Role of Ceramidases in Sphingolipid Metabolism and Human Diseases
Abstract
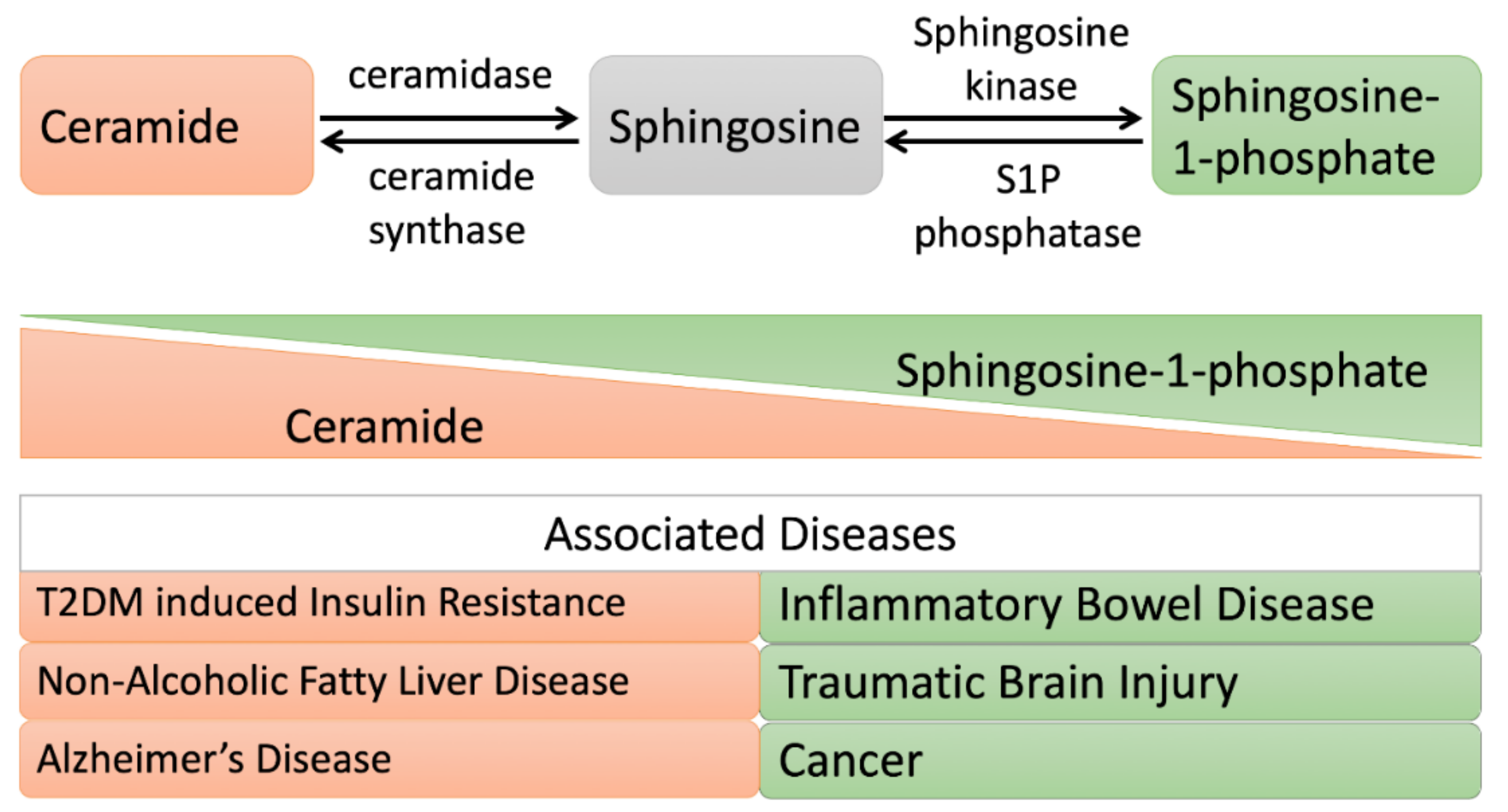
1. Introduction
2. General Features of Ceramidases
2.1. Nomenclature and Gene Loci
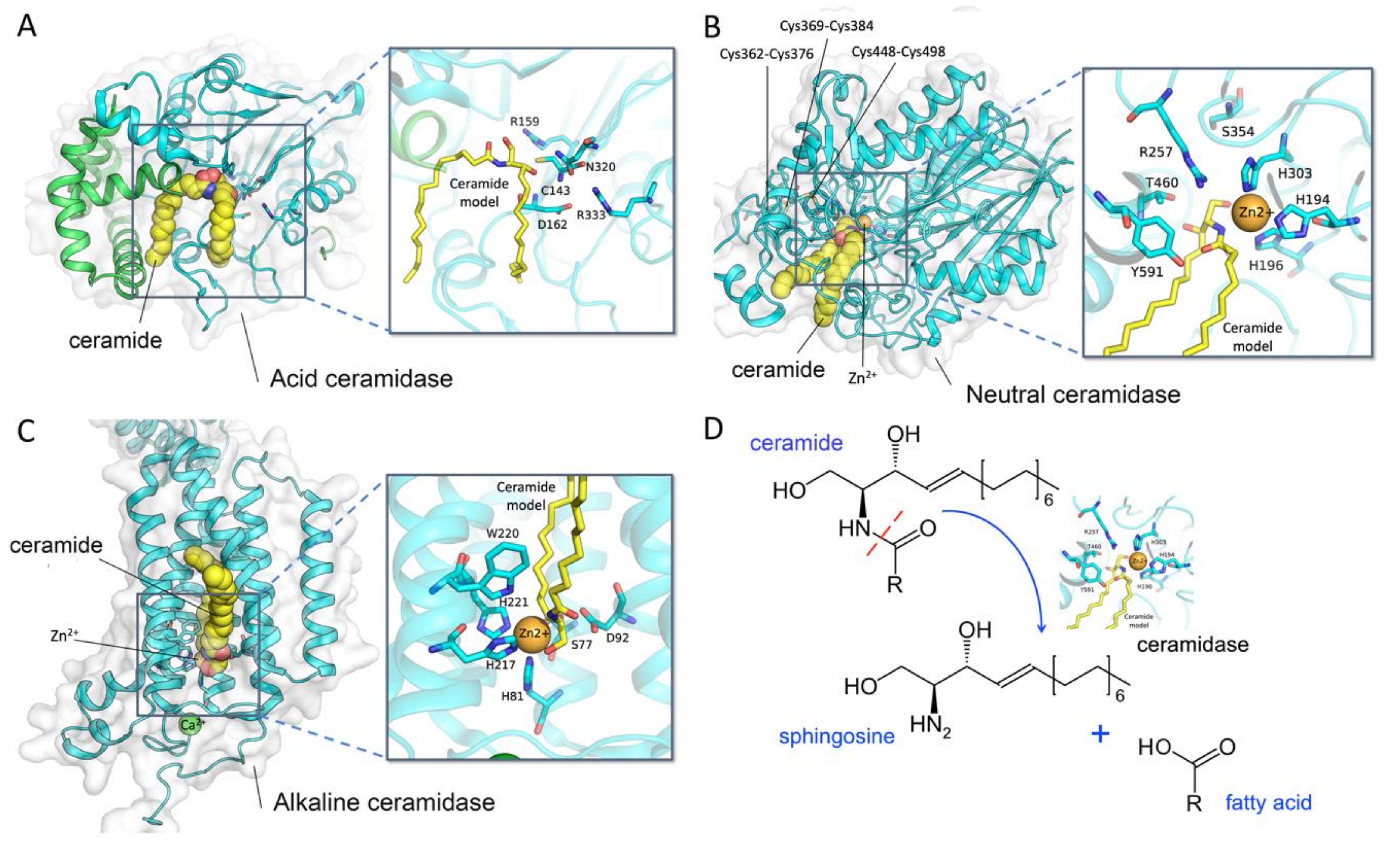
2.2. Acid Ceramidases
2.3. Neutral Ceramidases
2.4. Alkaline Ceramidases
3. Physiology of Neutral Ceramidase ASAH2
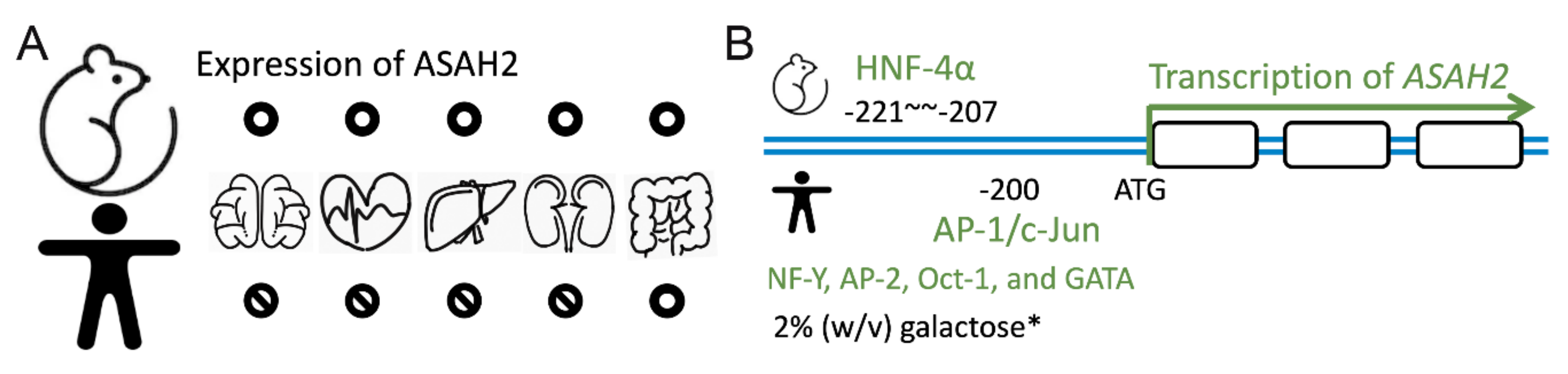
3.1. Tissue Distribution Levels of Neutral Ceramidases in Rats and Mices
3.2. Human Expression of Neutral Ceramidases
3.3. Factors Manipulating ASAH2 Expression and Activity
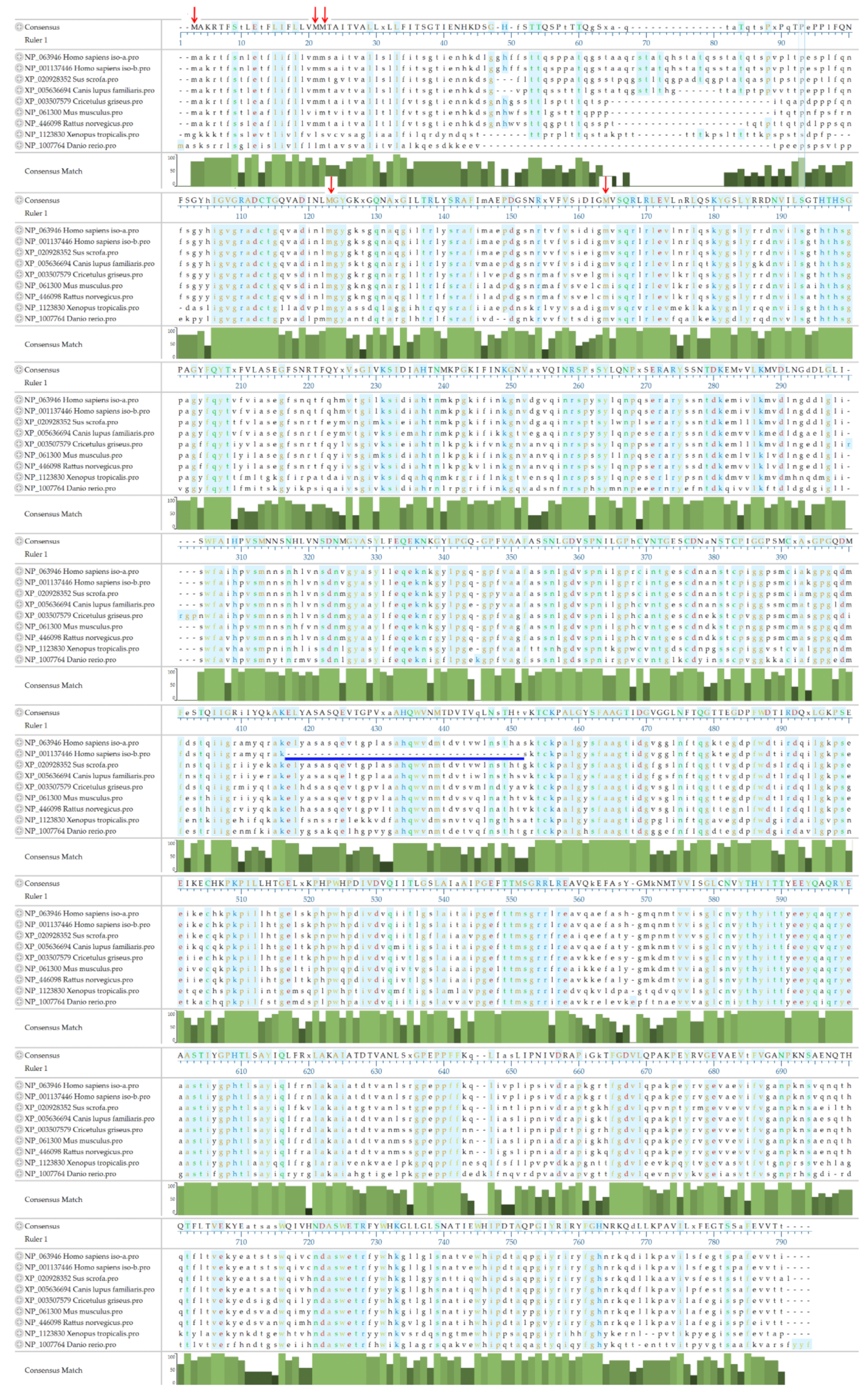
3.4. The Amino Acid Sequence of ASAH2
3.5. Structure of Human ASAH2
4. Role of Ceramidases in Pathological Conditions
4.1. Genetic Disorders Related to Ceramidases
4.1.1. Genetic Variations of ASAH1
4.1.2. Genetic Variations of ASAH2
4.1.3. Genetic Variations of ACER3
4.2. Metabolic Disorders
4.2.1. Insulin Resistance
4.2.2. Cardiovascular Disease
4.2.3. Alzheimer’s Disease
4.2.4. Traumatic Brain Injury
4.2.5. Cancer
4.2.6. Inflammatory Bowel Disease
5. Ceramidase Inhibitors
6. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| S1P | Sphingosine-1 phosphate |
| AD | Alzheimer’s disease |
| ASAH1 ASAH2 ACER1 ACER2 ACER3 | Acid ceramidase; N-acylsphingosine amidohydrolase 1 Neutral ceramidase; N-Acylsphingosine Amidohydrolase 2 Alkaline ceramidase 1 Alkaline ceramidase 2 Alkaline ceramidase 3 |
| BODIPY CHO ER FFA IBD NAFLD T2DM | boron-dipyrromethene fluorescent dye Chinese hamster ovary Endoplasmic reticulum Free fatty acid Inflammatory bowel disease Nonalcoholic fatty liver disease Type 2 diabetes mellitus |
| TBI | Traumatic brain injury |
References
- Simons, K.; Ikonen, E. Functional rafts in cell membranes. Nature 1997, 387, 569–572. [Google Scholar] [CrossRef] [PubMed]
- Perry, D.K.; Obeid, L.M.; Hannun, Y.A. Ceramide and the regulation of apoptosis and the stress response. Trends Cardiovasc. Med. 1996, 6, 158–162. [Google Scholar] [CrossRef]
- Spiegel, S.; Cuvillier, O.; Edsall, L.; Kohama, T.; Menzeleev, R.; Olivera, A.; Thomas, D.; Tu, Z.; Van Brocklyn, J.; Wang, F. Roles of sphingosine-1-phosphate in cell growth, differentiation, and death. Biochem. Biokhimiia 1998, 63, 69–73. [Google Scholar]
- Sassoli, C.; Pierucci, F.; Zecchi-Orlandini, S.; Meacci, E. Sphingosine 1-Phosphate (S1P)/ S1P Receptor Signaling and Mechanotransduction: Implications for Intrinsic Tissue Repair/Regeneration. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Van Brocklyn, J.R.; Williams, J.B. The control of the balance between ceramide and sphingosine-1-phosphate by sphingosine kinase: Oxidative stress and the seesaw of cell survival and death. Comp. Biochem. Physiol. Part BBiochem. Mol. Biol. 2012, 163, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Hannun, Y.A.; Obeid, L.M. Principles of bioactive lipid signalling: Lessons from sphingolipids. Nat. Rev. Mol. Cell Biol. 2008, 9, 139–150. [Google Scholar] [CrossRef] [PubMed]
- Filippov, V.; Song, M.A.; Zhang, K.; Vinters, H.V.; Tung, S.; Kirsch, W.M.; Yang, J.; Duerksen-Hughes, P.J. Increased ceramide in brains with Alzheimer’s and other neurodegenerative diseases. J. Alzheimer’s Dis. Jad 2012, 29, 537–547. [Google Scholar] [CrossRef]
- Das, U.N. Is There a Role for Bioactive Lipids in the Pathobiology of Diabetes Mellitus? Front. Endocrinol. 2017, 8, 182. [Google Scholar] [CrossRef]
- Laaksonen, R.; Ekroos, K.; Sysi-Aho, M.; Hilvo, M.; Vihervaara, T.; Kauhanen, D.; Suoniemi, M.; Hurme, R.; Marz, W.; Scharnagl, H.; et al. Plasma ceramides predict cardiovascular death in patients with stable coronary artery disease and acute coronary syndromes beyond LDL-cholesterol. Eur. Heart J. 2016, 37, 1967–1976. [Google Scholar] [CrossRef]
- Maceyka, M.; Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature 2014, 510, 58–67. [Google Scholar] [CrossRef]
- Sugita, M.; Dulaney, J.T.; Moser, H.W. Ceramidase deficiency in Farber’s disease (lipogranulomatosis). Science 1972, 178, 1100–1102. [Google Scholar] [CrossRef] [PubMed]
- Spiegel, S.; Milstien, S. The outs and the ins of sphingosine-1-phosphate in immunity. Nat. Rev. Immunol. 2011, 11, 403–415. [Google Scholar] [CrossRef] [PubMed]
