1. Introduction
Various acquired and inherited diseases of the lung, liver, intestine, and pancreas are heavy burdens for the affected patients. A better understanding of normal and disease processes and cellular pathomechanisms of functional cells might provide possible new targets for medical interventions. In addition, patients could benefit from cell replacement therapies. It has been estimated that more than 10
9 pancreatic islet cells are needed for effective treatment of type I diabetes mellitus [
1]. For liver-based metabolic diseases and acute liver failure, most clinics use up to 2 × 10
8 hepatocyte cells/kg to treat patients [
2]. As these required cell numbers can scarcely be generated from autologous, primary cells, alternative cell sources for clinical applications are of great interest. Human pluripotent stem cells (hPSCs) have great potential as a universal therapeutic cell source for different diseases because they can be grown indefinitely while maintaining their potential to differentiate into the three germ layers [
3]. In addition, they have been shown to be useful tools for basic research, in vitro disease modelling and high throughput drug screening assays [
4,
5]. In one example, 1.3 × 10
9 differentiated cystic fibrosis patient-specific iPSCs were used to screen 42,500 compounds for modulators of CFTR activity [
5].
As both the routine use of hPSC-based cell therapies and organotypic in vitro cell applications become more important [
6], new systems are needed to reproducibly deliver large quantities of cells that meet the required standards of “good manufacturing practice” (GMP) [
7,
8]. Currently, many laboratories are still performing adherent 2D cell culture for expansion and differentiation of hPSCs. However, when it comes to large scale cell production in 2D, there are two major limitations: these systems are restricted to culture surface area and require a lot of manual work. Culturing cells in dynamic 3D suspension using platforms such as a bioreactor can overcome these restraints, as such systems have a high surface area-to-volume ratio. We and others have already shown that undifferentiated hPSCs can be maintained and expanded in suspension cultures up to clinically relevant cell numbers [
1,
7,
9] and that further differentiation to cardiomyocytes [
10] and endothelial cells [
11] is applicable in this format. From a technical point of view, bioreactors allow linear upscaling processes with multi-parametric monitoring possibilities (pH, metabolics, feeding, etc.) combined with versatile options for process intervention and optimization, and for the generation of homogeneous production batches. The combination with an efficient cryopreservation protocol would further advance the process for off-the-shelf allocation of definitive endoderm (DE) as starting material for production of different DE lineages.
It has been previously shown that DE differentiation in suspension is feasible [
12,
13,
14,
15,
16,
17,
18]. According to Yabe et al., differentiation into DE in suspension may be superior to adherent culture differentiation, since DE markers appear faster and are more strongly expressed [
12]. Despite promising prospects to use suspension cultures for DE differentiation, most of these protocols do not use chemically-defined, xeno-free media. In addition, the capability of these aggregates to differentiate into most lineages of the DE has not yet been demonstrated. Considering the technical and biological requirements of industrial scale cell production, we established a straightforward protocol for chemically-defined, xeno-free generation of DE from hiPSCs and hESCs. We tested two different methods for DE generation to see if both conditions could give rise to viable DE. We first used Erlenmeyer flasks for proof-of principle purposes, and then successfully scaled up to a controlled, stirred bioreactor system. Finally, we showed that hPSC-derived DE cells maintain their multi-lineage differentiation potential after cryopreservation and thawing. This work is a valuable step towards efficient disease modeling, drug discovery, and clinical scale up of therapeutic DE cell products.
2. Materials and Methods
2.1. Stem Cell Culture
The hESC line, HES3, and the hiPSC cell line, MHHi001-A [
19], generated from CD34+ cells of a healthy patient using Sendai virus, were used for all experiments. The third cell line (MHHi006-A) was generated from a healthy patient using lenti-viral based reprogramming. Undifferentiated hPSCs were cultivated on Geltrex™ (Gibco #A1413202) using E8 medium (in house) for up to 12 passages. The cells were passaged with StemPro
® Accutase
® (Gibco #A1110501) twice a week.
2.2. DE Differentiation Using CHIR99021 and Activin A (CA)
At day 1, a 90% confluent flask of hPSCs was dissociated with StemPro® Accutase® (Gibco #A1110501), and inoculated in 125 mL Erlenmeyer flasks (Corning #431143) containing 20 mL of E8 medium (in house) with 10 µM Y-27632 (Tocris #1254) at a cell density of 3.33 × 105 cells/mL for hiPSCs and 6.67 × 105 cells/mL for hESCs. Flasks were placed in an incubator on an orbital shaker (Celltron, Infors HT) rotating at 70 rotations per minute (rpm). Medium was changed every 24 h (h). For each medium change, the aggregates were collected and centrifuged (Multifuge 3SR+, Thermo) at 300 g for 2 min before placing them in fresh medium. The following day (day 0), the media was changed to RPMI 1640 (Gibco #21875034), 100 ng/mL activin A (PeproTech #120-14E), and 3 µM CHIR99021 (provided by the Institute of Organic Chemistry, Leibniz University, Hannover, Germany). On day 1, the medium was changed to 100 ng/mL activin A, and 0.8% Knockout™ Serum Replacement (KSR; Gibco #10828028) in RPMI 1640. On day 2, the medium was changed to 100 ng/mL activin A, 8% KSR in RPMI 1640. On day 3 of differentiation, aggregates were dissociated with Accutase for 3 min in a water bath at 37 °C, analyzed for DE markers, and further differentiated into liver, pancreatic, intestinal, and lung progenitor cells. Dissociated day 3 cells were also frozen in CryoStor® CS10 Freezing Media (BioLife Solutions #210102).
