. Analysis of lncRNA-Protein Interactions by RNA-Protein Pull-Down Assays and RNA Immunoprecipitation (RIP). Cell. Quiescence 2018, 1686, 241–250. [Google Scholar]
- Mermaz, B.; Liu, F.; Song, J. RNA Immunoprecipitation Protocol to Identify Protein-RNA Interactions in Arabidopsis thaliana. Plant Chromatin Dyn. 2018, 1675, 331–343. [Google Scholar]
- Liu, X.; Hao, L.; Li, D.; Zhu, L.; Hu, S. Long non-coding RNAs and their biological roles in plants. Genom. Proteom. Bioinform. 2015, 13, 137–147. [Google Scholar] [CrossRef]
- Han, S.; Liang, Y.; Ma, Q.; Xu, Y.; Zhang, Y.; Du, W.; Wang, C.; Li, Y. LncFinder: An integrated platform for long non-coding RNA identification utilizing sequence intrinsic composition, structural information and physicochemical property. Briefings Bioinform. 2018. [Google Scholar] [CrossRef]
- Singh, U.; Khemka, N.; Rajkumar, M.S.; Garg, R.; Jain, M. PLncPRO for prediction of long non-coding RNAs (lncRNAs) in plants and its application for discovery of abiotic stress-responsive lncRNAs in rice and chickpea. Nucleic Acids Res. 2017, 45, e183. [Google Scholar] [CrossRef]
- Vieira, L.M.; Grativol, C.; Thiebaut, F.; Carvalho, T.G.; Hardoim, P.R.; Hemerly, A.; Lifschitz, S.; Ferreira, P.C.G.; Walter, M.E.M.T. PlantRNA_Sniffer: A SVM-Based Workflow to Predict Long Intergenic Non-Coding RNAs in Plants. Non-Coding RNA 2017, 3, 11. [Google Scholar] [CrossRef]
- Yang, C.; Yang, L.; Zhou, M.; Xie, H.; Zhang, C.; Wang, M.D.; Zhu, H. LncADeep: an ab initio lncRNA identification and functional annotation tool based on deep learning. Bioinformatics 2018, 34, 3825–3834. [Google Scholar] [CrossRef]
- Zhang, W.; Yue, X.; Tang, G.; Wu, W.; Huang, F.; Zhang, X. SFPEL-LPI: Sequence-based feature projection ensemble learning for predicting LncRNA-protein interactions. PLOS Comput. Boil. 2018, 14, e1006616. [Google Scholar] [CrossRef]
- Bellucci, M.; Agostini, F.; Masin, M.; Tartaglia, G.G. Predicting protein associations with long noncoding RNAs. Nat. Methods 2011, 8, 444–445. [Google Scholar] [CrossRef]
- Lu, Q.; Ren, S.; Lu, M.; Zhang, Y.; Zhu, D.; Zhang, X.; Li, T. Computational prediction of associations between long non-coding RNAs and proteins. BMC Genom. 2013, 14, 651. [Google Scholar] [CrossRef]
- Suresh, V.; Liu, L.; Adjeroh, D.; Zhou, X. RPI-Pred: Predicting ncRNA-protein interaction using sequence and structural information. Nucleic Acids Res. 2015, 43, 1370–1379. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Zhu, C.; Ai, H.; Zhang, L.; Zhao, J.; Zhao, Q.; Liu, H. LPI-ETSLP: lncRNA–protein interaction prediction using eigenvalue transformation-based semi-supervised link prediction. Mol. BioSyst. 2017, 13, 1781–1787. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Zhang, Y.; Hu, H.; Ren, G.; Zhang, W.; Liu, H. IRWNRLPI: Integrating Random Walk and Neighborhood Regularized Logistic Matrix Factorization for lncRNA-Protein Interaction Prediction. Front. Genet. 2018, 9, 239. [Google Scholar] [CrossRef]
- Zhang, Y.; Wang, M.; Li, A.; Ge, M.; Peng, C. Predicting Long Noncoding RNA and Protein Interactions Using Heterogeneous Network Model. BioMed Res. Int. 2015, 2015. [Google Scholar] [CrossRef]