- Gault, C.R.; Obeid, L.M.; Hannun, Y.A. An overview of sphingolipid metabolism: From synthesis to breakdown. Adv. Exp. Med. Biol. 2010, 688, 1–23. [Google Scholar] [PubMed]
- Merrill, A.H., Jr. De novo sphingolipid biosynthesis: A necessary, but dangerous, pathway. J. Biol. Chem. 2002, 277, 25843–25846. [Google Scholar] [CrossRef] [PubMed]
- Kitatani, K.; Idkowiak-Baldys, J.; Hannun, Y.A. The sphingolipid salvage pathway in ceramide metabolism and signaling. Cell. Signal. 2008, 20, 1010–1018. [Google Scholar] [CrossRef]
- Xia, J.Y.; Holland, W.L.; Kusminski, C.M.; Sun, K.; Sharma, A.X.; Pearson, M.J.; Sifuentes, A.J.; McDonald, J.G.; Gordillo, R.; Scherer, P.E. Targeted Induction of Ceramide Degradation Leads to Improved Systemic Metabolism and Reduced Hepatic Steatosis. Cell Metab. 2015, 22, 266–278. [Google Scholar] [CrossRef]
- Bruce, C.R.; Risis, S.; Babb, J.R.; Yang, C.; Kowalski, G.M.; Selathurai, A.; Lee-Young, R.S.; Weir, J.M.; Yoshioka, K.; Takuwa, Y.; et al. Overexpression of sphingosine kinase 1 prevents ceramide accumulation and ameliorates muscle insulin resistance in high-fat diet-fed mice. Diabetes 2012, 61, 3148–3155. [Google Scholar] [CrossRef]
- Mao, C.; Obeid, L.M. Ceramidases: Regulators of cellular responses mediated by ceramide, sphingosine, and sphingosine-1-phosphate. Biochim. Et Biophys. Acta 2008, 1781, 424–434. [Google Scholar] [CrossRef]
- Schulze, H.; Schepers, U.; Sandhoff, K. Overexpression and mass spectrometry analysis of mature human acid ceramidase. Biol. Chem. 2007, 388, 1333–1343. [Google Scholar] [CrossRef]
- Gatt, S. Enzymatic hydrolysis of sphingolipids. I. Hydrolysis and synthesis of ceramides by an enzyme from rat brain. J. Biol. Chem. 1966, 241, 3724–3730. [Google Scholar]
- Okino, N.; He, X.; Gatt, S.; Sandhoff, K.; Ito, M.; Schuchman, E.H. The reverse activity of human acid ceramidase. J. Biol. Chem. 2003, 278, 29948–29953. [Google Scholar] [CrossRef] [PubMed]
- Ferlinz, K.; Kopal, G.; Bernardo, K.; Linke, T.; Bar, J.; Breiden, B.; Neumann, U.; Lang, F.; Schuchman, E.H.; Sandhoff, K. Human acid ceramidase: Processing, glycosylation, and lysosomal targeting. J. Biol. Chem. 2001, 276, 35352–35360. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Okino, N.; Dhami, R.; Dagan, A.; Gatt, S.; Schulze, H.; Sandhoff, K.; Schuchman, E.H. Purification and characterization of recombinant, human acid ceramidase. Catalytic reactions and interactions with acid sphingomyelinase. J. Biol. Chem. 2003, 278, 32978–32986. [Google Scholar] [CrossRef] [PubMed]
- Azuma, N.; O’Brien, J.S.; Moser, H.W.; Kishimoto, Y. Stimulation of acid ceramidase activity by saposin D. Arch. Biochem. Biophys. 1994, 311, 354–357. [Google Scholar] [CrossRef] [PubMed]
- Yildiz, E.P.; Yesil, G.; Bektas, G.; Caliskan, M.; Tatli, B.; Aydinli, N.; Ozmen, M. Spinal muscular atrophy with progressive myoclonic epilepsy linked to mutations in ASAH1. Clin. Neurol. Neurosurg. 2018, 164, 47–49. [Google Scholar] [CrossRef]
- Huang, Y.; Tanimukai, H.; Liu, F.; Iqbal, K.; Grundke-Iqbal, I.; Gong, C.X. Elevation of the level and activity of acid ceramidase in Alzheimer’s disease brain. Eur. J. Neurosci. 2004, 20, 3489–3497. [Google Scholar] [CrossRef]
- Realini, N.; Palese, F.; Pizzirani, D.; Pontis, S.; Basit, A.; Bach, A.; Ganesan, A.; Piomelli, D. Acid Ceramidase in Melanoma: expression, localization, and effects of pharmacological inhibition. J. Biol. Chem. 2016, 291. [Google Scholar] [CrossRef]
- Mehta, S.; Blackinton, D.; Omar, I.; Kouttab, N.; Myrick, D.; Klostergaard, J.; Wanebo, H. Combined cytotoxic action of paclitaxel and ceramide against the human Tu138 head and neck squamous carcinoma cell line. Cancer Chemother. Pharmacol. 2000, 46, 85–92. [Google Scholar] [CrossRef]
- Liu, X.; Cheng, J.C.; Turner, L.S.; Elojeimy, S.; Beckham, T.H.; Bielawska, A.; Keane, T.E.; Hannun, Y.A.; Norris, J.S. Acid ceramidase upregulation in prostate cancer: Role in tumor development and implications for therapy. Expert Opin. Targets 2009, 13, 1449–1458. [Google Scholar] [CrossRef]
- Chavez, J.A.; Holland, W.L.; Bar, J.; Sandhoff, K.; Summers, S.A. Acid ceramidase overexpression prevents the inhibitory effects of saturated fatty acids on insulin signaling. J. Biol. Chem. 2005, 280, 20148–20153. [Google Scholar] [CrossRef]
