Regulation of Survivin Isoform Expression by GLI Proteins in Ovarian Cancer
Abstract
1. Introduction
2. Materials and Methods
2.1. Generation of Knockout Cell Lines
2.2. Cell Culture Experiments
2.3. Clinical Samples
2.4. Western Blot
2.5. Expression Analysis
2.6. Genotyping
2.7. Statistical Analysis
3. Results and Discussion
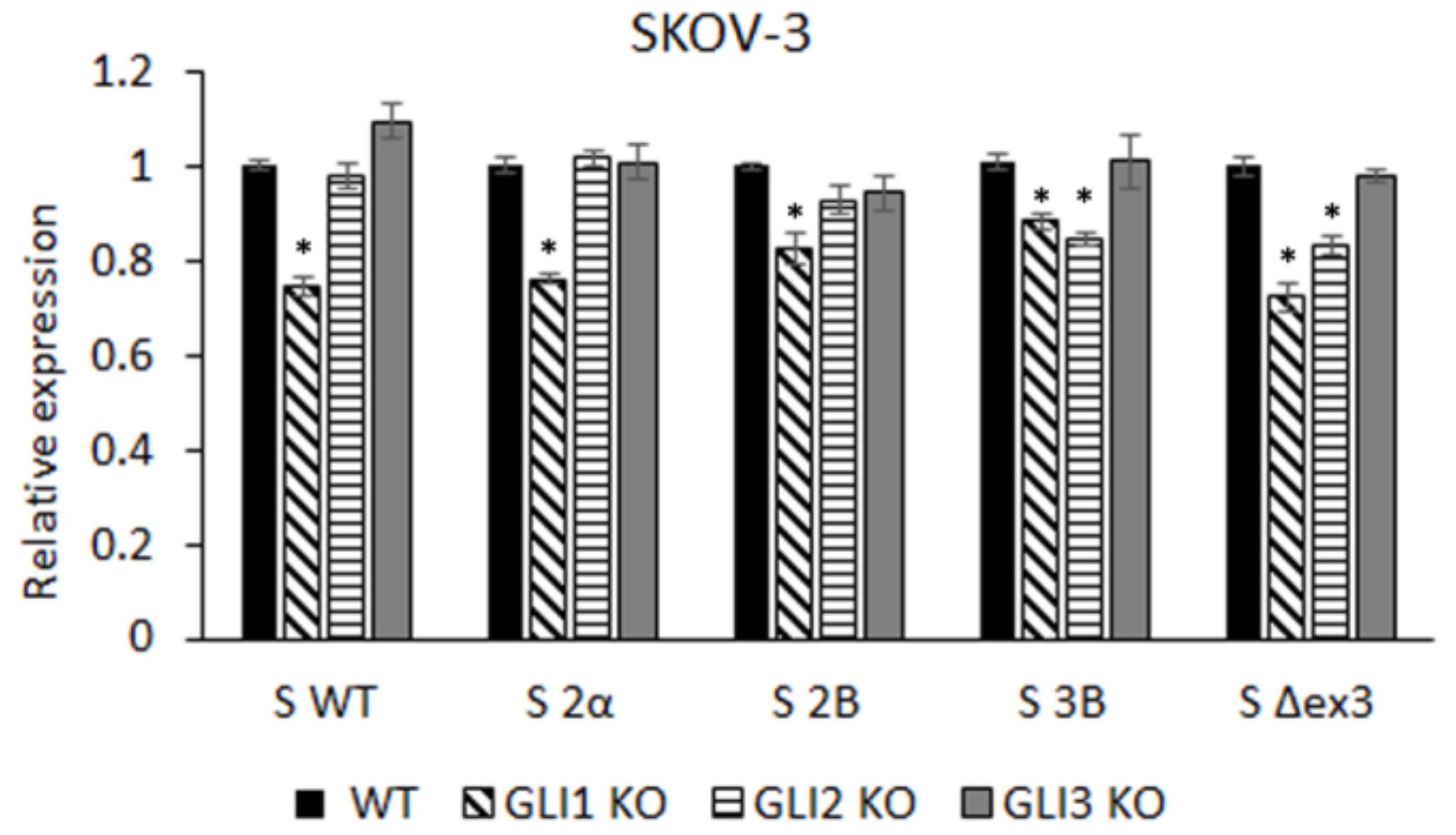
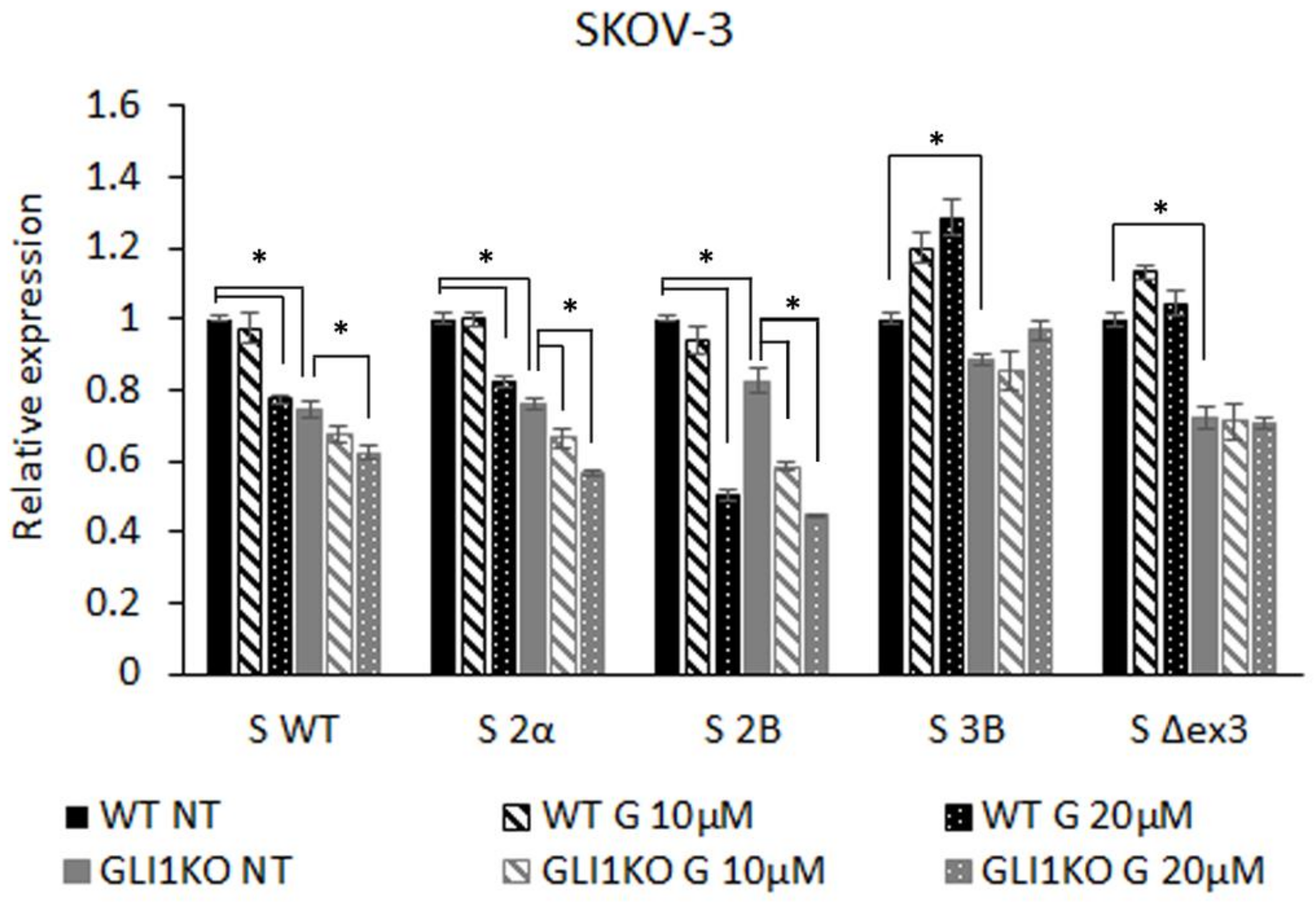
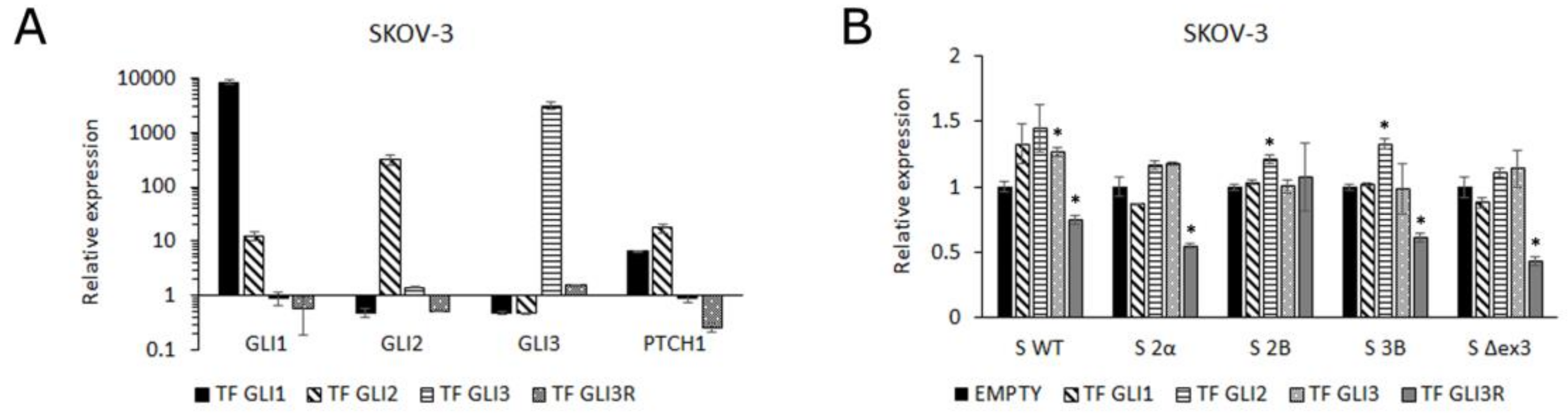
3.1. GLI Regulation of Survivin Isoform Expression
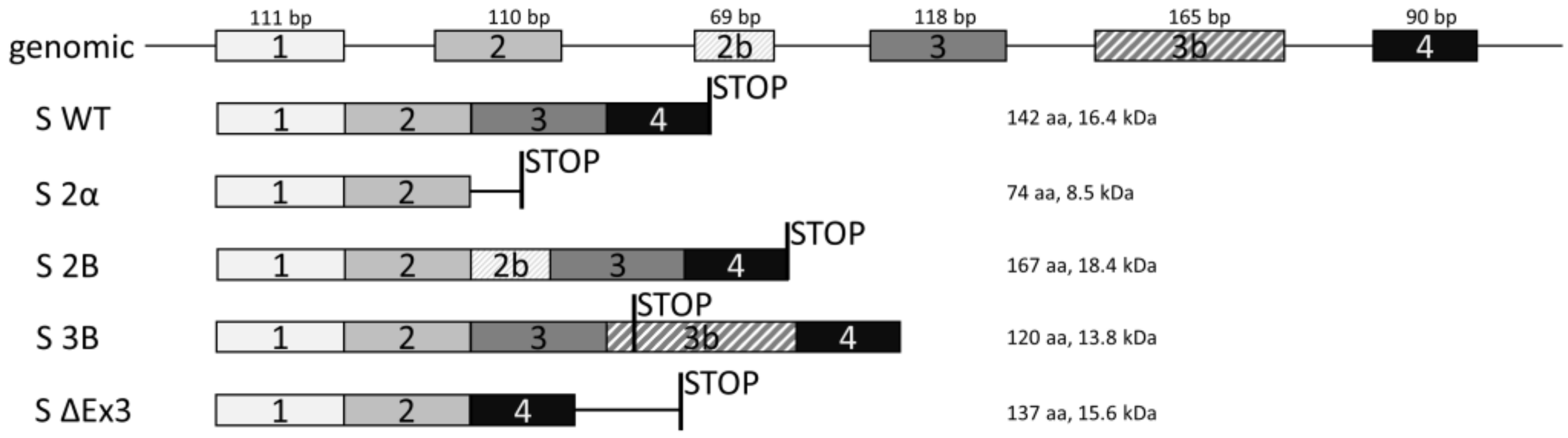
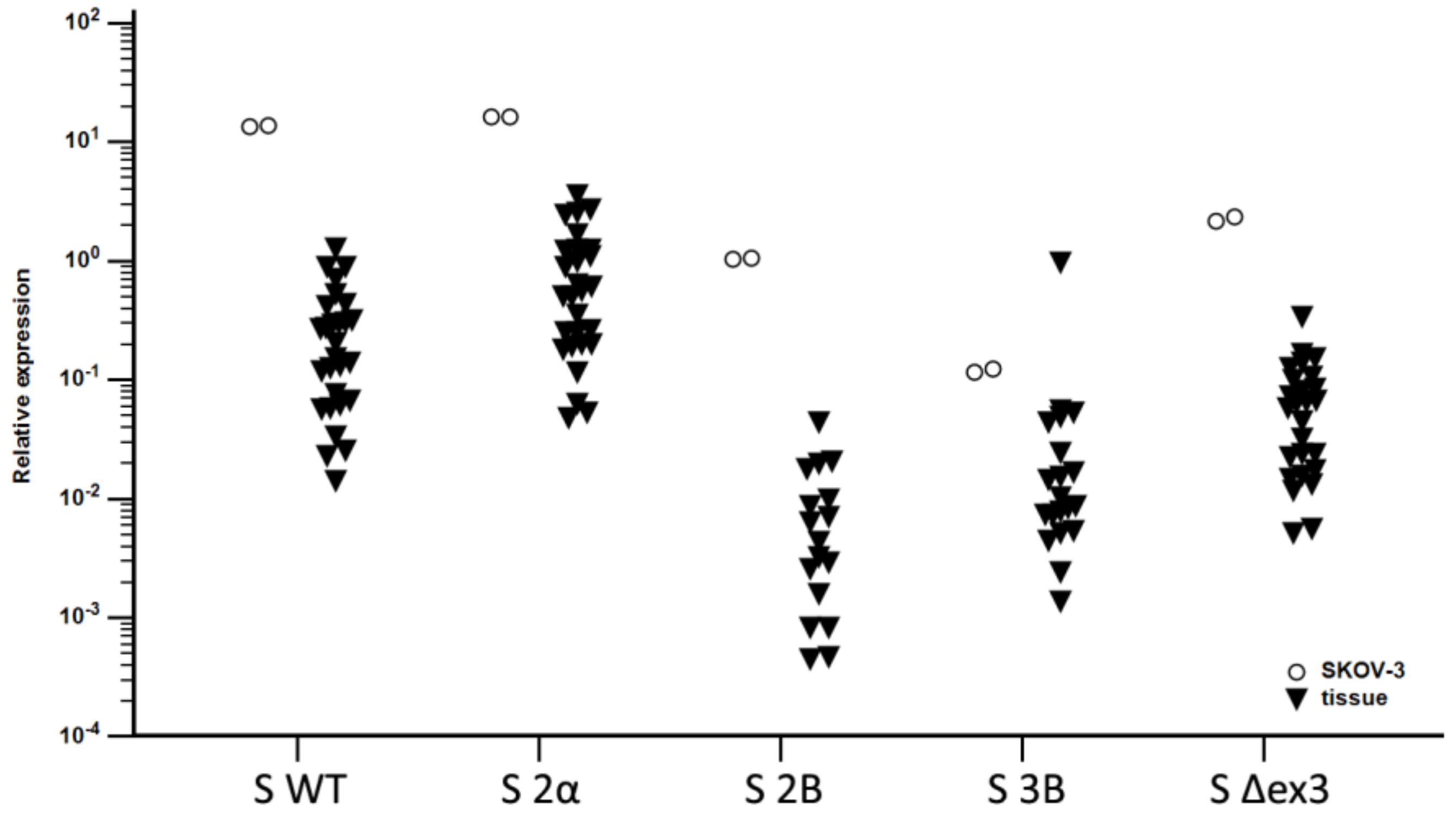
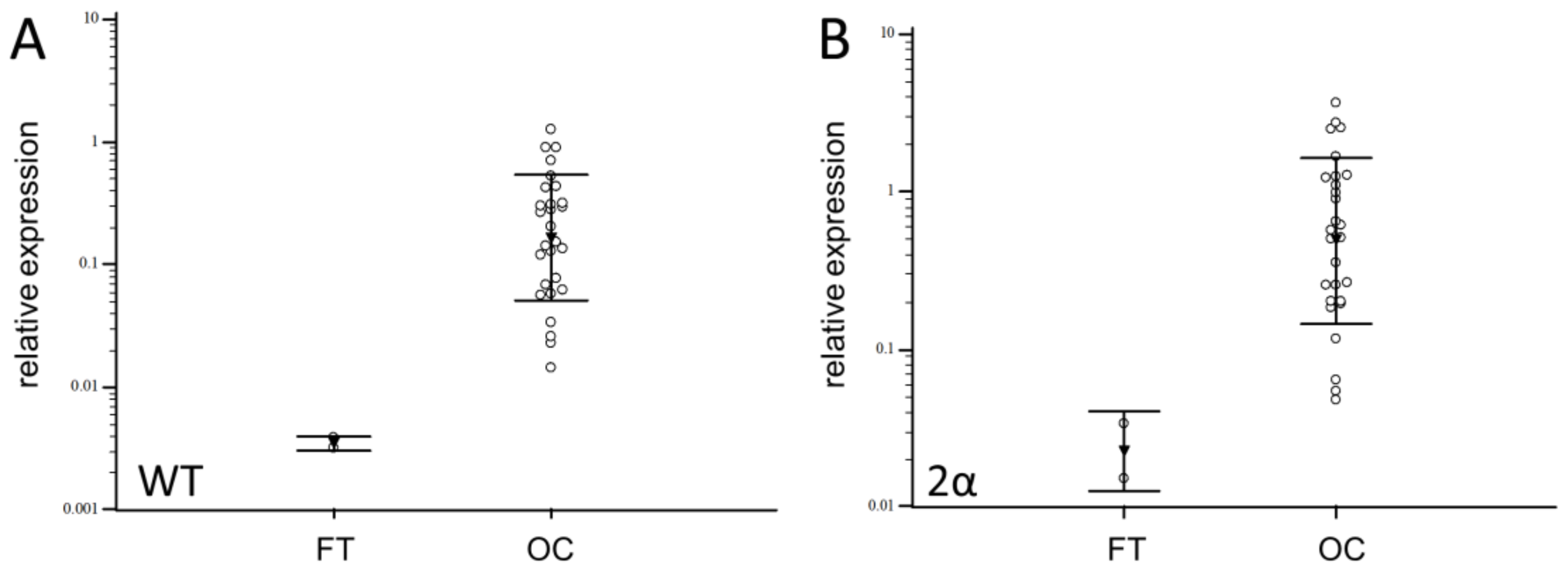
3.2. Expression of Splice Variants
3.3. Genotyping
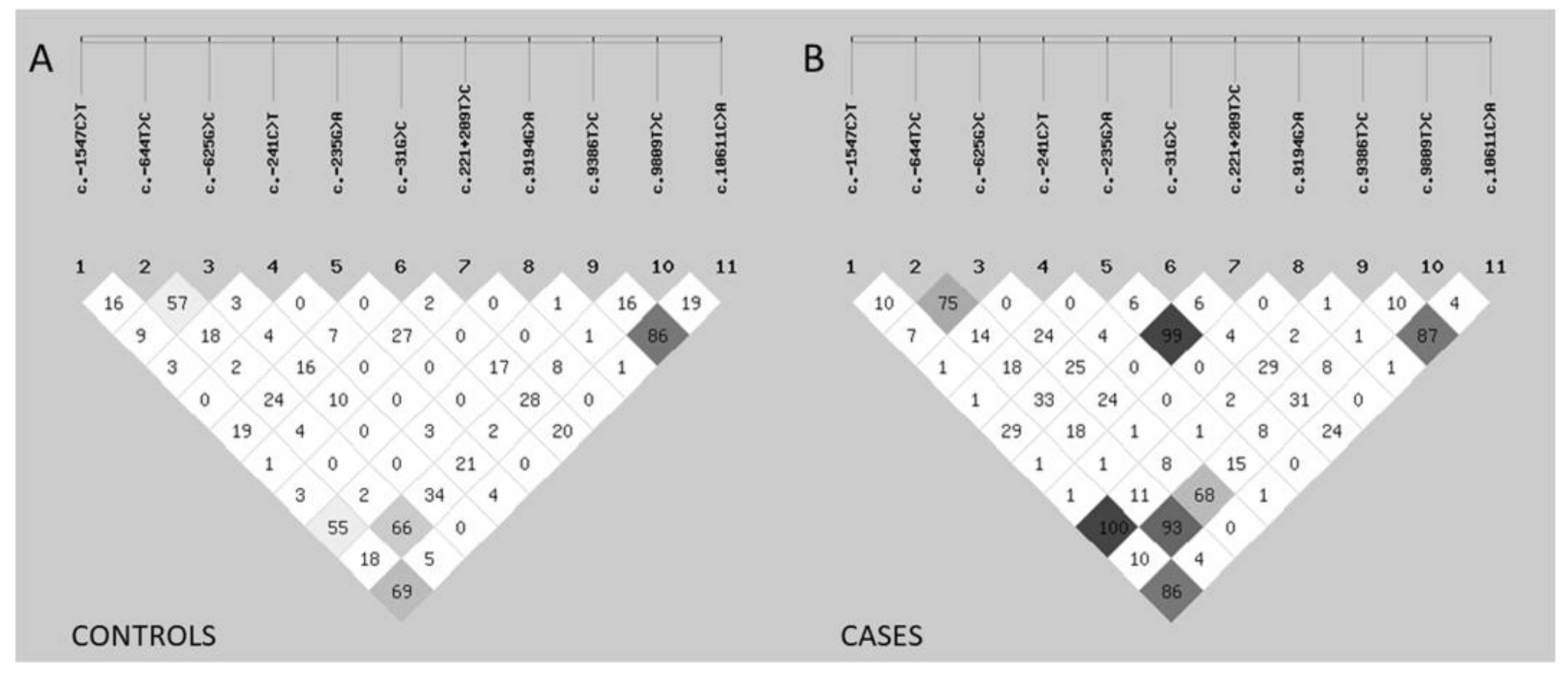
3.4. Linkage Disequilibrium
3.5. Correlation of Splice Variant Expression with Genetic Polymorphisms in OC Patients
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bell, S.M.; Schreiner, C.M.; Goetz, J.A.; Robbins, D.J.; Scott, W.J. Shh signaling in limb bud ectoderm: Potential role in teratogen-induced postaxial ectrodactyly. Dev. Dyn. 2005, 233, 313–325. [Google Scholar] [CrossRef]
- Sabol, M.; Trnski, D.; Musani, V.; Ozretić, P.; Levanat, S. Role of GLI Transcription Factors in Pathogenesis and Their Potential as New Therapeutic Targets. Int. J. Mol. Sci. 2018, 19, 2562. [Google Scholar] [CrossRef]
- Sabol, M.; Car, D.; Musani, V.; Ozretic, P.; Oreskovic, S.; Weber, I.; Levanat, S. The Hedgehog signaling pathway in ovarian teratoma is stimulated by Sonic Hedgehog which induces internalization of Patched. Int. J. Oncol. 2012, 41, 1411–1418. [Google Scholar] [CrossRef] [PubMed]
- Vlčková, K.; Ondrušová, L.; Vachtenheim, J.; Réda, J.; Dundr, P.; Zadinová, M.; Žáková, P.; Poučková, P. Survivin, a novel target of the Hedgehog/GLI signaling pathway in human tumor cells. Cell Death Dis. 2016, 7, e2048. [Google Scholar] [CrossRef]
- Brun, S.N.; Markant, S.L.; Esparza, L.A.; Garcia, G.; Terry, D.; Huang, J.-M.; Pavlyukov, M.S.; Li, X.-N.; Grant, G.A.; Crawford, J.R.; et al. Survivin as a therapeutic target in Sonic hedgehog-driven medulloblastoma. Oncogene 2015, 34, 3770–3779. [Google Scholar] [CrossRef] [PubMed]