2.3. DE Differentiation Using STEMDiff™ Definitive Endoderm Kit (SD)
Six days before the start of differentiation, pluripotent stem cells were passaged in flasks at a low density, 1.2 × 104 cells/cm2, using E8 medium and 10 µM Y-27632 (Tocris #1254). Medium was changed every 24 h. At day 3, medium was changed to E8 medium and Supplement 1, supplied in the SD kit (STEMCELL Tech. #05115). On day 1 of differentiation, cells were dissociated with StemPro® Accutase® (Gibco #A1110501) and inoculated in shaker flasks, at a cell density of 3.33 × 105 cells/mL for hiPSCs and 6.67 × 105 cells/mL for hESCs, in E8 medium, E8 supplement (STEMCELL Tech. #05116), and 10 µM Y-27632. For each medium change, the aggregates were collected and centrifuged (Multifuge 3SR+, Thermo) at 300× g for 2 min before placing them in fresh medium. On day 0, medium was changed to STEMDiff™ Endoderm Basal Media containing Supplement MR and CJ. On day 1 and day 2, aggregates were fed with STEMDiff™ Endoderm Basal Media containing Supplement CJ only. On day 3, aggregates were dissociated and analyzed for DE markers, and also further differentiated into liver, pancreatic, intestinal, and lung progenitor cells. Dissociated cells were also frozen in CryoStor® CS10 Freezing Media (BioLife Solutions #210102) at 6 × 106 cells/vial.
2.4. Differentiation into the Hepatic Lineage
For hepatic differentiation, aggregates on day 3 of DE differentiation were adapted to hepatic differentiation media [
20]. In short, the medium was changed to hepatocyte culture medium (Lonza #CC-3198) with 30 ng/mL of fibroblast growth factor 4 (FGF-4, Peprotech #100-31), 20 ng/mL of bone morphogenetic protein 2 (BMP-2, Peprotech #120-02), and 10 µM SB431542 (Sigma Aldrich #S4317), 0.5 µg/mL of secreted frizzled-related protein 5 (sFRP-5, R&D Systems #6266-SF) for 24 h in Erlenmeyer flasks rotating at 70 rpm. Aggregates were then dissociated into single cells using TrypLE (Thermo Fisher #12604013) and plated on Matrigel
® (Corning #356231) coated plates with a density of 45,000 cells/cm
2 in hepatic differentiation media containing 10 µM Y-27632 (Tocris #1254). The cells were cultured for three more days with daily medium changes. On day 5 of differentiation, the medium was changed to hepatocyte culture medium supplemented with 20 ng/mL hepatocyte growth factor (HGF, Peprotech #100-39) for a further four days with daily medium change. On day 9 of differentiation, the medium was changed to hepatocyte culture medium (Lonza) with 20 ng/mL HGF (Peprotech #100-39), 10 ng/mL Oncostatin M (OSM; Peprotech #300-10) and 10 ng/mL dexamethasone (Sigma Aldrich #D4902) for an additional four days. Cells were analyzed on day 14 of differentiation.
2.5. Differentiation into the Pancreatic Lineage
For differentiation into pancreatic PDX1+ cells [
21], aggregates were dissociated using Accutase (Capricorn #ACC-1B), counted, and seeded on plates coated with Matrigel
® (Corning #354277)at a density of 2.6 × 10
5 cells/cm
2 in Advanced RPMI 1640 medium (Gibco #12-633-012) supplemented with 1 µM all-trans retinoic acid (Sigma Aldrich #302-79-4), 0.5 µM LDN 193,189 (Selleckchem #DM-3189), 2 µM IWR-1 (Selleckchem #S7086), 5 ng/mL FGF7 (Reliatech #100-163-L), 0.5× B27 (Gibco #17-504-044), 1% L-glutamine (Sigma Aldrich #G7513), and 1% penicillin/streptomycin (Santa Cruz #sc-391048, Sigma Aldrich # S9137). 10 µM Y-27632 (Selleckchem #S1049) was added for the first 24 h. Differentiation was performed in 12-well plates and 4-well slides (SPL Life Sciences) for immunofluorescent (IF) staining. The medium was changed daily for an additional 7 days (day 10), and then harvested for qRT-PCR analysis or fixed for IF staining.
2.6. Differentiation into the Intestinal Lineage
A previously established protocol [
22] was adapted where WNT3A was substituted with CHIR99021. On day 3 of DE differentiation, aggregates were dissociated with Accutase (Gibco #A1110501) and plated down at 2 × 10
5 cells/cm
2 in intestinal medium: DMEM/F12 (Gibco #11330032), 2% fetal bovine serum (PAA #A11-101), 500 ng/mL FGF4 (PeproTech #100-31), 3 µM CHIR99021 (provided by the Institute of Organic Chemistry, Leibniz University, Hannover, Germany), and 1% penicillin/streptomycin (Thermo Fisher #15140122). Medium was changed every other day until day 7, when cells were analyzed.