- El Bawab, S.; Roddy, P.; Qian, T.; Bielawska, A.; Lemasters, J.J.; Hannun, Y.A. Molecular cloning and characterization of a human mitochondrial ceramidase. J. Biol. Chem. 2000, 275, 21508–21513. [Google Scholar] [CrossRef] [PubMed]
- Hwang, Y.H.; Tani, M.; Nakagawa, T.; Okino, N.; Ito, M. Subcellular localization of human neutral ceramidase expressed in HEK293 cells. Biochem. Biophys Res. Commun. 2005, 331, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Kono, M.; Dreier, J.L.; Ellis, J.M.; Allende, M.L.; Kalkofen, D.N.; Sanders, K.M.; Bielawski, J.; Bielawska, A.; Hannun, Y.A.; Proia, R.L. Neutral ceramidase encoded by the Asah2 gene is essential for the intestinal degradation of sphingolipids. J. Biol. Chem. 2006, 281, 7324–7331. [Google Scholar] [CrossRef] [PubMed]
- Tani, M.; Iida, H.; Ito, M. O-glycosylation of mucin-like domain retains the neutral ceramidase on the plasma membranes as a type II integral membrane protein. J. Biol. Chem. 2003, 278, 10523–10530. [Google Scholar] [CrossRef]
- Galadari, S.; Wu, B.X.; Mao, C.; Roddy, P.; El Bawab, S.; Hannun, Y.A. Identification of a novel amidase motif in neutral ceramidase. Biochem. J. 2006, 393, 687–695. [Google Scholar] [CrossRef]
- Liakath-Ali, K.; Vancollie, V.E.; Lelliott, C.J.; Speak, A.O.; Lafont, D.; Protheroe, H.J.; Ingvorsen, C.; Galli, A.; Green, A.; Gleeson, D.; et al. Alkaline ceramidase 1 is essential for mammalian skin homeostasis and regulating whole-body energy expenditure. J. Pathol. 2016, 239, 374–383. [Google Scholar] [CrossRef]
- Sun, W.; Xu, R.; Hu, W.; Jin, J.; Crellin, H.A.; Bielawski, J.; Szulc, Z.M.; Thiers, B.H.; Obeid, L.M.; Mao, C. Upregulation of the human alkaline ceramidase 1 and acid ceramidase mediates calcium-induced differentiation of epidermal keratinocytes. J. Investig. Dermatol. 2008, 128, 389–397. [Google Scholar] [CrossRef]
- El Bawab, S.; Bielawska, A.; Hannun, Y.A. Purification and characterization of a membrane-bound nonlysosomal ceramidase from rat brain. J. Biol. Chem. 1999, 274, 27948–27955. [Google Scholar] [CrossRef]
- Xu, R.; Jin, J.; Hu, W.; Sun, W.; Bielawski, J.; Szulc, Z.; Taha, T.; Obeid, L.M.; Mao, C. Golgi alkaline ceramidase regulates cell proliferation and survival by controlling levels of sphingosine and S1P. Faseb J. 2006, 20, 1813–1825. [Google Scholar] [CrossRef]
- Sun, W.; Jin, J.; Xu, R.; Hu, W.; Szulc, Z.M.; Bielawski, J.; Obeid, L.M.; Mao, C. Substrate specificity, membrane topology, and activity regulation of human alkaline ceramidase 2 (ACER2). J. Biol. Chem. 2010, 285, 8995–9007. [Google Scholar] [CrossRef]
- Xu, R.; Wang, K.; Mileva, I.; Hannun, Y.A.; Obeid, L.M.; Mao, C. Alkaline ceramidase 2 and its bioactive product sphingosine are novel regulators of the DNA damage response. Oncotarget 2016, 7, 18440–18457. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhang, C.; Jin, Y.; He, Q.; Liu, Z.; Ai, Q.; Lei, Y.; Li, Y.; Song, F.; Bu, Y. Alkaline ceramidase 2 is a novel direct target of p53 and induces autophagy and apoptosis through ROS generation. Sci. Rep. 2017, 7, 44573. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Garcia-Barros, M.; Wen, S.; Li, F.; Lin, C.L.; Hannun, Y.A.; Obeid, L.M.; Mao, C. Tumor suppressor p53 links ceramide metabolism to DNA damage response through alkaline ceramidase 2. Cell Death Differ. 2018, 25, 841–856. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhang, Y.; Wang, P.; Zhang, S.Y.; Dong, Y.; Zeng, G.; Yan, Y.; Sun, L.; Wu, Q.; Liu, H.; et al. Adipocyte Hypoxia-Inducible Factor 2alpha Suppresses Atherosclerosis by Promoting Adipose Ceramide Catabolism. Cell Metab. 2019, 30, 937–951. [Google Scholar] [CrossRef]
- Mao, C.; Xu, R.; Szulc, Z.M.; Bielawska, A.; Galadari, S.H.; Obeid, L.M. Cloning and characterization of a novel human alkaline ceramidase. A mammalian enzyme that hydrolyzes phytoceramide. J. Biol. Chem. 2001, 276, 26577–26588. [Google Scholar] [CrossRef]
- Hu, W.; Xu, R.; Sun, W.; Szulc, Z.M.; Bielawski, J.; Obeid, L.M.; Mao, C. Alkaline ceramidase 3 (ACER3) hydrolyzes unsaturated long-chain ceramides, and its down-regulation inhibits both cell proliferation and apoptosis. J. Biol. Chem. 2010, 285, 7964–7976. [Google Scholar] [CrossRef]