- Altieri, D.C. Validating survivin as a cancer therapeutic target. Nat. Rev. Cancer 2003, 3, 46–54. [Google Scholar] [CrossRef] [PubMed]
- Han, C.H.; Wei, Q.; Lu, K.K.; Liu, Z.; Mills, G.B.; Wang, L.-E. Polymorphisms in the survivin promoter are associated with age of onset of ovarian cancer. Int. J. Clin. Exp. Med. 2009, 2, 289–299. [Google Scholar]
- Antonacopoulou, A.G.; Floratou, K.; Bravou, V.; Kottorou, A.; Dimitrakopoulos, F.-I.; Marousi, S.; Stavropoulos, M.; Koutras, A.K.; Scopa, C.D.; Kalofonos, H.P. The survivin -31 snp in human colorectal cancer correlates with survivin splice variant expression and improved overall survival. Anal. Cell Pathol. 2010, 33, 177–189. [Google Scholar] [CrossRef]
- Ambrosini, G.; Adida, C.; Altieri, D.C. A novel anti-apoptosis gene, survivin, expressed in cancer and lymphoma. Nat. Med. 1997, 3, 917–921. [Google Scholar] [CrossRef]
- Caldas, H.; Honsey, L.E.; Altura, R.A. Survivin 2α: A novel Survivin splice variant expressed in human malignancies. Mol. Cancer 2005, 4, 11. [Google Scholar] [CrossRef]
- Boidot, R.; Vegran, F.; Lizard-Nacol, S. Predictive value of survivin alternative transcript expression in locally advanced breast cancer patients treated with neoadjuvant chemotherapy. Int. J. Mol. Med. 2009, 23, 285–291. [Google Scholar]
- Zaki Dizaji, M.; Ghaffari, S.H.; Hosseini, E.; Alizadeh, N.; Rostami, S.; Momeny, M.; Alimoghaddam, K.; Ghavamzadeh, A. Survivin isoform expression in arsenic trioxide-treated acute promyelocytic leukemia cell line and patients: The odd expression pattern of survivin-2α. Asia Pac. J. Clin. Oncol. 2017, 13, e21–e30. [Google Scholar] [CrossRef] [PubMed]
- Mahotka, C.; Wenzel, M.; Springer, E.; Gabbert, H.E.; Gerharz, C.D. Survivin-deltaEx3 and survivin-2B: Two novel splice variants of the apoptosis inhibitor survivin with different antiapoptotic properties. Cancer Res. 1999, 59, 6097–6102. [Google Scholar] [PubMed]