- Ge, M.; Li, A.; Wang, M. A Bipartite Network-based Method for Prediction of Long Non-coding RNA-protein Interactions. Genom. Proteom. Bioinform. 2016, 14, 62–71. [Google Scholar] [CrossRef]
- Liu, H.; Ren, G.; Hu, H.; Zhang, L.; Ai, H.; Zhang, W.; Zhao, Q. LPI-NRLMF: lncRNA-protein interaction prediction by neighborhood regularized logistic matrix factorization. Oncotarget 2017, 8, 103975–103984. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Li, R.; Zeng, T.; Sun, Q.; Kumar, S.; Ye, J.; Ji, S. Deep Model Based Transfer and Multi-Task Learning for Biological Image Analysis. IEEE Trans. Big Data 2016. [Google Scholar] [CrossRef]
- Zhang, L.; Lv, C.; Jin, Y.; Cheng, G.; Fu, Y.; Yuan, D.; Tao, Y.; Guo, Y.; Ni, X.; Shi, T. Deep Learning-Based Multi-Omics Data Integration Reveals Two Prognostic Subtypes in High-Risk Neuroblastoma. Front. Genet. 2018, 9, 477. [Google Scholar] [CrossRef]
- Fuentes, A.F.; Yoon, S.; Lee, J.; Park, D.S. High-Performance Deep Neural Network-Based Tomato Plant Diseases and Pests Diagnosis System With Refinement Filter Bank. Front. Plant Sci. 2018, 9, 1162. [Google Scholar] [CrossRef]
- Mohanty, S.P.; Hughes, D.P.; Salathé, M. Using Deep Learning for Image-Based Plant Disease Detection. Front. Plant Sci. 2016, 7, 346. [Google Scholar] [CrossRef]
- Liu, G.; Bao, H.; Han, B. A Stacked Autoencoder-Based Deep Neural Network for Achieving Gearbox Fault Diagnosis. Math. Probl. Eng. 2018, 2018, 1–10. [Google Scholar] [CrossRef]
- LeCun, Y.; Bengio, Y.; Hinton, G. Deep learning. Nature 2015, 521, 436–444. [Google Scholar] [CrossRef] [PubMed]
- Alipanahi, B.; Delong, A.; Weirauch, M.T.; Frey, B.J. Predicting the sequence specificities of DNA- and RNA-binding proteins by deep learning. Nat. Biotechnol. 2015, 33, 831–838. [Google Scholar] [CrossRef]
- Zhou, J.; Troyanskaya, O.G. Predicting effects of noncoding variants with deep learning-based sequence model. Nat. Methods 2015, 12, 931–934. [Google Scholar] [CrossRef]
- Wang, Y.; You, Z.-H.; Yang, S.; Li, X.; Jiang, T.-H.; Zhou, X. A High Efficient Biological Language Model for Predicting Protein-Protein Interactions. Cells 2019, 8, 122. [Google Scholar] [CrossRef]
- Pan, X.; Shen, H.-B. Predicting RNA-protein binding sites and motifs through combining local and global deep convolutional neural networks. Bioinformatics 2018, 34, 3427–3436. [Google Scholar] [CrossRef]
- Hosseini-Asl, E.; Zurada, J.M.; Nasraoui, O. Deep Learning of Part-Based Representation of Data Using Sparse Autoencoders With Nonnegativity Constraints. IEEE Trans. Neural Networks Learn. Syst. 2016, 27, 1–13. [Google Scholar] [CrossRef]
- Halkias, X.; Paris, S.; Glotin, H. Sparse Penalty in Deep Belief Networks: Using the Mixed Norm Constraint. Available online: https://arxiv.org/abs/1301.3533 (accessed on 27 December 2018).
- Vincent, P.; LaRochelle, H.; Bengio, Y.; Manzagol, P.-A. Extracting and composing robust features with denoising autoencoders. In Proceedings of the 25th international conference on Machine learning, Helsinki, Finland, 5–9 July 2008; pp. 1096–1103. [Google Scholar]
- Van Der Laan, M.J.; Polley, E.C.; Hubbard, A.E. Super learner in prediction. U.C. Berkeley Division of Biostatistics Working Paper Series. Available online: https://biostats.bepress.com/ucbbiostat/paper222/ (accessed on 1 November 2018).