- Wang, K.; Xu, R.; Schrandt, J.; Shah, P.; Gong, Y.Z.; Preston, C.; Wang, L.; Yi, J.K.; Lin, C.L.; Sun, W.; et al. Alkaline Ceramidase 3 Deficiency Results in Purkinje Cell Degeneration and Cerebellar Ataxia Due to Dyshomeostasis of Sphingolipids in the Brain. PLoS Genet. 2015, 11, e1005591. [Google Scholar] [CrossRef]
- Li, C.M.; Park, J.H.; He, X.; Levy, B.; Chen, F.; Arai, K.; Adler, D.A.; Disteche, C.M.; Koch, J.; Sandhoff, K. The human acid ceramidase gene (ASAH): Structure, chromosomal location, mutation analysis, and expression. Genomics 1999, 62, 223–231. [Google Scholar] [CrossRef]
- Kernohan, K.D.; Fresard, L.; Zappala, Z.; Hartley, T.; Smith, K.S.; Wagner, J.; Xu, H.; McBride, A.; Bourque, P.R.; Consortium, C.R.C.; et al. Whole-transcriptome sequencing in blood provides a diagnosis of spinal muscular atrophy with progressive myoclonic epilepsy. Hum. Mutat. 2017, 38, 611–614. [Google Scholar] [CrossRef]
- Avramopoulos, D.; Wang, R.; Valle, D.; Fallin, M.D.; Bassett, S.S. A novel gene derived from a segmental duplication shows perturbed expression in Alzheimer’s disease. Neurogenetics 2007, 8, 111–120. [Google Scholar] [CrossRef]
- Novgorodov, S.A.; Riley, C.L.; Yu, J.; Borg, K.T.; Hannun, Y.A.; Proia, R.L.; Kindy, M.S.; Gudz, T.I. Essential roles of neutral ceramidase and sphingosine in mitochondrial dysfunction due to traumatic brain injury. J. Biol. Chem. 2014, 289, 13142–13154. [Google Scholar] [CrossRef] [PubMed]
- Coant, N.; Garcia-Barros, M.; Zhang, Q.; Obeid, L.M.; Hannun, Y.A. AKT as a key target for growth promoting functions of neutral ceramidase in colon cancer cells. Oncogene 2018. [Google Scholar] [CrossRef] [PubMed]
- Snider, A.J.; Wu, B.X.; Jenkins, R.W.; Sticca, J.A.; Kawamori, T.; Hannun, Y.A.; Obeid, L.M. Loss of neutral ceramidase increases inflammation in a mouse model of inflammatory bowel disease. Prostaglandins Other Lipid Mediat. 2012, 99, 124–130. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, W.; Coant, N.; Canals, D.; Obeid, L.M.; Hannun, Y.A. Functions of neutral ceramidase in the Golgi apparatus. J. Lipid Res. 2018, 59, 2116–2125. [Google Scholar] [CrossRef] [PubMed]
- Van der Knaap, M.S.; Bugiani, M. Leukodystrophies: A proposed classification system based on pathological changes and pathogenetic mechanisms. Acta Neuropathol. 2017, 134, 351–382. [Google Scholar] [CrossRef]
- Edvardson, S.; Yi, J.K.; Jalas, C.; Xu, R.; Webb, B.D.; Snider, J.; Fedick, A.; Kleinman, E.; Treff, N.R.; Mao, C. Deficiency of the alkaline ceramidase ACER3 manifests in early childhood by progressive leukodystrophy. J. Med. Genet. 2016, 53, 389–396. [Google Scholar] [CrossRef]
- Tani, M.; Okino, N.; Mori, K.; Tanigawa, T.; Izu, H.; Ito, M. Molecular cloning of the full-length cDNA encoding mouse neutral ceramidase. A novel but highly conserved gene family of neutral/alkaline ceramidases. J. Biol. Chem. 2000, 275, 11229–11234. [Google Scholar] [CrossRef]
- Mitsutake, S.; Tani, M.; Okino, N.; Mori, K.; Ichinose, S.; Omori, A.; Iida, H.; Nakamura, T.; Ito, M. Purification, characterization, molecular cloning, and subcellular distribution of neutral ceramidase of rat kidney. J. Biol. Chem. 2001, 276, 26249–26259. [Google Scholar] [CrossRef]
- Duan, R.D.; Cheng, Y.; Yang, L.; Ohlsson, L.; Nilsson, A. Evidence for specific ceramidase present in the intestinal contents of rats and humans. Lipids 2001, 36, 807–812. [Google Scholar] [CrossRef]
- Ohlsson, L.; Palmberg, C.; Duan, R.D.; Olsson, M.; Bergman, T.; Nilsson, A. Purification and characterization of human intestinal neutral ceramidase. Biochimie 2007, 89, 950–960. [Google Scholar] [CrossRef]
- Tani, M.; Okino, N.; Sueyoshi, N.; Ito, M. Conserved amino acid residues in the COOH-terminal tail are indispensable for the correct folding and localization and enzyme activity of neutral ceramidase. J. Biol. Chem. 2004, 279, 29351–29358. [Google Scholar] [CrossRef] [PubMed]
- Maltesen, H.R.; Troelsen, J.T.; Olsen, J. Identification of a functional hepatocyte nuclear factor 4 binding site in the neutral ceramidase promoter. J. Cell Biochem. 2010, 111, 1330–1336. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, S.M.; Houck, K.L.; Yun, J.K.; Fox, T.E.; Kester, M. AP-1 binding transcriptionally regulates human neutral ceramidase. Arch. Biochem. Biophys. 2011, 511, 31–39. [Google Scholar] [CrossRef] [PubMed]