- Krieg, A.; Mahotka, C.; Krieg, T.; Grabsch, H.; Müller, W.; Takeno, S.; Suschek, C.V.; Heydthausen, M.; Gabbert, H.E.; Gerharz, C.D. Expression of different survivin variants in gastric carcinomas: First clues to a role of survivin-2B in tumour progression. Br. J. Cancer 2002, 86, 737–743. [Google Scholar] [CrossRef] [PubMed]
- Ling, X.; Cheng, Q.; Black, J.D.; Li, F. Forced expression of survivin-2B abrogates mitotic cells and induces mitochondria-dependent apoptosis by blockade of tubulin polymerization and modulation of Bcl-2, Bax, and survivin. J. Biol. Chem. 2007, 282, 27204–27214. [Google Scholar] [CrossRef]
- Cho, G.-S.; Ahn, T.S.; Jeong, D.; Kim, J.-J.; Kim, C.-J.; Cho, H.-D.; Park, D.-K.; Baek, M.-J. Expression of the survivin-2B splice variant related to the progression of colorectal carcinoma. J. Korean Surg. Soc. 2011, 80, 404–411. [Google Scholar] [CrossRef] [PubMed]
- Shi, K.; An, J.; Shan, L.; Jiang, Q.; Li, F.; Ci, Y.; Wu, P.; Duan, J.; Hui, K.; Yang, Y.; et al. Survivin-2B promotes autophagy by accumulating IKK alpha in the nucleus of selenite-treated NB4 cells. Cell Death Dis. 2014, 5, e1071. [Google Scholar] [CrossRef] [PubMed]
- Vivas-Mejia, P.E.; Rodriguez-Aguayo, C.; Han, H.-D.; Shahzad, M.M.K.; Valiyeva, F.; Shibayama, M.; Chavez-Reyes, A.; Sood, A.K.; Lopez-Berestein, G. Silencing survivin splice variant 2B leads to antitumor activity in taxane--resistant ovarian cancer. Clin. Cancer Res. 2011, 17, 3716–3726. [Google Scholar] [CrossRef]
- Badran, A.; Yoshida, A.; Ishikawa, K.; Goi, T.; Yamaguchi, A.; Ueda, T.; Inuzuka, M. Identification of a novel splice variant of the human anti-apoptopsis gene survivin. Biochem. Biophys. Res. Commun. 2004, 314, 902–907. [Google Scholar] [CrossRef]
- Knauer, S.K.; Bier, C.; Schlag, P.; Fritzmann, J.; Dietmaier, W.; Rödel, F.; Klein-Hitpass, L.; Kovács, A.F.; Döring, C.; Hansmann, M.-L.; et al. The survivin isoform survivin-3B is cytoprotective and can function as a chromosomal passenger complex protein. Cell Cycle 2007, 6, 1502–1509. [Google Scholar] [CrossRef]