2.7. Differentiation into Lung Progenitor Cells
On day 3 of DE differentiation, aggregates were dissociated with Accutase (Gibco #A1110501) and plated at 1 × 105 cells/cm2 for hiPSC-derived SD condition, 2.0 × 105 cells/cm2 for hESC-derived SD condition, 2.0 × 105 cells/cm2 for hiPSC-derived CA, and 2.57 × 105 cells/cm2 for hESC-derived CA condition, in anterior foregut medium 1 (AFE1): lung basal medium (LBM; Knockout™ DMEM (Gibco #12660012), 5% KSR (Gibco #10828028), 1% l-glutamine (Gibco #25030081), 1% non-essential amino acids (Gibco #11140035), 1% penicillin/streptomycin (Gibco #25030081), 0.46 mM 1-thioglycerol (Sigma-Aldrich #M6145)), 3 µM dorsomorphin (Sigma-Aldrich #P5499), and 10 µM SB435142 (provided by the Institute of Organic Chemistry, Leibniz University, Hannover, Germany) with 10 µM Y-27632 (Tocris #1254). On day 5, medium was changed to anterior foregut medium 2 (AFE2): LBM containing 2 µM IWP-2 (Tocris #3533) and 10 µM SB435142. On day 7, cells in the SD condition were fed with B4CF10 medium: LBM containing 10 ng/mL BMP-4 (R&D Systems #314-BP), 3 µM CHIR99021 (provided by the Institute of Organic Chemistry, Leibniz University, Hannover, Germany), and 10 ng/mL FGF10 (R&D Systems #345-FG), while cells in the CA condition were passed with TrypLE (Thermo Fisher #12604013) at a ratio of 1:2 in B4CF10 with 10 µM Y-27632. B4CF10 medium was changed every 2–3 days and cells were collected between days 10–14 for analysis.
2.8. Thawing of Dissociated DE Aggregates for Further Differentiation
When differentiating cryopreserved DE cells, the same protocols as described above were used to differentiate into each lineage, with some changes to cell seeding density. For hepatic differentiation, cells were seeded at 4 × 104 cells/cm2. For pancreatic differentiation, cells were seeded at 5.26 × 105 cells/cm2. For intestinal differentiation, cells were seeded at 2 × 105 cells/cm2. Media was changed the following day to remove cell debris, and then fed every other day. For lung differentiation, cells were seeded at a density of 1.8 × 105 cells/cm2 for the SD condition and 2.85 × 105 cells/cm2 for the CA condition. After 24 h, the cells were fed with fresh AFE1 and the same protocol was used as described in the lung progenitor differentiation section above.
2.9. DE Differentiation in Controlled, Stirred Bioreactor
DE differentiation in the bioreactor was performed using a DASbox Mini Bioreactor System (Eppendorf). Parallel operation comprised of two independently controlled DASbox Mini bioreactor vessels for cell culture applications equipped with an eight-blade impeller (60° pitch) optimized for hPSC expansion [
9]. An overhead drive allowed for smooth agitation. Sensors for pH and DO monitoring as well as temperature control ensured a tight monitoring and control of critical process parameters. Sensors were calibrated as previously described [
10]. The bioreactors were inoculated with hiPSCs at a density of 5 × 10
5 cells/mL in E8 (in house) and 10 µM Y-27632 (Tocirs #1254) in a final culture volume of 150 mL (75 × 10
6 cells per vessel). Cells were cultivated at 37 °C, stirred at 60 rpm, headspace aerated with 3 sl/hour with 21% O
2 and 5% CO
2. The same feeding protocol was applied as described in the DE differentiation section above. Differentiation into all the lineages was performed using the same protocol used for differentiation in the Erlenmeyer flask.
2.10. Cryosectioning and IF Staining of Day 3 Aggregates
On day 3, aggregates were placed in Tissue Tek
® (Sakura #SA62550-01), frozen on dry ice and stored at −80 °C. Samples were sectioned (10 µm slices) using a microtom, mounted on microscope slides, air-dried for 1 h, and stored at −80 °C. IF staining was performed as described in Supplemental Experimental Procedures. Antibodies used can be found in
Supplementary Materials (Table S1). After staining procedures were performed, slides were covered with cover slips using fluorescent mounting medium (Dako #S3023).
2.11. Immunofluorescent Staining and Quantification
Detailed immunostaining procedures are described in the Supplemental Experimental Procedures. For quantification of hepatic differentiation, 10 grey-scale images were taken in three independent experiments. The DAPI/HNF4α positive area was evaluated using ImageJ with the threshold set between 9–12 for the left limit and 50 for the right for all images within one experiment. For intestinal, lung, and pancreatic progenitor cell quantification, 4–6 images from three independent experiments were quantified using the plugin image-based tool for counting nuclei (ITCN) on ImageJ.
2.12. Flow Cytometry
Flow cytometry was performed on aggregates at day 3 of DE differentiation. Aggregates were dissociated with Accutase (Gibco #A1110501), washed with FACS buffer (0.5% BSA, 3 mM EDTA, PBS), and stained with CXCR4-APC (eBioscience #17-999-42), c-Kit-PE (eBioscience #12-1178-42), and EpCam-PE (BD #347198) at the company-recommended dilution on ice for 30 min. The cells were then washed with FACS buffer and analyzed using the MACSQuant® Analyzer 10. FlowJo V10 software was used to analyze the data.
2.13. Quantitative Real-Time PCR
Total-RNA isolation, reverse transcription, and qRT-PCR procedures are described in detail in the Supplemental Experimental Procedures.
2.14. Aggregate Size and Cell Density Measurement
To monitor aggregate formation and diameters, four independent light microscopic images were captured for each sample (Axiovert A1; Zeiss) followed by diameter analysis via ImageJ. Mean diameters represent arithmetic averages from 76–470 single aggregates. Cell density measurements were taken with Vi-Cell™ XR Cell Viability Analyzer (Beckman Coulter).
2.15. Statistics
Unpaired, parametric, two-tailed t-test was performed on all data sets via GraphPad Prism.
4. Discussion
In the present study, we report the development of a versatile, xeno-free, and scalable protocol for the generation of multipotent DE from hPSCs using industry-scale bioreactors. We also provide evidence that cells can be cryopreserved at the DE stage for later off-the-shelf differentiation and retain their multi-lineage potential.