- O’Neill, S.M.; Yun, J.K.; Fox, T.E.; Kester, M. Transcriptional regulation of the human neutral ceramidase gene. Arch. Biochem. Biophys. 2011, 511, 21–30. [Google Scholar] [CrossRef]
- Airola, M.V.; Allen, W.J.; Pulkoski-Gross, M.J.; Obeid, L.M.; Rizzo, R.C.; Hannun, Y.A. Structural Basis for Ceramide Recognition and Hydrolysis by Human Neutral Ceramidase. Structure 2015, 23, 1482–1491. [Google Scholar] [CrossRef]
- Malinina, L.; Brown, R.E. Catalytic Mechanism of Eukaryotic Neutral Ceramidase. Structure 2015, 23, 1371–1372. [Google Scholar] [CrossRef][Green Version]
- Bar, J.; Linke, T.; Ferlinz, K.; Neumann, U.; Schuchman, E.H.; Sandhoff, K. Molecular analysis of acid ceramidase deficiency in patients with Farber disease. Hum. Mutat. 2001, 17, 199–209. [Google Scholar] [CrossRef]
- Levade, T.; Moser, H.W.; Fensom, A.H.; Harzer, K.; Moser, A.B.; Salvayre, R. Neurodegenerative course in ceramidase deficiency (Farber disease) correlates with the residual lysosomal ceramide turnover in cultured living patient cells. J. Neurol. Sci. 1995, 134, 108–114. [Google Scholar] [CrossRef]
- Pellissier, J.F.; Berard-Badier, M.; Pinsard, N. Farber’s disease in two siblings, sural nerve and subcutaneous biopsies by light and electron microscopy. Acta Neuropathol. 1986, 72, 178–188. [Google Scholar] [CrossRef]
- Haraoka, G.; Muraoka, M.; Yoshioka, N.; Wakami, S.; Hayashi, I. First case of surgical treatment of Farber’s disease. Ann. Plast. Surg. 1997, 39, 405–410. [Google Scholar] [CrossRef]
- Li, C.M.; Park, J.H.; Simonaro, C.M.; He, X.; Gordon, R.E.; Friedman, A.H.; Ehleiter, D.; Paris, F.; Manova, K.; Hepbildikler, S.; et al. Insertional mutagenesis of the mouse acid ceramidase gene leads to early embryonic lethality in homozygotes and progressive lipid storage disease in heterozygotes. Genomics 2002, 79, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Bi, F.C.; Yin, J.; Wu, J.X.; Rong, C.; Wu, J.L.; Yao, N. An Arabidopsis neutral ceramidase mutant ncer1 accumulates hydroxyceramides and is sensitive to oxidative stress. Front. Plant Sci. 2015, 6, 460. [Google Scholar] [CrossRef] [PubMed]
- Vasiliauskaite-Brooks, I.; Healey, R.D.; Rochaix, P.; Saint-Paul, J.; Sounier, R.; Grison, C.; Waltrich-Augusto, T.; Fortier, M.; Hoh, F.; Saied, E.M.; et al. Structure of a human intramembrane ceramidase explains enzymatic dysfunction found in leukodystrophy. Nat. Commun. 2018, 9, 5437. [Google Scholar] [CrossRef] [PubMed]
- Ghareeb, D.A.; Hafez, H.S.; Hussien, H.M.; Kabapy, N.F. Non-alcoholic fatty liver induces insulin resistance and metabolic disorders with development of brain damage and dysfunction. Metab. Brain Dis. 2011, 26, 253–267. [Google Scholar] [CrossRef]
- Tojima, H.; Kakizaki, S.; Kosone, T.; Horiguchi, N.; Yamazaki, Y.; Sato, K.; Takagi, H.; Mori, M. Hepatocyte growth factor overexpression ameliorates liver inflammation and fibrosis in a mouse model of nonalcoholic steatohepatitis. Hepatol. Int. 2012, 6, 620–630. [Google Scholar] [CrossRef]
- Daugherity, E.K.; Balmus, G.; Al Saei, A.; Moore, E.S.; Abi Abdallah, D.; Rogers, A.B.; Weiss, R.S.; Maurer, K.J. The DNA damage checkpoint protein ATM promotes hepatocellular apoptosis and fibrosis in a mouse model of non-alcoholic fatty liver disease. Cell Cycle 2012, 11, 1918–1928. [Google Scholar] [CrossRef]
- Chavez, J.A.; Summers, S.A. Lipid oversupply, selective insulin resistance, and lipotoxicity: Molecular mechanisms. Biochim. Et Biophys. Acta 2010, 1801, 252–265. [Google Scholar] [CrossRef]
- Matsuzawa, N.; Takamura, T.; Kurita, S.; Misu, H.; Ota, T.; Ando, H.; Yokoyama, M.; Honda, M.; Zen, Y.; Nakanuma, Y.; et al. Lipid-induced oxidative stress causes steatohepatitis in mice fed an atherogenic diet. Hepatology 2007, 46, 1392–1403. [Google Scholar] [CrossRef]
- Nakamura, S.; Takamura, T.; Matsuzawa-Nagata, N.; Takayama, H.; Misu, H.; Noda, H.; Nabemoto, S.; Kurita, S.; Ota, T.; Ando, H.; et al. Palmitate induces insulin resistance in H4IIEC3 hepatocytes through reactive oxygen species produced by mitochondria. J. Biol. Chem. 2009, 284, 14809–14818. [Google Scholar] [CrossRef]
- Summers, S.A. Could Ceramides Become the New Cholesterol? Cell Metab. 2018, 27, 276–280. [Google Scholar] [CrossRef]