- Sawai, K.; Goi, T.; Hirono, Y.; Katayama, K.; Yamaguchi, A. Survivin-3B gene decreases the invasion-inhibitory effect of colon cancer cells with 5-fluorouracil. Oncol. Res. 2010, 18, 541–547. [Google Scholar] [CrossRef]
- Végran, F.; Boidot, R.; Bonnetain, F.; Cadouot, M.; Chevrier, S.; Lizard-Nacol, S. Apoptosis gene signature of Survivin and its splice variant expression in breast carcinoma. Endocr. Relat. Cancer 2011, 18, 783–792. [Google Scholar] [CrossRef] [PubMed]
- Mahotka, C.; Liebmann, J.; Wenzel, M.; Suschek, C.V.; Schmitt, M.; Gabbert, H.E.; Gerharz, C.D. Differential subcellular localization of functionally divergent survivin splice variants. Cell Death Differ. 2002, 9, 1334–1342. [Google Scholar] [CrossRef] [PubMed]
- Lopergolo, A.; Tavecchio, M.; Lisanti, S.; Ghosh, J.C.; Dohi, T.; Faversani, A.; Vaira, V.; Bosari, S.; Tanigawa, N.; Delia, D.; et al. Chk2 phosphorylation of survivin-DeltaEx3 contributes to a DNA damage-sensing checkpoint in cancer. Cancer Res. 2012, 72, 3251–3259. [Google Scholar] [CrossRef] [PubMed]
- Vandghanooni, S.; Eskandani, M.; Montazeri, V.; Halimi, M.; Babaei, E.; Feizi, M.A.H. Survivin-deltaEx3: A novel biomarker for diagnosis of papillary thyroid carcinoma. J. Cancer Res. Ther. 2011, 7, 325–330. [Google Scholar] [PubMed]
- Altieri, D.C. Targeted therapy by disabling crossroad signaling networks: The survivin paradigm. Mol. Cancer Ther. 2006, 5, 478–482. [Google Scholar] [CrossRef] [PubMed]
- Stobiecka, M.; Chalupa, A.; Dworakowska, B. Piezometric biosensors for anti-apoptotic protein survivin based on buried positive-potential barrier and immobilized monoclonal antibodies. Biosens. Bioelectron. 2016, 84, 37–43. [Google Scholar] [CrossRef]
- Ratajczak, K.; Krazinski, B.E.; Kowalczyk, A.E.; Dworakowska, B.; Jakiela, S.; Stobiecka, M. Optical Biosensing System for the Detection of Survivin mRNA in Colorectal Cancer Cells Using a Graphene Oxide Carrier-Bound Oligonucleotide Molecular Beacon. Nanomaterials 2018, 8, 510. [Google Scholar] [CrossRef]
- Jang, J.S.; Kim, K.M.; Kang, K.H.; Choi, J.E.; Lee, W.K.; Kim, C.H.; Kang, Y.M.; Kam, S.; Kim, I.-S.; Jun, J.E.; et al. Polymorphisms in the survivin gene and the risk of lung cancer. Lung Cancer 2008, 60, 31–39. [Google Scholar] [CrossRef]