In the past, one impeding obstacle which prevented the use of dynamic cultures for hPSC applications was hPSC sensitivity for enzymatic dissociation and failure to form proliferating aggregates or single cell-derived clones. This was overcome by the advent of the small molecule Rho kinase inhibitor Y-27632 [
33]. We have shown that with this molecule, undifferentiated hPSC form floating aggregates and can be expanded under defined conditions over several passages while maintaining their three-lineage differentiation potential [
34]. As a logical continuation of our research, we have now addressed the questions as to whether undifferentiated hPSC 3D aggregates can be directly differentiated into DE and its derivatives in dynamic suspension, if industrial scale bioreactor culture is possible, and whether multi-lineage differentiation potential is maintained after cryopreservation at the DE stage.
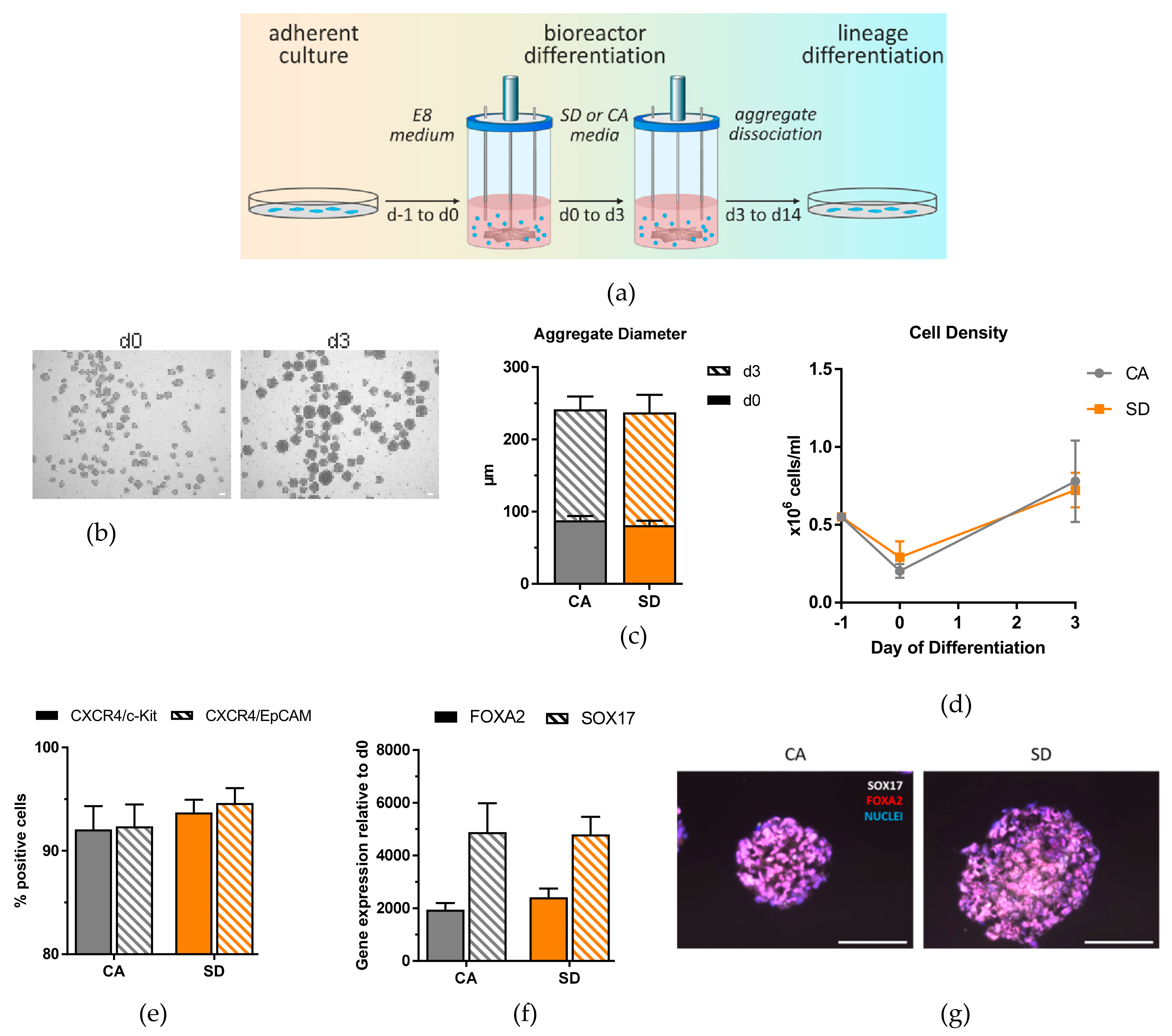
For proof-of-principle, we used Erlenmeyer flasks stirred by an orbital shaker as the main suspension culture system. This system has been well-established, and can be affordably transferred to any laboratory [
35]. Suspension aggregate formation was performed in E8 medium plus Y-27632, representing a defined condition for the culture of undifferentiated hPSCs. This procedure eliminates the need for xeno-matrix substances, like Matrigel
®, which are commonly used as substrates for cell attachment in 2D. Even though both hiPSCs and hESCs were capable of generating stable floating aggregates, the HES3 line was more sensitive to dissociation and aggregate formation during the first 24 h of suspension culture. This suggests that it is important to optimize certain parameters such as cell inoculation density based on the cell line being used. Further optimization in the direction of protecting singularized cells from apoptosis during aggregate formation in dynamic suspension would provide the opportunity to include cell lines that are delicate in this regard, but this was not further addressed in this study.