- Muppirala, U.K.; Honavar, V.G.; Dobbs, D. Predicting RNA-Protein Interactions Using Only Sequence Information. BMC Bioinform. 2011, 12, 489. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, D.; Chen, R.; Chen, L.; Wang, Y.; Chen, X.; Liu, Z.-P.; Huang, Q.; Zhang, X.-S. De novo prediction of RNA–protein interactions from sequence information. Mol. BioSyst. 2013, 9, 133–142. [Google Scholar] [CrossRef]
- Pan, X.; Fan, Y.-X.; Yan, J.; Shen, H.-B. IPMiner: hidden ncRNA-protein interaction sequential pattern mining with stacked autoencoder for accurate computational prediction. BMC Genom. 2016, 17, 861. [Google Scholar] [CrossRef]
- Yi, H.-C.; You, Z.-H.; Huang, D.-S.; Li, X.; Jiang, T.-H.; Li, L.-P. A Deep Learning Framework for Robust and Accurate Prediction of ncRNA-Protein Interactions Using Evolutionary Information. Mol. Ther. Nucleic Acids 2018, 11, 337–344. [Google Scholar] [CrossRef]
- Cao, Z.; Pan, X.; Yang, Y.; Huang, Y.; Shen, H.-B.; Zhen, C. The lncLocator: a subcellular localization predictor for long non-coding RNAs based on a stacked ensemble classifier. Bioinformatics 2018, 34, 2185–2194. [Google Scholar] [CrossRef]
- Hu, H.; Zhang, L.; Ai, H.; Zhang, H.; Fan, Y.; Zhao, Q.; Liu, H. HLPI-Ensemble: Prediction of human lncRNA-protein interactions based on ensemble strategy. RNA Boil. 2018, 15, 1–10. [Google Scholar] [CrossRef]
- Xiao, Y.; Wu, J.; Lin, Z.; Zhao, X. A deep learning-based multi-model ensemble method for cancer prediction. Comput. Methods Programs Biomed. 2018, 153, 1–9. [Google Scholar] [CrossRef]
- Liu, Z.-P.; Miao, H. Prediction of protein-RNA interactions using sequence and structure descriptors. Neurocomputing 2016, 206, 28–34. [Google Scholar] [CrossRef]
- Liu, B. BioSeq-Analysis: A platform for DNA, RNA and protein sequence analysis based on machine learning approaches. Briefings Bioinform. 2017, 165. [Google Scholar] [CrossRef] [PubMed]
- Le, Q.V. Building high-level features using large scale unsupervised learning. In Proceedings of the 2013 IEEE International Conference on Acoustics, Speech and Signal Processing, Vancouver, BC, Canada, 26–31 May 2013; pp. 8595–8598. [Google Scholar]
- Sankaran, A.; Vatsa, M.; Singh, R.; Majumdar, A. Group sparse autoencoder. Image Vision Comput. 2017, 60, 64–74. [Google Scholar] [CrossRef]
- Zhang, X.; Liu, S. RBPPred: Predicting RNA-binding proteins from sequence using SVM. Bioinformatics 2017, 33, 854–862. [Google Scholar] [CrossRef] [PubMed]
- Ke, G.; Meng, Q.; Finley, T.; Wang, T.; Chen, W.; Ma, W.; Ye, Q.; Liu, T.-Y. Lightgbm: A highly efficient gradient boosting decision tree. In Proceedings of the Advances in Neural Information Processing Systems 30 (NIPS 2017), Long Beach, CA, USA, 4–9 December 2017; Curran Associates Inc: Red Hook, NY, USA, 2017; pp. 3146–3154. [Google Scholar]
- Zhan, Z.H.; You, Z.H.; Li, L.P.; Zhou, Y.; Yi, H.C. Accurate Prediction of ncRNA-Protein Interactions From the Integration of Sequence and Evolutionary Information. Front Genet. 2018, 9, 458. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Wang, H.; Liu, C.; Deng, L. Enhanced Prediction of Hot Spots at Protein-Protein Interfaces Using Extreme Gradient Boosting. Sci. Rep. 2018, 8, 14285. [Google Scholar] [CrossRef]
- Abadi, M.; Agarwal, A.; Barham, P.; Brevdo, E.; Chen, Z.; Citro, C.; Corrado, G.S.; Davis, A.; Dean, J.; Devin, M.; et al. TensorFlow: Large-Scale Machine Learning on Heterogeneous Distributed Systems. Available online: https://www.usenix.org/conference/osdi16/technical-sessions/presentation/abadi (accessed on 30 October 2018).