- Cheng, Z.-J.; Hu, L.-H.; Huang, S.-J. [Correlation of -31G/C polymorphisms of survivin promoter to tumorigenesis of gastric carcinoma]. Ai Zheng 2008, 27, 258–263. [Google Scholar]
- Kawata, N.; Tsuchiya, N.; Horikawa, Y.; Inoue, T.; Tsuruta, H.; Maita, S.; Satoh, S.; Mitobe, Y.; Narita, S.; Habuchi, T. Two survivin polymorphisms are cooperatively associated with bladder cancer susceptibility. Int. J. Cancer 2011, 129, 1872–1880. [Google Scholar] [CrossRef] [PubMed]
- Weng, C.J.; Hsieh, Y.H.; Chen, M.K.; Tsai, C.M.; Lin, C.W.; Yang, S.F. Survivin SNP-carcinogen interactions in oral cancer. J. Dent. Res. 2012, 91, 358–363. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, Y.-S.; Tsai, C.-M.; Yeh, C.-B.; Yang, S.-F.; Hsieh, Y.-H.; Weng, C.-J. Survivin T9809C, an SNP located in 3’-UTR, displays a correlation with the risk and clinicopathological development of hepatocellular carcinoma. Ann. Surg. Oncol. 2012, 19, S625–S633. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.; Bevier, M.; Johansson, R.; Enquist-Olsson, K.; Henriksson, R.; Hemminki, K.; Lenner, P.; Försti, A. Prognostic impact of polymorphisms in the MYBL2 interacting genes in breast cancer. Breast Cancer Res. Treat. 2012, 131, 1039–1047. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex Genome Engineering Using CRISPR/Cas Systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Ozretić, P.; Trnski, D.; Musani, V.; Maurac, I.; Kalafatić, D.; Orešković, S.; Levanat, S.; Sabol, M. Non-canonical Hedgehog signaling activation in ovarian borderline tumors and ovarian carcinomas. Int. J. Oncol. 2017, 51, 1869–1877. [Google Scholar] [CrossRef] [PubMed]
- Hofstetter, G.; Berger, A.; Fiegl, H.; Slade, N.; Zorić, A.; Holzer, B.; Schuster, E.; Mobus, V.J.; Reimer, D.; Daxenbichler, G.; et al. Alternative splicing of p53 and p73: The novel p53 splice variant p53delta is an independent prognostic marker in ovarian cancer. Oncogene 2010, 29, 1997–2004. [Google Scholar] [CrossRef]