Characterization of DE aggregates produced in the Erlenmeyer flask revealed that we obtained highly homogeneous DE populations with greater than 92% of cells positive for CXCR4/c-Kit, CXCR4/EpCAM under all conditions for all three cell lines. In addition, the DE transcription factors
SOX17 and
FOXA2 were highly expressed. This is a clear improvement in differentiation efficiencies compared to other suspension protocols [
14,
17], and was an ideal starting point to efficiently derive different downstream lineages. The growth rate of the two conditions and total number of cells retrieved after differentiation were significantly higher than those from 2D adherent cultures, which supports the use of suspension culture differentiation over 2D adherent culture for generating large quantities of DE cells.
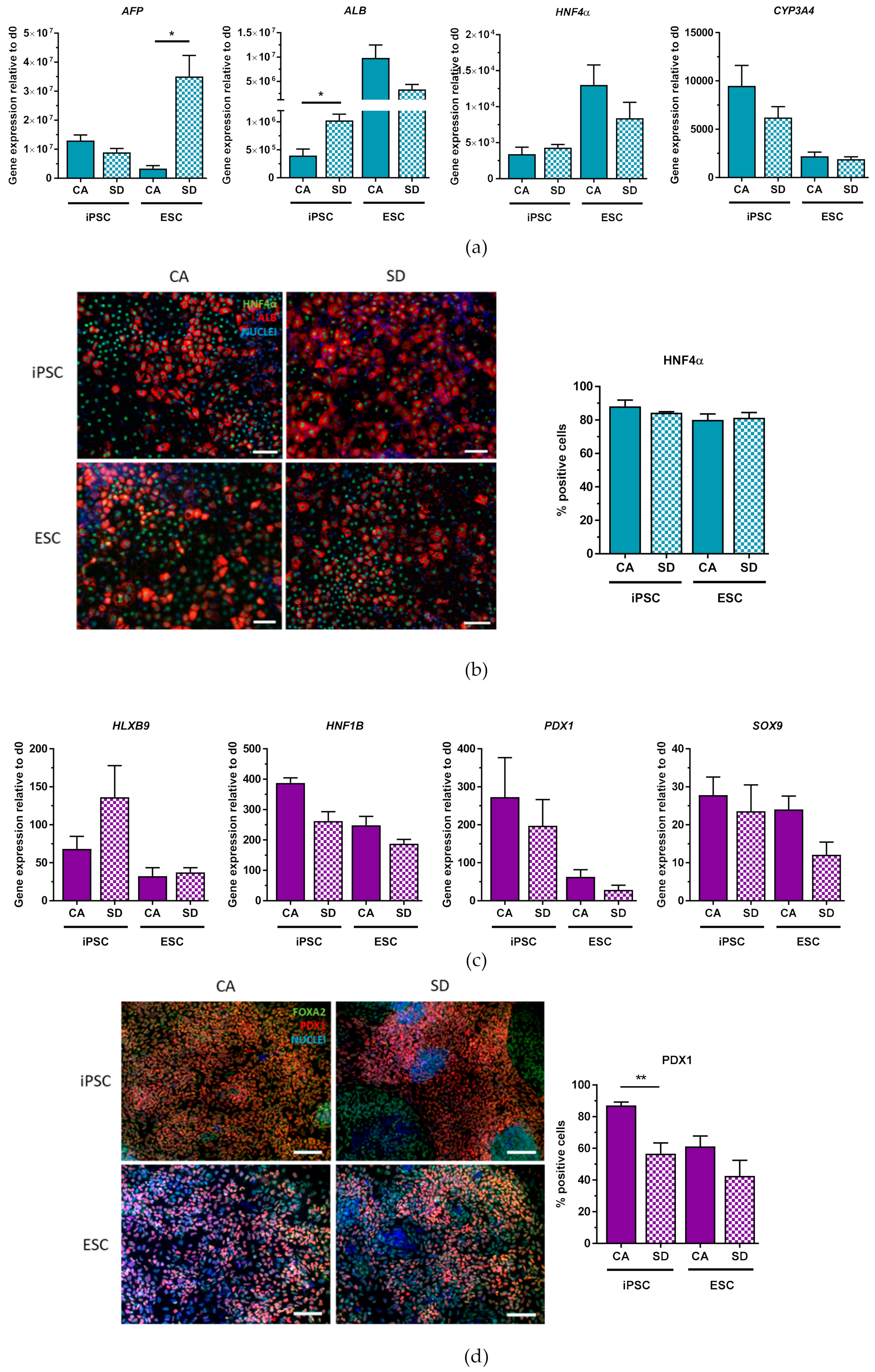
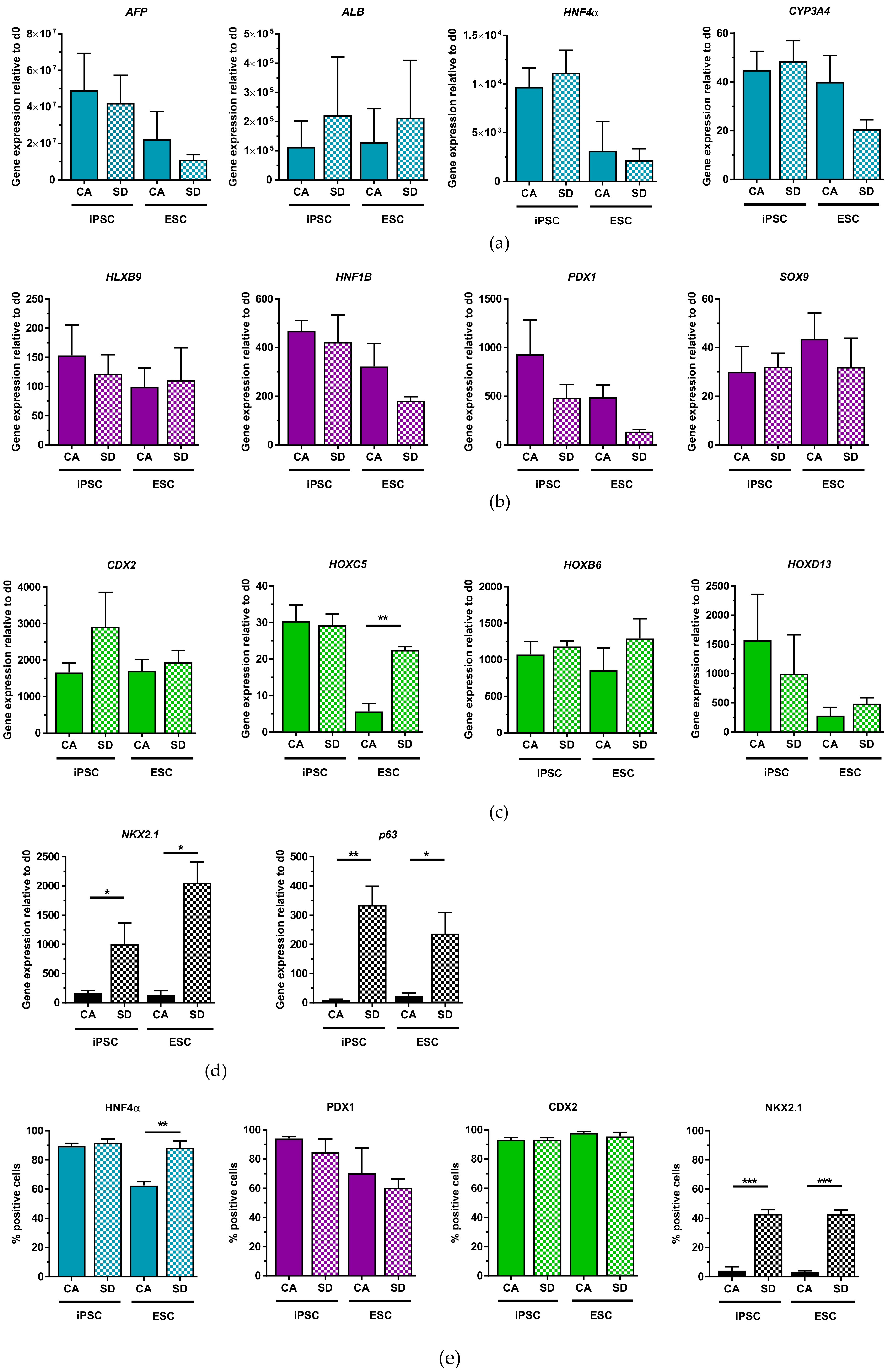
Dissociated DE aggregates generated in Erlenmeyer flasks were capable of differentiating into the different lineages of the DE with similar efficiencies as to what has been published. In general, the HES3 line was slightly less efficient during differentiation. We have not further investigated this observation, but the lower differentiation efficiency may be due to a higher vulnerability of this cell line to the forces acting on the aggregates under dynamic culture conditions. The efficiency of hepatic differentiation achieved was relatively high with 84–88% of cells that were positive for HNF4α, which is an essential transcription factor in the transition of endoderm towards hepatic fate [
36]. This efficiency is even better than results others recently achieved with defined, small molecule-based hepatic differentiation protocol (72% AFP+HN4A+ cells, [
37], 60% ALB, [