- Summers, S.A. Ceramides in insulin resistance and lipotoxicity. Prog. Lipid Res. 2006, 45, 42–72. [Google Scholar] [CrossRef] [PubMed]
- Sokolowska, E.; Blachnio-Zabielska, A. The Role of Ceramides in Insulin Resistance. Front Endocrinol. (Lausanne) 2019, 10, 577. [Google Scholar] [CrossRef] [PubMed]
- Bandet, C.L.; Tan-Chen, S.; Bourron, O.; Le Stunff, H.; Hajduch, E. Sphingolipid Metabolism: New Insight into Ceramide-Induced Lipotoxicity in Muscle Cells. Int. J. Mol. Sci. 2019, 20. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.; Zhu, R.; Jin, J. Neutral ceramidase-enriched exosomes prevent palmitic acid-induced insulin resistance in H4IIEC3 hepatocytes. Febs Open Bio 2016, 6, 1078–1084. [Google Scholar] [CrossRef] [PubMed]
- Tang, S.; Luo, F.; Feng, Y.M.; Wei, X.; Miao, H.; Lu, Y.B.; Tang, Y.; Ding, D.F.; Jin, J.F.; Zhu, Q. Neutral Ceramidase Secreted Via Exosome Protects Against Palmitate-Induced Apoptosis in INS-1 Cells. Exp. Clin. Endocrinol. Diabetes 2017, 125, 130–135. [Google Scholar] [CrossRef] [PubMed]
- Chavez, J.A.; Summers, S.A. Characterizing the effects of saturated fatty acids on insulin signaling and ceramide and diacylglycerol accumulation in 3T3-L1 adipocytes and C2C12 myotubes. Arch. Biochem. Biophys. 2003, 419, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, X.; Lau, W.B.; Yuan, Y.; Booth, D.; Li, J.J.; Scalia, R.; Preston, K.; Gao, E.; Koch, W. Adiponectin inhibits tumor necrosis factor-alpha-induced vascular inflammatory response via caveolin-mediated ceramidase recruitment and activation. Circ. Res. 2014, 114, 792–805. [Google Scholar] [CrossRef]
- He, X.; Huang, Y.; Li, B.; Gong, C.X.; Schuchman, E.H. Deregulation of sphingolipid metabolism in Alzheimer’s disease. Neurobiol. Aging 2010, 31, 398–408. [Google Scholar] [CrossRef]
- Yamauchi, T.; Kamon, J.; Waki, H.; Terauchi, Y.; Kubota, N.; Hara, K.; Mori, Y.; Ide, T.; Murakami, K.; Tsuboyama-Kasaoka, N.; et al. The fat-derived hormone adiponectin reverses insulin resistance associated with both lipoatrophy and obesity. Nat. Med. 2001, 7, 941–946. [Google Scholar] [CrossRef]
- Achari, A.E.; Jain, S.K. Adiponectin, a Therapeutic Target for Obesity, Diabetes, and Endothelial Dysfunction. Int. J. Mol. Sci. 2017, 18, 1321. [Google Scholar]
- Maruyoshi, H.; Kojima, S.; Otsuka, F.; Funahashi, T.; Kaikita, K.; Sugiyama, S.; Sakamoto, T.; Yoshimura, M.; Shimomura, I.; Ogawa, H. Hypoadiponectinemia is associated with coronary artery spasm in men. Circ. J. Off. J. Jpn. Circ. Soc. 2005, 69, 1154–1156. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Goldstein, B.J.; Scalia, R.G.; Ma, X.L. Protective vascular and myocardial effects of adiponectin. Nat. Clin. Pract. Cardiovasc. Med. 2009, 6, 27–35. [Google Scholar] [CrossRef]
- Yamauchi, T.; Kamon, J.; Ito, Y.; Tsuchida, A.; Yokomizo, T.; Kita, S.; Sugiyama, T.; Miyagishi, M.; Hara, K.; Tsunoda, M.; et al. Cloning of adiponectin receptors that mediate antidiabetic metabolic effects. Nature 2003, 423, 762–769. [Google Scholar] [CrossRef] [PubMed]
- Holland, W.L.; Miller, R.A.; Wang, Z.V.; Sun, K.; Barth, B.M.; Bui, H.H.; Davis, K.E.; Bikman, B.T.; Halberg, N.; Rutkowski, J.M.; et al. Receptor-mediated activation of ceramidase activity initiates the pleiotropic actions of adiponectin. Nat. Med. 2011, 17, 55–63. [Google Scholar] [CrossRef] [PubMed]
- Vasiliauskaite-Brooks, I.; Sounier, R.; Rochaix, P.; Bellot, G.; Fortier, M.; Hoh, F.; De Colibus, L.; Bechara, C.; Saied, E.M.; Arenz, C.; et al. Structural insights into adiponectin receptors suggest ceramidase activity. Nature 2017, 544, 120–123. [Google Scholar] [CrossRef] [PubMed]
- Jazvinscak Jembrek, M.; Hof, P.R.; Simic, G. Ceramides in Alzheimer’s Disease: Key Mediators of Neuronal Apoptosis Induced by Oxidative Stress and Abeta Accumulation. Oxidative Med. Cell. Longev. 2015, 2015, 346783. [Google Scholar] [CrossRef]
- Jana, A.; Hogan, E.L.; Pahan, K. Ceramide and neurodegeneration: Susceptibility of neurons and oligodendrocytes to cell damage and death. J. Neurol. Sci. 2009, 278, 5–15. [Google Scholar] [CrossRef]
- Cutler, R.G.; Kelly, J.; Storie, K.; Pedersen, W.A.; Tammara, A.; Hatanpaa, K.; Troncoso, J.C.; Mattson, M.P. Involvement of oxidative stress-induced abnormalities in ceramide and cholesterol metabolism in brain aging and Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 2004, 101, 2070–2075. [Google Scholar] [CrossRef]