- McCall, M.N.; McMurray, H.R.; Land, H.; Almudevar, A. On non-detects in qPCR data. Bioinformatics 2014, 30, 2310–2316. [Google Scholar] [CrossRef]
- Untergasser, A.; Cutcutache, I.; Koressaar, T.; Ye, J.; Faircloth, B.C.; Remm, M.; Rozen, S.G. Primer3--new capabilities and interfaces. Nucleic Acids Res. 2012, 40, e115. [Google Scholar] [CrossRef]
- Cvok, M.L.; Cretnik, M.; Musani, V.; Ozretic, P.; Levanat, S. New sequence variants in BRCA1 and BRCA2 genes detected by high-resolution melting analysis in an elderly healthy female population in Croatia. Clin. Chem. Lab. Med. 2008, 46, 1376–1383. [Google Scholar] [CrossRef]
- Roessler, E.; Ermilov, A.N.; Grange, D.K.; Wang, A.; Grachtchouk, M.; Dlugosz, A.A.; Muenke, M. A previously unidentified amino-terminal domain regulates transcriptional activity of wild-type and disease-associated human GLI2. Hum. Mol. Genet. 2005, 14, 2181–2188. [Google Scholar] [CrossRef]
- Dai, J.; Jin, G.; Dong, J.; Chen, Y.; Xu, L.; Hu, Z.; Shen, H. Prognostic significance of survivin polymorphisms on non-small cell lung cancer survival. J. Thorac. Oncol. 2010, 5, 1748–1754. [Google Scholar] [CrossRef] [PubMed]
- Aynaci, E.; Coskunpinar, E.; Eren, A.; Kum, O.; Oltulu, Y.M.; Akkaya, N.; Turna, A.; Yaylim, I.; Yildiz, P. Association between survivin gene promoter -31G/C and -644C/T polymorphisms and non-small cell lung cancer. Genet. Mol. Res. 2013, 12, 3975–3982. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Xiong, G.; Chen, X.; Xu, X.; Wang, K.; Fu, Y.; Yang, K.; Bai, Y. Polymorphisms of survivin promoter are associated with risk of esophageal squamous cell carcinoma. J. Cancer Res. Clin. Oncol. 2009, 135, 1341–1349. [Google Scholar] [CrossRef] [PubMed]
- Hmeljak, J.; Erčulj, N.; Dolžan, V.; Kern, I.; Cör, A. BIRC5 promoter SNPs do not affect nuclear survivin expression and survival of malignant pleural mesothelioma patients. J. Cancer Res. Clin. Oncol. 2011, 137, 1641–1651. [Google Scholar] [CrossRef]