13]). Pancreatic lineage potential of our DE aggregates was demonstrated by differentiation towards PDX1+ cells. hiPSC-derived cells from the CA condition differentiated with similar efficiencies (87%) compared to what Yabe et al. reported (92.1%). Intestinal lineage differentiation was highly efficient, with 92–95% of cells expressing CDX2 by day 7 of differentiation. Lung progenitor differentiation of DE aggregates revealed that the SD condition led to significantly more NKX2.1+ cells compared to the CA condition. The reason for this was not addressed in further detail, but it indicates that although our characterization with standard DE markers (CXCR4, c-Kit, EpCAM, FOXA2, and SOX17) on day 3 led to similar and homogeneous results between the two conditions, the DE aggregates generated with the SD condition can more efficiently differentiate towards lung progenitor cells using our lung differentiation protocol. Differentiation towards lung progenitor cells from hiPSCs using the SD condition is comparable to another established adherent lung differentiation protocol which reported up to 40% NKX2.1+ cells using the SD kit [
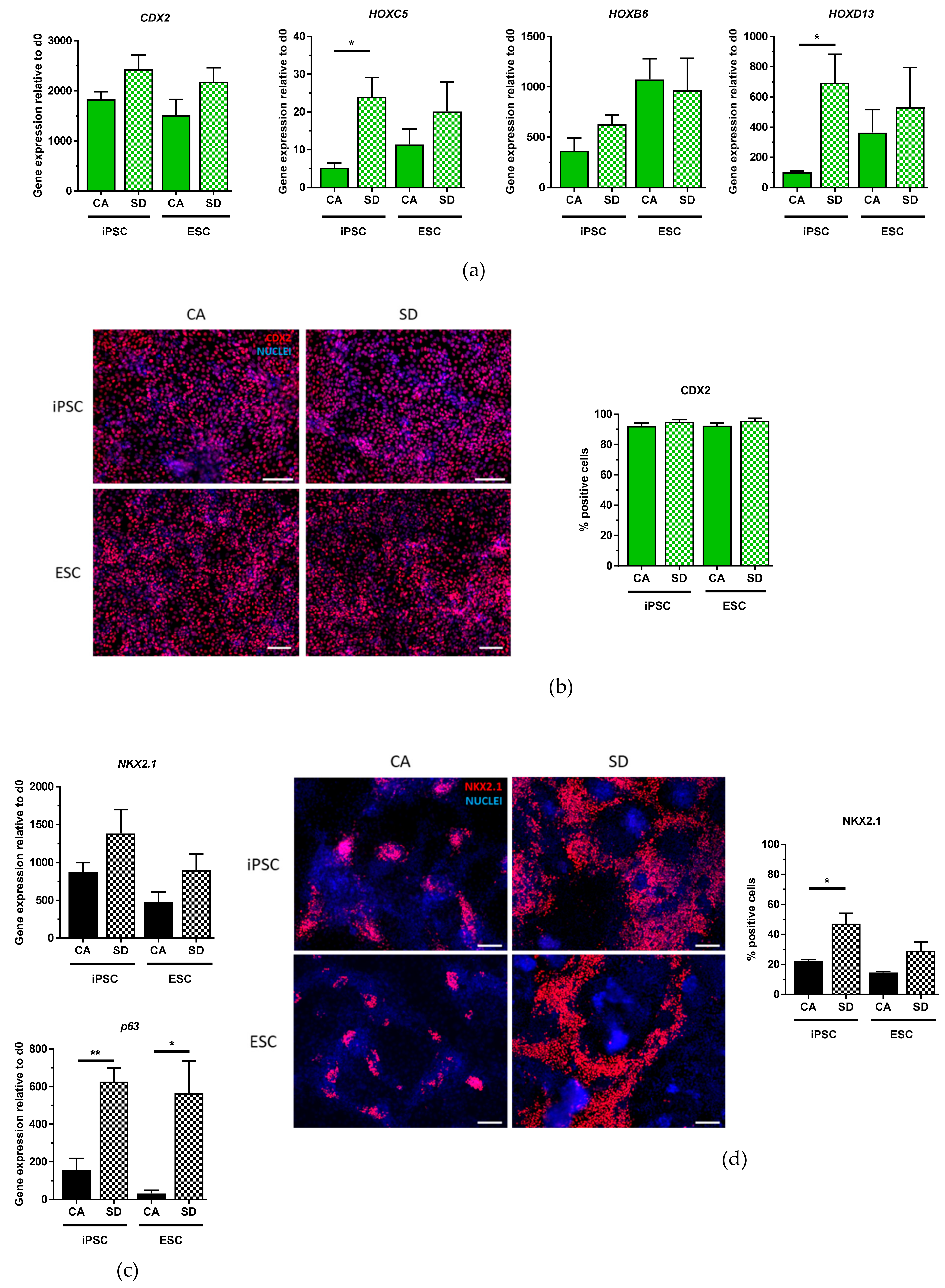
30]. This comparison suggests that our method for DE generation is a competitive alternative to other methods of DE generation not only because of good differentiation efficiency but also because there is the added advantage that only one DE protocol is needed to differentiate towards many lineages of the DE.
After testing the feasibility of suspension culture DE differentiation in Erlenmeyer flasks, the protocol was adapted to a stirred-tank bioreactor, which has been previously shown to maintain hPSCs in suspension [
7] and can be used to differentiate them to cardiomyocytes [
9], endothelial cells [
11], and macrophages [