- Qian, N. On the momentum term in gradient descent learning algorithms. Neural Networks 1999, 12, 145–151. [Google Scholar] [CrossRef]
- Arendsee, Z.W.; Li, L.; Wurtele, E.S. Coming of age: orphan genes in plants. Trends Plant. Sci. 2014, 19, 698–708. [Google Scholar] [CrossRef]
Sample Availability: The code and datasets for implementing the proposed method are available at https://github.com/Mjwl/PLRPIM. |

| Species | Dataset | Positive Samples | Negative Samples | Total |
|---|---|---|---|---|
| Arabidopsis thaliana | Training set | 758 | 758 | 1516 |
| Test set | 190 | 190 | 380 | |
| Zea mays | Training set | 17,706 | 17,706 | 35,412 |
| Test set | 4427 | 4427 | 8854 |
| Name | Settings |
|---|---|
| Learning rate | 0.5, 1, 2 |
| Parameter optimization | SGD, momentum, Adam |
| Batch size | 256, 128, 64 |
| Activation | ReLU |
| Loss | MSE, Cross Entropy |
| Dropout rate | 0.6 |
| Dataset | Test Set | ACC | PRE | SEN | SPE | MCC | AUC |
|---|---|---|---|---|---|---|---|
| Arabidopsis thaliana | 1 | 0.9000 | 0.9250 | 0.8506 | 0.9417 | 0.7995 | 0.9527 |
| 2 | 0.8865 | 0.9368 | 0.8359 | 0.9402 | 0.7784 | 0.9488 | |
| 3 | 0.8971 | 0.9344 | 0.8636 | 0.9337 | 0.7970 | 0.9590 | |
| 4 | 0.9050 | 0.9353 | 0.8641 | 0.9436 | 0.8117 | 0.9490 | |
| 5 | 0.9103 | 0.9223 | 0.9036 | 0.9176 | 0.8206 | 0.9615 | |
| Average | 0.8998 ± 0.008 | 0.9308 ± 0.006 | 0.8636 ± 0.02 | 0.9354 ± 0.009 | 0.8015 ± 0.01 | 0.9542 ± 0.005 | |
| Zea mays | 1 | 0.9372 | 0.9394 | 0.9338 | 0.9405 | 0.8744 | 0.9849 |
| 2 | 0.9340 | 0.9380 | 0.9313 | 0.9369 | 0.8681 | 0.9846 | |
| 3 | 0.9312 | 0.9318 | 0.9303 | 0.9321 | 0.8624 | 0.9817 | |
| 4 | 0.9310 | 0.9350 | 0.9259 | 0.9361 | 0.8620 | 0.9812 | |
| 5 | 0.9387 | 0.9367 | 0.9408 | 0.9366 | 0.8773 | 0.9833 | |
| Average | 0.9344 ± 0.003 | 0.9362 ± 0.003 | 0.9324 ± 0.005 | 0.9364 ± 0.003 | 0.8689 ± 0.006 | 0.9831 ± 0.001 |
| Dataset | Method | ACC | PRE | SEN | SPE | MCC | AUC |
|---|---|---|---|---|---|---|---|
| Arabidopsis thaliana | PLRPIM | 0.8998 | 0.9308 | 0.8636 | 0.9354 | 0.8015 | 0.9542 |
| LGBM | 0.8950 | 0.9182 | 0.8668 | 0.9230 | 0.7912 | 0.8949 | |
| XGB | 0.7452 | 0.7615 | 0.7661 | 0.7187 | 0.4993 | 0.8352 | |
| RF | 0.7088 | 0.6851 | 0.8164 | 0.5951 | 0.4294 | 0.8171 | |
| AdaBoost | 0.6962 | 0.6829 | 0.8234 | 0.5622 | 0.4214 | 0.8061 | |
| DT | 0.6233 | 0.6331 | 0.6060 | 0.6359 | 0.2478 | 0.6284 | |
| Zea mays | PLRPIM | 0.9344 | 0.9362 | 0.9324 | 0.9364 | 0.8688 | 0.9831 |
| LGBM | 0.9317 | 0.9331 | 0.9300 | 0.9333 | 0.8634 | 0.9317 | |
| XGB | 0.7936 | 0.7689 | 0.8426 | 0.7446 | 0.5909 | 0.8862 | |