- Boidot, R.; Vegran, F.; Jacob, D.; Chevrier, S.; Gangneux, N.; Taboureau, J.; Oudin, C.; Rainville, V.; Mercier, L.; Lizard-Nacol, S. The expression of BIRC5 is correlated with loss of specific chromosomal regions in breast carcinomas. Genes Chromosomes Cancer 2008, 47, 299–308. [Google Scholar] [CrossRef]
- Pu, F.; Shao, Z.; Yang, S.; Liu, J.; Lin, S.; Ma, X.; Yang, H. Association between functional variants in BIRC5/survivin gene 3’ untranslated region and mRNA expression in lymphoblastoid cell lines. Oncol. Lett. 2015, 10, 2319–2322. [Google Scholar] [CrossRef] [PubMed]
- Li, X.-B.; Li, S.-N.; Yang, Z.-H.; Cao, L.; Duan, F.-L.; Sun, X.-W. Polymorphisms of survivin and its protein expression are associated with colorectal cancer susceptibility in Chinese population. DNA Cell Biol. 2013, 32, 236–242. [Google Scholar] [CrossRef]
- Rosato, A.; Menin, C.; Boldrin, D.; Dalla Santa, S.; Bonaldi, L.; Scaini, M.C.; Del Bianco, P.; Zardo, D.; Fassan, M.; Cappellesso, R.; et al. Survivin expression impacts prognostically on NSCLC but not SCLC. Lung Cancer 2013, 79, 180–186. [Google Scholar] [CrossRef]
- Kafadar, D.; Yaylim, I.; Kafadar, A.M.; Cacina, C.; Ergen, A.; Kaynar, M.Y.; Isbir, T. Investigation of Survivin Gene Polymorphism and Serum Survivin Levels in Patients with Brain Tumors. Anticancer Res. 2018, 38, 5991–5998. [Google Scholar] [CrossRef]
- Zu, Y.; Ban, J.; Xia, Z.; Wang, J.; Cai, Y.; Ping, W.; Sun, W. Genetic variation in a miR-335 binding site in BIRC5 alters susceptibility to lung cancer in Chinese Han populations. Biochem. Biophys. Res. Commun. 2013, 430, 529–534. [Google Scholar] [CrossRef] [PubMed]
| S 2α | S 2B | S 3B | S Δex3 | - | - |
|---|---|---|---|---|---|
| 0.92 | 0.69 | 0.81 | 0.75 | S WT | ϱ |
| <0.0001 | 0.002 | <0.0001 | <0.0001 | p | |
| - | 0.67 | 0.80 | 0.81 | S 2α | ϱ |
| - | 0.005 | <0.0001 | <0.0001 | p | |
| - | - | 0.55 | 0.67 | S 2B | ϱ |
| - | - | 0.021 | 0.004 | p | |
| - | - | - | 0.66 | S 3B | ϱ |
| - | - | - | 0.002 | p | |
| - | - | - | - | - | - |
| Spearman rank correlation coefficient (ϱ) | |||||
| Gene Region | SNP ID Number | Nucleotide Change | MAF in OC (%) | MAF in Controls (%) | p-Value |
|---|---|---|---|---|---|