38]. Interestingly, no further optimization to the media composition was necessary to generate DE in the bioreactor. An average of 1.12 × 10
8 iPSC-derived DE cells could be generated in one 150 mL bioreactor, which equates to 7.5 × 10
5 cells/mL, and is slightly lower than cell numbers achieved in our Erlenmeyer flasks differentiation experiments (1.2 × 10
6 cells/mL). Generated cell numbers in the bioreactor are comparable to what was reported so far. Lock et al. reported 4–5 × 10
5 cells/mL and Shinohara et al. reported 5–8 × 10
5 cell/mL using their suspension culture system [
15,
16]. With our current system, a 2 L bioreactor could provide a sufficient amount of DE cells for human islet differentiation and transplantation [
1], while upscaling to 19 L bioreactor could provide a sufficient amount of DE cells that could be differentiated into hepatic cells to treat a 70 kg patient [
2]. The differences in aggregate diameter, cell density, and marker gene expression between Erlenmeyer flask and bioreactor could be attributed to the physical differences between each platform including shape of vessel, method of stirring, and shear stress as a result of both those factors. Despite these differences, this bioreactor setup is suitable to efficiently generate multipotent DE. There may be further potential for optimizing parameters like stirring speed and impeller design and for harmonizing pH control, dissolved oxygen, and feeding to generate a further increase in cell density and reduce the volume of media needed to attain clinically relevant cell numbers. These variables have to be addressed in future work. With our current bioreactor system and protocol, we can successfully generate large quantities of DE cells that can be used for multi-lineage differentiation that can potentially be used for cell therapy, drug screening, and disease-modelling purposes.