| AdaBoost | 0.7849 | 0.7676 | 0.8182 | 0.7516 | 0.5725 | 0.8693 | |
| RF | 0.7536 | 0.7407 | 0.7972 | 0.7103 | 0.5111 | 0.8641 | |
| DT | 0.6523 | 0.6500 | 0.6676 | 0.6368 | 0.3049 | 0.6894 |
| Dataset | Method | ACC | PRE | SEN | SPE | MCC | AUC |
|---|---|---|---|---|---|---|---|
| Arabidopsis thaliana | PLRPIM | 0.8998 | 0.9308 | 0.8636 | 0.9354 | 0.8015 | 0.9546 |
| IPMiner | 0.8275 | 0.8930 | 0.7448 | 0.9107 | 0.6646 | 0.8823 | |
| RPISeq-RF | 0.8059 | 0.8144 | 0.7922 | 0.8200 | 0.6124 | 0.8761 | |
| RPI-SAN | 0.7579 | 0.7955 | 0.6966 | 0.8199 | 0.5210 | 0.8164 | |
| Zea mays | PLRPIM | 0.9344 | 0.9362 | 0.9324 | 0.9364 | 0.8688 | 0.9823 |
| IPMiner | 0.8127 | 0.8142 | 0.8106 | 0.8148 | 0.6258 | 0.9034 | |
| RPISeq-RF | 0.8069 | 0.7993 | 0.8192 | 0.7945 | 0.6142 | 0.8980 | |
| RPI-SAN | 0.7890 | 0.7909 | 0.7869 | 0.7911 | 0.5784 | 0.8792 |
| Species | lncRNAs | Biological Functions |
|---|---|---|
| Arabidopsis thaliana | TCONS_00011717 | GO:0006913—Nucleocytoplasmic transport |
| TCONS_00008833 | GO:0006083—Acetate metabolic process | |
| TCONS_00012080 | GO:0009867—Regulation of jasmonic acid mediated signaling pathway | |
| Zea mays | GRMZM2G374777 | PO:0001052—2 leaf expansion stage PO:0006339—juvenile vascular leaf PO:0009089—endosperm |
| GRMZM2G097084 | GO:0005524—ATP binding GO:0006183—GTP biosynthetic process GO:0006952—defense response | |
| GRMZM2G078523 | PO:0001052—2 leaf expansion stage PO:0009084—pericarp PO:0009089—endosperm | |
| GRMZM2G543070 | PO:0001052—2 leaf expansion stage PO:0001095—true leaf formation stage PO:0007016—4 flowering stage | |
| GRMZM2G147020 | PO:0001052—2 leaf expansion stage PO:0001095—true leaf formation stage PO:0007016—4 flowering stage PO:0001009—D pollen mother cell meiosis stage |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wekesa, J.S.; Luan, Y.; Chen, M.; Meng, J. A Hybrid Prediction Method for Plant lncRNA-Protein Interaction. Cells 2019, 8, 521. https://doi.org/10.3390/cells8060521
Wekesa JS, Luan Y, Chen M, Meng J. A Hybrid Prediction Method for Plant lncRNA-Protein Interaction. Cells. 2019; 8(6):521. https://doi.org/10.3390/cells8060521
Chicago/Turabian StyleWekesa, Jael Sanyanda, Yushi Luan, Ming Chen, and Jun Meng. 2019. "A Hybrid Prediction Method for Plant lncRNA-Protein Interaction" Cells 8, no. 6: 521. https://doi.org/10.3390/cells8060521
APA StyleWekesa, J. S., Luan, Y., Chen, M., & Meng, J. (2019). A Hybrid Prediction Method for Plant lncRNA-Protein Interaction. Cells, 8(6), 521. https://doi.org/10.3390/cells8060521