- Patil, S.; Melrose, J.; Chan, C. Involvement of astroglial ceramide in palmitic acid-induced Alzheimer-like changes in primary neurons. Eur. J. Neurosci. 2007, 26, 2131–2141. [Google Scholar] [CrossRef]
- Puglielli, L.; Ellis, B.C.; Saunders, A.J.; Kovacs, D.M. Ceramide stabilizes beta-site amyloid precursor protein-cleaving enzyme 1 and promotes amyloid beta-peptide biogenesis. J. Biol. Chem. 2003, 278, 19777–19783. [Google Scholar] [CrossRef]
- Sullivan, P.G.; Keller, J.N.; Mattson, M.P.; Scheff, S.W. Traumatic brain injury alters synaptic homeostasis: Implications for impaired mitochondrial and transport function. J. Neurotrauma 1998, 15, 789–798. [Google Scholar] [CrossRef] [PubMed]
- Bryan, L.; Kordula, T.; Spiegel, S.; Milstien, S. Regulation and functions of sphingosine kinases in the brain. Biochim. Et Biophys. Acta 2008, 1781, 459–466. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Barros, M.; Coant, N.; Kawamori, T.; Wada, M.; Snider, A.J.; Truman, J.P.; Wu, B.X.; Furuya, H.; Clarke, C.J.; Bialkowska, A.B.; et al. Role of neutral ceramidase in colon cancer. Faseb J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2016, 30, 4159–4171. [Google Scholar] [CrossRef] [PubMed]
- Fakhoury, M.; Negrulj, R.; Mooranian, A.; Al-Salami, H. Inflammatory bowel disease: Clinical aspects and treatments. J. Inflamm. Res. 2014, 7, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Bock, J.; Liebisch, G.; Schweimer, J.; Schmitz, G.; Rogler, G. Exogenous sphingomyelinase causes impaired intestinal epithelial barrier function. World J. Gastroenterol. 2007, 13, 5217–5225. [Google Scholar] [CrossRef] [PubMed]
- Liu, T.C.; Stappenbeck, T.S. Genetics and Pathogenesis of Inflammatory Bowel Disease. Annu. Rev. Pathol. 2016, 11, 127–148. [Google Scholar] [CrossRef]
- Axelrad, J.E.; Lichtiger, S.; Yajnik, V. Inflammatory bowel disease and cancer: The role of inflammation, immunosuppression, and cancer treatment. World J. Gastroenterol. 2016, 22, 4794–4801. [Google Scholar] [CrossRef]
- Espaillat, M.P.; Snider, A.J.; Qiu, Z.; Channer, B.; Coant, N.; Schuchman, E.H.; Kew, R.R.; Sheridan, B.S.; Hannun, Y.A.; Obeid, L.M. Loss of acid ceramidase in myeloid cells suppresses intestinal neutrophil recruitment. Faseb J. 2018, 32, 2339–2353. [Google Scholar] [CrossRef]
- Zeidan, Y.H.; Jenkins, R.W.; Korman, J.B.; Liu, X.; Obeid, L.M.; Norris, J.S.; Hannun, Y.A. Molecular targeting of acid ceramidase: Implications to cancer therapy. Curr. Drug Targets 2008, 9, 653–661. [Google Scholar] [CrossRef] [PubMed]
- Pizzirani, D.; Pagliuca, C.; Realini, N.; Branduardi, D.; Bottegoni, G.; Mor, M.; Bertozzi, F.; Scarpelli, R.; Piomelli, D.; Bandiera, T. Discovery of a new class of highly potent inhibitors of acid ceramidase: Synthesis and structure-activity relationship (SAR). J. Med. Chem. 2013, 56, 3518–3530. [Google Scholar] [CrossRef]
- Delgado, A.; Casas, J.; Llebaria, A.; Abad, J.L.; Fabrias, G. Inhibitors of sphingolipid metabolism enzymes. Biochim. Et Biophys. Acta 2006, 1758, 1957–1977. [Google Scholar] [CrossRef] [PubMed]
- Szulc, Z.M.; Mayroo, N.; Bai, A.; Bielawski, J.; Liu, X.; Norris, J.S.; Hannun, Y.A.; Bielawska, A. Novel analogs of D-e-MAPP and B13. Part 1: Synthesis and evaluation as potential anticancer agents. Bioorganic Med. Chem. 2008, 16, 1015–1031. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bhabak, K.P.; Kleuser, B.; Huwiler, A.; Arenz, C. Effective inhibition of acid and neutral ceramidases by novel B-13 and LCL-464 analogues. Bioorganic Med. Chem. 2013, 21, 874–882. [Google Scholar] [CrossRef] [PubMed]
- Abuhusain, H.J.; Matin, A.; Qiao, Q.; Shen, H.; Kain, N.; Day, B.W.; Stringer, B.W.; Daniels, B.; Laaksonen, M.A.; Teo, C.; et al. A metabolic shift favoring sphingosine 1-phosphate at the expense of ceramide controls glioblastoma angiogenesis. J. Biol. Chem. 2013, 288, 37355–37364. [Google Scholar] [CrossRef] [PubMed]
- Saad, A.F.; Meacham, W.D.; Bai, A.; Anelli, V.; Elojeimy, S.; Mahdy, A.E.; Turner, L.S.; Cheng, J.; Bielawska, A.; Bielawski, J.; et al. The functional effects of acid ceramidase overexpression in prostate cancer progression and resistance to chemotherapy. Cancer Biol. Ther. 2007, 6, 1455–1460. [Google Scholar] [CrossRef]