| promoter | rs3764383 | c.-1547C>T * | 23.1 | 25.0 | 0.871 |
| promoter | rs143396310 | c.-1458C>T | 1.3 | 1.4 | 1.000 |
| promoter | rs8073903 | c.-644T>C | 26.3 | 33.1 | 0.297 |
| promoter | rs8073069 | c.-625G>C | 21.3 | 22.3 | 1.000 |
| promoter | rs17878731 | c.-267G>A | 1.3 | 0.7 | 1.000 |
| promoter | rs17878467 | c.-241C>T | 5.0 | 10.8 | 0.219 |
| promoter | rs17887126 | c.-235G>A | 6.3 | 1.4 | 0.053 |
| 5′UTR | rs9904341 | c.-31G>C | 48.8 | 37.2 | 0.093 |
| intron 2 | rs4789551 | c.221+209T>C | 6.4 | 4.7 | 0.756 |
| exon 4 | rs2071214 | c.9194G>A ** | 3.8 | 3.4 | 1.000 |
| 3′UTR | rs17885521 | c.9288G>C | 1.3 | 2.0 | 1.000 |
| 3′UTR | rs17882627 | c.9342G>A | 2.5 | 1.4 | 0.614 |
| 3′UTR | rs2239680 | c.9386T>C | 23.8 | 23.0 | 1.000 |
| 3′UTR | rs1042489 | c.9809T>C | 25.0 | 35.8 | 0.104 |
| 3′UTR | rs2661694 | c.10611C>A | 26.3 | 25.7 | 1.000 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Trnski, D.; Gregorić, M.; Levanat, S.; Ozretić, P.; Rinčić, N.; Vidaković, T.M.; Kalafatić, D.; Maurac, I.; Orešković, S.; Sabol, M.; et al. Regulation of Survivin Isoform Expression by GLI Proteins in Ovarian Cancer. Cells 2019, 8, 128. https://doi.org/10.3390/cells8020128
Trnski D, Gregorić M, Levanat S, Ozretić P, Rinčić N, Vidaković TM, Kalafatić D, Maurac I, Orešković S, Sabol M, et al. Regulation of Survivin Isoform Expression by GLI Proteins in Ovarian Cancer. Cells. 2019; 8(2):128. https://doi.org/10.3390/cells8020128
Chicago/Turabian StyleTrnski, Diana, Maja Gregorić, Sonja Levanat, Petar Ozretić, Nikolina Rinčić, Tajana Majić Vidaković, Držislav Kalafatić, Ivana Maurac, Slavko Orešković, Maja Sabol, and et al. 2019. "Regulation of Survivin Isoform Expression by GLI Proteins in Ovarian Cancer" Cells 8, no. 2: 128. https://doi.org/10.3390/cells8020128
APA StyleTrnski, D., Gregorić, M., Levanat, S., Ozretić, P., Rinčić, N., Vidaković, T. M., Kalafatić, D., Maurac, I., Orešković, S., Sabol, M., & Musani, V. (2019). Regulation of Survivin Isoform Expression by GLI Proteins in Ovarian Cancer. Cells, 8(2), 128. https://doi.org/10.3390/cells8020128