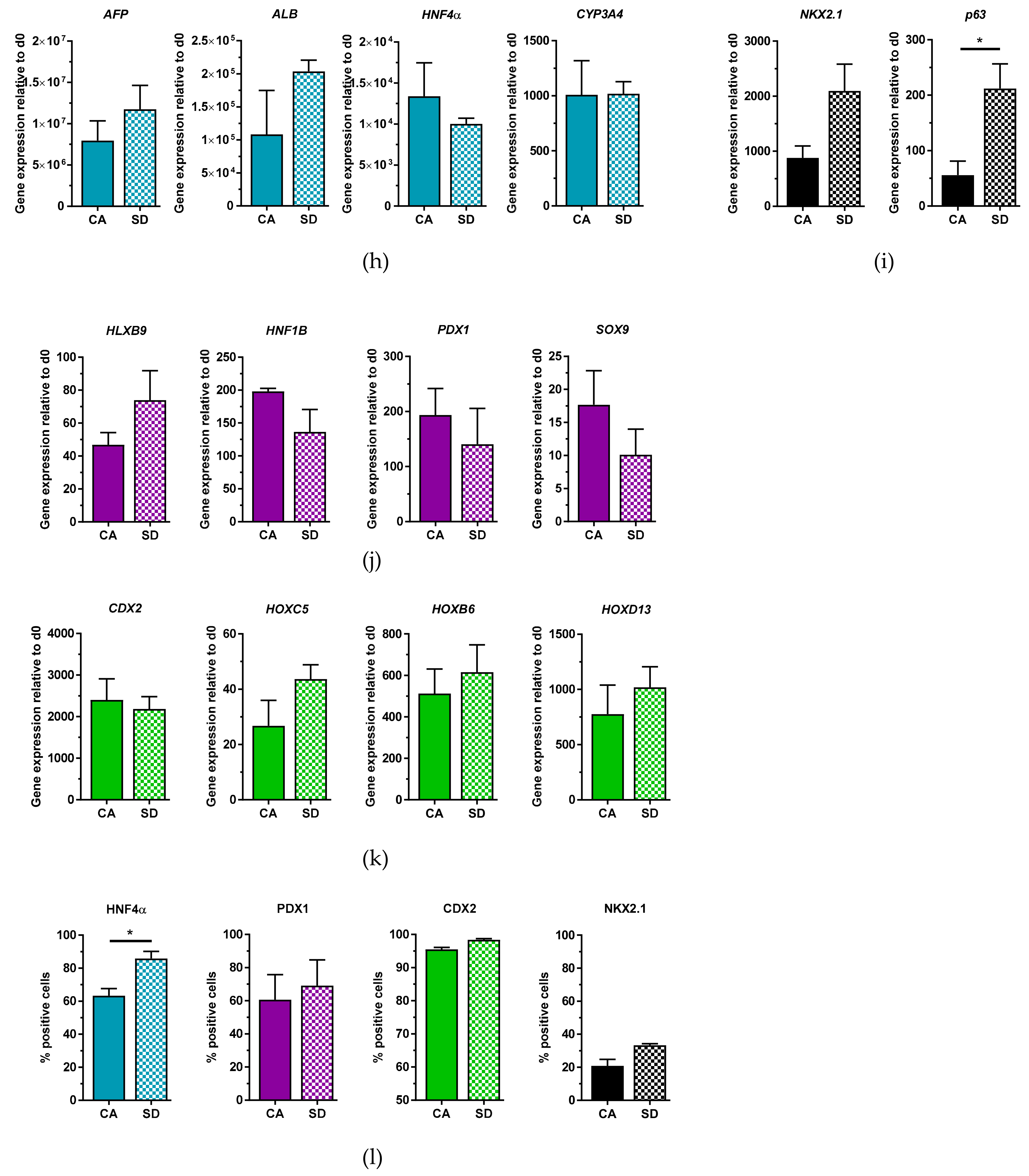
The last point we addressed was the ability to cryopreserve dissociated DE aggregates generated in the Erlenmeyer flask. Cryopreservation of DE cells allows for the storage of DE cells for future experiments, it reduces costs as one large stock of cells can be created, reducing the number of differentiations performed, and it can improve reproducibility when performing experiments. In addition, cryopreservation could separate the steps of mass cell production, cell-type-specific differentiation and final therapeutic or scientific destination. For these reasons, we sought to cryopreserve our DE cells and test whether they maintain their multi-lineage potential upon thawing. Our results show that cryopreserved DE cells from the CA and SD conditions were capable of differentiating towards hepatic-like, pancreatic, and intestinal cells in general with similar efficiency as non-cryopreserved DE cells. However, in the case of hepatic differentiation, expression levels of markers for more mature hepatic cells like
CYP3A4 and
ALB are lower than after differentiation of non-cryopreserved cells, suggesting a delay in differentiation or even a loss of a specific progenitor subpopulation due to cryopreservation. Differences in the thawed iPSC-derived DE cells from the CA condition were also observed during intestinal differentiation. These cells expressed higher levels of
HOXC5 and
HOXD13 compared to non-cryopreserved cells, suggesting a stimulating effect of cryopreservation that resulted in better differentiation. However, the same pattern of higher expression of
HOXB6 and
HOXD13 compared to
HOXC5 can be observed in the Erlenmeyer flask, the bioreactor, and upon thawing, which suggests that these cells retain their hindgut phenotype even after cryopreservation [
31]. While lung differentiation of thawed DE cells from the SD condition was as efficient as non-cryopreserved cells, differentiation of thawed cells from the CA condition was less efficient. One possible reason for this is that the differentiation protocol and media must be further optimized to support higher
NKX2.1 expression of cells from the CA condition. Seeding density of thawed cells may also need to be further optimized, as we have seen that this greatly affects differentiation efficiency. Furthermore, DE cells primed for anterior foregut differentiation may be more sensitive to cryopreservation. Despite the lower lung progenitor differentiation efficiency in the CA condition and a possible delayed maturation of hepatic cells, DE cells from both SD and CA conditions maintain their multi-lineage potential upon cryopreservation, making the process for DE generation and differentiation more flexible.
Taken together, our study provides a versatile, robust, and highly efficient protocol for the generation of DE in dynamic suspension cultures with industrial/clinical scale potential and enables off-the-shelf supply of DE cells for production of different DE lineages. The described protocols represent a major step towards the routine application of cellular therapies, toxicity screening, and disease modeling with endoderm cell types derived from hPSCs.