- Doan, N.B.; Alhajala, H.; Al-Gizawiy, M.M.; Mueller, W.M.; Rand, S.D.; Connelly, J.M.; Cochran, E.J.; Chitambar, C.R.; Clark, P.; Kuo, J.; et al. Acid ceramidase and its inhibitors: A de novo drug target and a new class of drugs for killing glioblastoma cancer stem cells with high efficiency. Oncotarget 2017, 8, 112662–112674. [Google Scholar] [CrossRef]
- Sato, S.; Ueyama, T.; Fukui, H.; Miyazaki, K.; Kuwano, M. Anti-tumor effects of carmofur on human 5-FU resistant cells. Gan Kagaku Ryoho. Cancer Chemother. 1999, 26, 1613–1616. [Google Scholar]
| Ceramidase | Encoded by Gene | Optimal pH | Expression | Subcellular Localization | Associated Diseases |
|---|---|---|---|---|---|
| Acid ceramidase | ASAH1, located at p22 of chromosome 8 | ∼4.0–4.5 | Heart, kidneys, lungs, placenta, etc. [48] | Lysosomes [48] | Farber’s disease, Alzheimer’s disease, cancer, diabetes, and spinal muscular atrophy [11,25,26,27,28,30,49] |
| Neutral ceramidase | ASAH2, located at q11.23 of chromosome 10 | ∼7.0–7.4 | Small intestine [33] | Mitochondria (HEK293T overexpressing cells) [31] | Alzheimer’s disease [50], various metabolic diseases [51,52,53] |
| Plasma membrane (HEK293T overexpressing cells) [32,34] | |||||
| Mitochondria and Golgi (HCT116 overexpressing cells) [54] | |||||
| Alkaline ceramidase | ACER1, located at p13.3 of chromosome 19 | ∼9.0 | Skin | ER | Progressive leukodystrophy [55,56] |
| ACER2, located at p22.1 of chromosome 9 | Placental tissue | Golgi complex [39,40] | |||
| ACER3, located at q13.5 of chromosome 11 | Ubiquitous, highly expressed in placental tissue | ER and Golgi complex [45,46] |
| Name | pH | Temp. | KM (µM) | Vmax (nmol/min mg) | Substrate | Note | References |
|---|---|---|---|---|---|---|---|
| Acid ceramidase | 4.5 | 37 | 389 | 462.97 | N-lauroylsphingosine | pH 4.5, 37 °C, recombinant ceramidase expressed in CHO cells, 14C-labeled substrate | [23] |
| 413.2 | 33.33 | pH 4.5, 37 °C, recombinant ceramidase expressed in CHO cells, BODIPY-conjugated substrate | |||||
| Neutral ceramidase | 7.5 | 37 | 60.1 33.41 | 0.68 N/A | D-erythro-C12-4nitrobenzo-2-oxa-1,3-diazole-ceramide NBD-C12-ceramide | pH 7.5, 37 °C, recombinant ceramidase 28 °C, recombinant enzyme (mutant, residues 99–780) | [35] [57] |
| Alkaline ceramidase (ACER2)* | 9.0 | 37 | 98.5 94.8 | 0.0237 0.0261 | C16:0-ceramide C18:0-ceramide | pH 9.0, 37 °C | [40] |
| Disease | ASAH2 Expression | Ceramide Level | S1P Level | Tissue/Cells | References |
|---|---|---|---|---|---|
| Insulin resistance | Decreased | Increased | Not known | INS-1 pancreatic β cells, H4IIEC3 hepatocytes | [84,85] |
| Cardiovascular diseases | Activity decreased | Not known | Not Known | Human umbilical vein endothelial cells (HUVECs) | [87] |
| Alzheimer’s disease | Not known | Ceramide accumulation | Decreased | Human brain | [88] |
| Traumatic brain injury | Increased activity | Decreased | No difference | Mouse brain, mitochondria dysfunction | [51] |
| Cancer | Increased expression | Not known | Not known | CaCo2BBe, HCT116 colon cancer cells | [52] |
| Inflammatory bowel diseases | Decreased activity | Increased in colon epithelium | Increased upon ASAH2 knockout | Gut from DSS-treated C57BL/6 mice | [53] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Parveen, F.; Bender, D.; Law, S.-H.; Mishra, V.K.; Chen, C.-C.; Ke, L.-Y. Role of Ceramidases in Sphingolipid Metabolism and Human Diseases. Cells 2019, 8, 1573. https://doi.org/10.3390/cells8121573
Parveen F, Bender D, Law S-H, Mishra VK, Chen C-C, Ke L-Y. Role of Ceramidases in Sphingolipid Metabolism and Human Diseases. Cells. 2019; 8(12):1573. https://doi.org/10.3390/cells8121573
Chicago/Turabian StyleParveen, Farzana, Daniel Bender, Shi-Hui Law, Vineet Kumar Mishra, Chih-Chieh Chen, and Liang-Yin Ke. 2019. "Role of Ceramidases in Sphingolipid Metabolism and Human Diseases" Cells 8, no. 12: 1573. https://doi.org/10.3390/cells8121573
APA StyleParveen, F., Bender, D., Law, S.-H., Mishra, V. K., Chen, C.-C., & Ke, L.-Y. (2019). Role of Ceramidases in Sphingolipid Metabolism and Human Diseases. Cells, 8(12), 1573. https://doi.org/10.3390/cells8121573






